VI PMT meeting Feb 1 2011 Agenda Validation

VI PMT meeting – Feb 1, 2011 Agenda: - Validation vs Pathology - Congress presentations & Meetings - Paper publication
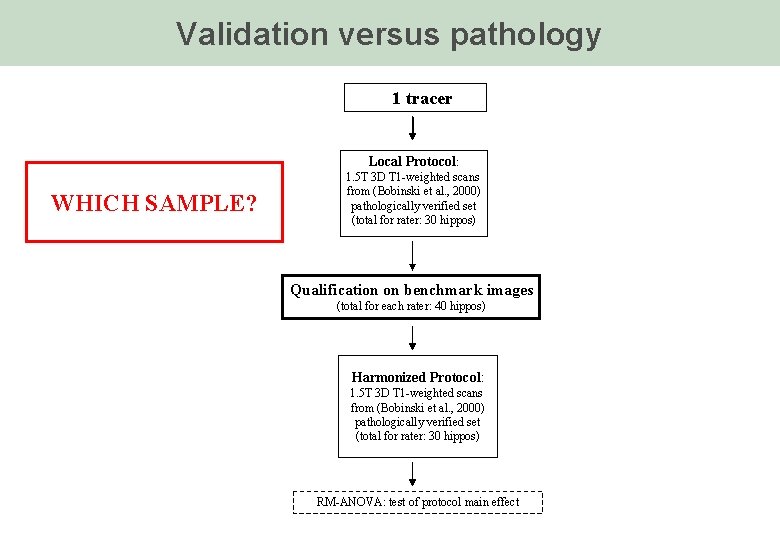
Validation versus pathology 1 tracer Local Protocol: WHICH SAMPLE? 1. 5 T 3 D T 1 -weighted scans from (Bobinski et al. , 2000) pathologically verified set (total for rater: 30 hippos) Qualification on benchmark images (total for each rater: 40 hippos) Harmonized Protocol: 1. 5 T 3 D T 1 -weighted scans from (Bobinski et al. , 2000) pathologically verified set (total for rater: 30 hippos) RM-ANOVA: test of protocol main effect

Congress presentations & Meetings

Congress presentations - AAN 2012, New Orleans Oral Presentation S 04. 003, April 24, Aging and Dementia: Therapeutic Interventions, 1. 30 pm M Boccardi, M Bocchetta, L Apostolova, J Barnes, G Bartzokis, G Corbetta, C De. Carli, L De. Toledo-Morrell, M Firbank, R Ganzola, L Gerritsen, W Henneman, R Killiany, N Malykhin, P Pasqualetti, J Pruessner, A Redolfi, N Robitaille, H Soininen, D Tolomeo, L Wang, C Watson, H Wolf, S Duchesne, CR Jack, GB Frisoni. Delphi consensus on landmarks for the manual segmentation of the hippocampus on MRI: preliminary results from the EADCADNI Harmonized Protocol working group. - AAIC 2012, Vancouver Abstract n. 26291, submitted to AAIC 2012 M Boccardi, M Bocchetta, A Redolfi, P Pasqualetti, R Ganzola, N Robitaille, S Duchesne, CR Jack, GB Frisoni and the EADC-ADNI Hippocampal Harmonization Group. Definition of Harmonized Protocol for Hippocampal Segmentation Abstract n. 26292, submitted to AIC 2012 M Boccardi, M Bocchetta, A Redolfi, P Pasqualetti, R Ganzola, N Robitaille, S Duchesne, CR Jack, GB Frisoni and the EADC-ADNI Hippocampal Harmonization Group. Definition of Harmonized Protocol for Hippocampal Segmentation

SOPs periodical meetings AAN – April 24, 2012 AGENDA: 1. Delphi Paper *; 2. Italian-ADNI Paper using prototype -protocol (? ) 3. Publication policy AAIC – July 2012 * AGENDA: 1. Presentation of Harmonized Protocol; 2. Reliability figures
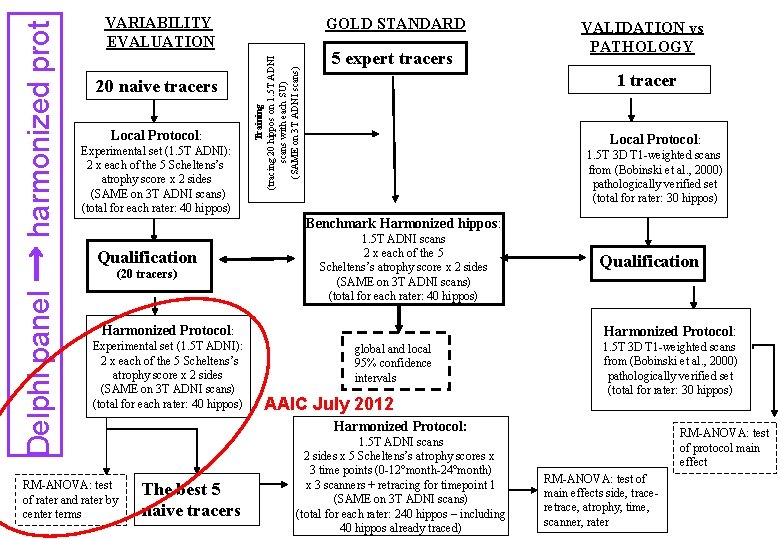
20 naive tracers Local Protocol: Experimental set (1. 5 T ADNI): 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) GOLD STANDARD Training (tracing 20 hippos on 1. 5 T ADNI scans with each SU) (SAME on 3 T ADNI scans) Delphi panel → harmonized prot VARIABILITY EVALUATION 5 expert tracers VALIDATION vs PATHOLOGY 1 tracer Local Protocol: 1. 5 T 3 D T 1 -weighted scans from (Bobinski et al. , 2000) pathologically verified set (total for rater: 30 hippos) Benchmark Harmonized hippos: Qualification (20 tracers) 1. 5 T ADNI scans 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) Harmonized Protocol: Experimental set (1. 5 T ADNI): 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) RM-ANOVA: test of rater and rater by center terms Qualification Harmonized Protocol: global and local 95% confidence intervals AAIC July 2012 1. 5 T 3 D T 1 -weighted scans from (Bobinski et al. , 2000) pathologically verified set (total for rater: 30 hippos) Harmonized Protocol: The best 5 naive tracers 1. 5 T ADNI scans 2 sides x 5 Scheltens’s atrophy scores x 3 time points (0 -12°month-24°month) x 3 scanners + retracing for timepoint 1 (SAME on 3 T ADNI scans) (total for each rater: 240 hippos – including 40 hippos already traced) RM-ANOVA: test of protocol main effect RM-ANOVA: test of main effects side, traceretrace, atrophy, time, scanner, rater

Papers publication

Next: Paper operationalization Operationalization of differences among protocols for manual hippocampal segmentation M Boccardi, M Bocchetta, R Ganzola, N Robitaille, A Redolfi, S Duchesne, CR Jack , GB Frisoni, and the Alzheimer's Disease Neuroimaging Initiative. Collaborators: G Bartzokis, JG Csernansky, MJ de Leon, L de. Toledo-Morrell, RJ Killiany, S Lehéricy, N Malykhin, J Pantel, JC Pruessner, H Soininen, C Watson To be submitted to Alzheimer’s and Dementia

Papers describing the project Survey of protocols (preliminary phase; published, JAD 2011) Operationalization (preliminary phase; to be completed) Delphi consensus (Brescia Team) Master tracers’ practice and reliability (Brescia Team) Development of certification platform (Duchesne and coll? ) Validation data and Protocol definition (Brescia Team) Validation vs pathology (TBD)
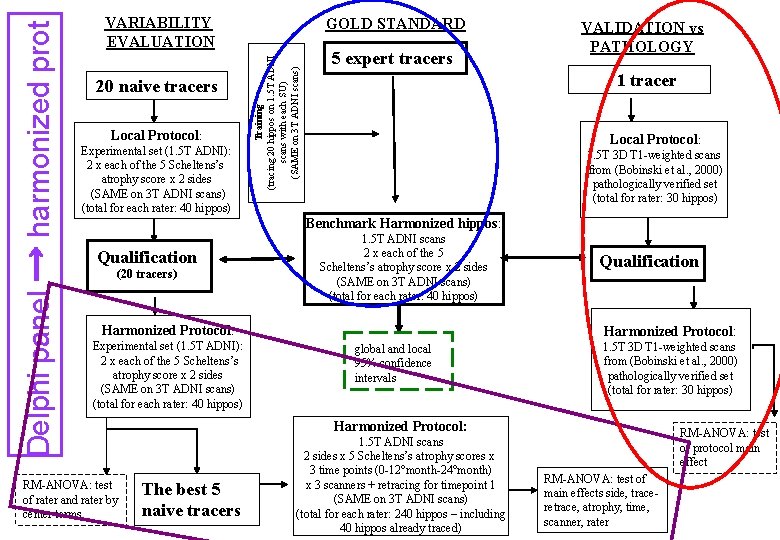
20 naive tracers Local Protocol: Experimental set (1. 5 T ADNI): 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) GOLD STANDARD Training (tracing 20 hippos on 1. 5 T ADNI scans with each SU) (SAME on 3 T ADNI scans) Delphi panel → harmonized prot VARIABILITY EVALUATION 5 expert tracers VALIDATION vs PATHOLOGY 1 tracer Local Protocol: 1. 5 T 3 D T 1 -weighted scans from (Bobinski et al. , 2000) pathologically verified set (total for rater: 30 hippos) Benchmark Harmonized hippos: Qualification (20 tracers) 1. 5 T ADNI scans 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) Harmonized Protocol: Experimental set (1. 5 T ADNI): 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) RM-ANOVA: test of rater and rater by center terms Qualification Harmonized Protocol: global and local 95% confidence intervals 1. 5 T 3 D T 1 -weighted scans from (Bobinski et al. , 2000) pathologically verified set (total for rater: 30 hippos) Harmonized Protocol: The best 5 naive tracers 1. 5 T ADNI scans 2 sides x 5 Scheltens’s atrophy scores x 3 time points (0 -12°month-24°month) x 3 scanners + retracing for timepoint 1 (SAME on 3 T ADNI scans) (total for each rater: 240 hippos – including 40 hippos already traced) RM-ANOVA: test of protocol main effect RM-ANOVA: test of main effects side, traceretrace, atrophy, time, scanner, rater
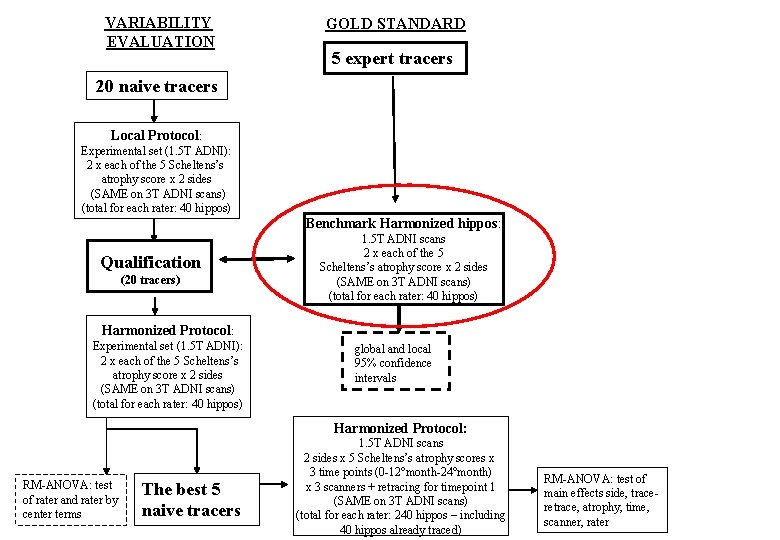
VARIABILITY EVALUATION GOLD STANDARD 5 expert tracers 20 naive tracers Local Protocol: Experimental set (1. 5 T ADNI): 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) Benchmark Harmonized hippos: Qualification (20 tracers) 1. 5 T ADNI scans 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) Harmonized Protocol: Experimental set (1. 5 T ADNI): 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) global and local 95% confidence intervals Harmonized Protocol: RM-ANOVA: test of rater and rater by center terms The best 5 naive tracers 1. 5 T ADNI scans 2 sides x 5 Scheltens’s atrophy scores x 3 time points (0 -12°month-24°month) x 3 scanners + retracing for timepoint 1 (SAME on 3 T ADNI scans) (total for each rater: 240 hippos – including 40 hippos already traced) RM-ANOVA: test of main effects side, traceretrace, atrophy, time, scanner, rater

GANTT
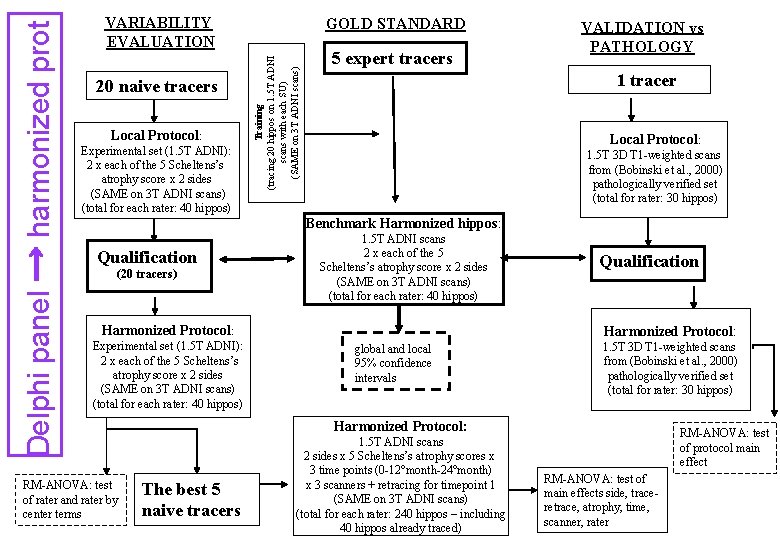
20 naive tracers Local Protocol: Experimental set (1. 5 T ADNI): 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) GOLD STANDARD Training (tracing 20 hippos on 1. 5 T ADNI scans with each SU) (SAME on 3 T ADNI scans) Delphi panel → harmonized prot VARIABILITY EVALUATION 5 expert tracers VALIDATION vs PATHOLOGY 1 tracer Local Protocol: 1. 5 T 3 D T 1 -weighted scans from (Bobinski et al. , 2000) pathologically verified set (total for rater: 30 hippos) Benchmark Harmonized hippos: Qualification (20 tracers) 1. 5 T ADNI scans 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) Harmonized Protocol: Experimental set (1. 5 T ADNI): 2 x each of the 5 Scheltens’s atrophy score x 2 sides (SAME on 3 T ADNI scans) (total for each rater: 40 hippos) RM-ANOVA: test of rater and rater by center terms Qualification Harmonized Protocol: global and local 95% confidence intervals 1. 5 T 3 D T 1 -weighted scans from (Bobinski et al. , 2000) pathologically verified set (total for rater: 30 hippos) Harmonized Protocol: The best 5 naive tracers 1. 5 T ADNI scans 2 sides x 5 Scheltens’s atrophy scores x 3 time points (0 -12°month-24°month) x 3 scanners + retracing for timepoint 1 (SAME on 3 T ADNI scans) (total for each rater: 240 hippos – including 40 hippos already traced) RM-ANOVA: test of protocol main effect RM-ANOVA: test of main effects side, traceretrace, atrophy, time, scanner, rater
- Slides: 13