Using a strand displacing polymerase to program chemical




















- Slides: 42

Using a strand displacing polymerase to program chemical reaction networks Shalin Shah

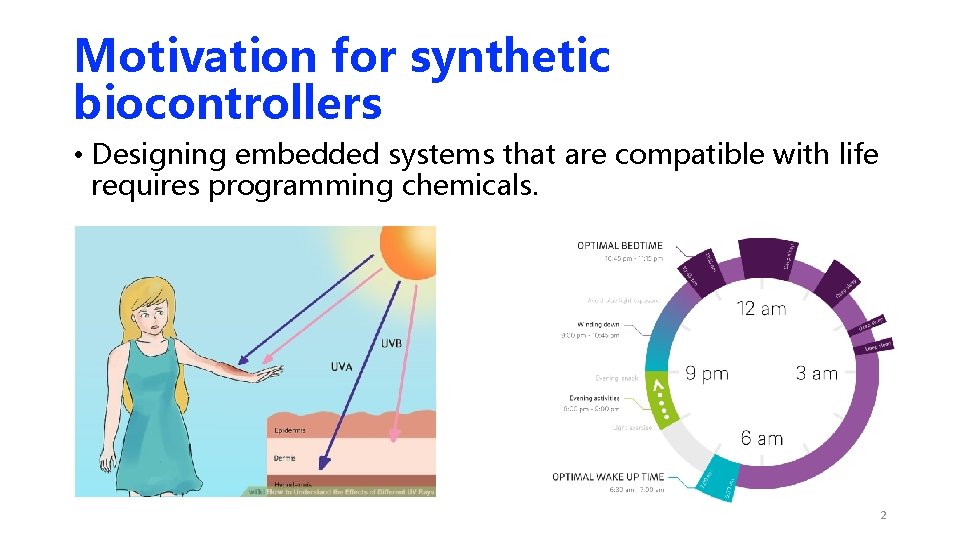
Motivation for synthetic biocontrollers • Designing embedded systems that are compatible with life requires programming chemicals. Intel 8085 COMP ARCH 101 2

Synthetic biocontrollers Soloveichik et al. PNAS (2010) Shah et al. DNA 25 (2018) • Inverse problem is to use CRN (Turing universal) as a highlevel programming language. • Use DNA systems (highly programmable) to implement arbitrary chemical programs. CRNs: Chemical reaction 3

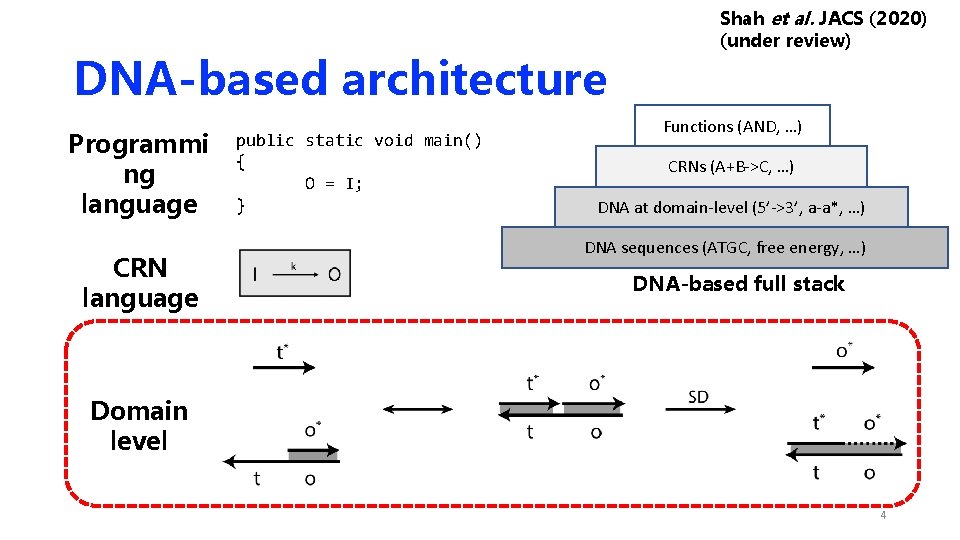
DNA-based architecture Programmi ng language CRN language public static void main() { O = I; } Shah et al. JACS (2020) (under review) Functions (AND, …) CRNs (A+B->C, …) DNA at domain-level (5’->3’, a-a*, …) DNA sequences (ATGC, free energy, …) DNA-based full stack Domain level 4

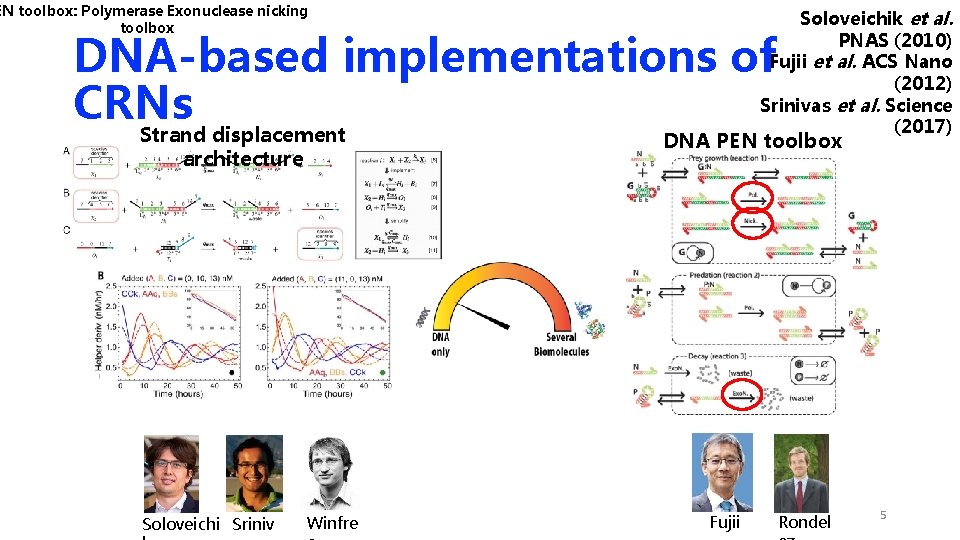
EN toolbox: Polymerase Exonuclease nicking toolbox Soloveichik et al. PNAS (2010) Fujii et al. ACS Nano (2012) Srinivas et al. Science (2017) DNA-based implementations of CRNs Strand displacement architecture Soloveichi Sriniv Winfre DNA PEN toolbox Fujii Rondel 5

Problem statement • Several difficulties with existing techniques: • DNA only systems are biologically simpler to design but they can be slow and leaky. • Multi-component systems are biologically more complex restricting the environmental conditions. • Problem statement: Design synthetic bio-controllers that are biologically simple and fast. • Our proposed solution: Polymerase strand displacement 6

Outline • Introduction to synthetic biocontrollers • CRN implementation: Model and theory • Towards in vitro implementation of PSD • Closing remarks on PSD and future work 7

What is strand displacement? t o t* o* Toehold mediated strand displacement (TMSD) o t* o* Polymerase strand Displacement (PSD) 8

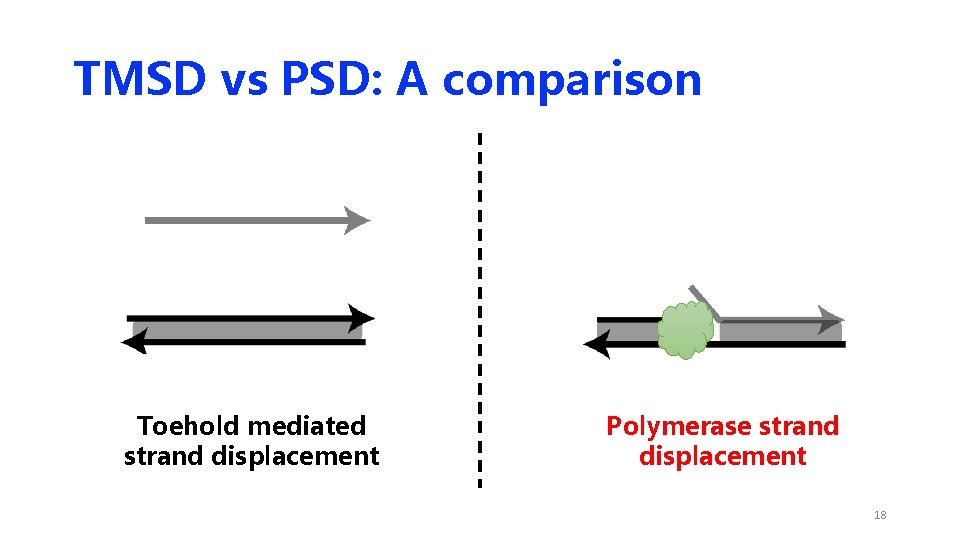
TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 9

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 10

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 11

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 12

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 13

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 14

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 15

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 16

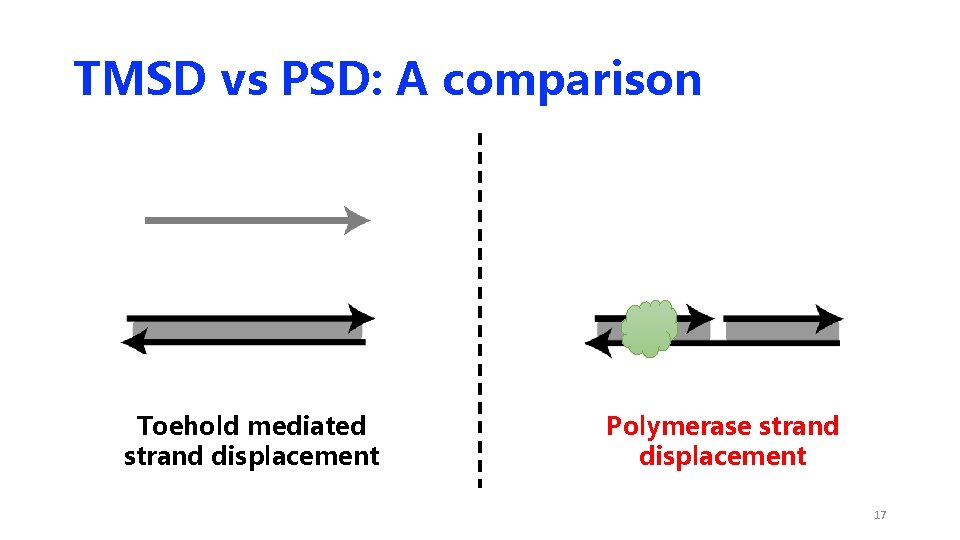
TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 17

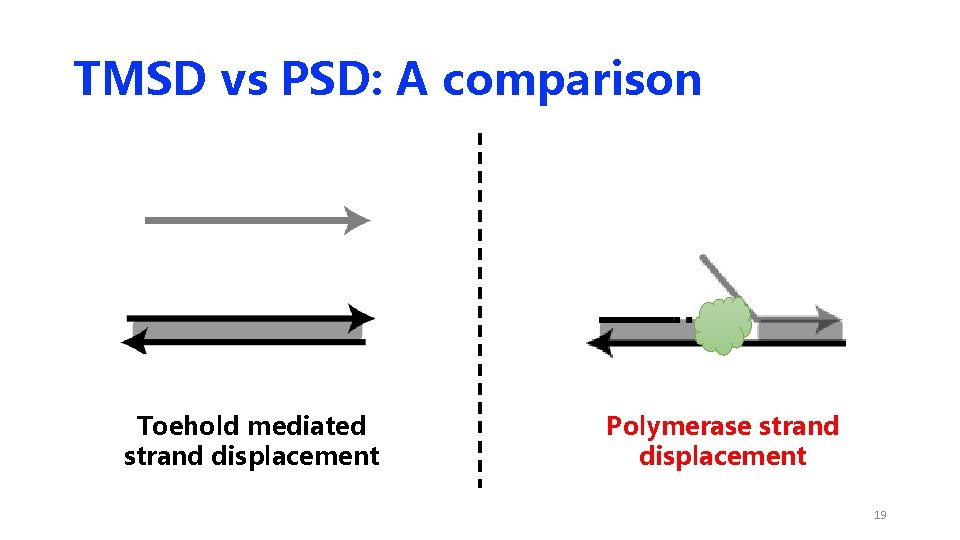
TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 18

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 19

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 20

TMSD vs PSD: A comparison Toehold mediated strand displacement Polymerase strand displacement 21

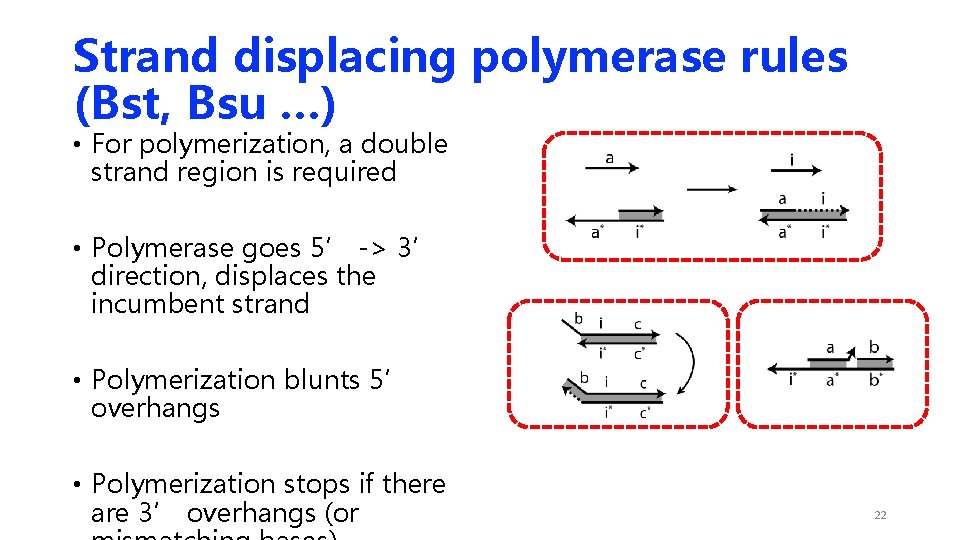
Strand displacing polymerase rules (Bst, Bsu …) • For polymerization, a double strand region is required • Polymerase goes 5’ -> 3’ direction, displaces the incumbent strand • Polymerization blunts 5’ overhangs • Polymerization stops if there are 3’ overhangs (or 22

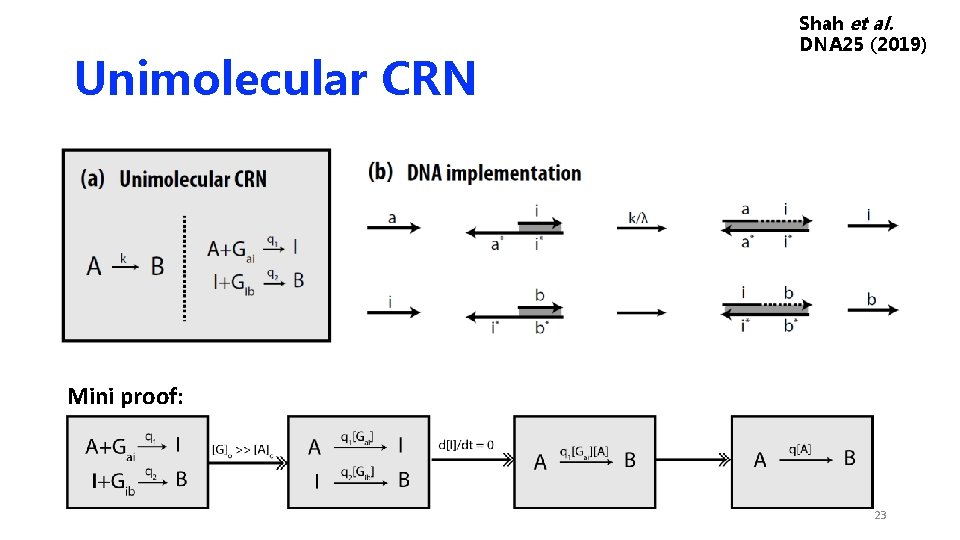
Unimolecular CRN Shah et al. DNA 25 (2019) Mini proof: 23

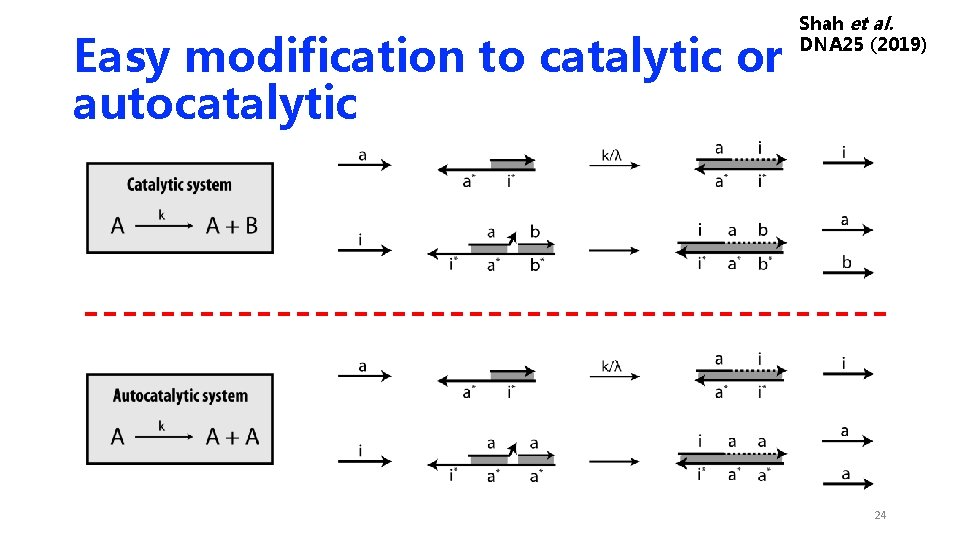
Easy modification to catalytic or autocatalytic Shah et al. DNA 25 (2019) 24

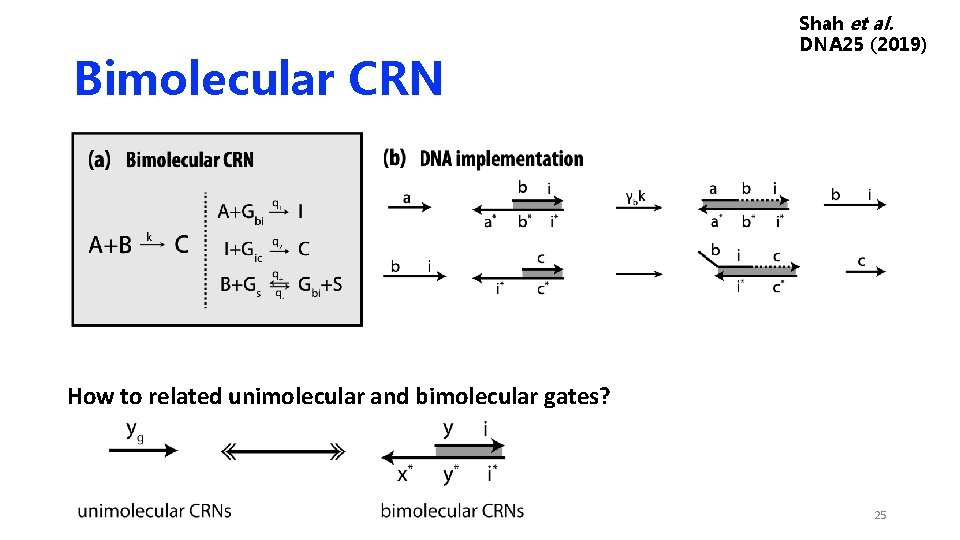
Bimolecular CRN Shah et al. DNA 25 (2019) How to related unimolecular and bimolecular gates? 25

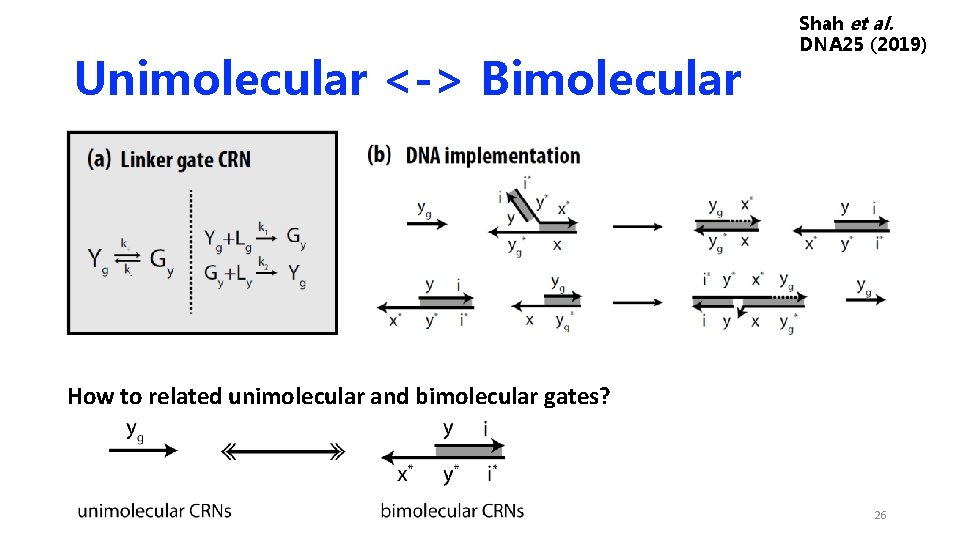
Unimolecular <-> Bimolecular Shah et al. DNA 25 (2019) How to related unimolecular and bimolecular gates? 26

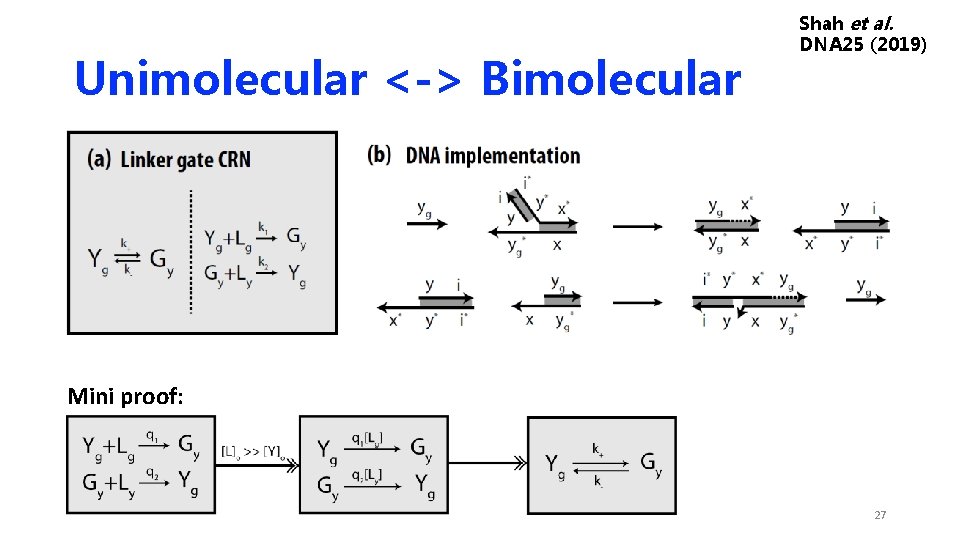
Unimolecular <-> Bimolecular Shah et al. DNA 25 (2019) Mini proof: 27

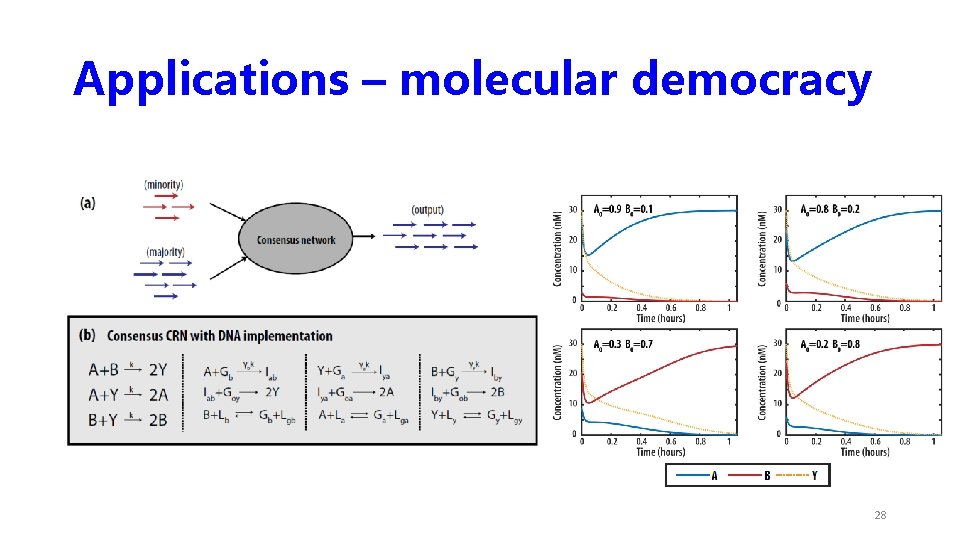
Applications – molecular democracy 28

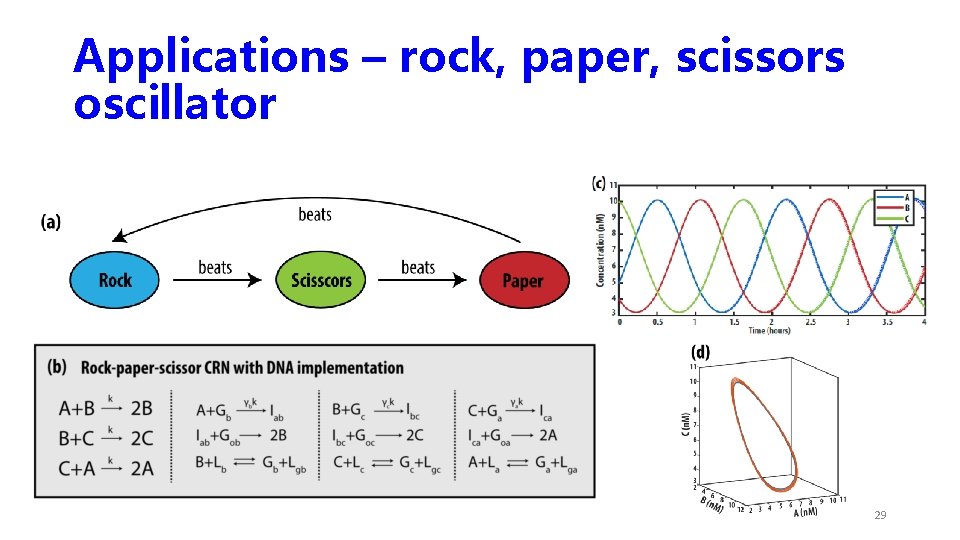
Applications – rock, paper, scissors oscillator 29

Outline • Introduction to synthetic biocontrollers • CRN implementation: Model and theory • Towards in vitro implementation of PSD • Closing remarks on PSD and future work 30

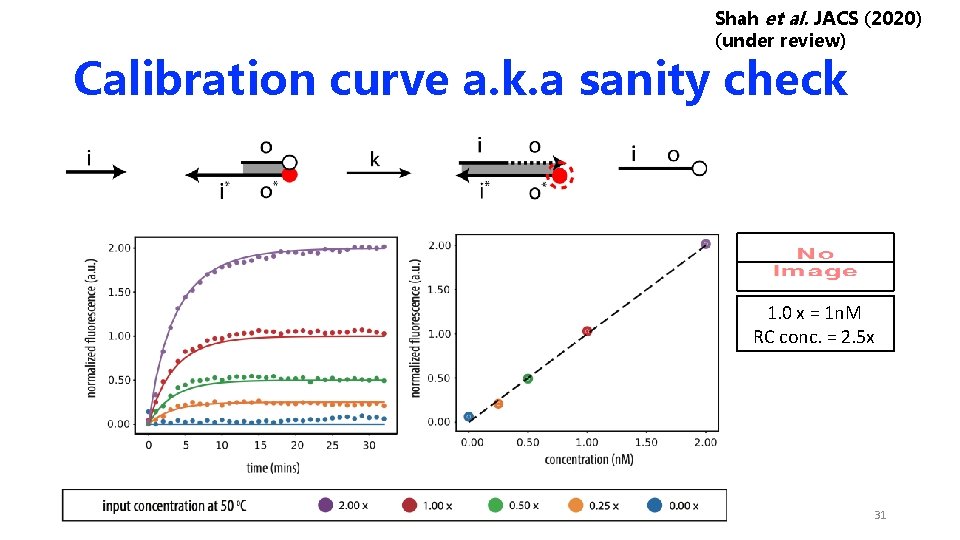
Shah et al. JACS (2020) (under review) Calibration curve a. k. a sanity check 1. 0 x = 1 n. M RC conc. = 2. 5 x 31

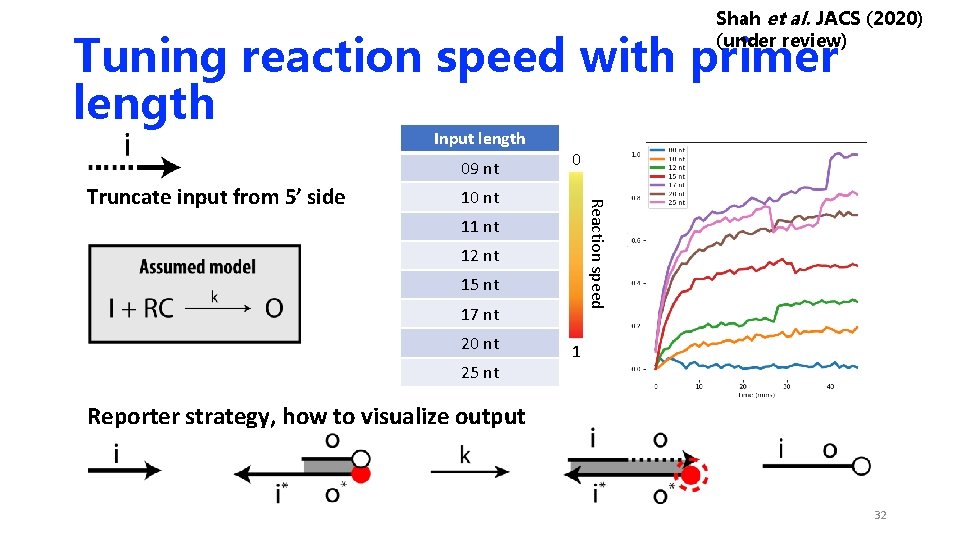
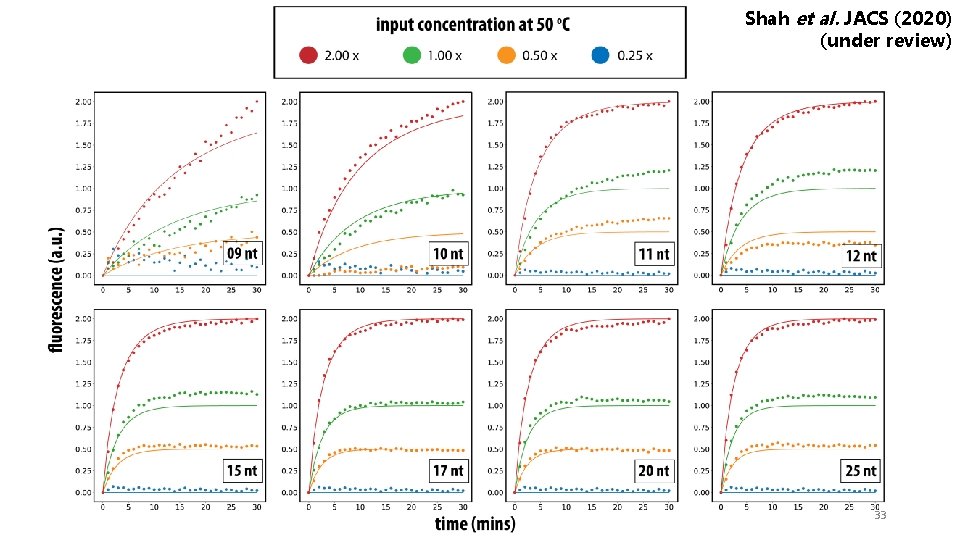
Shah et al. JACS (2020) (under review) Tuning reaction speed with primer length Input length 09 nt 10 nt Reaction speed Truncate input from 5’ side 0 11 nt 12 nt 15 nt 17 nt 20 nt 25 nt 1 Reporter strategy, how to visualize output 32

Shah et al. JACS (2020) (under review) 33

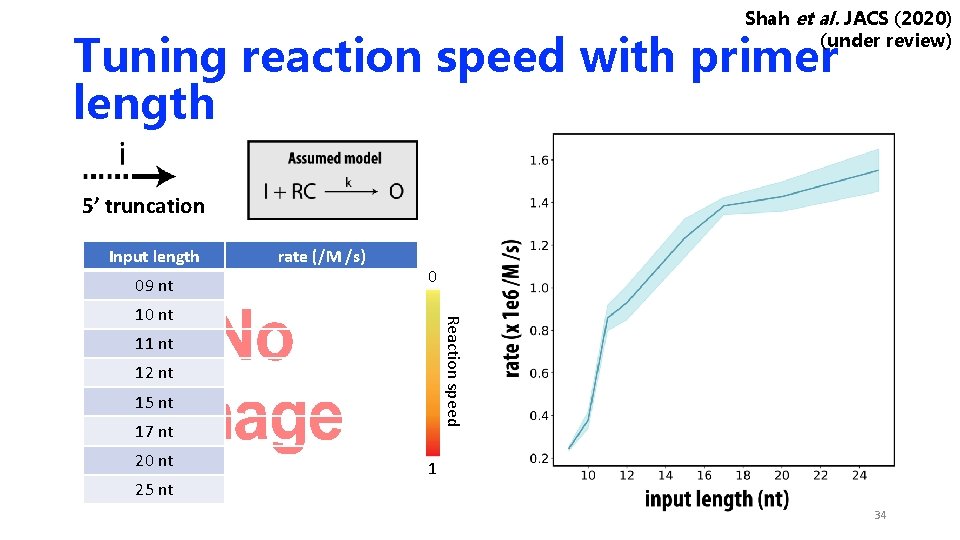
Shah et al. JACS (2020) (under review) Tuning reaction speed with primer length 5’ truncation Input length 09 nt rate (/M /s) 0 Reaction speed 10 nt 11 nt 12 nt 15 nt 17 nt 20 nt 25 nt 1 34

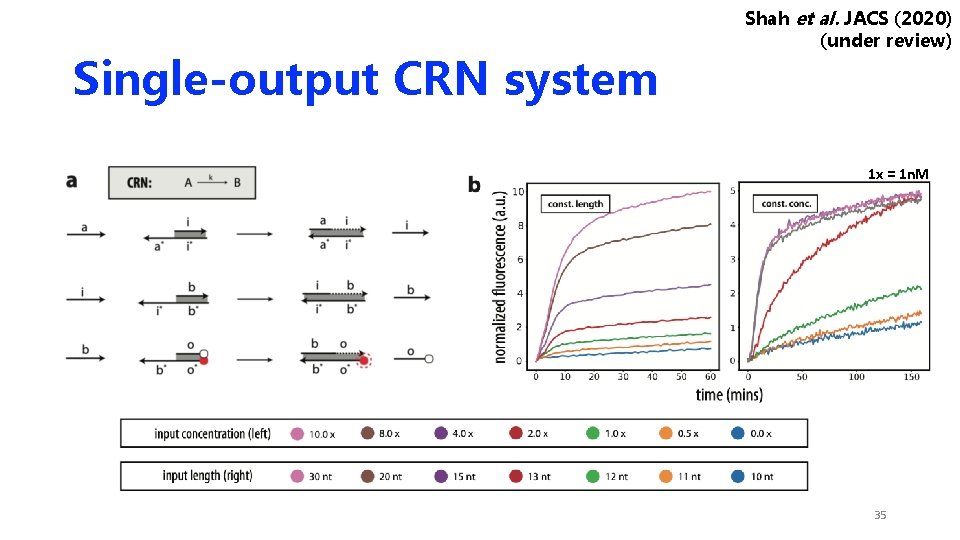
Single-output CRN system Shah et al. JACS (2020) (under review) 1 x = 1 n. M 35

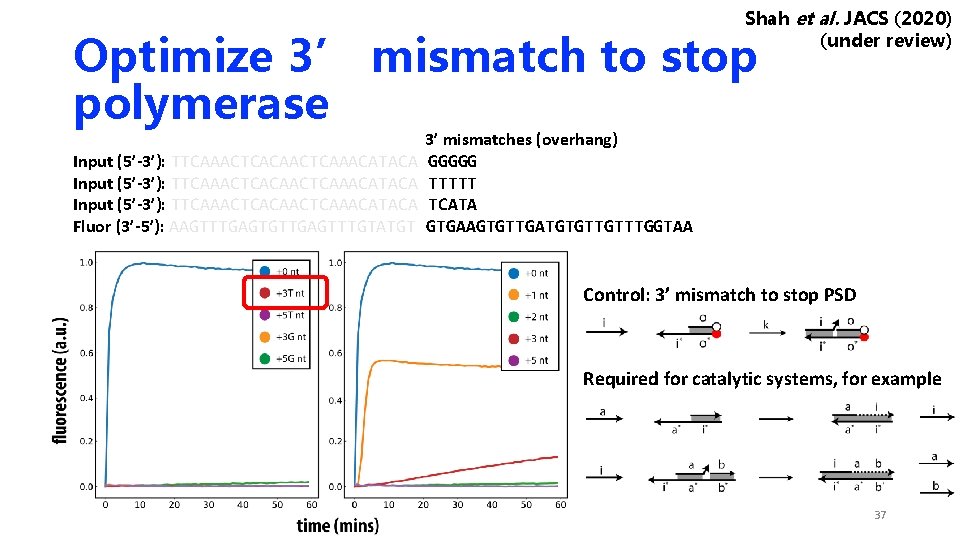
Optimize 3’ mismatch to stop polymerase • Such polymerase stopper strategy is required for any complex with more than one strand. For example, catalytic reaction A -> A + B • Two strategies to stop polymerase: • Random mismatch sequence with ~ 50% GC content • Poly-T tail/ poly-G tail 36

Shah et al. JACS (2020) (under review) Optimize 3’ mismatch to stop polymerase Input (5’-3’): TTCAAACTCACAACTCAAACATACA Fluor (3’-5’): AAGTTTGAGTGTTGAGTTTGTATGT 3’ mismatches (overhang) GGGGG TTTTT TCATA GTGAAGTGTTGATGTGTTGTTTGGTAA Control: 3’ mismatch to stop PSD Required for catalytic systems, for example 37

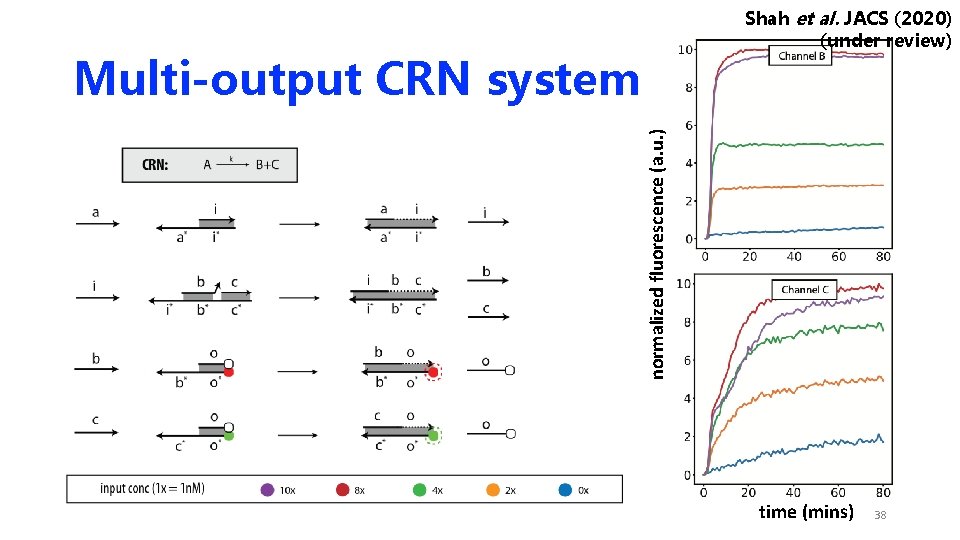
Shah et al. JACS (2020) (under review) normalized fluorescence (a. u. ) Multi-output CRN system time (mins) 38

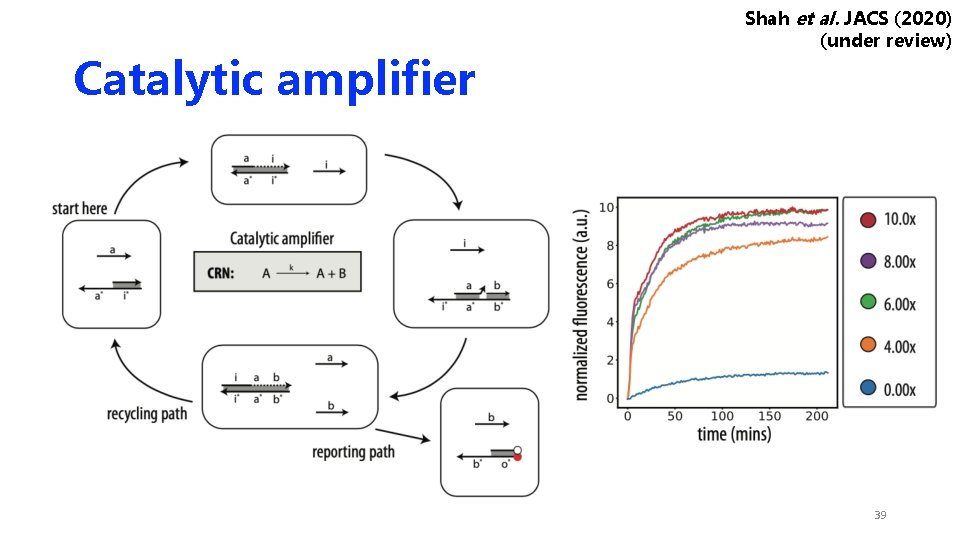
Catalytic amplifier Shah et al. JACS (2020) (under review) 39

Outline • Introduction to synthetic biocontrollers • CRN implementation: Model and theory • Towards in vitro implementation of PSD • Closing remarks on PSD and future work 40

Summary of our work • Proposed a new framework to implement arbitrary CRNs. • Our design uses a fast strand displacing polymerase enzyme as an energy source. • The reaction rate can be tuned by length of input (primer), concentration of input/ gates. • Clamps at the 5’ end can effectively reduce circuit leak due to fraying. 41

Discussion and future work • Experimental demonstration of complex reaction networks such as oscillators. • Development of a simple compiler that can convert a set of CRNs to our DNA-based CRNs. • Theoretical closed-form solution to test divergence error w. r. t gate conc. and time. 42