Vibrational Infrared Spectroscopy vibrational modes CO equilibrium bond
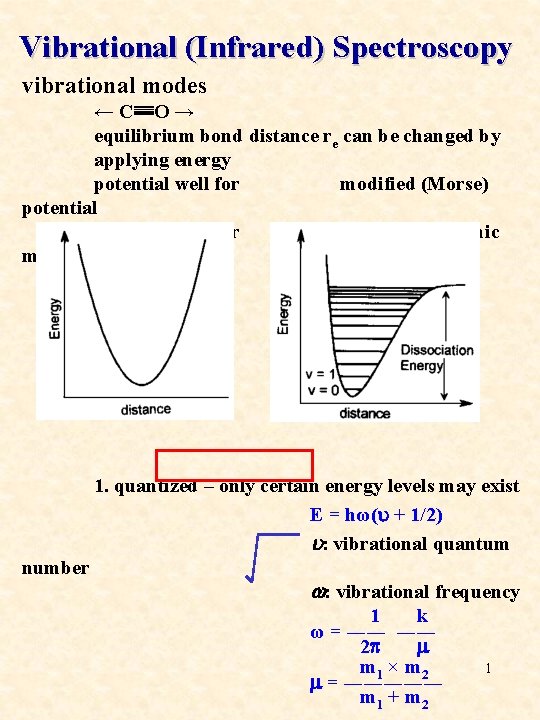
Vibrational (Infrared) Spectroscopy vibrational modes ← C≣≣O → equilibrium bond distance re can be changed by applying energy potential well for modified (Morse) potential classical vibrator well for a diatomic molecule 1. quantized – only certain energy levels may exist E = hω(u + 1/2) u: vibrational quantum number w: vibrational frequency 1 k ω = ―― ―― 2 p m m 1 × m 2 1 m = ――――― m 1 + m 2
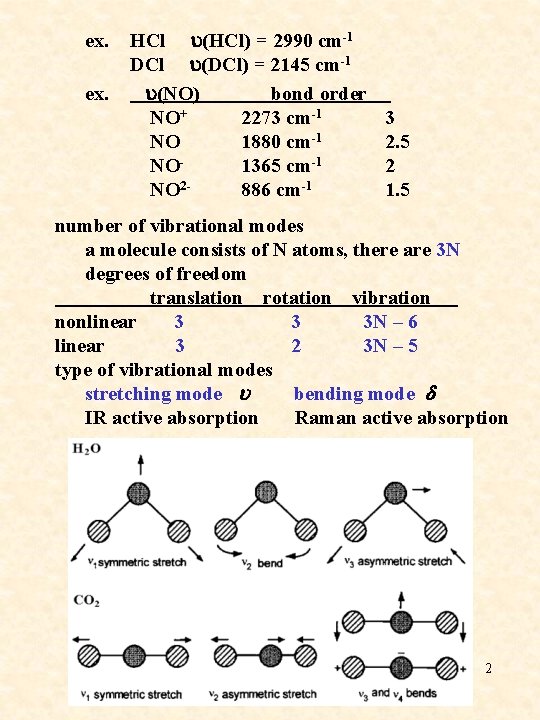
ex. HCl u(HCl) = 2990 cm-1 DCl u(DCl) = 2145 cm-1 u(NO) bond order NO+ 2273 cm-1 NO 1880 cm-1 NO 1365 cm-1 NO 2886 cm-1 3 2. 5 2 1. 5 number of vibrational modes a molecule consists of N atoms, there are 3 N degrees of freedom translation rotation vibration nonlinear 3 3 3 N – 6 linear 3 2 3 N – 5 type of vibrational modes stretching mode u bending mode d IR active absorption Raman active absorption 2
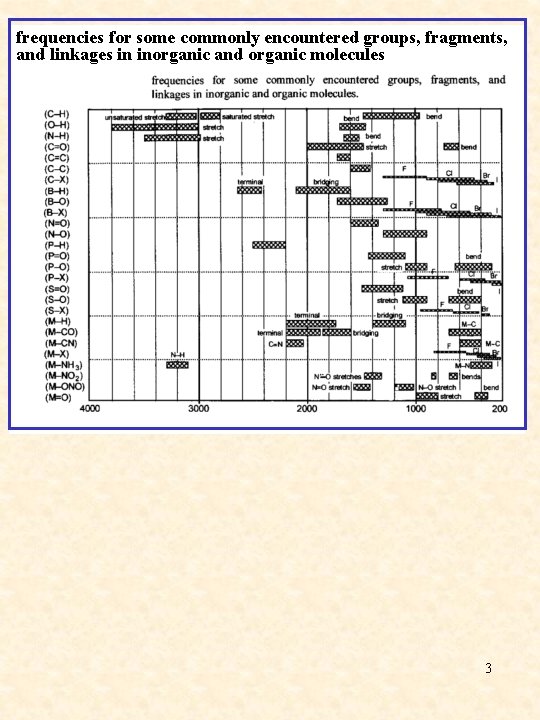
frequencies for some commonly encountered groups, fragments, and linkages in inorganic and organic molecules 3
![ex. W(CO)6 Mn(CO)5 Br compound [Ti(CO)6]2 [V(CO)6] Cr(CO)6 [Mn(CO)6]+ u(CO) (cm-1) 1740 1860 ex. W(CO)6 Mn(CO)5 Br compound [Ti(CO)6]2 [V(CO)6] Cr(CO)6 [Mn(CO)6]+ u(CO) (cm-1) 1740 1860](http://slidetodoc.com/presentation_image/bd3919b8e899548b1a504528a60ed367/image-4.jpg)
ex. W(CO)6 Mn(CO)5 Br compound [Ti(CO)6]2 [V(CO)6] Cr(CO)6 [Mn(CO)6]+ u(CO) (cm-1) 1740 1860 2000 2095 stretching modes of CO and IR frequencies (a) terminal (b) doubly bridging (c) triply bridging ex. 4
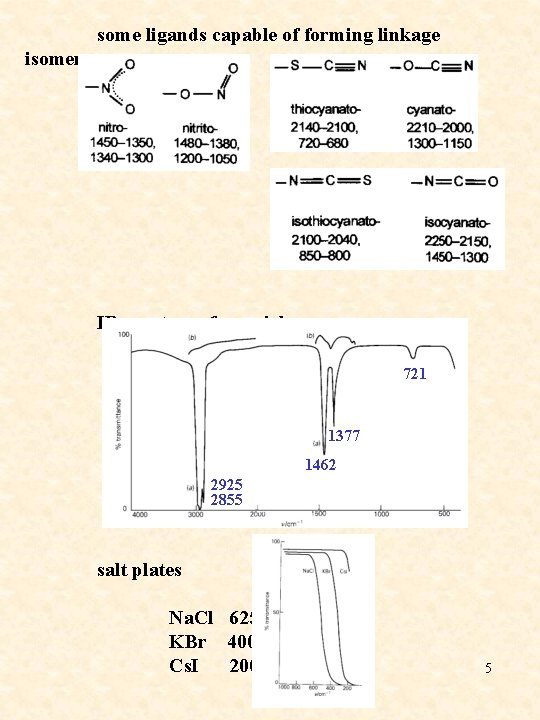
some ligands capable of forming linkage isomers IR spectrum for nujol 1377 1462 2925 2855 721 salt plates Na. Cl 625 cm-1 KBr 400 cm-1 Cs. I 200 cm-1 5
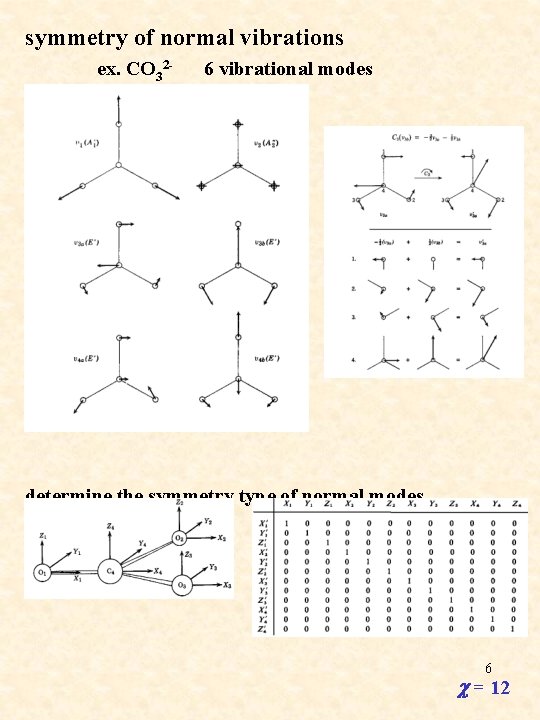
symmetry of normal vibrations ex. CO 32 - 6 vibrational modes C 3(u 3 a) = -1/2 u 3 a + 1/2 u 3 b C 3(u 3 b) = -3/2 u 3 a - 1/2 u 3 b determine the symmetry type of normal modes E 6 c = 12
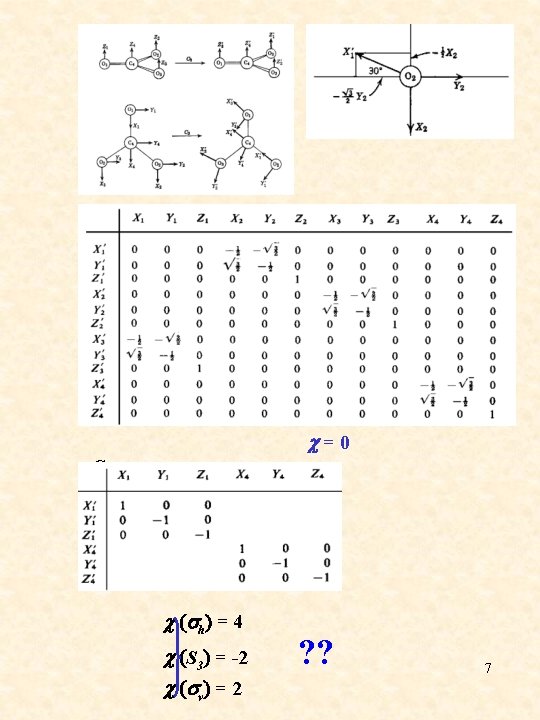
C 3 c = 0 C 2 c = -2 c (sh) = 4 c (S 3) = -2 c (sv) = 2 ? ? 7

+2 E’ + A 2” G = A 1’ + A 2’ + 3 E’ + 2 A 2” + E” 3 translatory modes: E’, A 2” 3 rotational modes: A 2’, E” genuine vibrational modes: Gg = A 1’ IR active: E’, A 2” (3 bands) Raman active: A 1’, E’ (3 bands) sv particular internal coordinates to normal modes C—O bonds E C 3 C 2 sh S 3 stretching bending plane bending G 3 0 1 3 0 1 GCO = A 1’ + E’ in-plane GOCO = A 1’ + E’ in-plane A 2” out-of 8
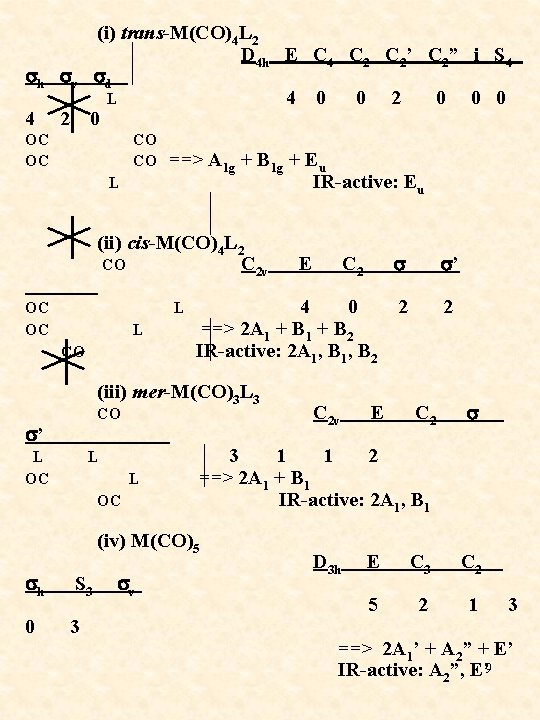
(i) trans-M(CO)4 L 2 D 4 h E C 4 C 2’ C 2” i S 4 sh sv sd L 4 0 0 2 0 0 0 4 2 0 OC OC CO L ==> A 1 g + B 1 g + Eu IR-active: Eu (ii) cis-M(CO)4 L 2 CO C 2 v E C 2 s s’ OC OC L CO L 4 0 2 2 ==> 2 A 1 + B 2 IR-active: 2 A 1, B 2 (iii) mer-M(CO)3 L 3 s’ CO L L OC 0 3 1 1 2 ==> 2 A 1 + B 1 IR-active: 2 A 1, B 1 (iv) M(CO)5 sh S 3 sv C 2 v E C 2 s D 3 h E C 3 C 2 5 2 1 3 ==> 2 A 1’ + A 2” + E’ IR-active: A 2”, E’ 9
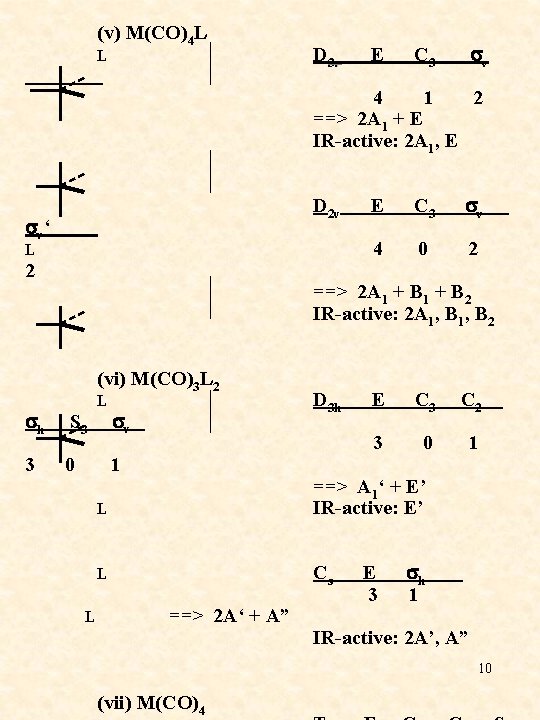
(v) M(CO)4 L L 4 1 2 ==> 2 A 1 + E IR-active: 2 A 1, E D 2 v E C 3 sv ‘ 4 0 2 L 2 ==> 2 A 1 + B 2 IR-active: 2 A 1, B 2 (vi) M(CO)3 L 2 L sh S 3 sv ==> A 1‘ + E’ IR-active: E’ L L D 3 h E C 3 C 2 3 0 1 3 0 1 L D 3 v E C 3 sv ==> 2 A‘ + A” Cs E sh 3 1 IR-active: 2 A’, A” 10 (vii) M(CO)4
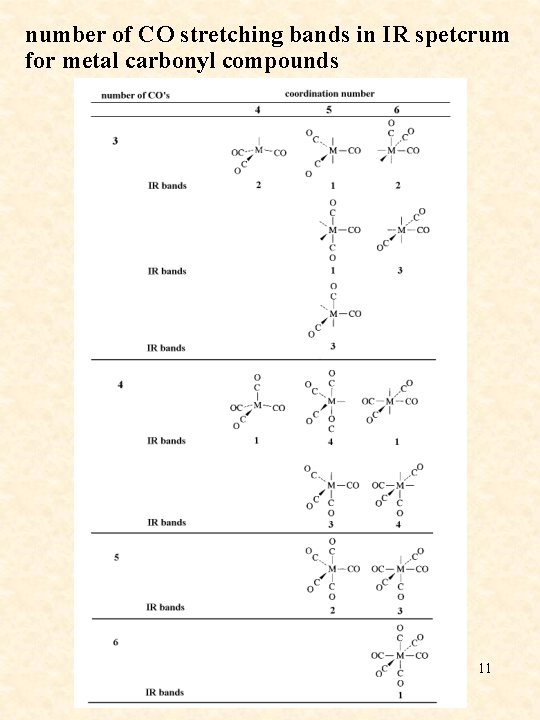
number of CO stretching bands in IR spetcrum for metal carbonyl compounds 11
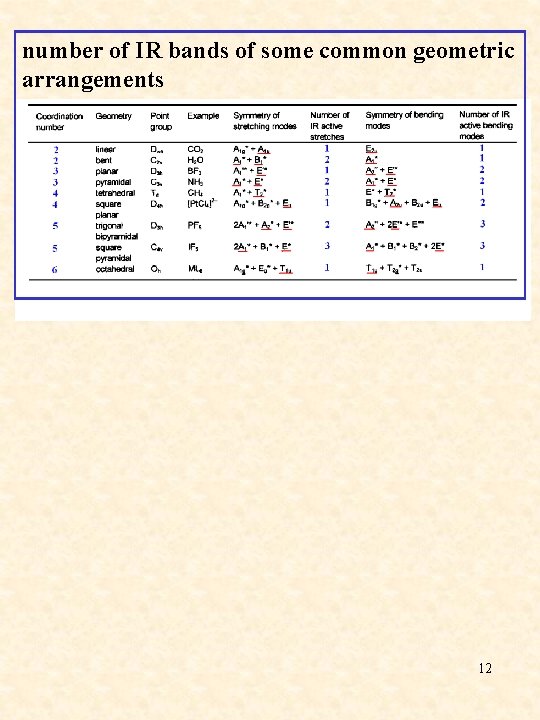
number of IR bands of some common geometric arrangements 12

calculation of force constants for diatomic molecule AB harmonic oscillator f • m-1 – l = 0 for polyatomic molecule Wilson’s method“The F and G matrix method” |FG – El| = 0 F: matrix of force constant (potential energy) G: matrix of masses and spatial relationship of atoms (kinetic energy) E: unit matrix e. g. + B 1 H 2 O Gg = 2 A 1 + B 1 2 O-H distance Dd 1, Dd 2 A 1 ∠HOH Dθ A 1 using projection operator to obtain complete set + Dd 2) of symmetry coordinates for vibrations A 1 : S 1 = Dθ S 2 = 1/√ 2(Dd 1 B 1 : Dd 2) S 3 = 1/√ 2(Dd 1 - 13 F matrix 2 V = Sfik si sk

fdd(Dd 1 Dd 2) fdθ Dd 1 2 V = fd(Dd 1)2 + fd(Dd 2)2 + fθ(Dθ)2 + 2 fdθ(Dd 1 Dθ) + 2 fdθ(Dd 2 Dθ) = [Dd 1 Dd 2 Dθ] fd fdd fd fdθ Dd 2 fθ Dθ fdθ = s’f s relationship between the internal coordinates and Dd 2 Dq the symmetry coordinates S = U s U matrix Dq 0 1 Dd 1 + Dd 2 = 1/√ 2 0 Dd 1 - Dd 2 1/√ 2 -1/√ 2 0 S = U s s = U’ S s’ = (U’ S)’ = S’U s’fs = S’FS (S’U)f(U’S) = S’FS S’(Uf. U’)S = S’FS ==> F = Uf. U’ 14 0 1/√ 2 0 1 fdd fdθ 0 1/√ 2
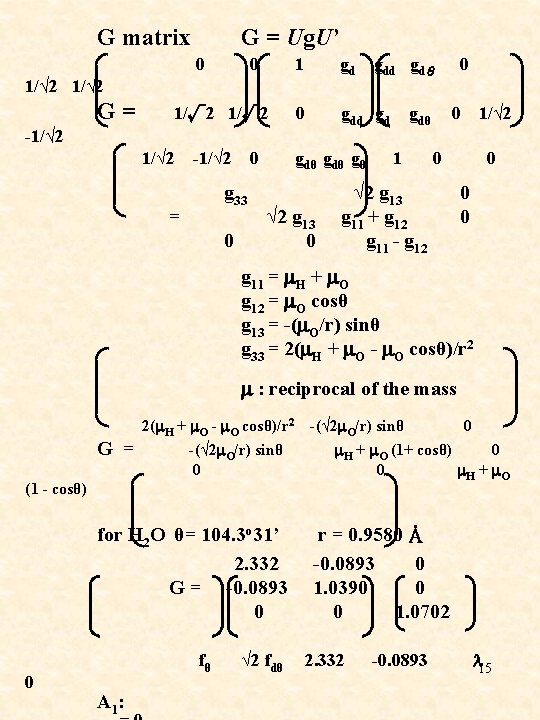
G matrix 0 1/√ 2 G = -1/√ 2 G = Ug. U’ 0 1 gdd gdθ 0 1/√ 2 0 gdd gd gdθ 0 1/√ 2 -1/√ 2 0 gdθ gθ 1 0 g 33 √ 2 g 13 = √ 2 g 13 g 11 + g 12 0 g 11 - g 12 0 0 g 11 = m. H + m. O g 12 = m. O cosθ g 13 = -(m. O/r) sinθ g 33 = 2(m. H + m. O - m. O cosθ)/r 2 m : reciprocal of the mass 2(m. H + m. O - m. O cosθ)/r 2 -(√ 2 m. O/r) sinθ 0 G = -(√ 2 m. O/r) sinθ m. H + m. O (1+ cosθ) 0 m. H + m. O 0 (1 - cosθ) for H 2 O θ= 104. 3 o 31’ 2. 332 G = -0. 0893 0 r = 0. 9580 Å -0. 0893 0 1. 0390 0 0 1. 0702 0 fθ √ 2 fdθ 2. 332 -0. 0893 l 15 A 1 :

elements of the g matrix mi: reciprocal mass of the ith atom rij: reciprocal of the distance between ith and jth 16
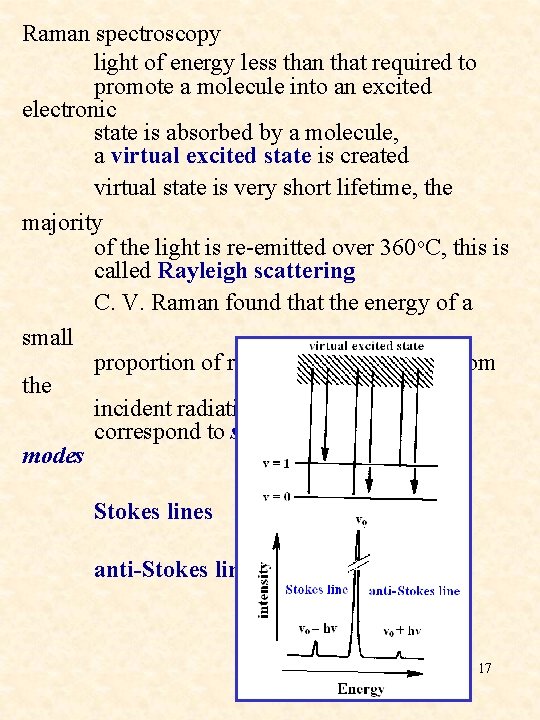
Raman spectroscopy light of energy less than that required to promote a molecule into an excited electronic state is absorbed by a molecule, a virtual excited state is created virtual state is very short lifetime, the majority of the light is re-emitted over 360 o. C, this is called Rayleigh scattering C. V. Raman found that the energy of a small proportion of re-emitted light differs from the incident radiation by energy gaps that correspond to some of the vibrational modes Stokes lines anti-Stokes line 17
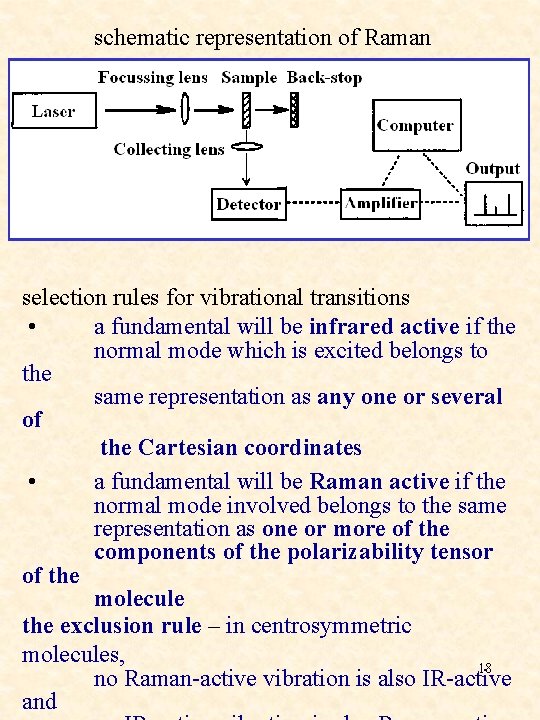
schematic representation of Raman spectrometer selection rules for vibrational transitions • a fundamental will be infrared active if the normal mode which is excited belongs to the same representation as any one or several of the Cartesian coordinates • a fundamental will be Raman active if the normal mode involved belongs to the same representation as one or more of the components of the polarizability tensor of the molecule the exclusion rule – in centrosymmetric molecules, 18 no Raman-active vibration is also IR-active and
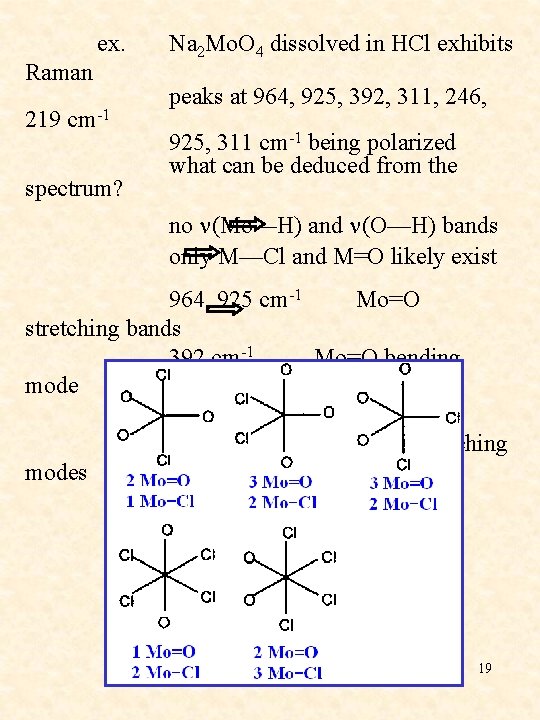
ex. Raman 219 cm-1 spectrum? Na 2 Mo. O 4 dissolved in HCl exhibits peaks at 964, 925, 392, 311, 246, 925, 311 cm-1 being polarized what can be deduced from the no n(Mo—H) and n(O—H) bands only M—Cl and M=O likely exist 964, 925 cm-1 Mo=O stretching bands 392 cm-1 Mo=O bending mode 311, 246, 219 cm-1 Mo—Cl stretching modes possible product: 19
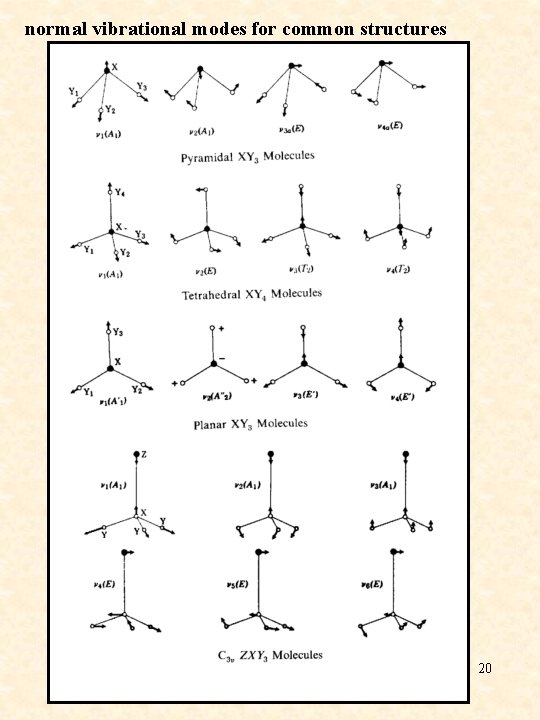
normal vibrational modes for common structures 20
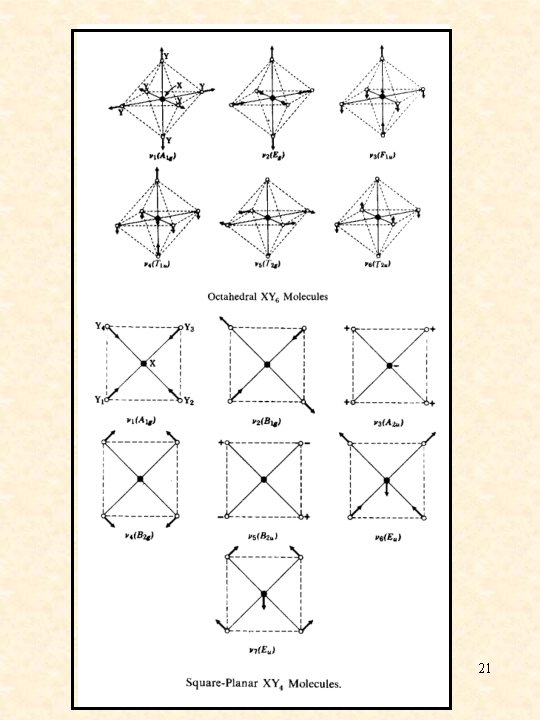
21
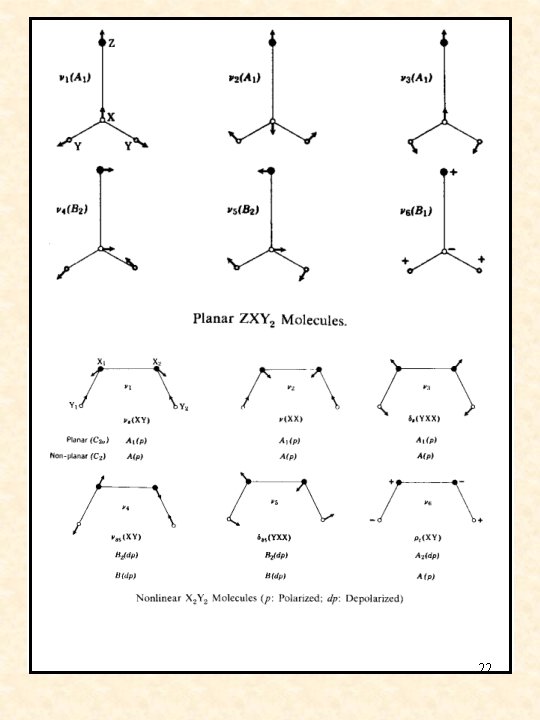
22
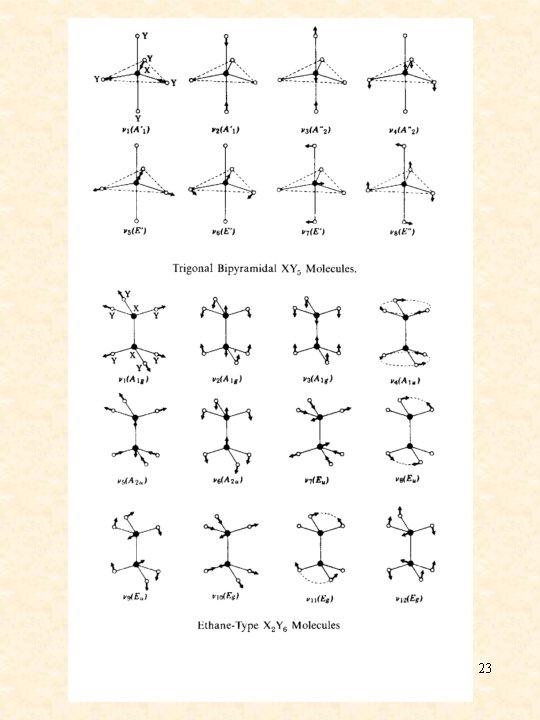
23

24

25

26
- Slides: 26