Vibrational Predissociation Spectra in the Shared Proton Region

Vibrational Predissociation Spectra in the Shared Proton Region of Protonated Formic Acid Wires: Characterizing Proton Motion in Linear H-Bonded Networks Helen K. Gerardi 6/24/2010 th The 65 International Symposium on Molecular Spectroscopy
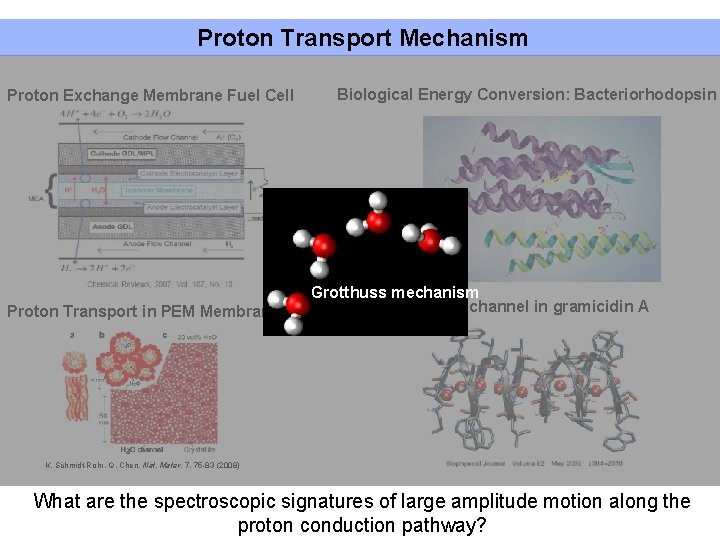
Proton Transport Mechanism Proton Exchange Membrane Fuel Cell Proton Transport in PEM Membrane Biological Energy Conversion: Bacteriorhodopsin Grotthuss mechanism Proton channel in gramicidin A K. Schmidt-Rohr, Q. Chen, Nat. Mater. 7, 75 -83 (2008) What are the spectroscopic signatures of large amplitude motion along the proton conduction pathway?
Outline of Talk Goal: Spectral characterization of mobile proton in H-bonded clusters • Method used to obtain resolved structure of intermolecular proton • Application to proton motion in protonated imidazole clusters (N–H·····N) • Application to proton motion in protonated formic acid clusters (O–H·····O) Ø Identification of low-frequency modes O C O • Effect of cluster size on vibrational features in spectra of protonated formic acid complexes.
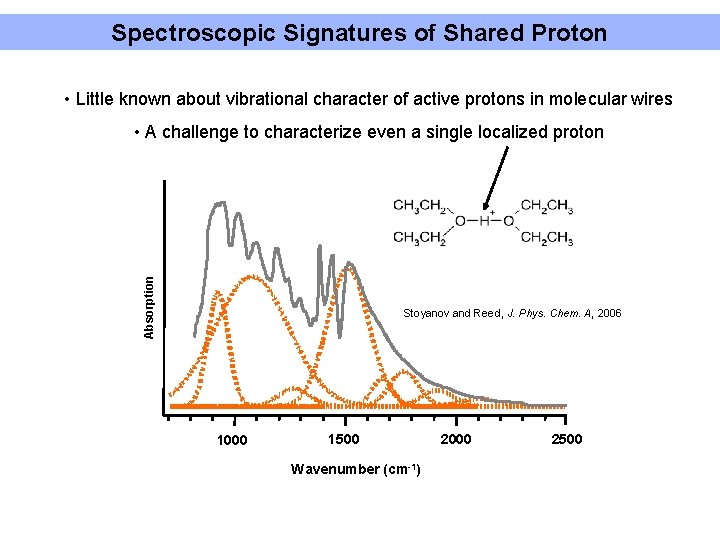
Spectroscopic Signatures of Shared Proton • Little known about vibrational character of active protons in molecular wires Absorption • A challenge to characterize even a single localized proton Stoyanov and Reed, J. Phys. Chem. A, 2006 1000 1500 Wavenumber (cm-1) 2000 2500
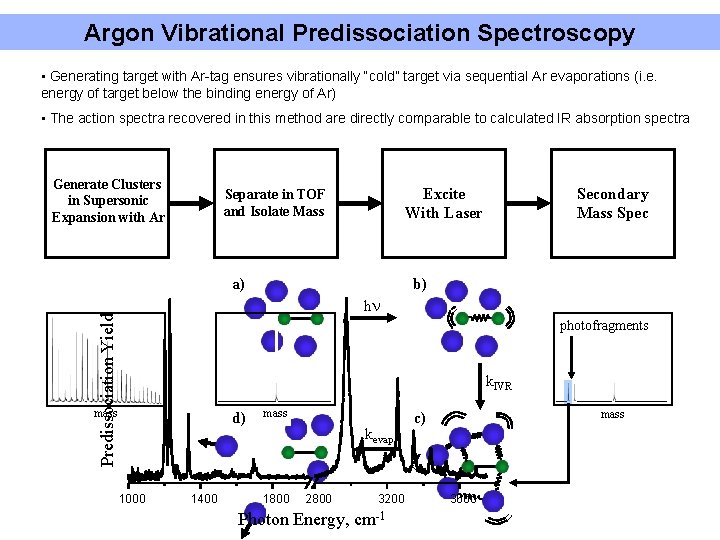
Argon Vibrational Predissociation Spectroscopy • Generating target with Ar-tag ensures vibrationally “cold” target via sequential Ar evaporations (i. e. energy of target below the binding energy of Ar) • The action spectra recovered in this method are directly comparable to calculated IR absorption spectra Generate Clusters in Supersonic Expansion with Ar Excite With Laser Separate in TOF and Isolate Mass a) Secondary Mass Spec b) Predissociation Yield hn photofragments k. IVR mass d) 1000 1400 mass kevap 1800 2800 Photon Energy, 3200 cm-1 mass c) 3600
![Spectroscopic Signatures of Shared Proton Vibrational Predissociation Spectroscopy Absorption Yield Ar Predissociation [RH+·Ar] + Spectroscopic Signatures of Shared Proton Vibrational Predissociation Spectroscopy Absorption Yield Ar Predissociation [RH+·Ar] +](http://slidetodoc.com/presentation_image/728b5c9db2719316ff6728c2ff3522d1/image-6.jpg)
Spectroscopic Signatures of Shared Proton Vibrational Predissociation Spectroscopy Absorption Yield Ar Predissociation [RH+·Ar] + h → [RH+] + Ar Stoyanov and Reed, J. Phys. Chem. A, 2006 Roscioli and Johnson, Science, 2007 1000 1500 2000 2500 Wavenumber (cm-1) 3000 3500
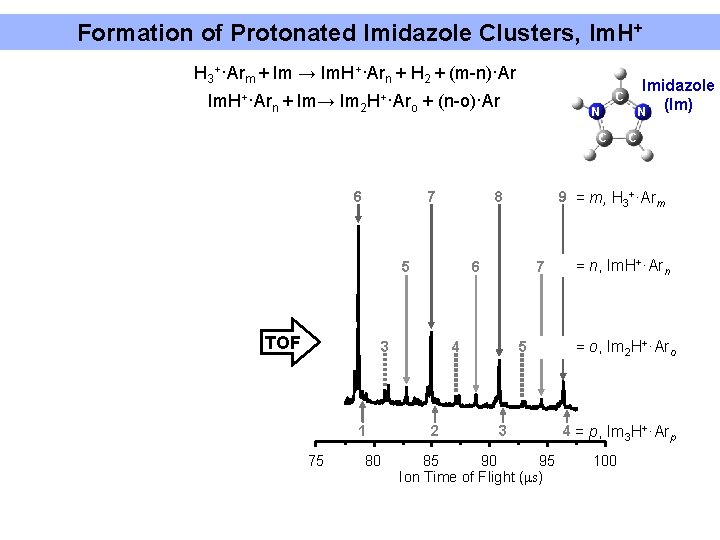
Formation of Protonated Imidazole Clusters, Im. H+ H 3+·Arm + Im → Im. H+·Arn + H 2 + (m-n)·Ar Im. H+·Ar n+ Im→ Im 2 H+·Ar o + (n-o)·Ar N C 6 7 TOF 6 3 1 75 80 7 4 2 C 9 = m, H 3+·Arm 8 5 Imidazole (Im) N C 5 3 85 90 95 Ion Time of Flight (μs) = n, Im. H+·Arn = o, Im 2 H+·Aro 4 = p, Im 3 H+·Arp 100
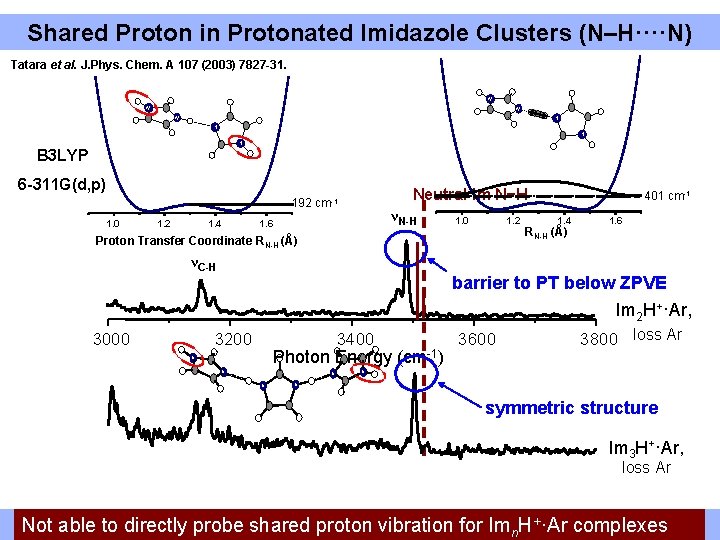
Shared Proton in Protonated Imidazole Clusters (N–H····N) Tatara et al. J. Phys. Chem. A 107 (2003) 7827 -31. N N N N B 3 LYP 6 -311 G(d, p) Neutral Im N−H 192 cm-1 1. 0 1. 2 1. 4 νN-H 1. 6 1. 0 1. 2 Proton Transfer Coordinate RN-H (Å) νC-H 3000 401 cm-1 1. 4 RN-H (Å) 1. 6 barrier to PT below ZPVE Im 2 H+·Ar, 3200 3400 Photon Energy (cm-1) N N 3600 3800 loss Ar N N symmetric structure Im 3 H+·Ar, loss Ar Freeto N-H stretching modes proton returnvibration to neutral energies Not able directly probe shared fortransition Imn. H+·Ar complexes
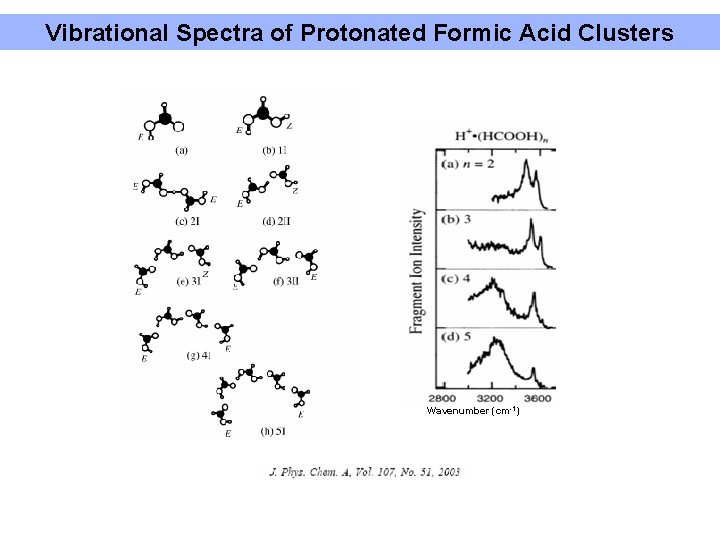
Vibrational Spectra of Protonated Formic Acid Clusters Wavenumber (cm-1)
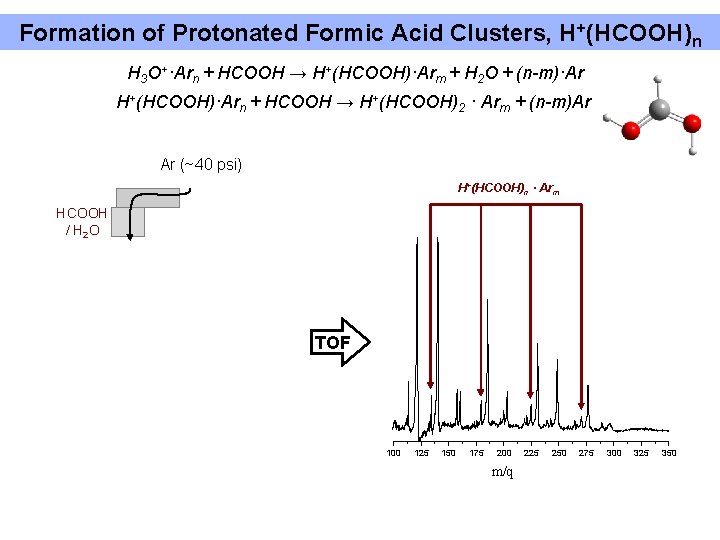
Formation of Protonated Formic Acid Clusters, H+(HCOOH)n H 3 O+·Arn + HCOOH → H+(HCOOH)·Arm + H 2 O + (n-m)·Ar H+(HCOOH)·Arn + HCOOH → H+(HCOOH)2 · Arm + (n-m)Ar Ar (~40 psi) H+(HCOOH)n · Arm HCOOH / H 2 O TOF 100 125 150 175 200 m/q 225 250 275 300 325 350
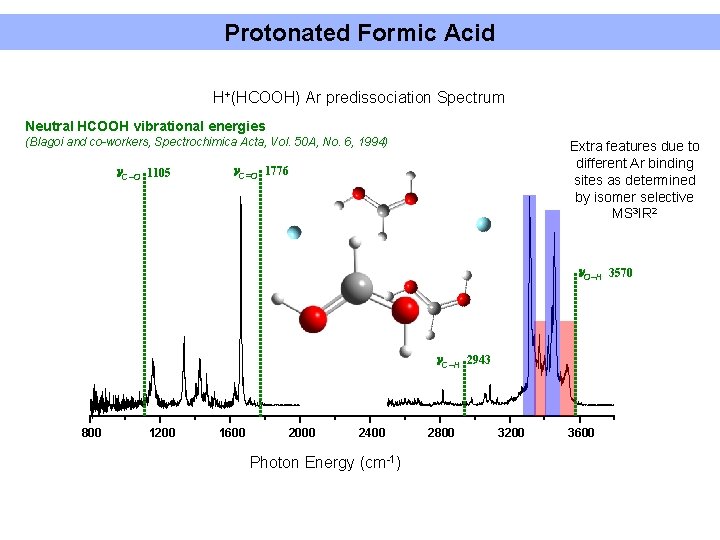
Protonated Formic Acid H+(HCOOH) Ar predissociation Spectrum Neutral HCOOH vibrational energies (Blagoi and co-workers, Spectrochimica Acta, Vol. 50 A, No. 6, 1994) C–O 1105 Extra features due to different Ar binding sites as determined by isomer selective MS 3 IR 2 C=O 1776 O–H 3570 C–H 2943 800 1200 1600 2000 2400 Photon Energy (cm-1) 2800 3200 3600
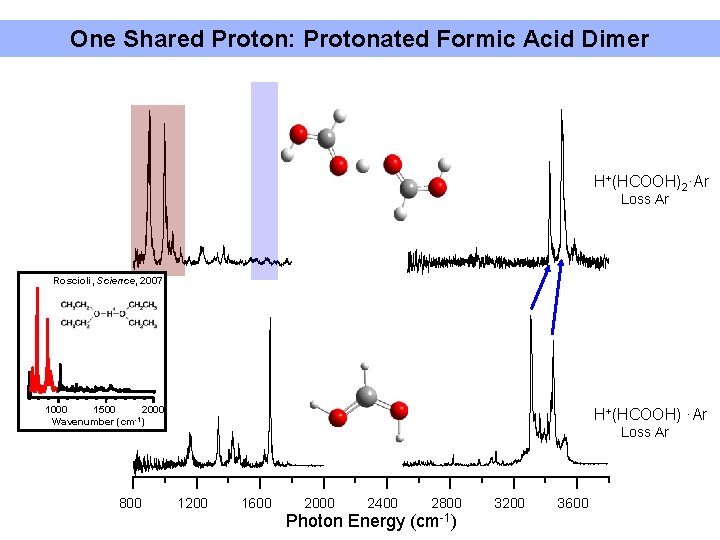
One Shared Proton: Protonated Formic Acid Dimer H+(HCOOH)2·Ar Loss Ar Roscioli, Science, 2007 1500 2000 1000 Wavenumber (cm-1) 800 H+(HCOOH) ·Ar Loss Ar 1200 1600 2000 2400 2800 Photon Energy (cm-1) 3200 3600
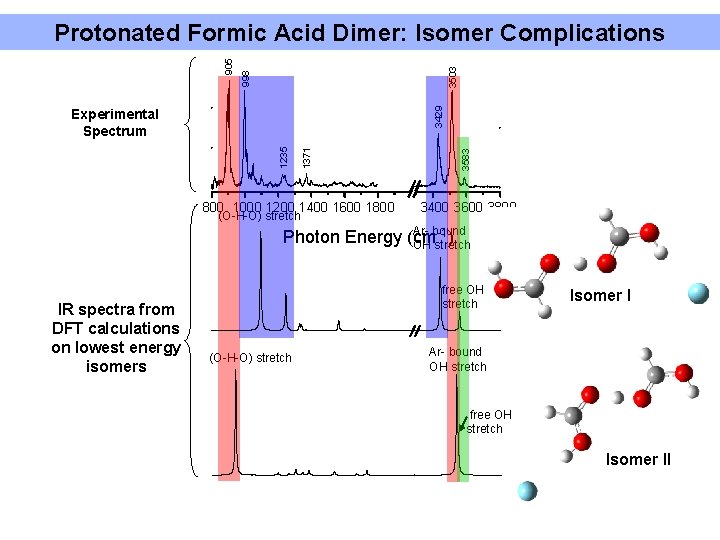
3503 1371 1235 Experimental Spectrum 800 1000 1200 1400 1600 1800 (O-H-O) stretch 3583 3429 998 905 Protonated Formic Acid Dimer: Isomer Complications 3400 3600 3800 Ar- bound -1) Photon Energy (cm OH stretch IR spectra from DFT calculations on lowest energy isomers free OH stretch (O-H-O) stretch Isomer I Ar- bound OH stretch free OH stretch Isomer II
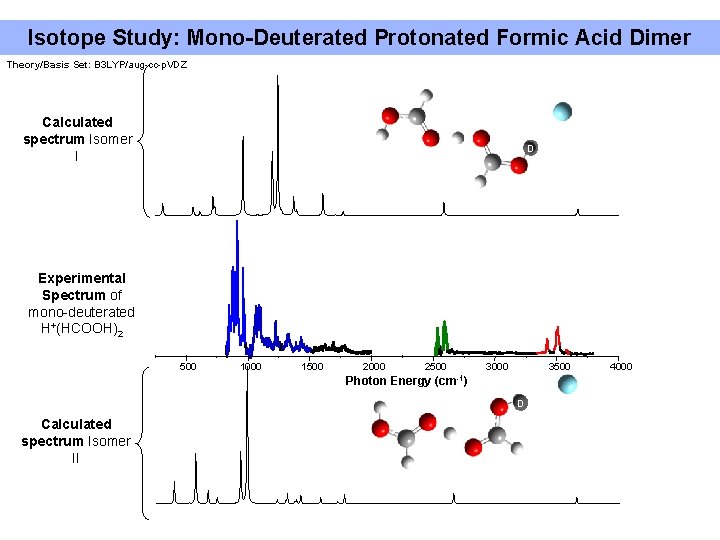
Isotope Study: Mono-Deuterated Protonated Formic Acid Dimer Theory/Basis Set: B 3 LYP/aug-cc-p. VDZ Calculated spectrum Isomer I D Experimental Spectrum of mono-deuterated H+(HCOOH)2 500 1000 1500 2000 2500 Photon Energy 3000 3500 (cm-1) D Calculated spectrum Isomer II 4000
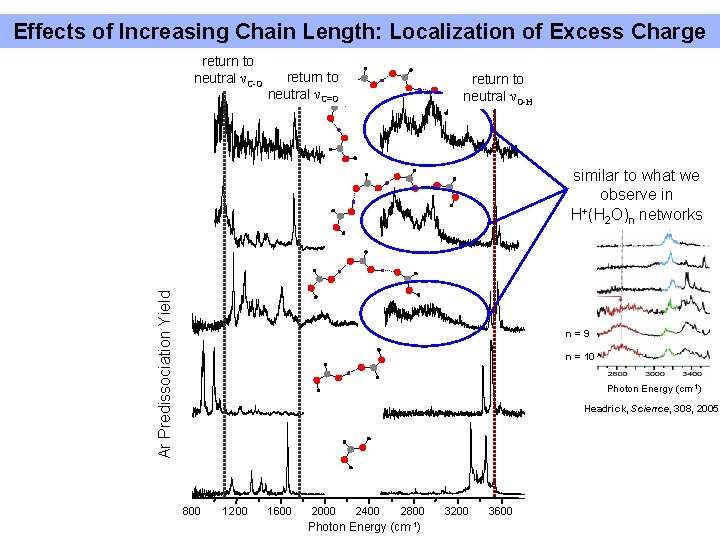
Effects of Increasing Chain Length: Localization of Excess Charge return to neutral νC-O return to neutral νC=O return to neutral νO-H Ar Predissociation Yield similar to what we observe in + H (H 2 O)n networks n=9 n = 10 Photon Energy (cm-1) Headrick, Science, 308, 2005 800 1200 1600 2000 2400 Photon Energy 2800 (cm-1) 3200 3600

Conclusions and Future Work From protonated imidazole wires: • Protonated imidazole dimer acts as a symmetric complex even though equilibrium structure is a double-minimum • Systematic blue-shift of N-H stretch to higher energies towards that of neutral imidazole • Make another attempt to obtain low-frequency spectra for these complexes From protonated formic acid wires: • Many isomers in play even for monomer, H+(HCOOH) • Sharp spectral features recovered in 800 -1000 cm-1 range for the dimer complex attributed to parallel stretching mode of shared proton. • Increasing chain length of the formic acid chains results in trend from n=3 -5 toward neutral formic acid spectrum, with broad features in 26003200 cm-1 region observed previously for large water networks isolated in gas phase

Acknowledgments The Johnson Group • Chris Leavitt • George Gardenier • Mark Johnson Collaborators at UMass: • Usha Viswanathan • Scott Auerbach
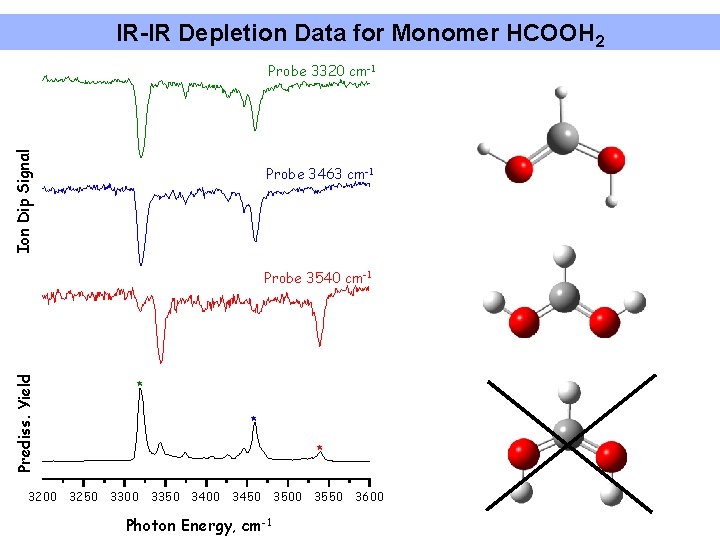
IR-IR Depletion Data for Monomer HCOOH 2 Ion Dip Signal Probe 3320 cm-1 Probe 3463 cm-1 Prediss. Yield Probe 3540 cm-1 * * * 3200 3250 3300 3350 3400 3450 3500 3550 3600 Photon Energy, cm-1
- Slides: 18