VIRAL GENETICS PATHOGENESIS LIFE CYCLES VACCINE DEVELOPMENT DRUG























- Slides: 37

VIRAL GENETICS • • PATHOGENESIS LIFE CYCLES VACCINE DEVELOPMENT DRUG RESISTANCE 1

VIRAL GENETICS “DNA chromosomes of eukaryotic host organisms generally require geologic time spans to evolve to the degree that their RNA viruses can achieve in a single human generation. ” 2

VIRAL GENETICS • VIRUSES GROW RAPIDLY • A SINGLE PARTICLE PRODUCES A LOT OF PROGENY • DNA VIRUSES SEEM TO HAVE ACCESS TO PROOF READING, RNA VIRUSES DO NOT SEEM TO 3

NATURE OF GENOMES • RNA or DNA • SEGMENTED OR NON-SEGMENTED 4

GENETIC CHANGE • MUTATION • RECOMBINATION 5

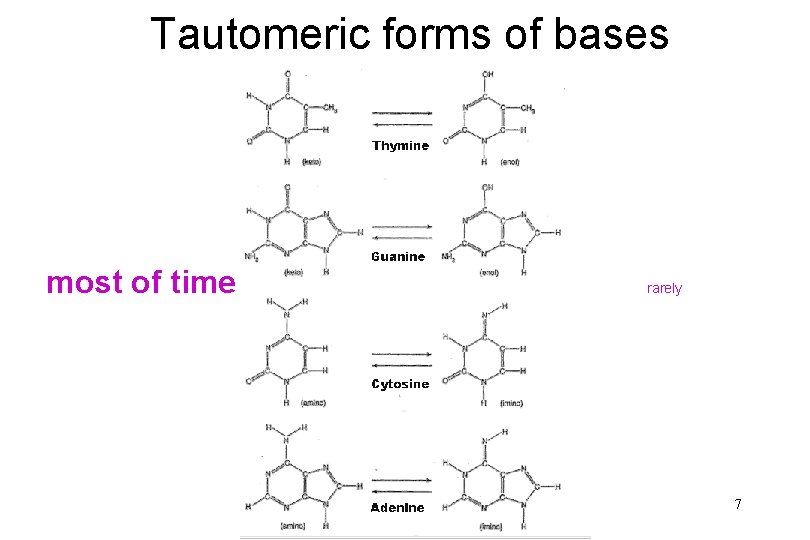
ORIGIN OF MUTATIONS • SPONTANEOUS – tautomeric form of bases – polymerase errors 6

Tautomeric forms of bases most of time rarely 7

ORIGIN OF MUTATIONS • SPONTANEOUS – tautomeric form of bases – polymerase errors mutation rates usually higher in RNA viruses (lack of proof reading) why do some viruses seem to alter very little, even though one would expect high mutation rates? 8

ORIGIN OF MUTATIONS • SPONTANEOUS • PHYSICALLY INDUCED – UV light , especially problem if no access to repair – X-rays • CHEMICALLY INDUCED 9

TYPES OF MUTATION • POINT • INSERTION • DELETION 10

PHENOTYPES PHENOTYPE – the observed properties of an organism 11

PHENOTYPIC CHANGES • CONDITIONAL LETHAL - multiply under some conditions but not others - wild-type (wt) grows under both sets of conditions • temperature-sensitive (ts) mutants do not grow at higher temperature (altered protein) • host-range mutants do not grow in all the cell types that the wt does 12

PHENOTYPIC CHANGES • PLAQUE SIZE – may show altered pathogenicity • DRUG RESISTANCE – important in the development of antiviral agents • ENZYME-DEFICIENT MUTANTS – some genes can be ‘optional’ in certain circumstances 13

PHENOTYPIC CHANGES • “HOT MUTANTS” – grow better at elevated temperature than wt – less susceptible to host fever response • ATTENUATED MUTANTS – milder (or no) symptoms – vaccine development – pathogenesis 14

GENETIC CHANGE • MUTATION • RECOMBINATION 15

RECOMBINATION Exchange of information between two genomes 16

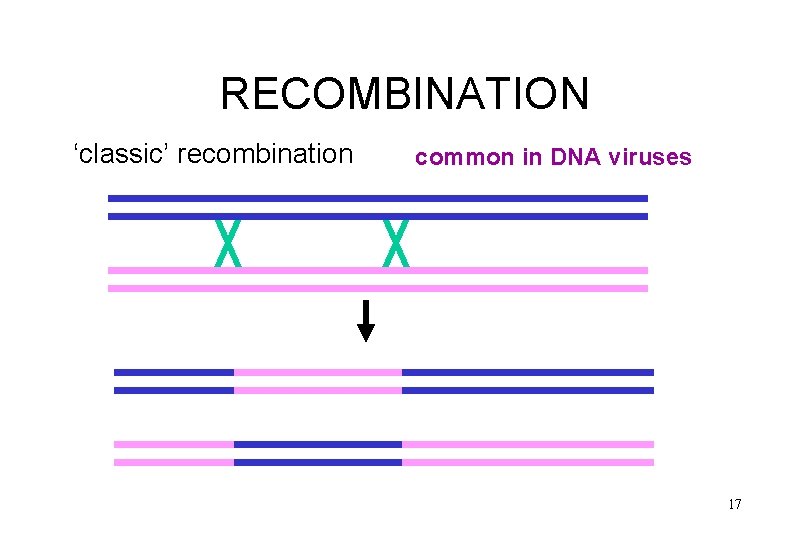
RECOMBINATION ‘classic’ recombination common in DNA viruses 17

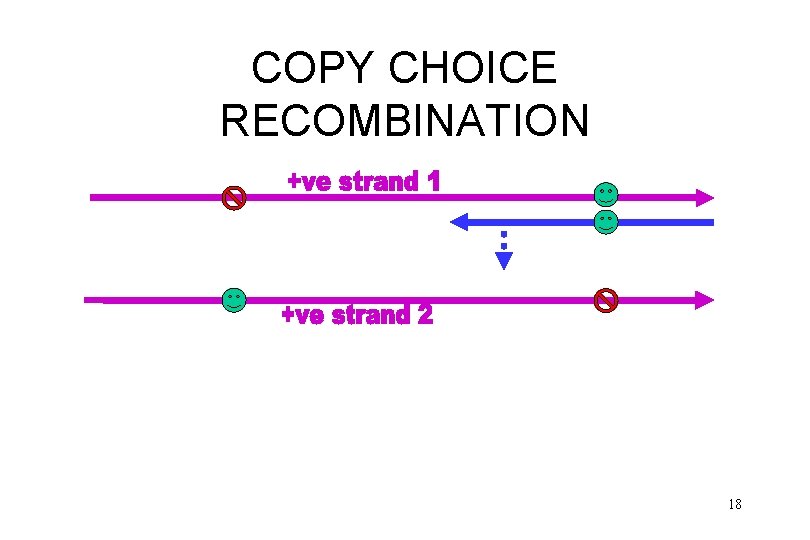
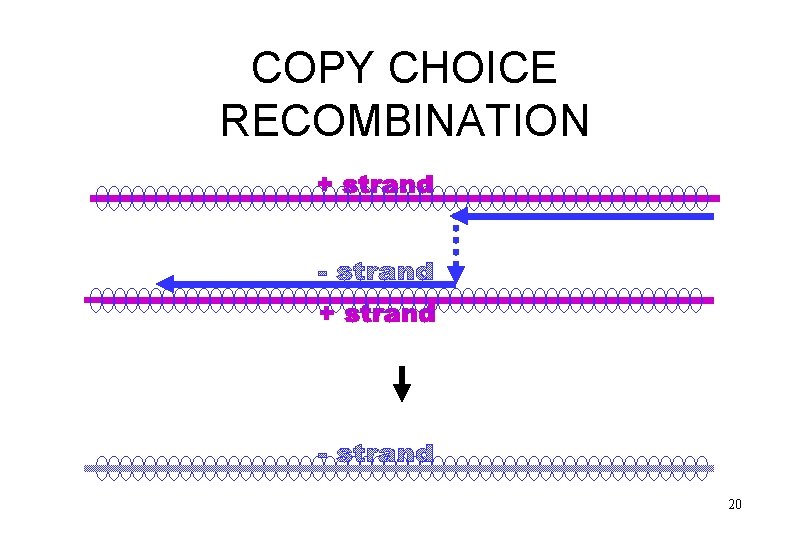
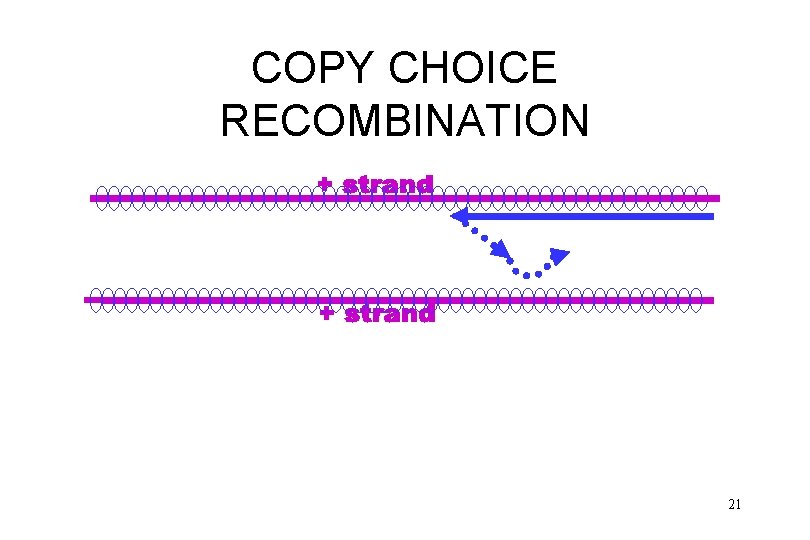
COPY CHOICE RECOMBINATION 18

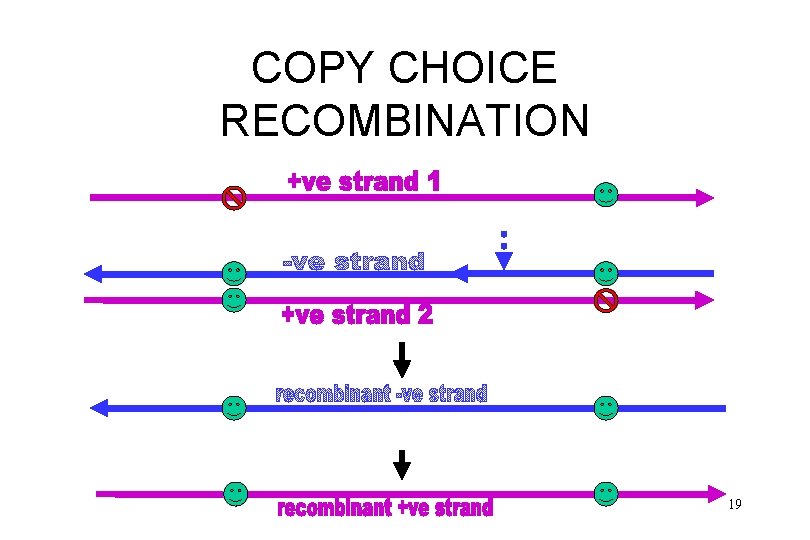
COPY CHOICE RECOMBINATION 19

COPY CHOICE RECOMBINATION 20

COPY CHOICE RECOMBINATION 21

Other methods recombination • Take advantage quirks in virus replication – eg. Coronaviruses (include SARS virus) 22

RECOMBINATION - SOME USES • mapping by recombination frequency • mapping by marker rescue 23

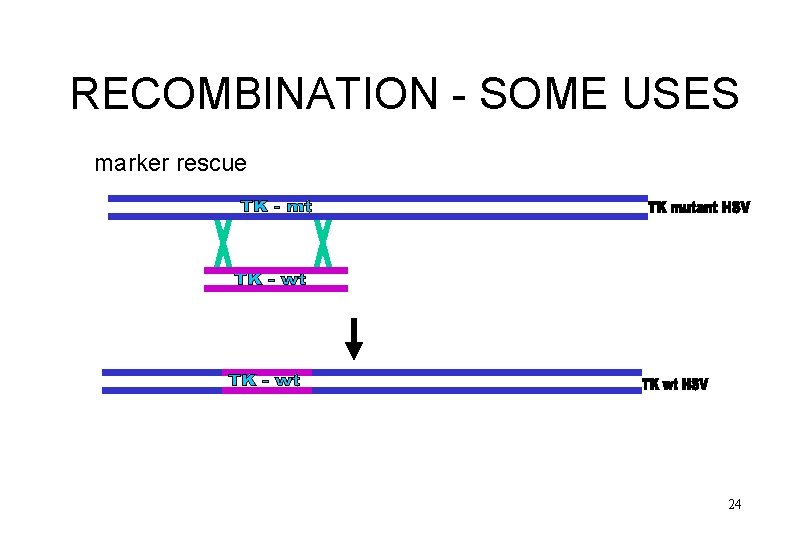
RECOMBINATION - SOME USES marker rescue 24

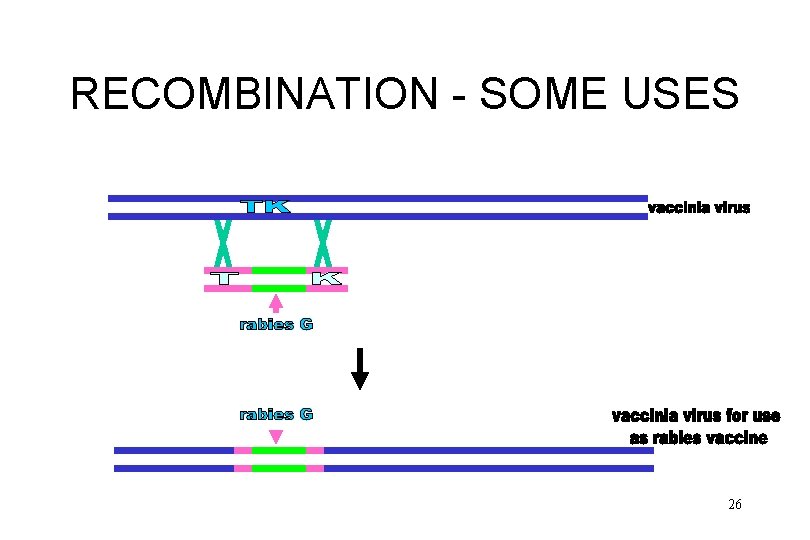
RECOMBINATION - SOME USES • mapping by recombination frequency • mapping by marker rescue • development of recombinant viruses for vaccines and therapeutic reasons 25

RECOMBINATION - SOME USES 26

raccoon eating bait with rabies vaccine in it 27

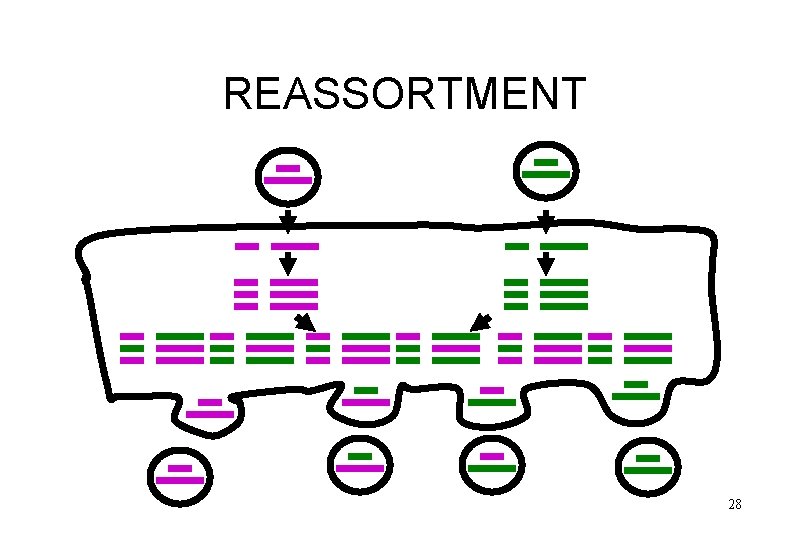
REASSORTMENT 28

REASSORTMENT • • form of recombination (non classical) very efficient segmented viruses only used in some new vaccines 29

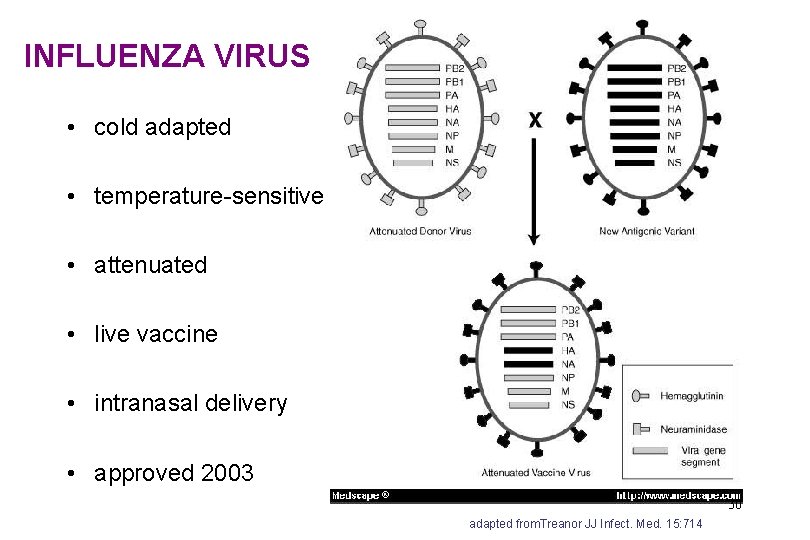
INFLUENZA VIRUS • cold adapted • temperature-sensitive • attenuated • live vaccine • intranasal delivery • approved 2003 30 adapted from. Treanor JJ Infect. Med. 15: 714

NON-SEGMENTED NEGATIVE STRAND RNA VIRUSES • no classical recombination • no copy choice • no reassortment – least ability to exchange genetic material 31

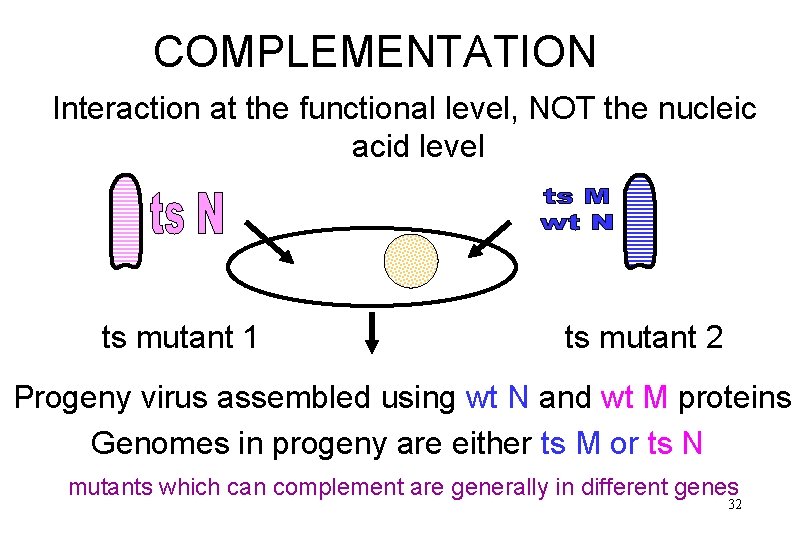
COMPLEMENTATION Interaction at the functional level, NOT the nucleic acid level ts mutant 1 ts mutant 2 Progeny virus assembled using wt N and wt M proteins Genomes in progeny are either ts M or ts N mutants which can complement are generally in different genes 32

DEFECTIVE VIRUSES • lack gene(s) necessary for a complete infectious cycle • ‘helper’ virus provides missing functions package me! copy me! 33

DEFECTIVE VIRUSES • some examples of defective viruses – some retroviruses (use related helper) – hepatitis delta virus (uses unrelated helper) 34

DEFECTIVE INTERFERING (DI) VIRUSES (PARTICLES) • decrease replication of helper virus – compete for viral precursors, etc. • may modulate wt infections • occur naturally eg. DI measles virus in subacute scelerosing panencephalitis - SSPE 35

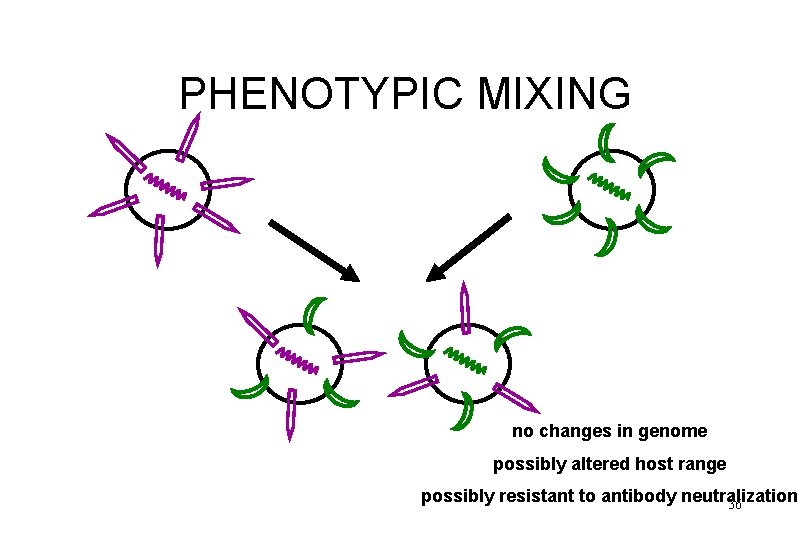
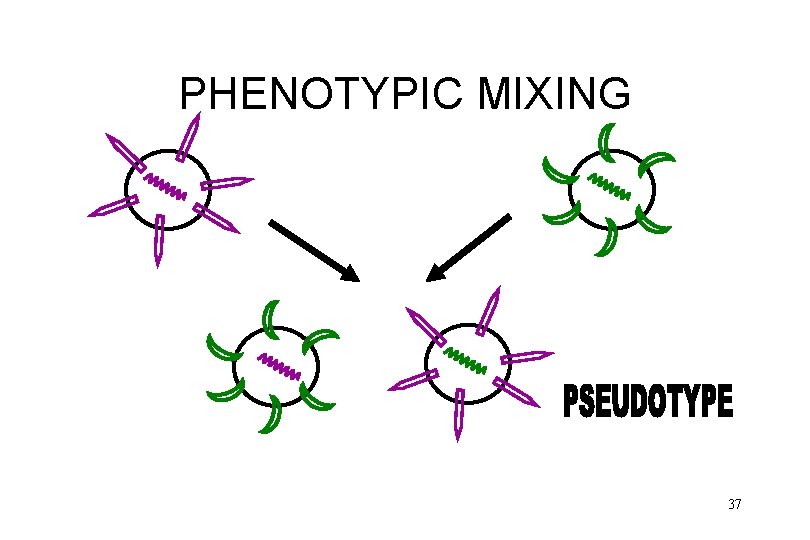
PHENOTYPIC MIXING no changes in genome possibly altered host range possibly resistant to antibody neutralization 36

PHENOTYPIC MIXING 37