Variationally optimized basis set for biological molecules Hiori

Variationally optimized basis set for biological molecules Hiori Kino Contents • Final target • Status of the openmx program • Purpose of this research • optimized basis set • applications and transferability • simplified (optimized) PAO • transferability
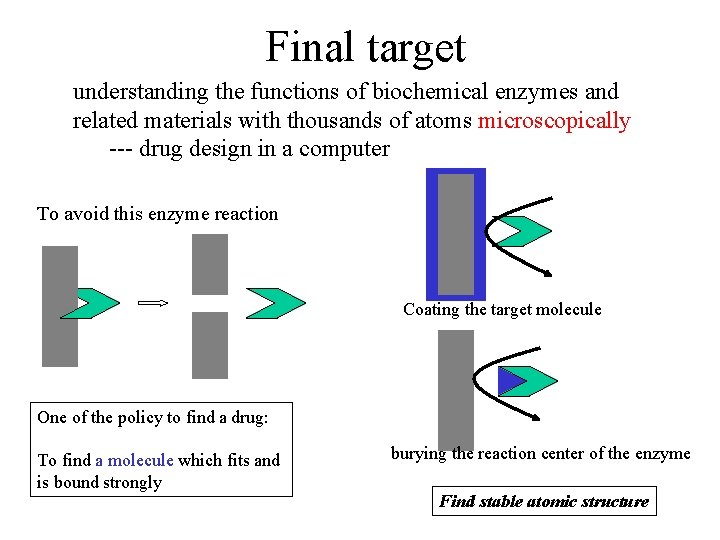
Final target understanding the functions of biochemical enzymes and related materials with thousands of atoms microscopically --- drug design in a computer To avoid this enzyme reaction Coating the target molecule One of the policy to find a drug: To find a molecule which fits and is bound strongly burying the reaction center of the enzyme Find stable atomic structure
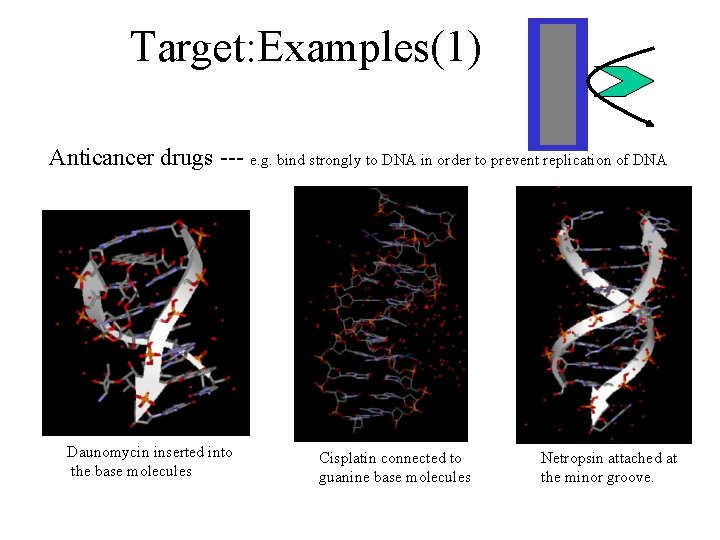
Target: Examples(1) Anticancer drugs --- e. g. bind strongly to DNA in order to prevent replication of DNA Daunomycin inserted into the base molecules Cisplatin connected to guanine base molecules Netropsin attached at the minor groove.
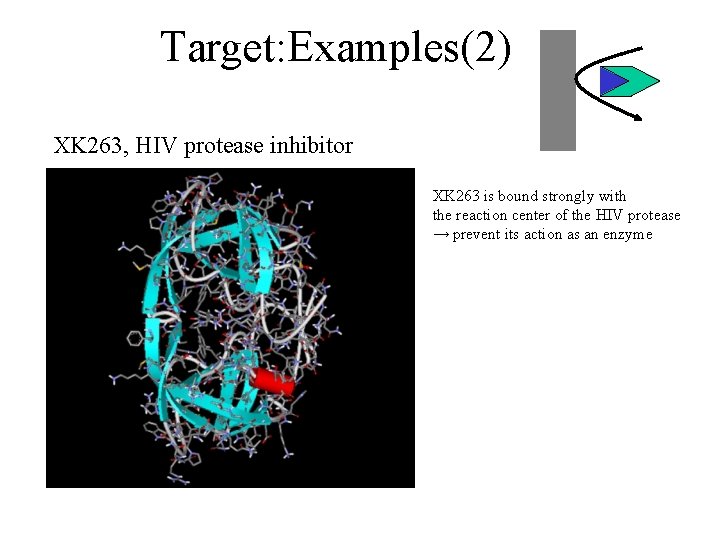
Target: Examples(2) XK 263, HIV protease inhibitor XK 263 is bound strongly with the reaction center of the HIV protease → prevent its action as an enzyme

Status of the Open. MX program Linear combination of Pseudo. Atomic. Orbital Pseudopotentials: TM, Schroedinger, dirac/scalar relativistic, partial core corrections XC-functional: LDA, PBE orbital optimization MPI- parallelized stable order(N) Download from http: //staff. aist. go. jp/t-ozaki/

Purpose of this research A number of order(N) programs But, how accurate is linear combination of (pseudo)atomic orbitals? More basis set → more accurate, but heavy calculation smaller basis set=more efficient and better accurary
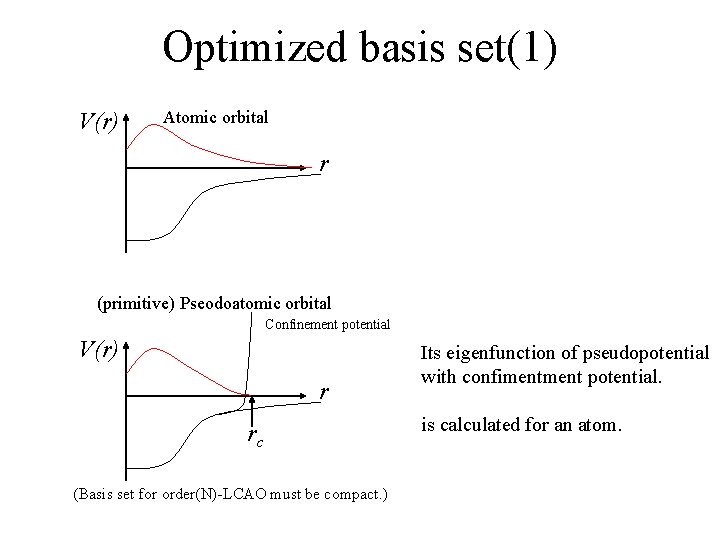
Optimized basis set(1) V(r) Atomic orbital r (primitive) Pseodoatomic orbital Confinement potential V(r) r rc (Basis set for order(N)-LCAO must be compact. ) Its eigenfunction of pseudopotential with confiment potential. is calculated for an atom.
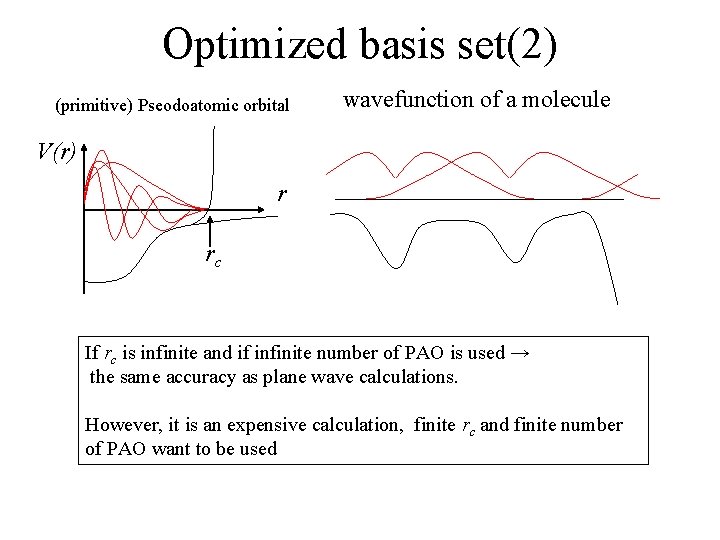
Optimized basis set(2) (primitive) Pseodoatomic orbital wavefunction of a molecule V(r) r rc If rc is infinite and if infinite number of PAO is used → the same accuracy as plane wave calculations. However, it is an expensive calculation, finite rc and finite number of PAO want to be used
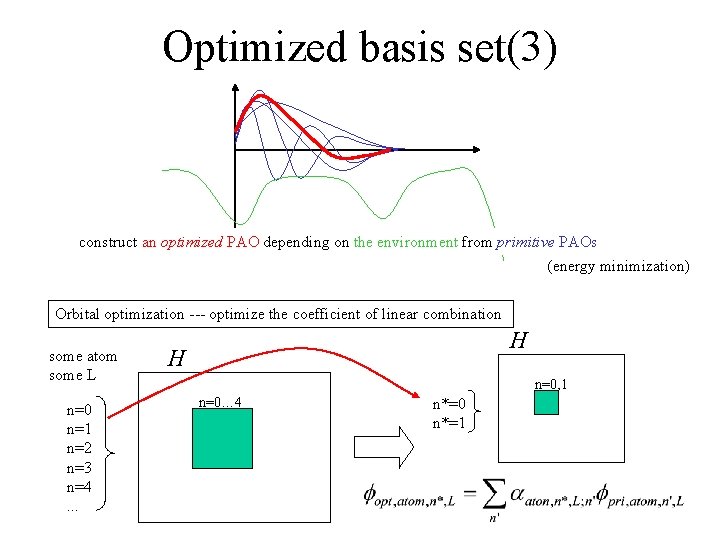
Optimized basis set(3) construct an optimized PAO depending on the environment from primitive PAOs (energy minimization) Orbital optimization --- optimize the coefficient of linear combination some atom some L n=0 n=1 n=2 n=3 n=4. . . H H n=0, 1 n=0… 4 n*=0 n*=1
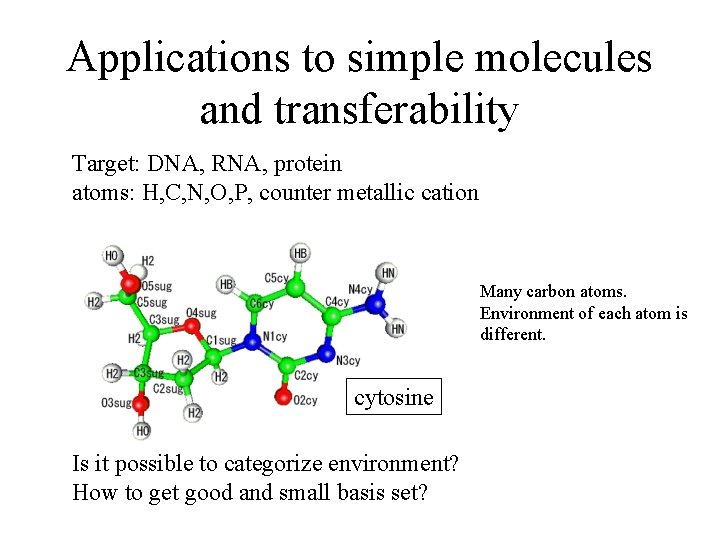
Applications to simple molecules and transferability Target: DNA, RNA, protein atoms: H, C, N, O, P, counter metallic cation Many carbon atoms. Environment of each atom is different. cytosine Is it possible to categorize environment? How to get good and small basis set?

Simplified orbital(1) 1. Calculate many molecules and optimize orbitals protein GGG GAG GVG GLG GIG GPG GFG GMG GWG GCG DKG DRG DHG EKG GNG GSG GTF GYG RNA, RNA B-G B-C B-T B-U A-AT A-GC B-AU (B-C)2 Na Saccaride arabinose D-Glico D-Glu. Ac fructose fucose glucose ribose Lipid DPPC oleic acid Acid citric acid lactic acid Atomic positions are optimized using amber 98 or mm 3 force field Others AMP ADP ATP

Simplified orbital(2) C, O, N, S, P: s, p --- 2 optimized orbitals from 5 primitive orbitals d --- 1 optimized orbitals from 5 primitive orbitals s 52 p 52 d 51 ( corresponding to double zeta plus polarized ) H: s --- 2 optimized orbitals from 5 primitive orbitals p --- 1 optimized orbitals from 5 primitive orbitals ( corresponding to double zeta plus polarized ) s 52 p 51
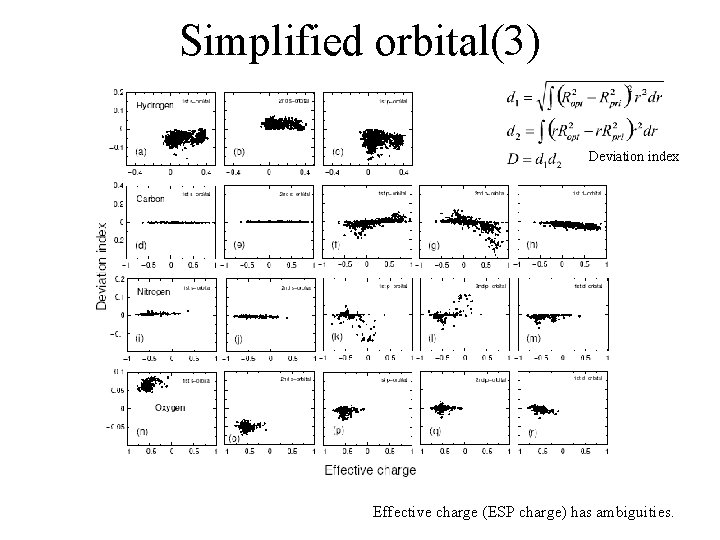
Simplified orbital(3) Deviation index Effective charge (ESP charge) has ambiguities.
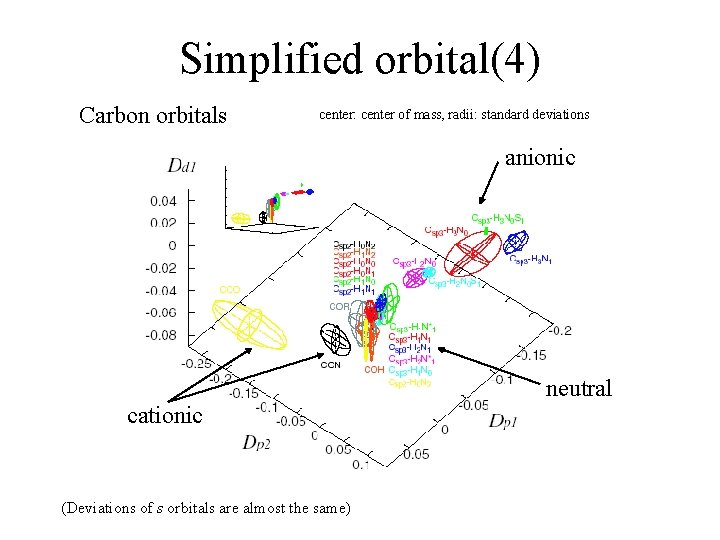
Simplified orbital(4) Carbon orbitals center: center of mass, radii: standard deviations anionic neutral cationic (Deviations of s orbitals are almost the same)
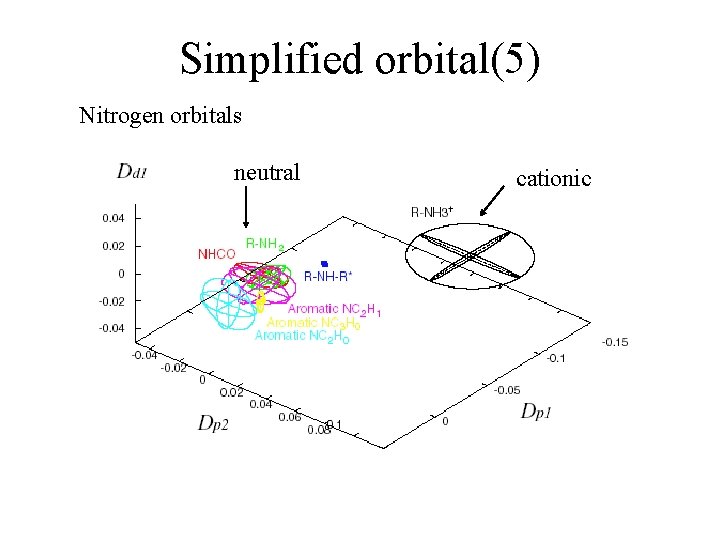
Simplified orbital(5) Nitrogen orbitals neutral cationic

Simplified orbital(6) a. u. /atom
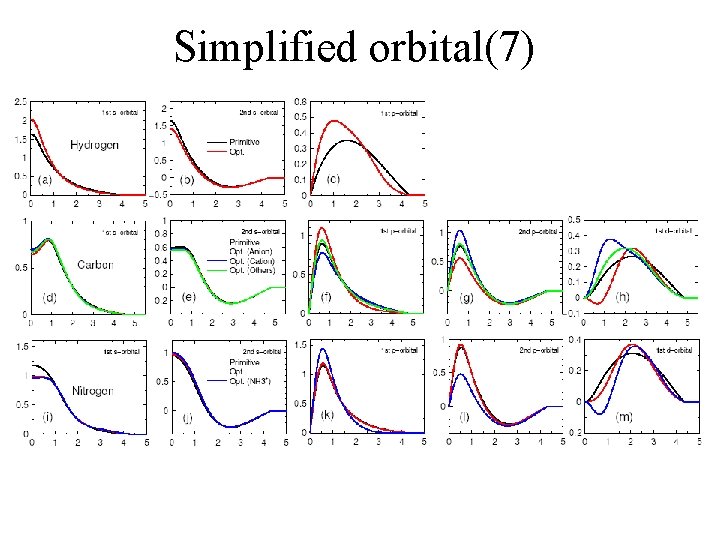
Simplified orbital(7)

Transferability(1): H 2 O 200 Ry, XC=PBE H 4. 5 -O 4. 5 full=s 5 p 5(d 5) PW theory: Sprik, J. Chem. Phys. 105, 1142 (1996). PW 91: Gaussian 03, 6 -311+G(d, p) Comment: Small dipole moment is due to finite truncation of PAO

Transferability(2): H 2 O dimer H 4. 5, O 4. 5, s 52 p 52 d 51 for O, s 52 p 51 for H, 200 Ry, PBE PW/… Sprik, J. Chem. Phys. 105, 1142(1996). PW/B 3 LYP: P. L. Silverstrelli amd M. Parrinello, J. Chem. Phys. 111, 3572(1999) so-DVZ gives good results for internal bonds, but it gives shorter bond length for hydrogen bonds. (more d orbitals are necessary for O. )

Transferability(3): acetic dimer H 4. 5 O 4. 5, 160 Ry H O O A. J. A. Aquino, et al. , J. Phys. Chem. A (2002), 106, 1862. BLYP? / SVP+sp? L. Turi, J. Phys. Chem. (1996) 100, 11285. MP 2/D 95++(d, p) A problem: so gives shorter length for O…O H O O
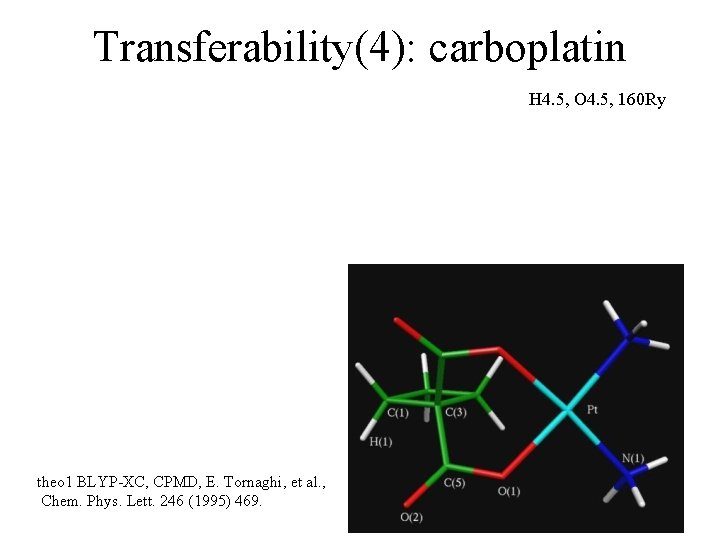
Transferability(4): carboplatin H 4. 5, O 4. 5, 160 Ry theo 1 BLYP-XC, CPMD, E. Tornaghi, et al. , Chem. Phys. Lett. 246 (1995) 469.

Transferability(5) CH 4, C 2 H 6, C 4 H 4, C 2 H 2, benzene, N 2, NH 3, HCN, O 2, CH 3 OH, formaldhyde, formie acid, formamide, NO 2, PH 3, H 2 S 2, H 2 SO 4, thioformamide, Glycine, CH 3 F, cisplatin, . . .

Transferability(6): problems H 2 O, H 4. 5 O 4. 5 Binding energy is 2 times as large as those of exp. and PW theory. ~SIESTA gives similar result to open. MX. D. Sanchez-Portal, et al. , Int. J. Quant. Chem. 65, 453 (1997). PW: Sprik, J. Chem. Phys. 105, 1142 (1996). about 2 times as large as those of exp. and PW theory in the cases of acetic dimer and Gp. C(DNA) ~SIESTA gives the result similar to exp and PW theories in Gp. C. M. Machado, P. Ordejon, condmat/9908022.

Summary • Good and efficient(small) optimized DVP basis set and high transferability for intramolecular parameters • insufficient DVP for hydrogen bond • too large binding energy for hydrogen bond
- Slides: 24