Using Synthetic Biology and p Clone Red for

Using Synthetic Biology and p. Clone Red for Authentic Research on Promoter Function: Genetics (analyzing mutant promoters) Todd T. Eckdahl and A. Malcolm Campbell

Timeline Learn about bacterial promoters Develop promoter mutation hypotheses Order oligonucleotides for mutant promoters Use Golden Gate assembly to clone mutant promoters upstream of RFP reporter gene • Measure output of reporter gene • Present results to class and enter into Registry of Functional Reporters (RFP) • •

What is the consensus sequence for the two elements of a bacterial promoter? -10 element = T 80 A 95 T 45 A 60 A 50 T 96 Optimal position is -10, but position varies from -18 to -9 from center of -10 element to TSS at +1 -35 element = T 82 T 84 G 78 A 65 C 54 A 45 Position varies, but spacing between -35 and -10 elements is 16 -18 bp in 90% of known promoters Optimal spacing is 17 bp Source: Genes IX, 2008, by Benjamin Lewin

Transcription Initiation • RNAP binds to DNA and engages in direct exchange of one sequence for another • Affinity of RNAP for nonspecific DNA decreased by Sigma factor • RNAP + Sigma “touches down” at -35 element • Contact is extended to -10 element, covering ~77 bp in “closed” binary complex • Melting is facillitated by A+T content of -10 element - of ~12 bp from -10 element to +1 produces “open” binary complex • Incorporation of NTPs forms ternary complex, and multiple rounds of abortive initiation occur • Change in RNAP structure occurs, Sigma factor is released or changes form, and it covers ~50 bp • RNAP clears the promoter, shortens to cover only 30 -40 bp, and elongation occurs at a rate of ~40 nt/sec

p. Tac Promoter • Hybrid promoter constructed from p. Trp and p. Lac promoters in E. coli • Efficiency of promotion measure to be 2 -3 times great than p. Trp and 7 -11 times greater than p. Lac • Perfect matches to -1 - and -35 consensus sequences and 16 bp between them -35 -10 De. Boer HA, Comstock LJ, Vasser M (1982) PNAS 80, 21 -25. +1

Mutations of p. Tac • Substitutions -10 element -35 element • Deletions – Decrease spacing between -35 and -10 – Decrease spacing beteen -10 and +1 • Insertions – Increase spacing between -35 and -10 – Increase spacing beteen -10 and +1

GGA Method with p. Clone Red by A. Malcolm Campbell and Todd Eckdahl

Eco RI G AATTC CTTAA G type II palindrome
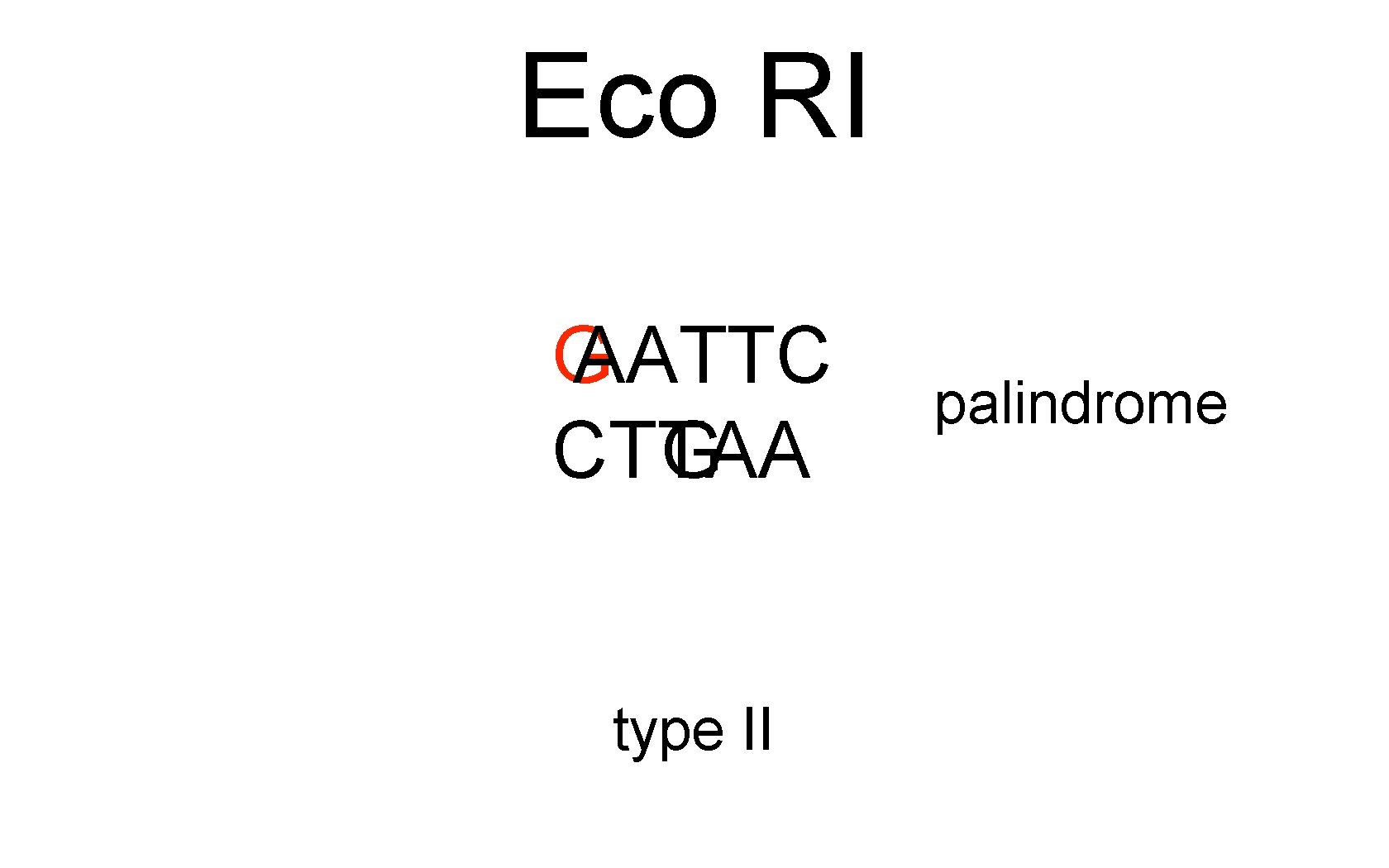
Eco RI G AATTC CTTAA G type II palindrome

Eco RI G AATTC CTTAA G type II
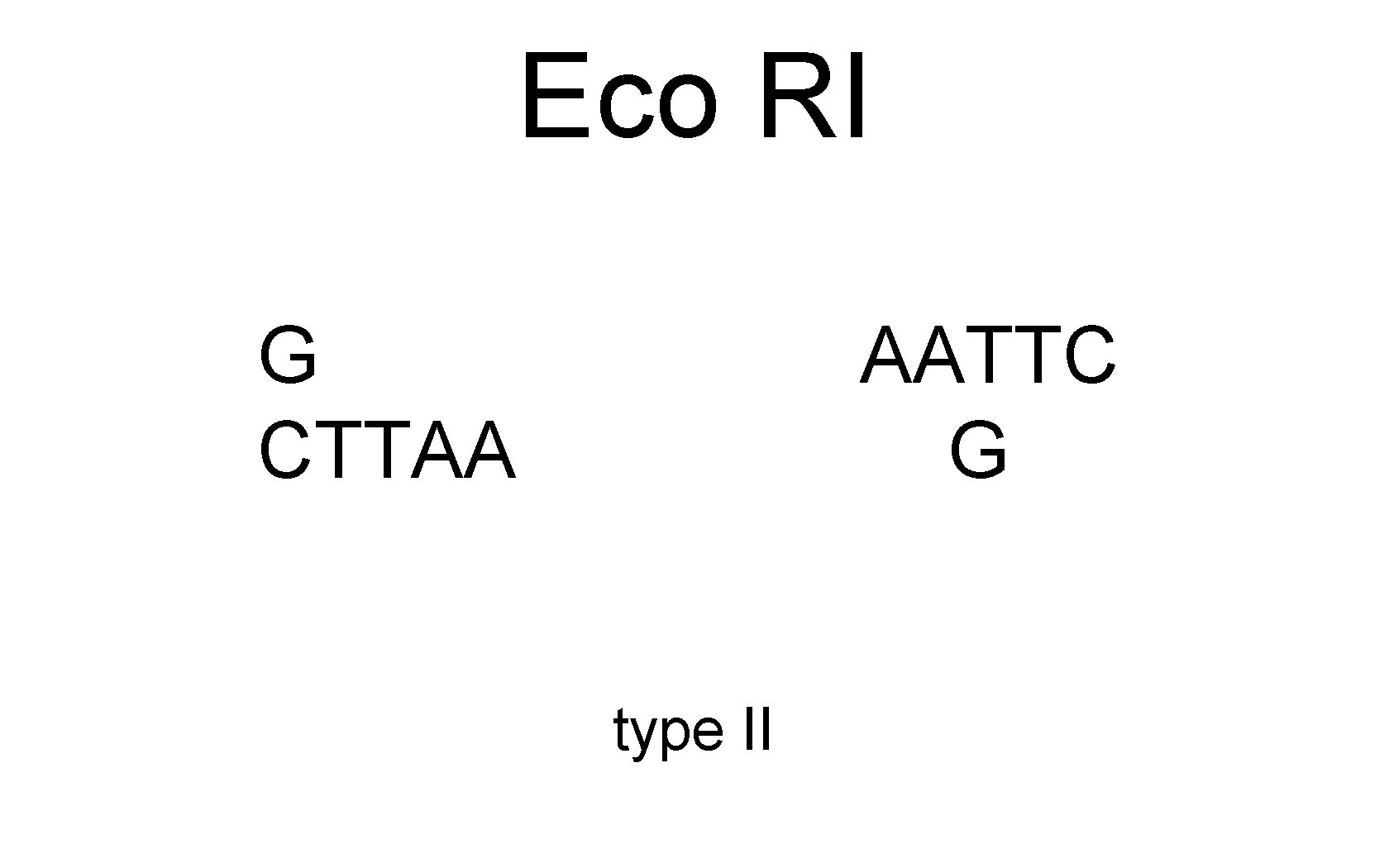
Eco RI G CTTAA AATTC G type II
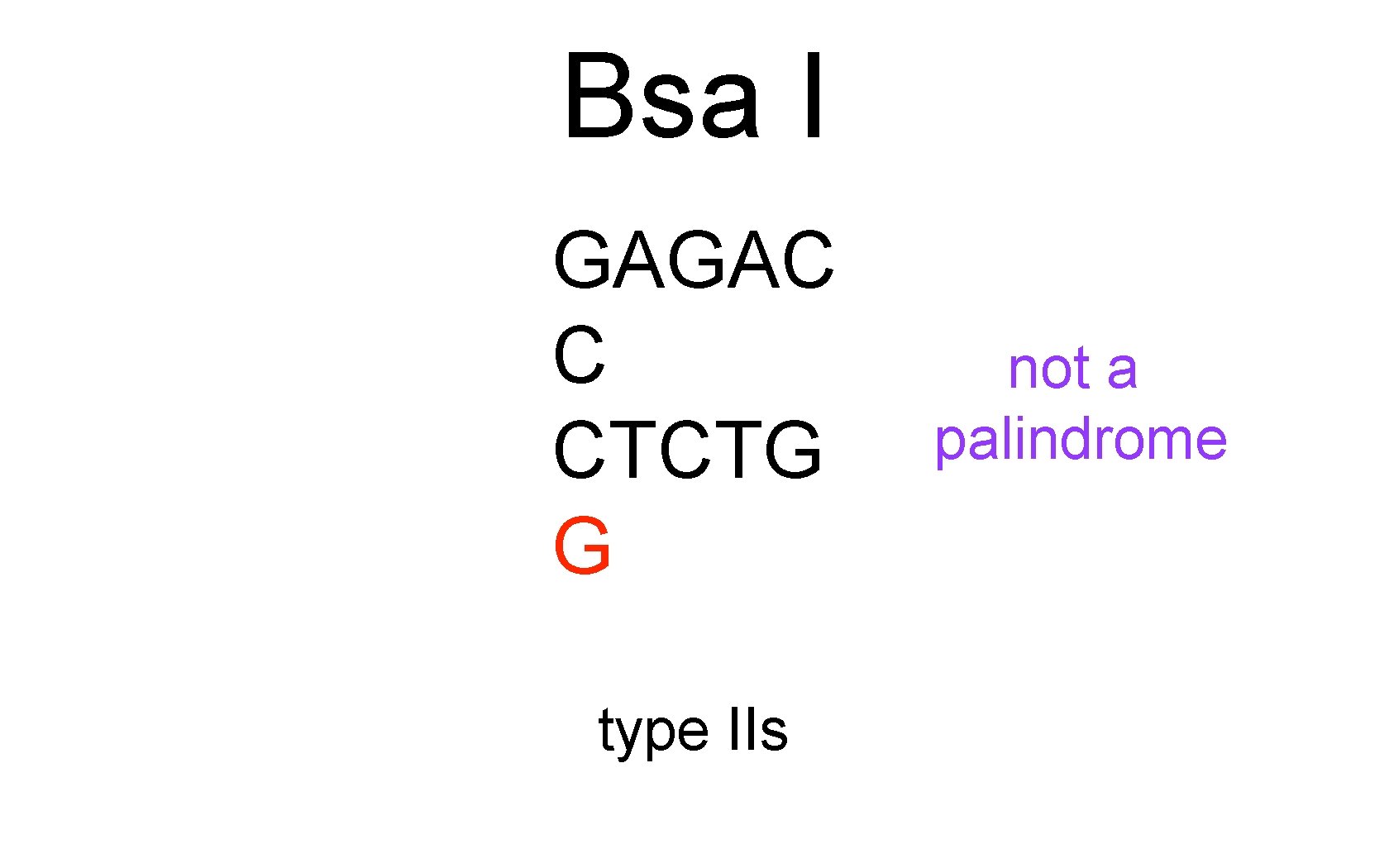
Bsa I GAGAC C CTCTG G type IIs not a palindrome
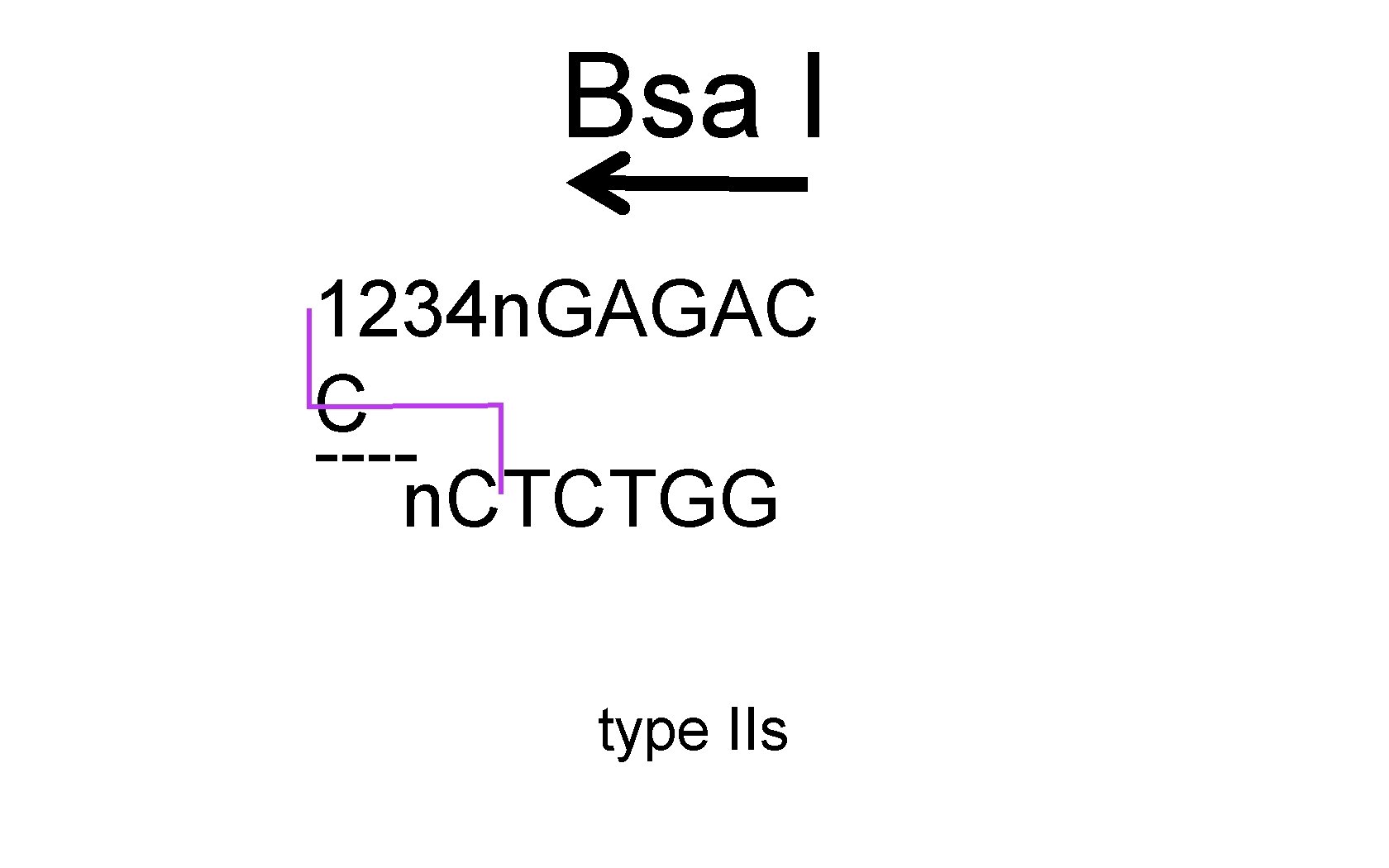
Bsa I 1234 n. GAGAC C ---n. CTCTGG type IIs
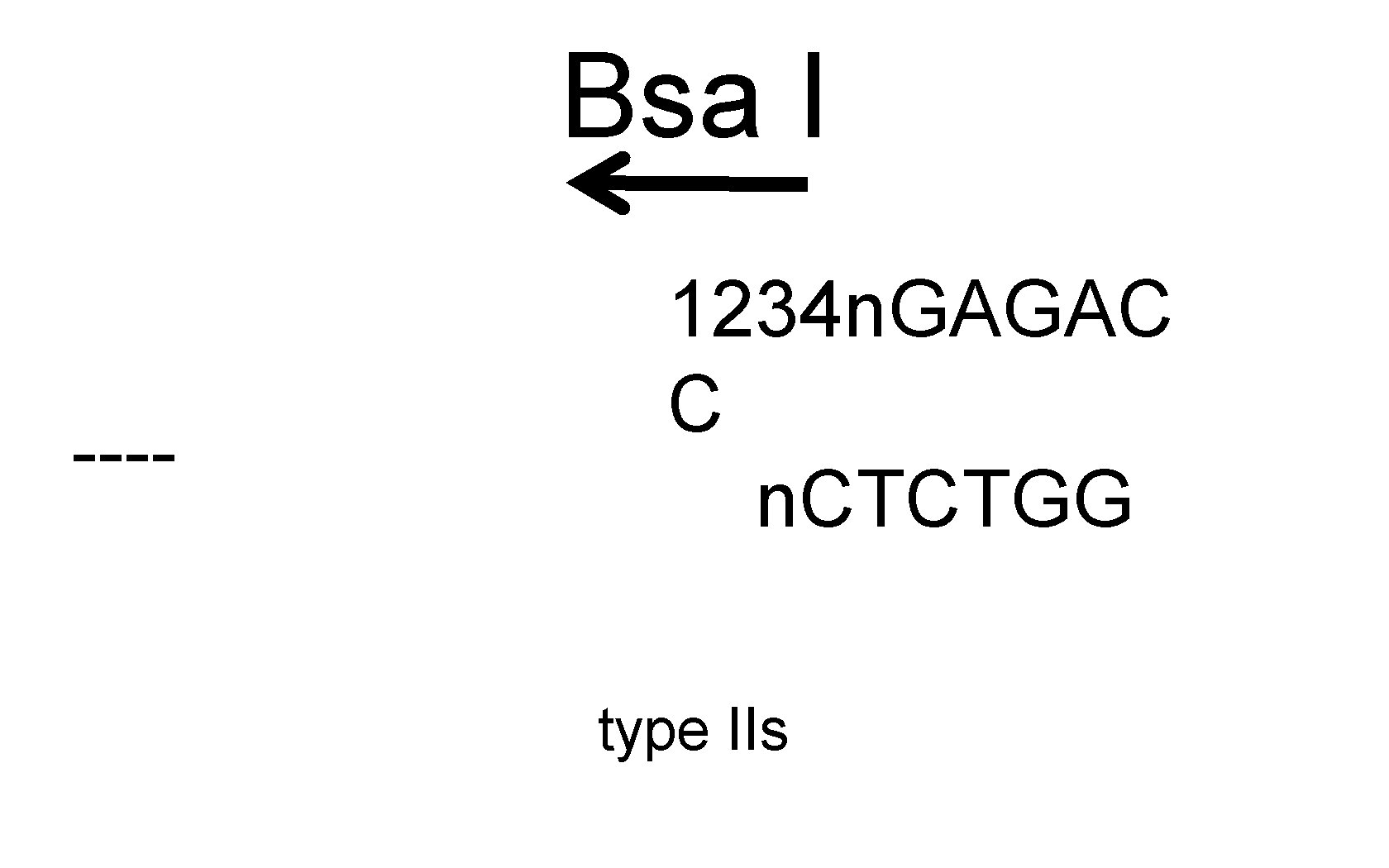
Bsa I ---- 1234 n. GAGAC C n. CTCTGG type IIs
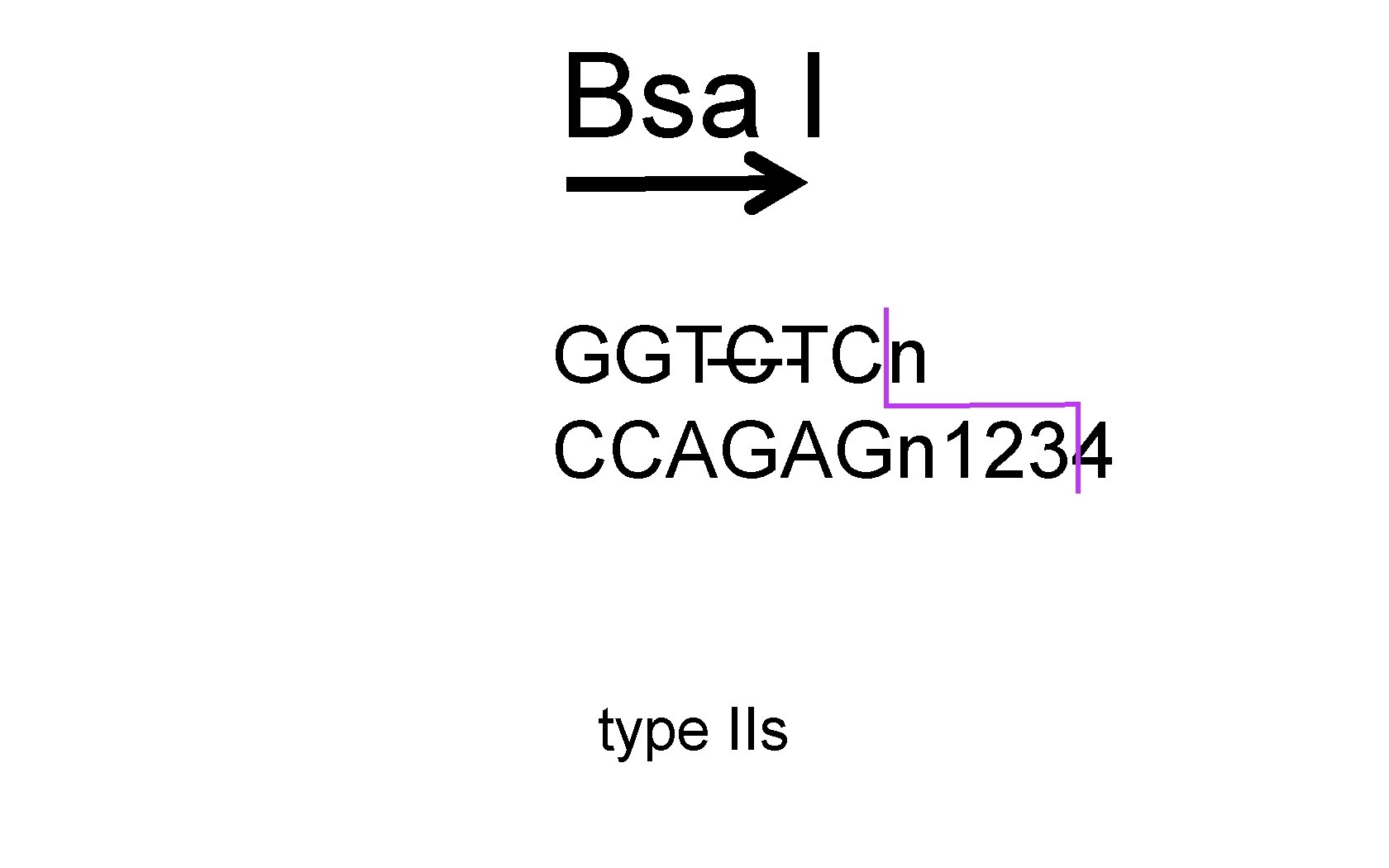
Bsa I GGTCTCn ---CCAGAGn 1234 type IIs
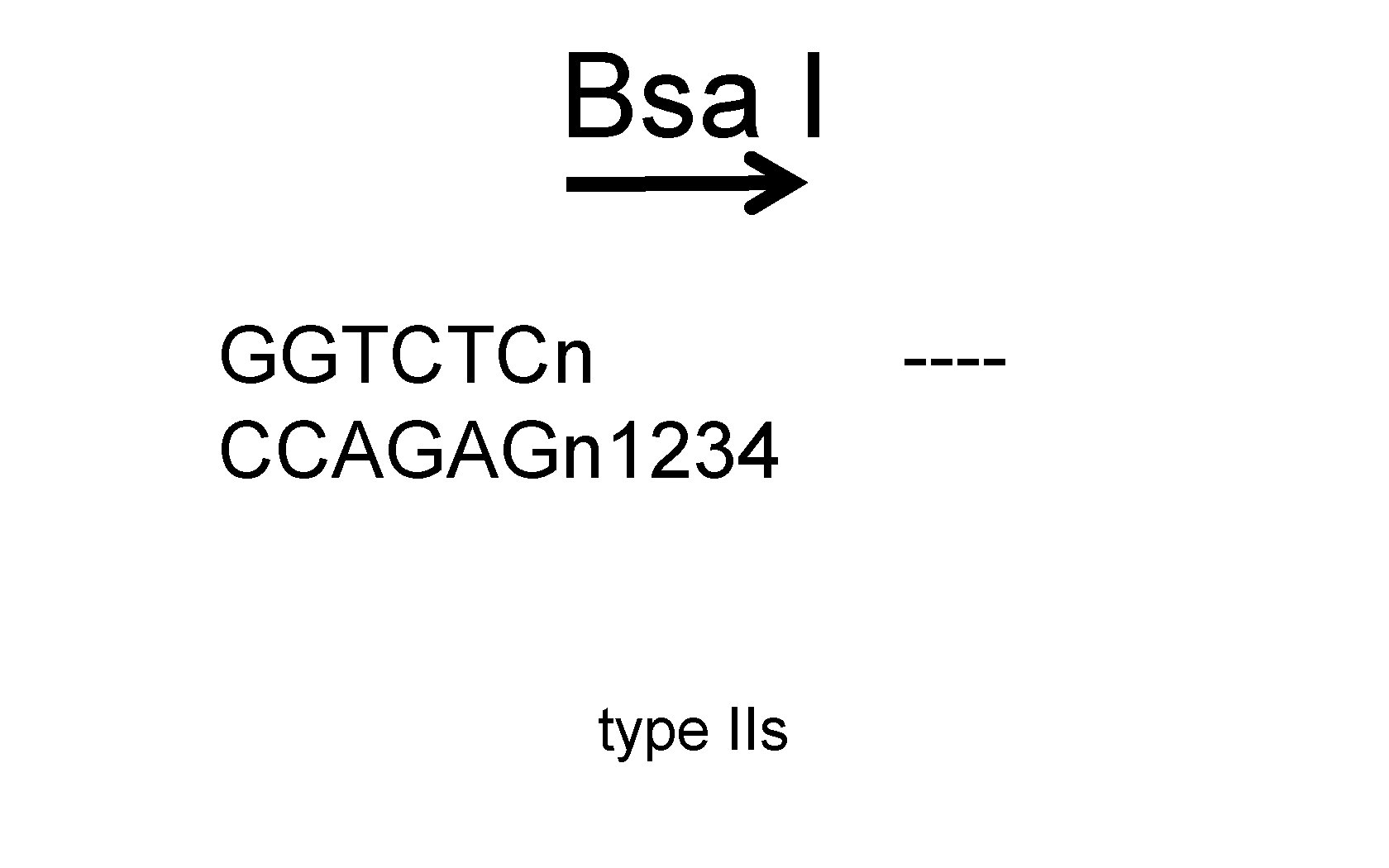
Bsa I GGTCTCn CCAGAGn 1234 type IIs ----
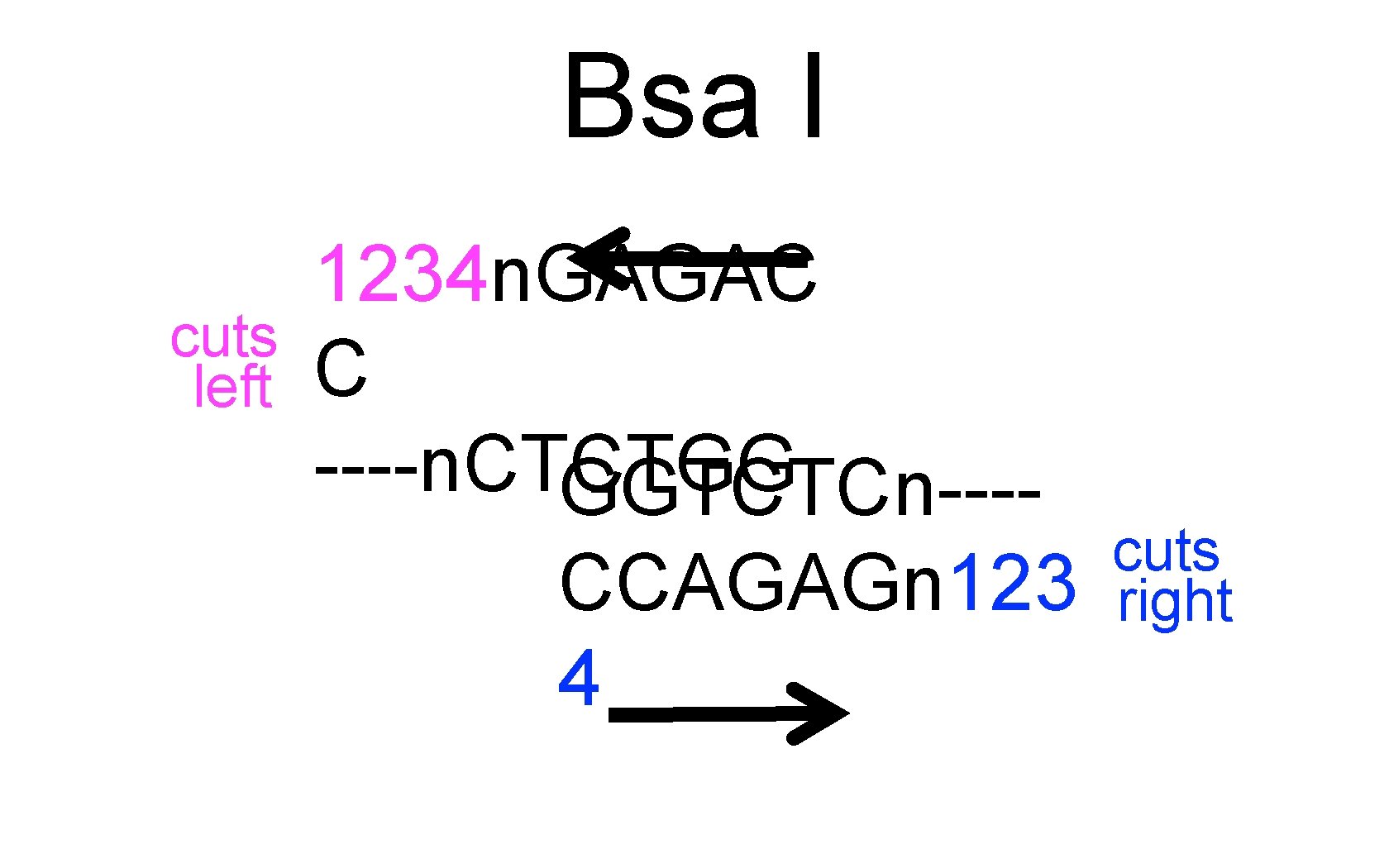
Bsa I cuts left 1234 n. GAGAC C ----n. CTCTGG GGTCTCn---CCAGAGn 123 4 cuts right
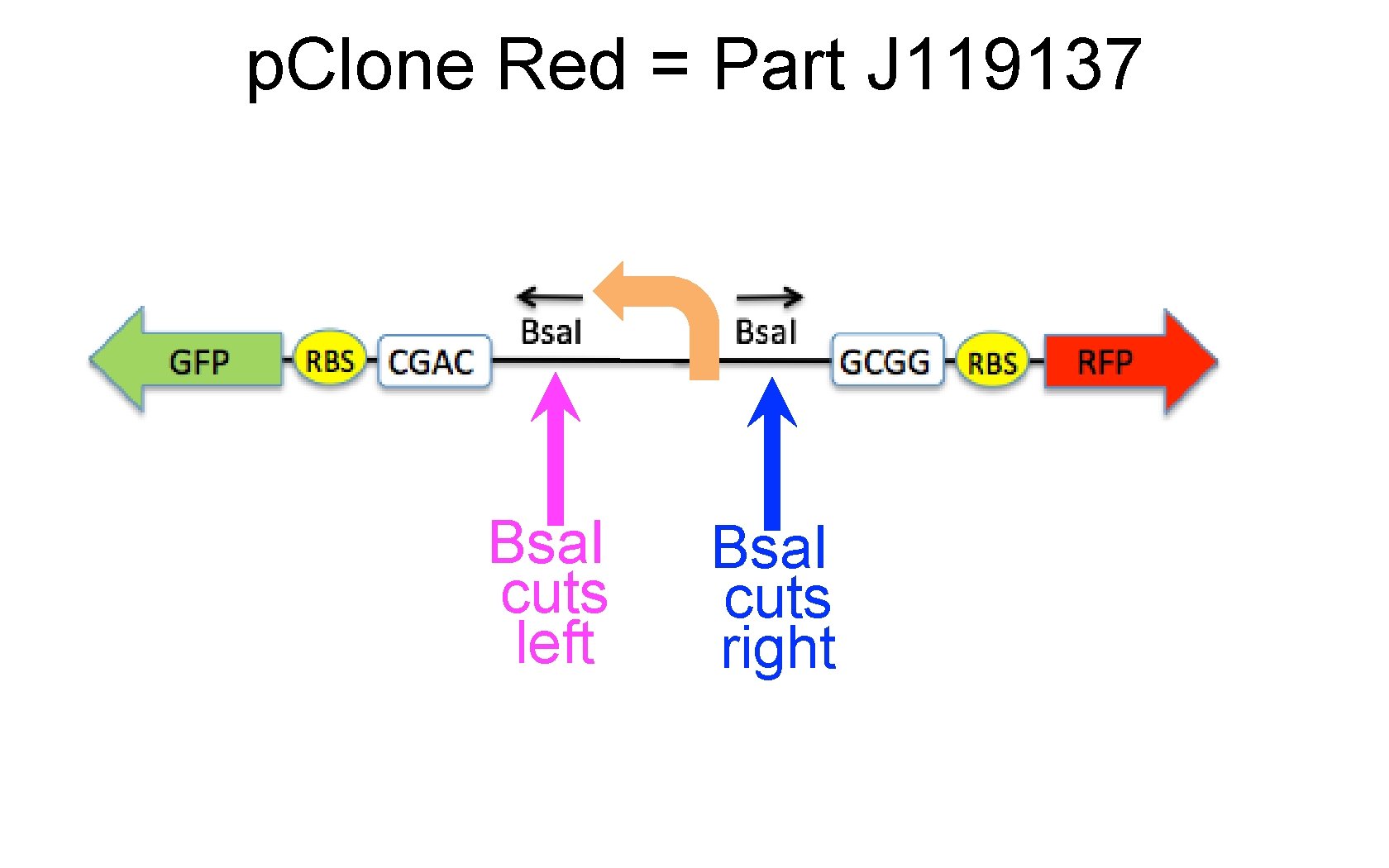
p. Clone Red = Part J 119137 Bsa. I cuts left Bsa. I cuts right
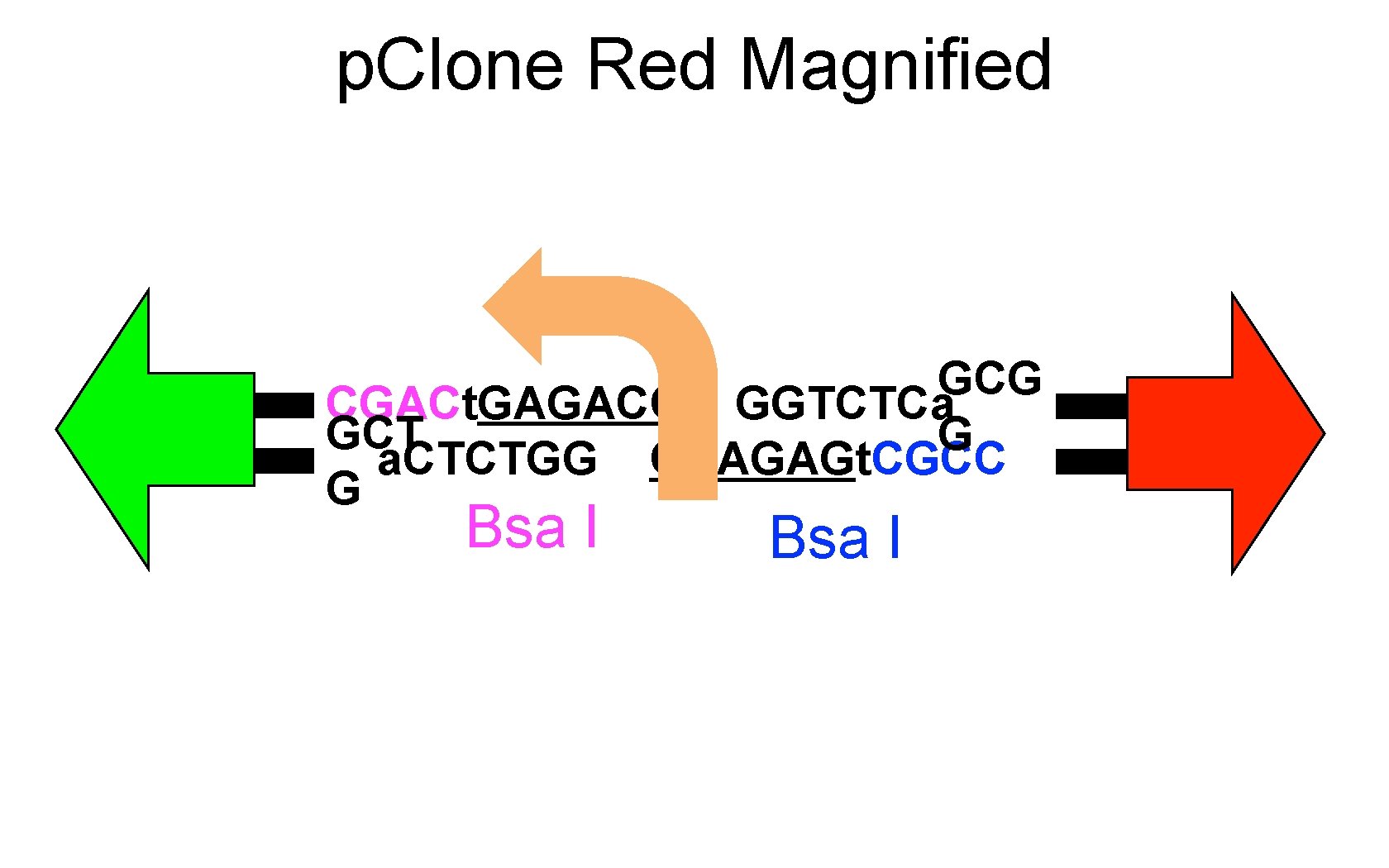
p. Clone Red Magnified GCG CGACt. GAGACC GGTCTCa GCT G a. CTCTGG CCAGAGt. CGCC G Bsa I
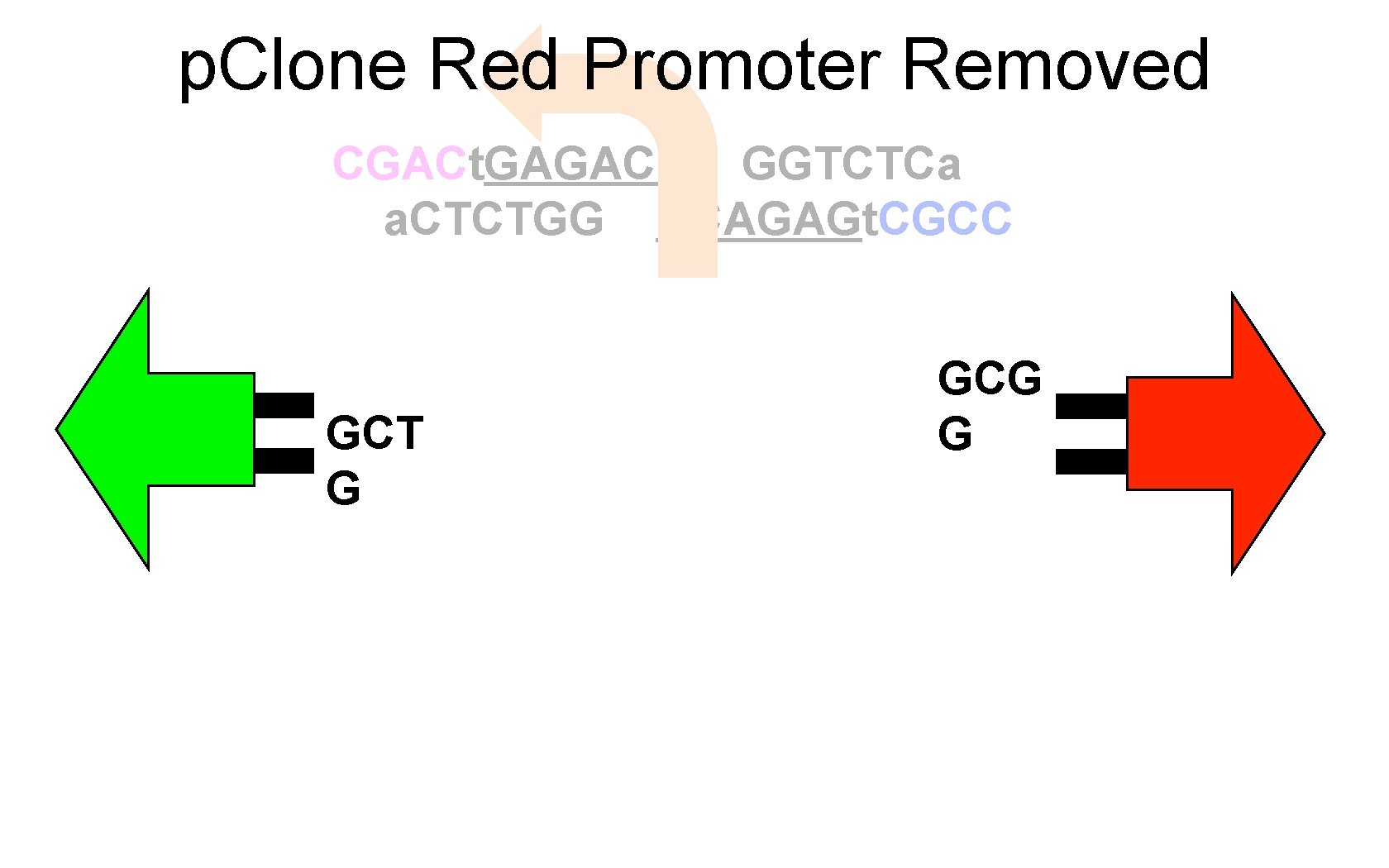
p. Clone Red Promoter Removed CGACt. GAGACC GGTCTCa a. CTCTGG CCAGAGt. CGCC GCT G GCG G
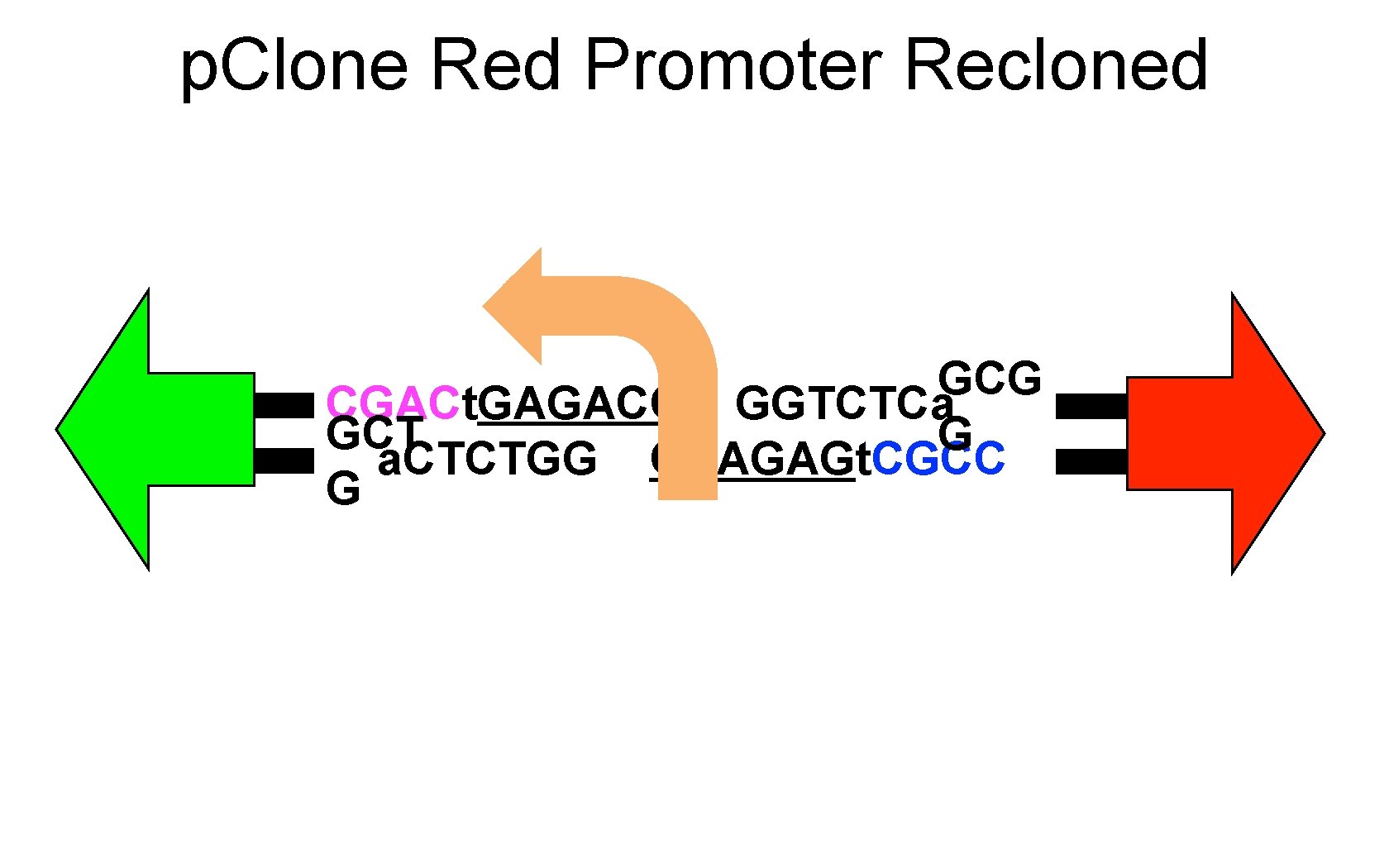
p. Clone Red Promoter Recloned GCG CGACt. GAGACC GGTCTCa GCT G a. CTCTGG CCAGAGt. CGCC G
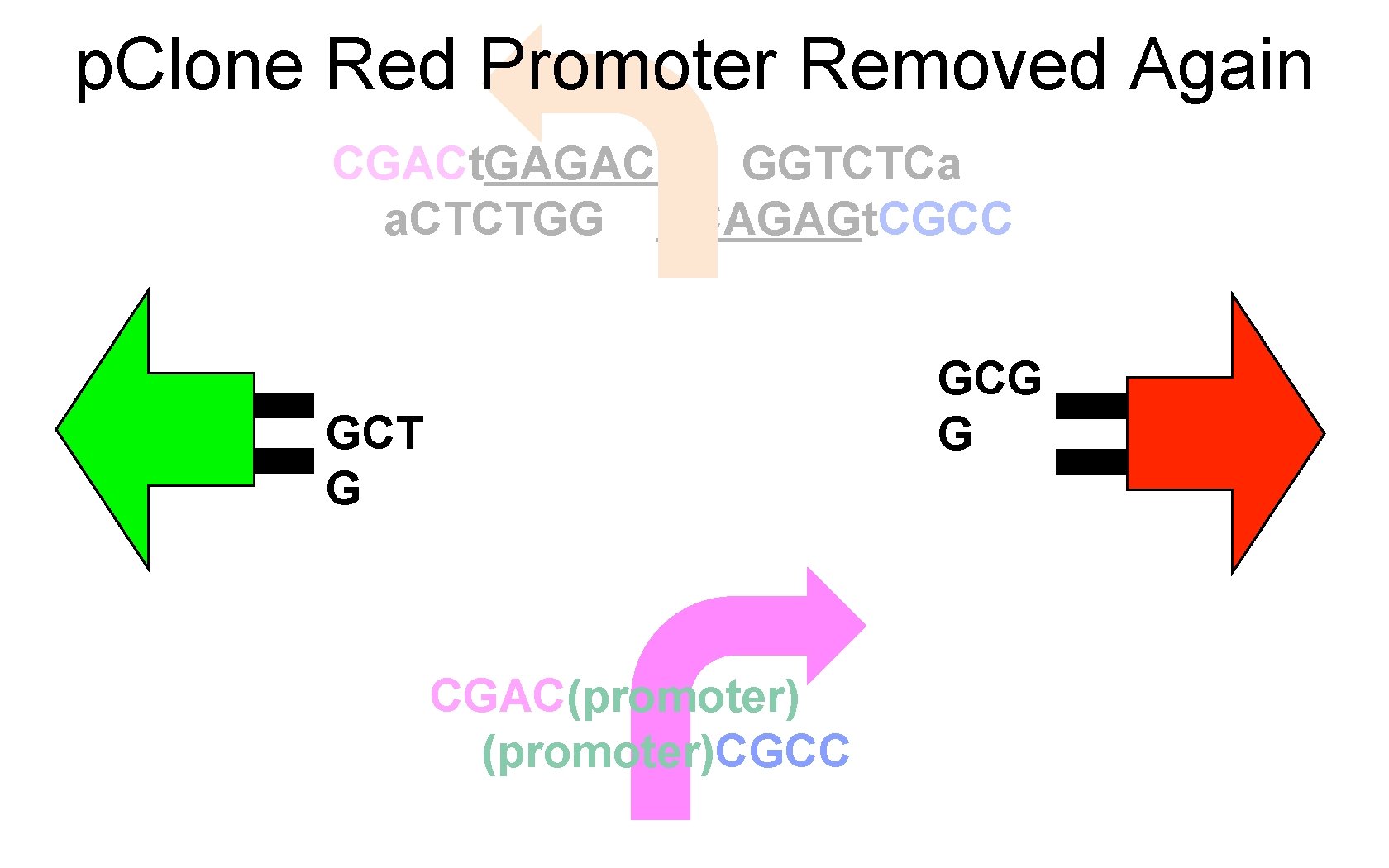
p. Clone Red Promoter Removed Again CGACt. GAGACC GGTCTCa a. CTCTGG CCAGAGt. CGCC GCG G GCT G CGAC(promoter)CGCC
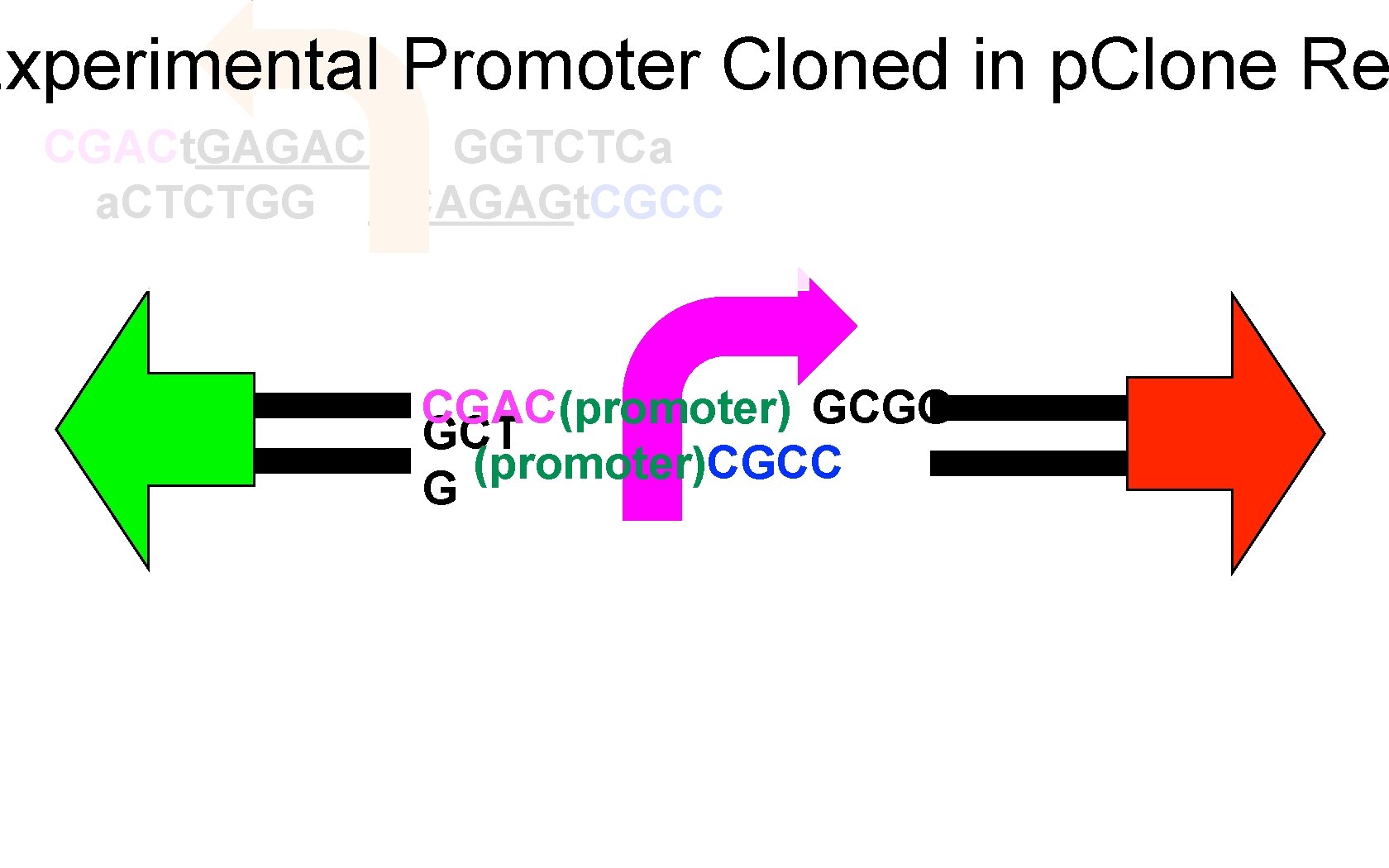
Experimental Promoter Cloned in p. Clone Re CGACt. GAGACC GGTCTCa a. CTCTGG CCAGAGt. CGCC CGAC(promoter) GCGG GCT (promoter)CGCC G
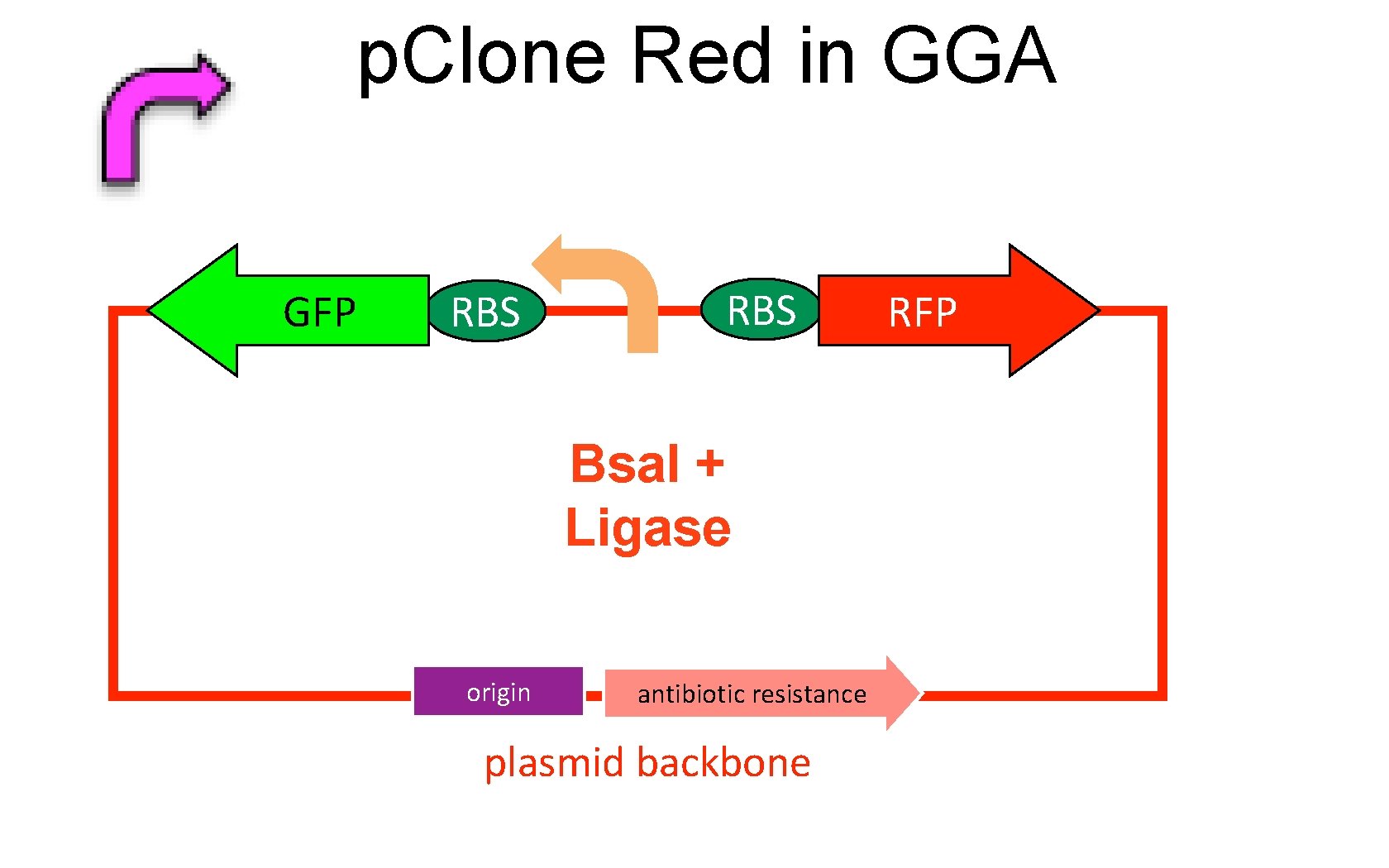
p. Clone Red in GGA GFP RBS Bsa. I + Ligase origin antibiotic resistance plasmid backbone RFP
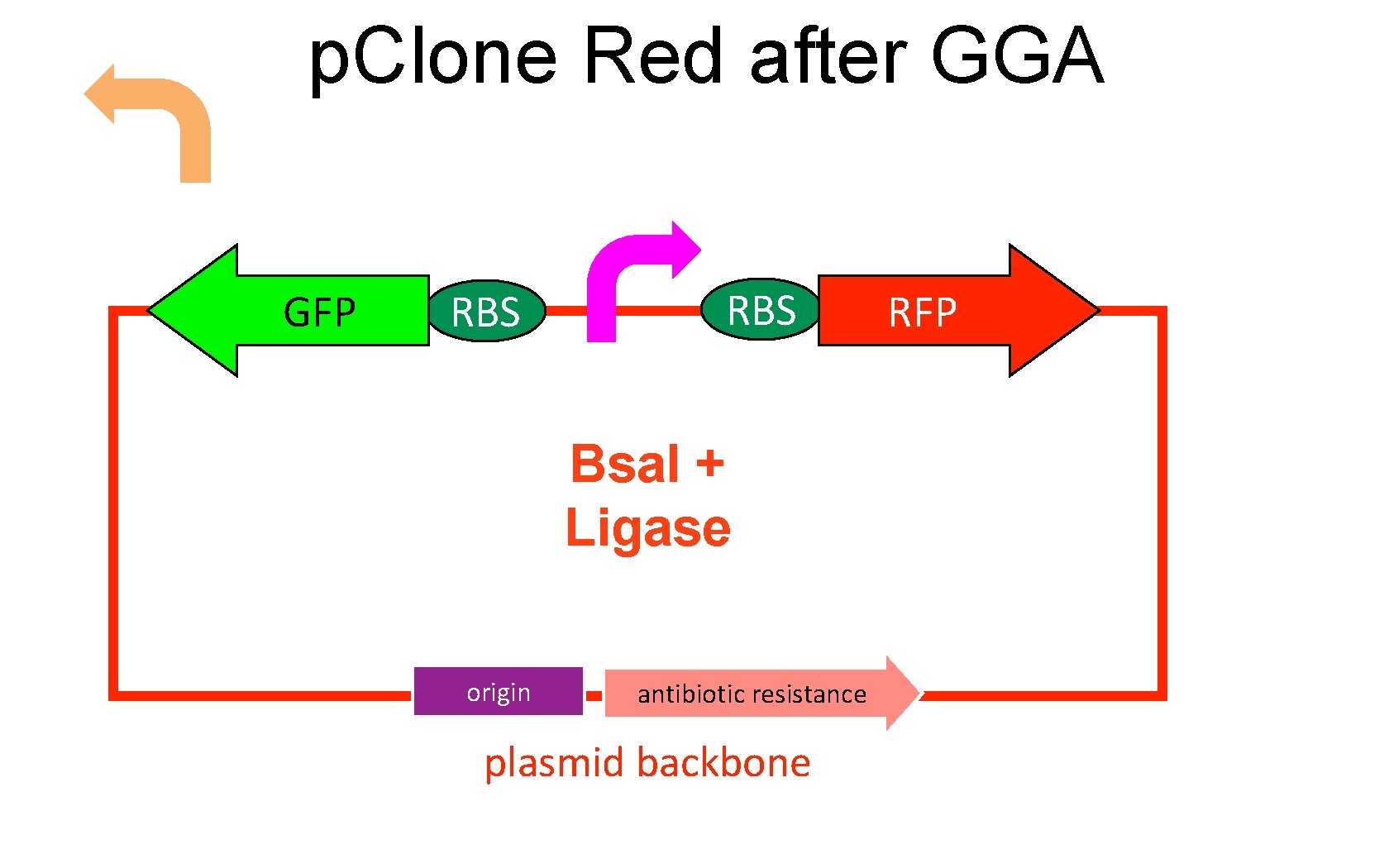
p. Clone Red after GGA GFP RBS Bsa. I + Ligase origin antibiotic resistance plasmid backbone RFP

p. Clone Red + No Promoter p. Clone Red only notice only green colonies
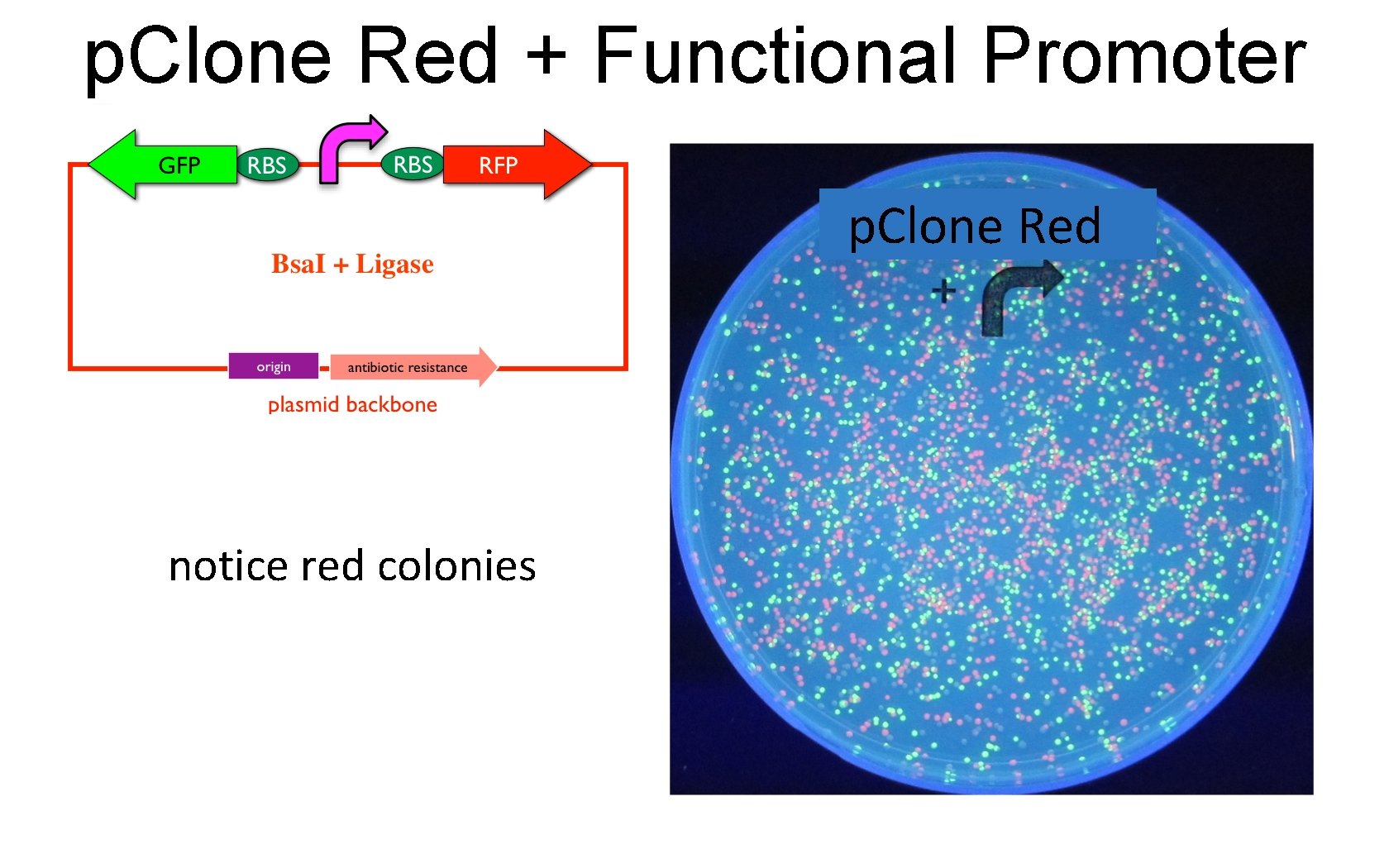
p. Clone Red + Functional Promoter p. Clone Red notice red colonies
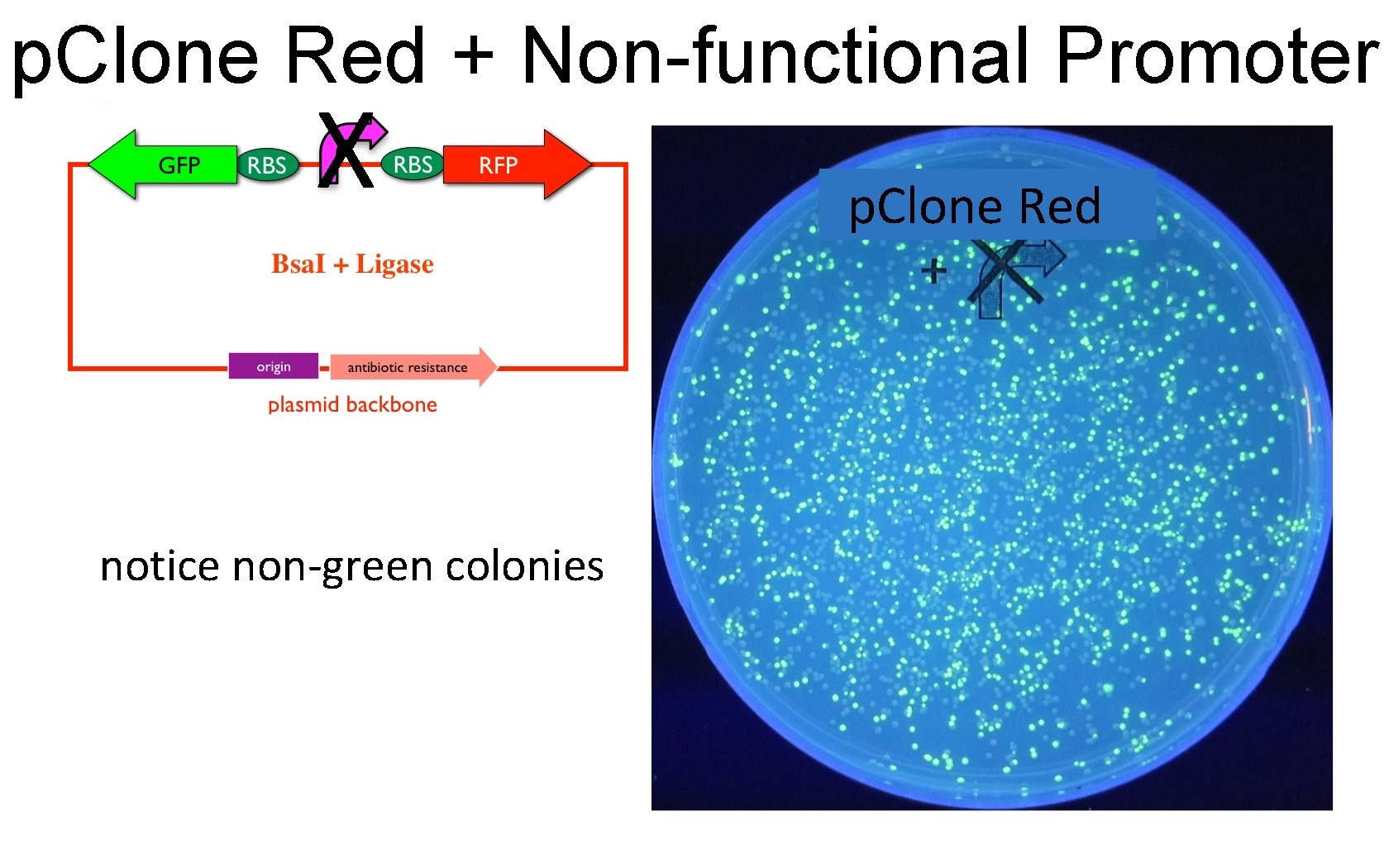
p. Clone Red + Non-functional Promoter X notice non-green colonies p. Clone Red

Registry of Standard Parts
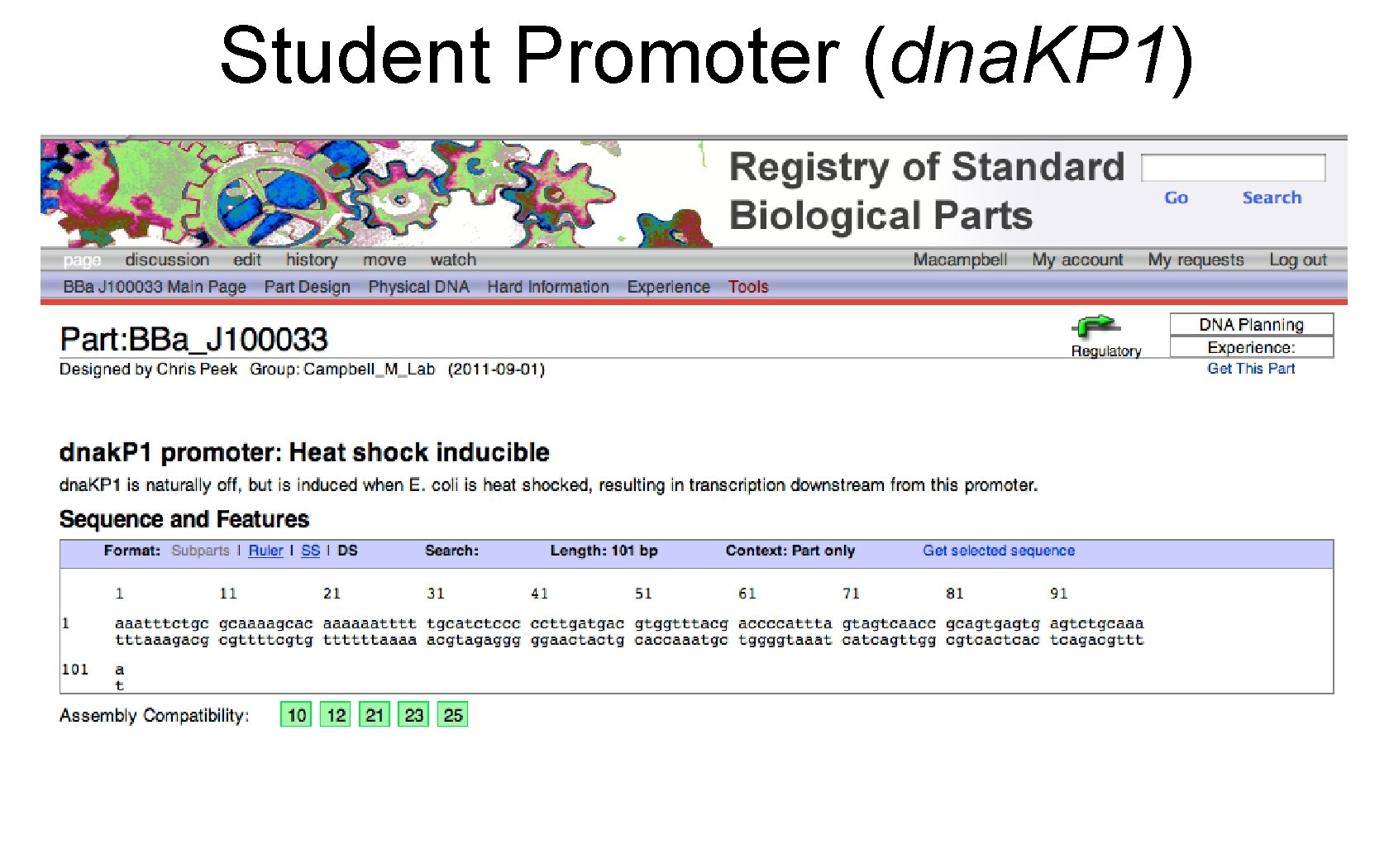
Student Promoter (dna. KP 1)
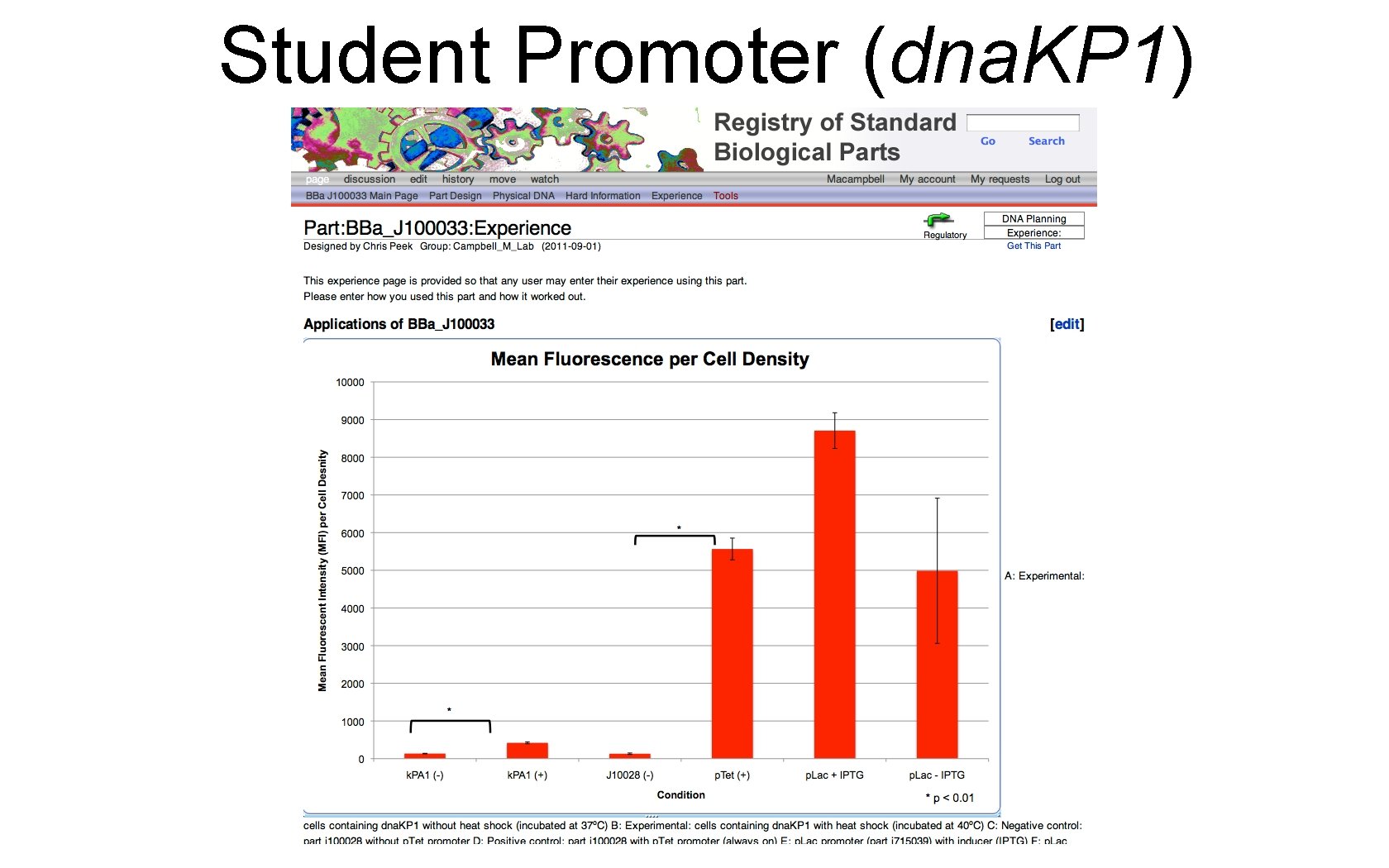
Student Promoter (dna. KP 1)
- Slides: 31