Uses of population genetic data Species identification taxonomy























- Slides: 41

Uses of population genetic data Species identification (taxonomy) Population/stock differentiation Individual identification

Genetics questions relevant to conservation biology: are x and y different species? are populations genetically different? how much variation is present in a population? how much variation has been lost? which parents contributed to the breeding population? how much migration is occurring between populations?

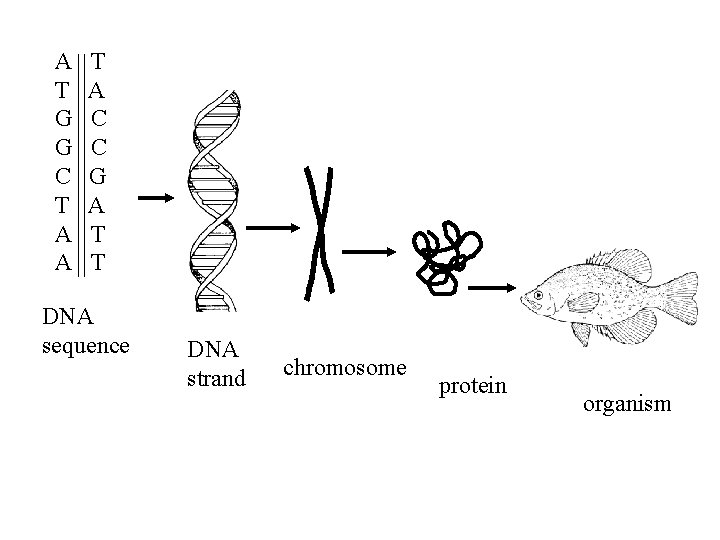
A T G G C T A A T A C C G A T T DNA sequence DNA strand chromosome protein organism

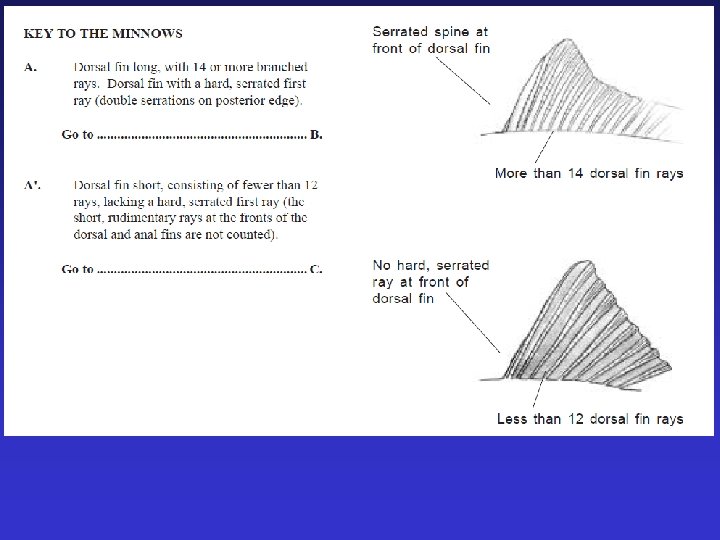
morphometric and meristic counts ~ 40 characters Often lethal


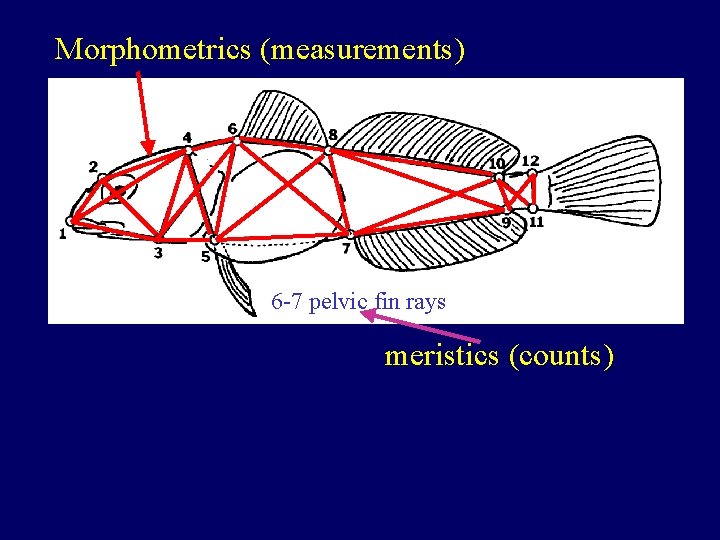
Morphometrics (measurements) 6 -7 pelvic fin rays meristics (counts)

Issues with morphometric and meristic data • Are they different enough? • What is the significance of the differences? (do they represent genetic differences? ) • Do the differences indicate reproductive isolation?

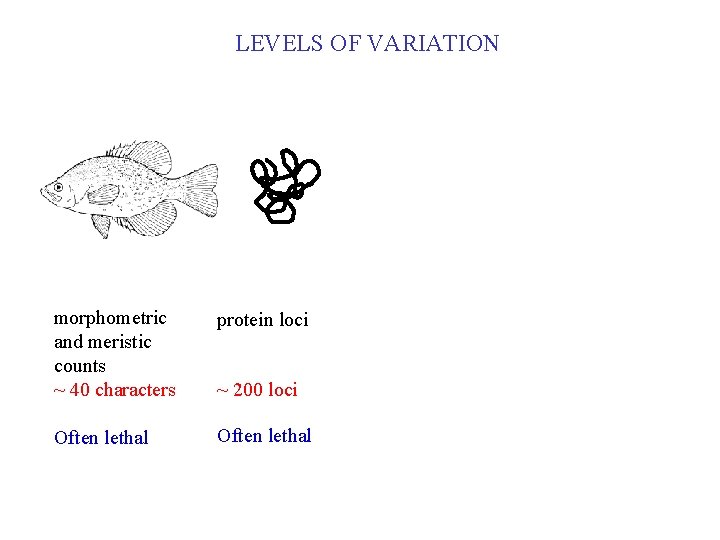
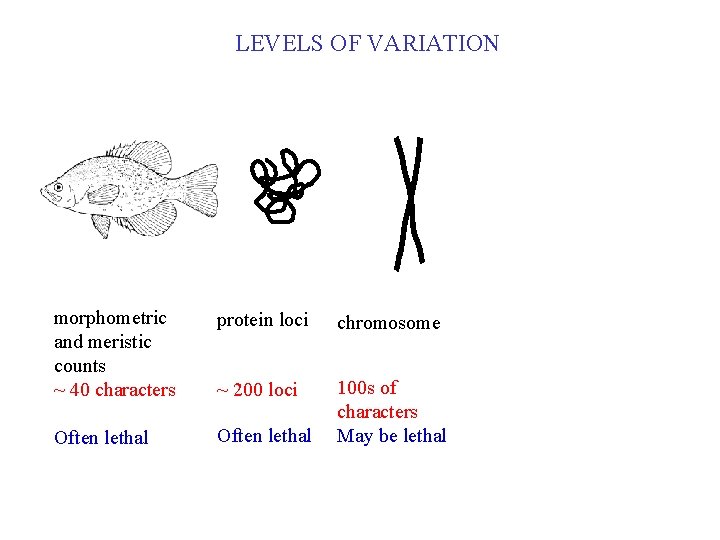
LEVELS OF VARIATION morphometric and meristic counts ~ 40 characters protein loci Often lethal ~ 200 loci

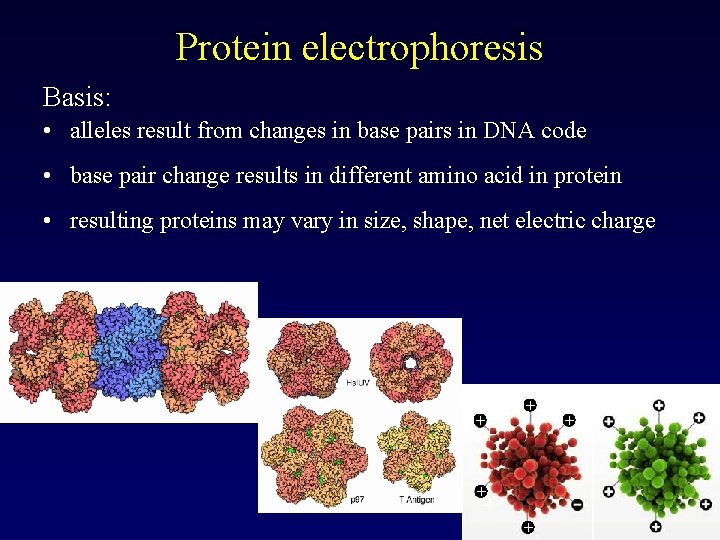
Protein electrophoresis Basis: • alleles result from changes in base pairs in DNA code • base pair change results in different amino acid in protein • resulting proteins may vary in size, shape, net electric charge + + + + +

Protein electrophoresis Basis: • electrophoresis separates proteins on the basis of their net electric charge, and size/shape • thus, alleles coding for proteins with different size or charge will be detected as different • genotype is deduced from protein types • co-dominant, low variability

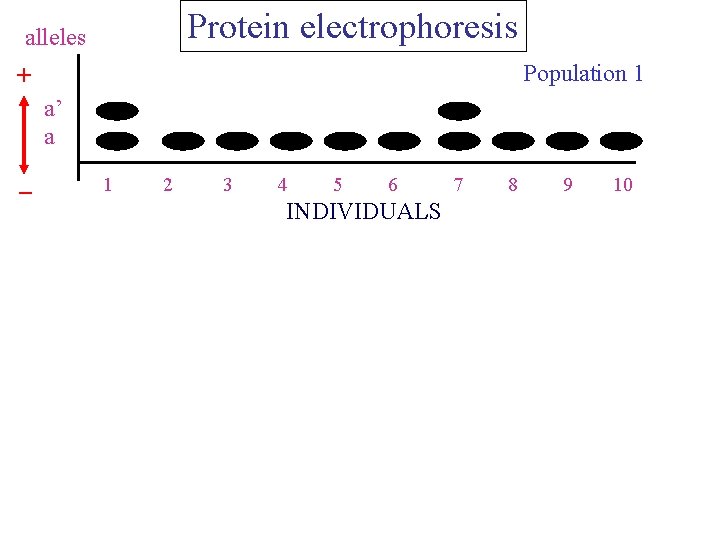
Protein electrophoresis alleles Population 1 + a’ a _ 1 2 3 4 5 6 INDIVIDUALS 7 8 9 10

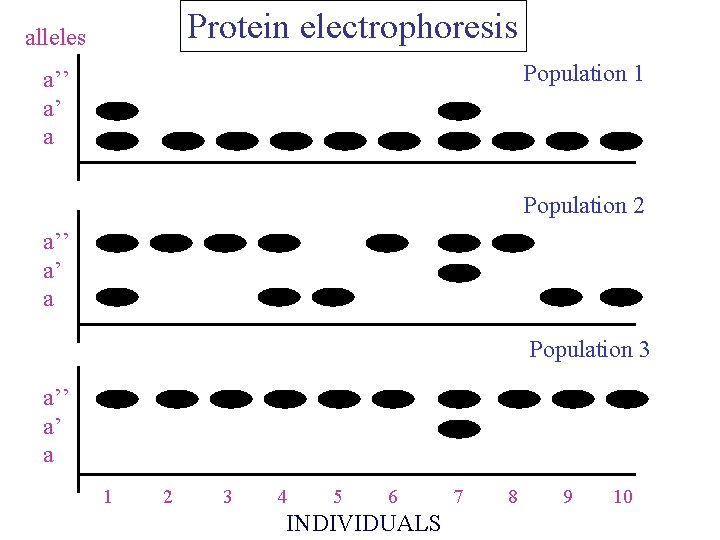
Protein electrophoresis alleles Population 1 a’’ a’ a Population 2 a’’ a’ a Population 3 a’’ a’ a 1 2 3 4 5 6 INDIVIDUALS 7 8 9 10

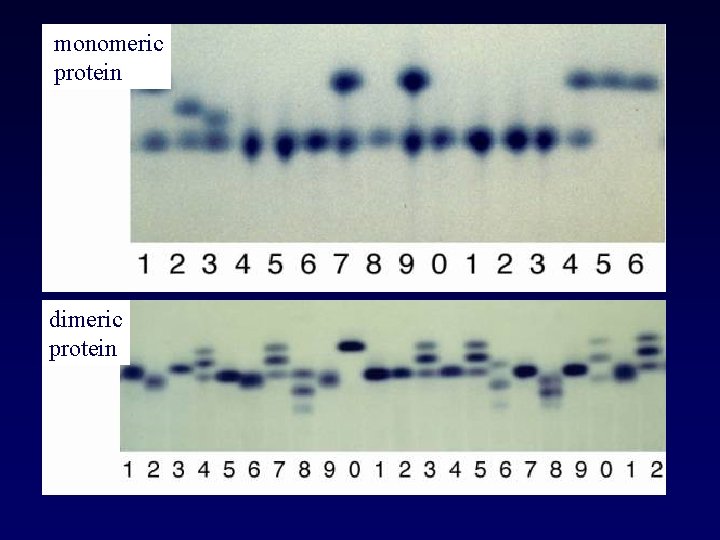
monomeric protein dimeric protein

LEVELS OF VARIATION morphometric and meristic counts ~ 40 characters protein loci chromosome ~ 200 loci Often lethal 100 s of characters May be lethal

Cytogenetics – information from chromosome number, shape, banding patterns

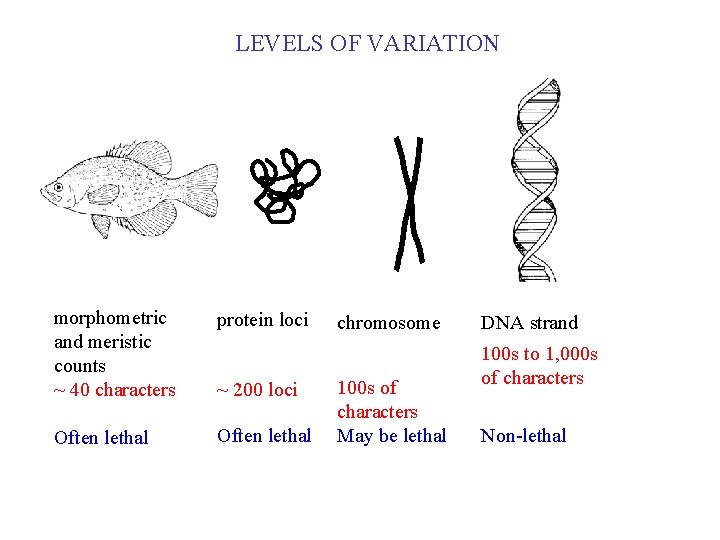
LEVELS OF VARIATION morphometric and meristic counts ~ 40 characters protein loci Often lethal ~ 200 loci chromosome 100 s of characters May be lethal DNA strand 100 s to 1, 000 s of characters Non-lethal

DNA Mitochondrial Nuclear • • • circular small - 16, 000 -20, 000 bp many copies maternal inheritance moderate variation mostly coded loci linear 3 billion + base pairs two copies per cell bi-parental inheritance high variation much non-coded DNA

‘Problems’ with DNA • not very much of it (2 -100 copies per cell) - need to ‘amplify’ it • too much of it (3 billion base pairs) - need to look at small pieces at a time • different areas of DNA have different variation

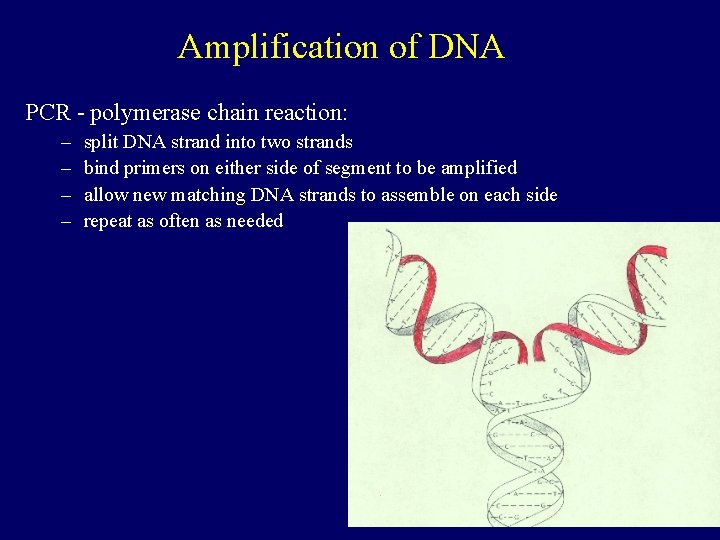
Amplification of DNA PCR - polymerase chain reaction: – – split DNA strand into two strands bind primers on either side of segment to be amplified allow new matching DNA strands to assemble on each side repeat as often as needed

Polymerase Chain Reaction (PCR) - DNA replication in a test tube Requires ingredients: DNA template DNA polymerase free nucleotide bases DNA template must be separated (denatured, unwound) must be a primer for DNA polymerase to add free nucleotides to

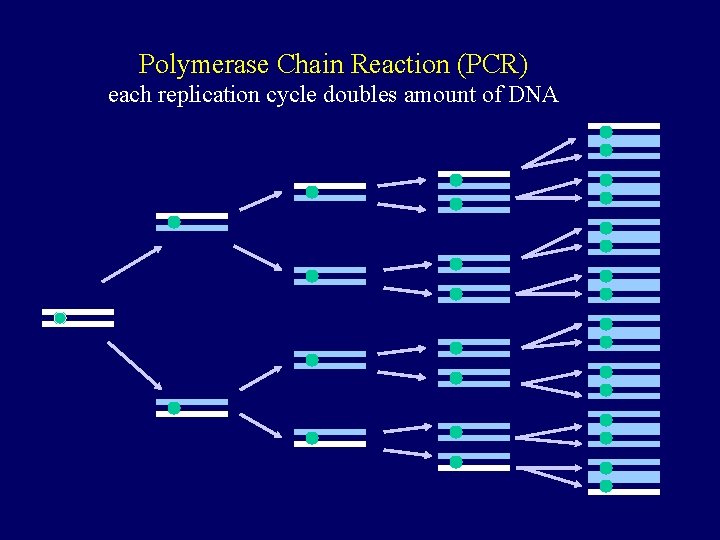
Polymerase Chain Reaction (PCR) each replication cycle doubles amount of DNA

Polymerase Chain Reaction (PCR) Requires ingredients: DNA template DNA polymerase free nucleotide bases DNA template must be separated (denatured, unwound) heat will denature DNA, but deactivates protein enzymes use Taq DNA polymerase (from bacterium Thermus aquaticus from thermal vents)


Polymerase Chain Reaction (PCR) Requires ingredients: DNA template DNA polymerase free nucleotide bases DNA template must be separated (denatured, unwound) must be a primer for DNA polymerase to add free nucleotides to short sequence of DNA that binds ‘upstream’ of area to be replicated

DNA analyses • mt. DNA (mitochondrial DNA analysis) • RFLP (restriction fragment length polymorphism) • RAPD (randomly amplified polymorphic DNA) • AFLP (amplified fragment length polymorphism) • microsatellites (SSR, simple sequence repeats) • single nucleotide polymorphisms (SNPs)

Restriction Fragment Length Polymorphism (RFLP) - cut DNA with restriction enzymes - isolate cut fragments based on length (electrophoresis) - deduce length of fragments - individuals differ based on mutations at restriction sites A T T G A C T T A A G C G T A A C T G A A T T C G C A T C cleavage site (palindrome)

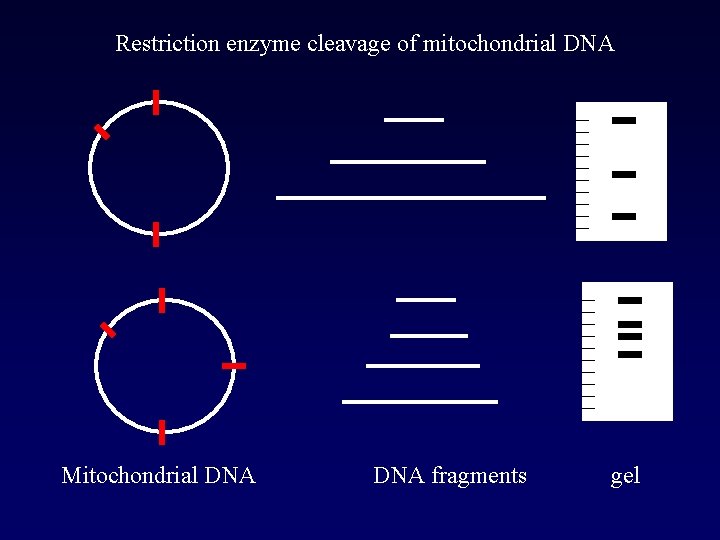
Restriction enzyme cleavage of mitochondrial DNA Mitochondrial DNA fragments gel

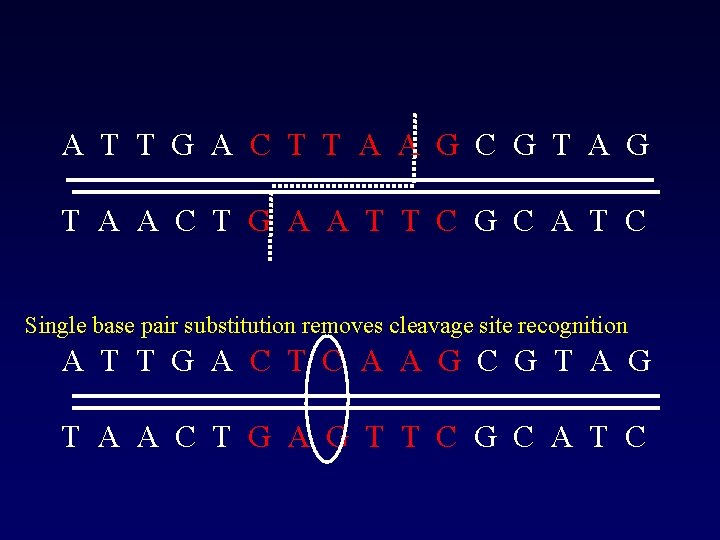
A T T G A C T T A A G C G T A A C T G A A T T C G C A T C Single base pair substitution removes cleavage site recognition A T T G A C T C A A G C G T A A C T G A G T T C G C A T C

Microsatellite DNA • tandem repeats of short DNA sequences (e. g. ACACAC…) • number of repeats is highly variable – easy cross-over ‘mistake” • co-dominant

Microsatellite DNA • tandem repeats of short DNA sequences (e. g. ACACAC…) • number of repeats is highly variable – easy cross-over ‘mistake” • co-dominant • isolate portions of DNA with primers (time-consuming to develop) • separate fragments by length ~ number of repeats

• SNPs (single nucleotide polymorphisms) • DNA sequencing • Genome sequencing • Barcode of Life Database (BOLD)

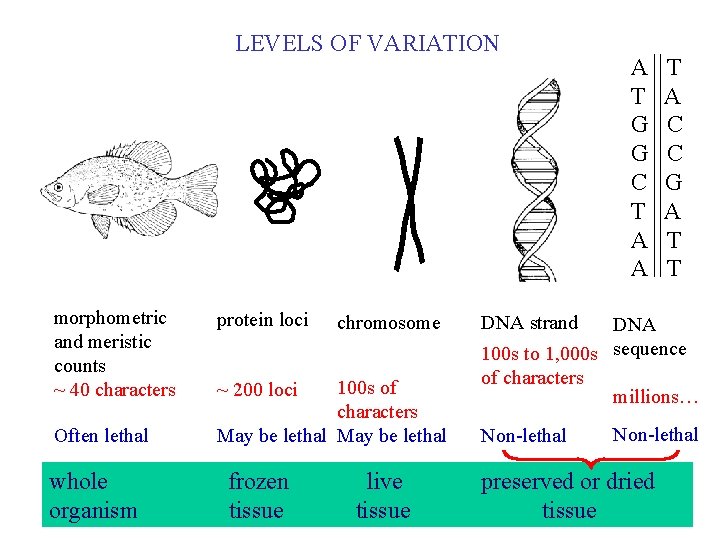
LEVELS OF VARIATION morphometric and meristic counts ~ 40 characters Often lethal whole organism protein loci chromosome 100 s of characters May be lethal ~ 200 loci frozen tissue live tissue A T G G C T A A T A C C G A T T DNA strand DNA 100 s to 1, 000 s sequence of characters millions… Non-lethal preserved or dried tissue

Step 1: find markers ‘survey’ species/population for polymorphisms before conducting full study = # of loci, # alleles/locus Step 1(b): find/develop primers for PCR often available on web databases from related taxa Step 2: determine how many markers are needed depends on question(s) of interest

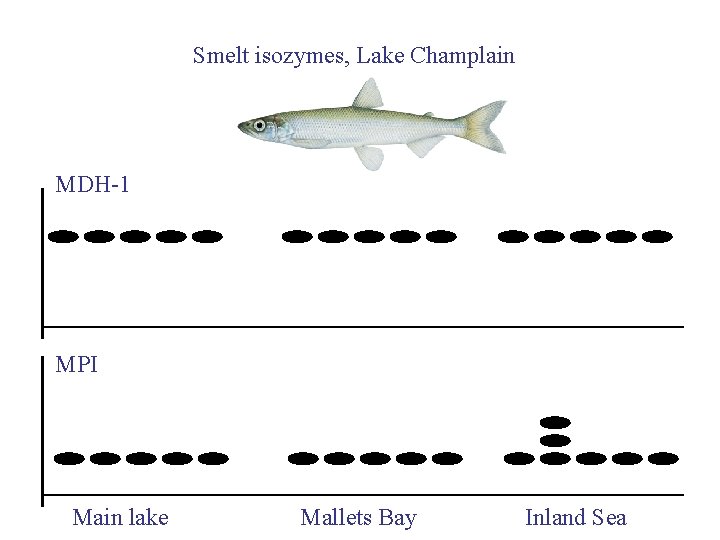
Smelt isozymes, Lake Champlain MDH-1 MPI Main lake Mallets Bay Inland Sea

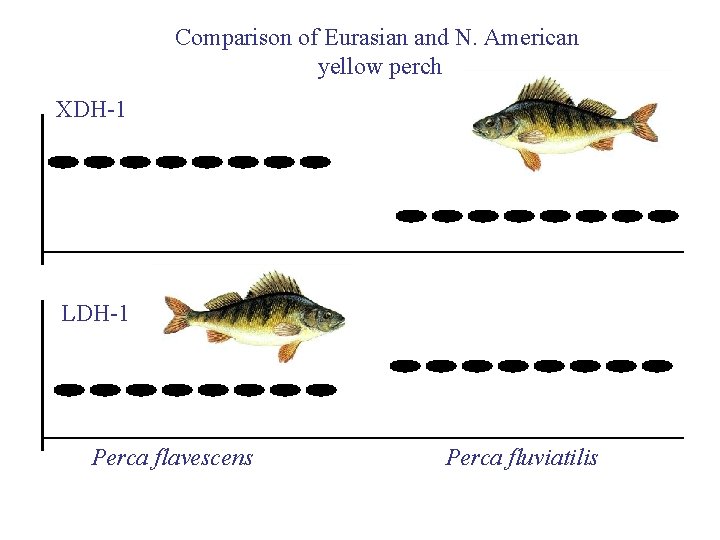
Comparison of Eurasian and N. American yellow perch XDH-1 LDH-1 Perca flavescens Perca fluviatilis

Cheetah (Acinonyx jubatus): O’Brien et al. 1983. The cheetah is depauperate in genetic variation - using protein electrophoresis - assumed to be result of small N, bottleneck, then inbreeding - highly vulnerable to disease outbreaks (50% mortality in one captive population)

Cheetah (Acinonyx jubatus): O’Brien et al. 1983. The cheetah is depauperate in genetic variation - using protein electrophoresis Species Drosophila Mus Homo sapiens Felis catus Cheetah # popns. N 43 >100 2 87 many >100 1 56 2 55 # loci 24 46 104 55 47 % poly. loci 43. 1 20. 5 31. 7 22 0. 0 av. H 0. 14 0. 088 0. 063 0. 076 0. 0

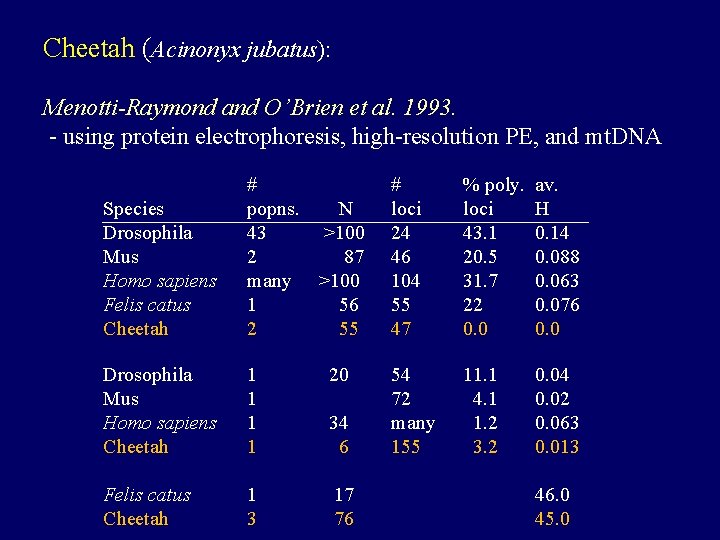
Cheetah (Acinonyx jubatus): Menotti-Raymond and O’Brien et al. 1993. - using protein electrophoresis, high-resolution PE, and mt. DNA Species Drosophila Mus Homo sapiens Felis catus Cheetah # popns. N 43 >100 2 87 many >100 1 56 2 55 # loci 24 46 104 55 47 % poly. loci 43. 1 20. 5 31. 7 22 0. 0 av. H 0. 14 0. 088 0. 063 0. 076 0. 0 Drosophila Mus Homo sapiens Cheetah 1 1 20 54 72 many 155 11. 1 4. 1 1. 2 3. 2 0. 04 0. 02 0. 063 0. 013 Felis catus Cheetah 1 3 17 76 34 6 46. 0 45. 0

Merola, 1994. A reassessment of homozygosity …. - carnivores tend to show levels of genetic variation (several have lower levels of H and P than cheetah) - measures of fluctuating asymmetry indicate cheetah is not suffering from low homozygosity or genetic stress - sperm deformities – do not affect fertility, may be normal in felids - low litter sizes – in captivity (high in wild) - susceptibility to disease – may be due to captive contact (in wild, cheetahs avoid conspecifics) Concluded that conservation is better directed at habitat

Spotted owls vs. barred owls (Haig et al. 2004, Cons. Biol. 18: 1347 -1357) Northern spotted owl – endangered Barred owl – rapid range expansion has led to overlap with spotted owl - potential competition - potential for hybridization mt. DNA and AFLP analysis: - species are distinct - no evidence of previous gene flow - hybrids occur with male spotted and female barred owls - hybrids can be identified until F 2 generation

e-DNA (environmental DNA) as a method for detection of rare/elusive species