Unit 3 The Central Dogma of Molecular Biology

Unit 3: The Central Dogma of Molecular Biology Chapter 12 - Processing of RNA Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
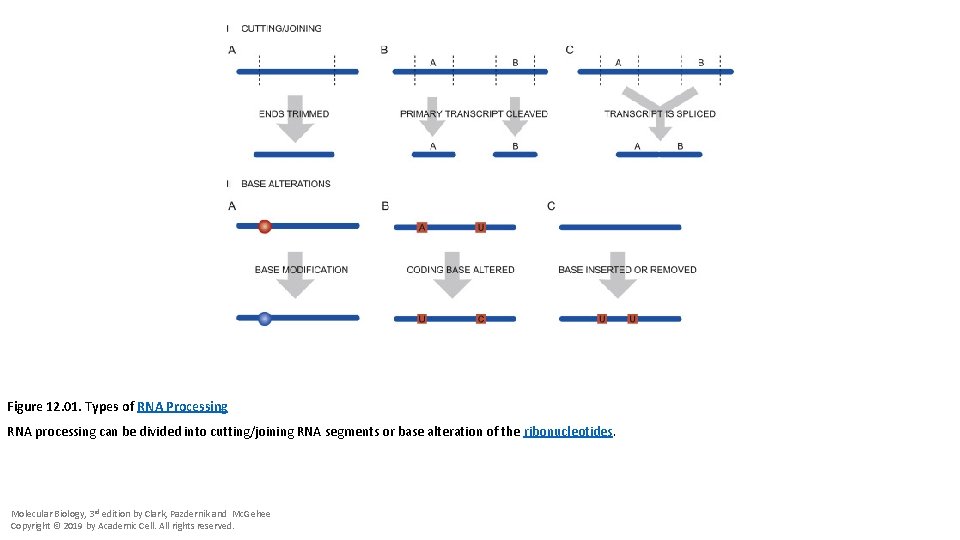
Figure 12. 01. Types of RNA Processing RNA processing can be divided into cutting/joining RNA segments or base alteration of the ribonucleotides. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
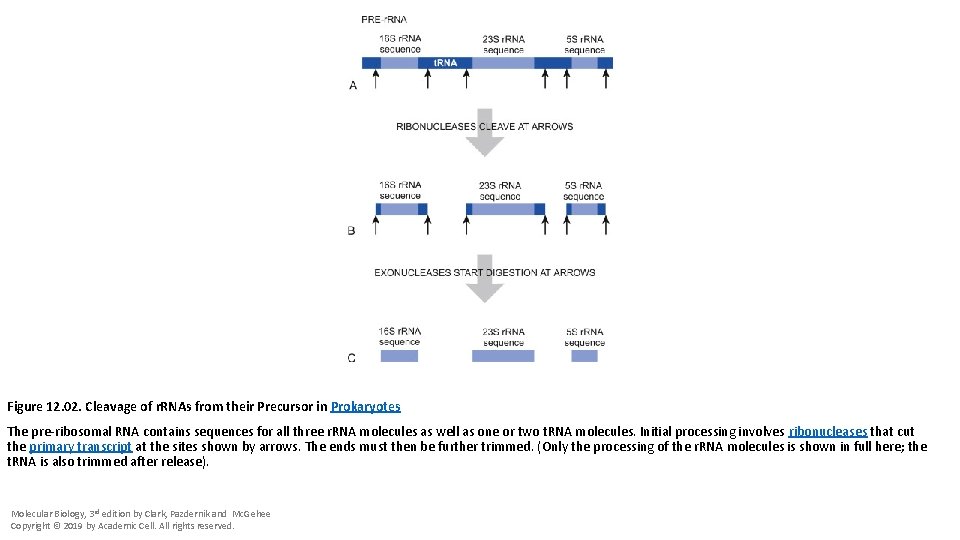
Figure 12. 02. Cleavage of r. RNAs from their Precursor in Prokaryotes The pre-ribosomal RNA contains sequences for all three r. RNA molecules as well as one or two t. RNA molecules. Initial processing involves ribonucleases that cut the primary transcript at the sites shown by arrows. The ends must then be further trimmed. (Only the processing of the r. RNA molecules is shown in full here; the t. RNA is also trimmed after release). Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
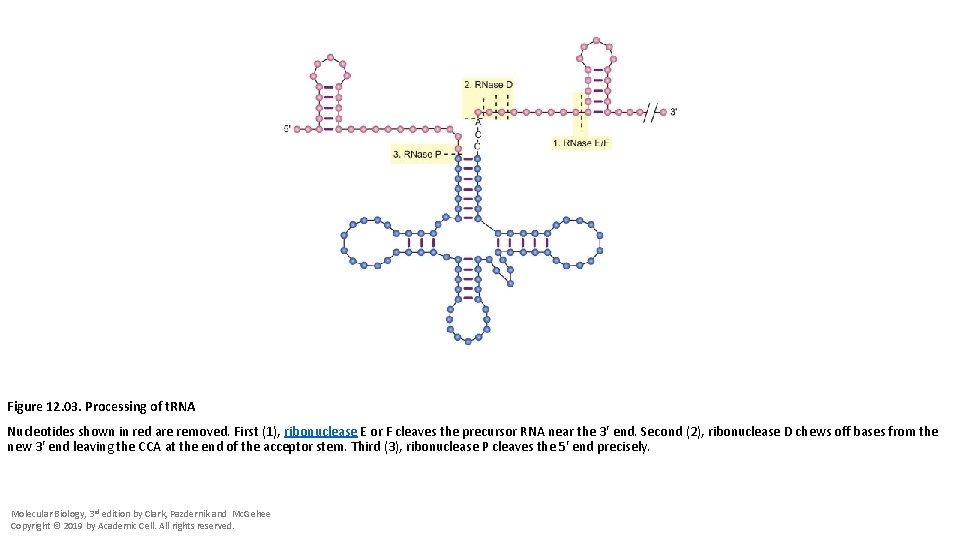
Figure 12. 03. Processing of t. RNA Nucleotides shown in red are removed. First (1), ribonuclease E or F cleaves the precursor RNA near the 3′ end. Second (2), ribonuclease D chews off bases from the new 3′ end leaving the CCA at the end of the acceptor stem. Third (3), ribonuclease P cleaves the 5′ end precisely. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
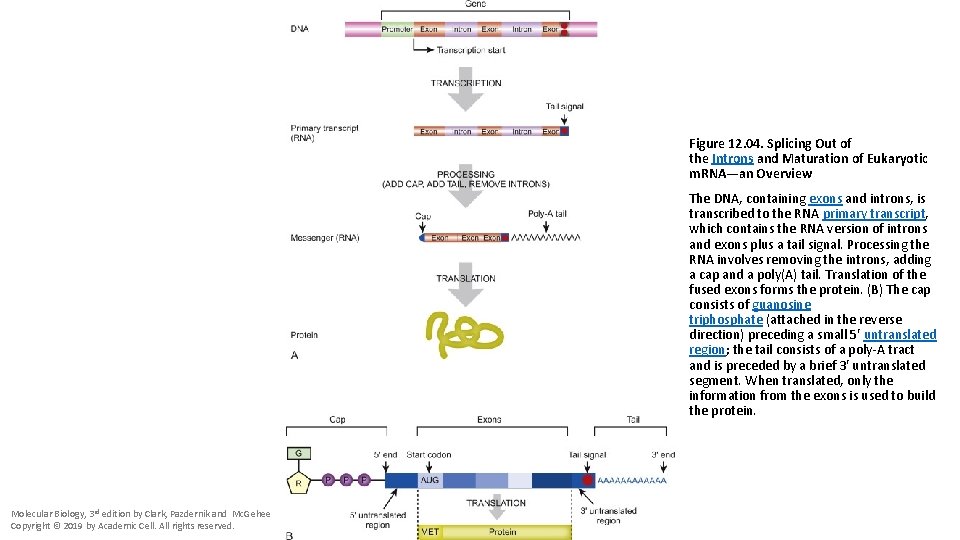
Figure 12. 04. Splicing Out of the Introns and Maturation of Eukaryotic m. RNA—an Overview The DNA, containing exons and introns, is transcribed to the RNA primary transcript, which contains the RNA version of introns and exons plus a tail signal. Processing the RNA involves removing the introns, adding a cap and a poly(A) tail. Translation of the fused exons forms the protein. (B) The cap consists of guanosine triphosphate (attached in the reverse direction) preceding a small 5′ untranslated region; the tail consists of a poly-A tract and is preceded by a brief 3′ untranslated segment. When translated, only the information from the exons is used to build the protein. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
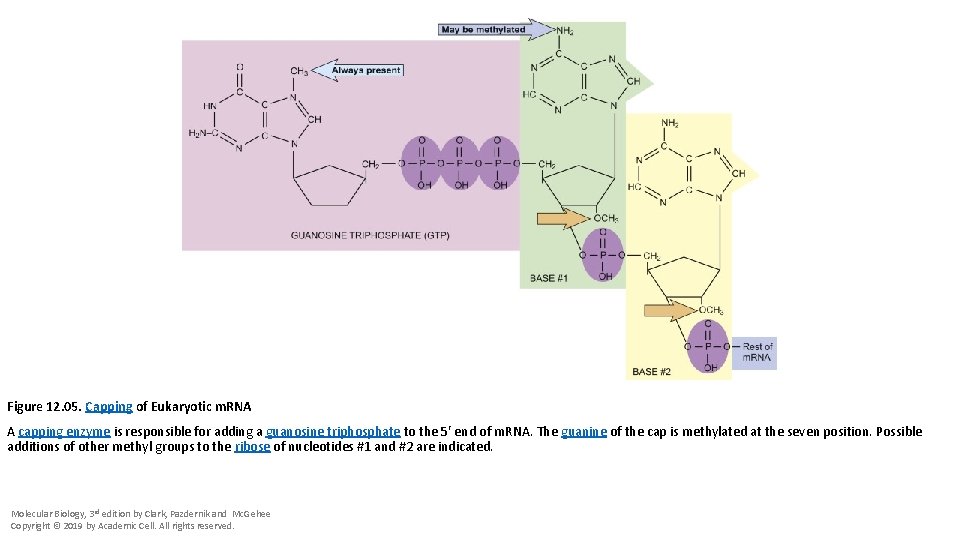
Figure 12. 05. Capping of Eukaryotic m. RNA A capping enzyme is responsible for adding a guanosine triphosphate to the 5′ end of m. RNA. The guanine of the cap is methylated at the seven position. Possible additions of other methyl groups to the ribose of nucleotides #1 and #2 are indicated. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
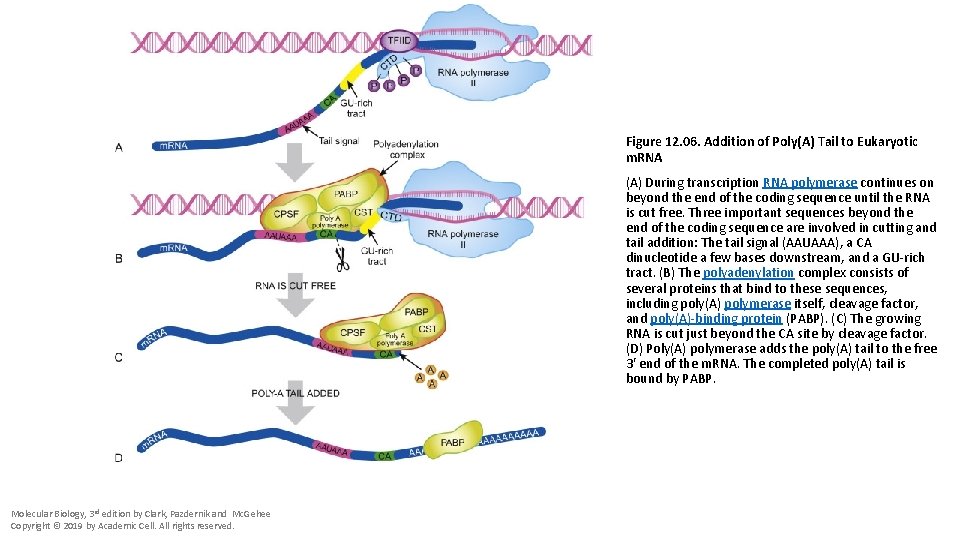
Figure 12. 06. Addition of Poly(A) Tail to Eukaryotic m. RNA (A) During transcription RNA polymerase continues on beyond the end of the coding sequence until the RNA is cut free. Three important sequences beyond the end of the coding sequence are involved in cutting and tail addition: The tail signal (AAUAAA), a CA dinucleotide a few bases downstream, and a GU-rich tract. (B) The polyadenylation complex consists of several proteins that bind to these sequences, including poly(A) polymerase itself, cleavage factor, and poly(A)-binding protein (PABP). (C) The growing RNA is cut just beyond the CA site by cleavage factor. (D) Poly(A) polymerase adds the poly(A) tail to the free 3′ end of the m. RNA. The completed poly(A) tail is bound by PABP. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
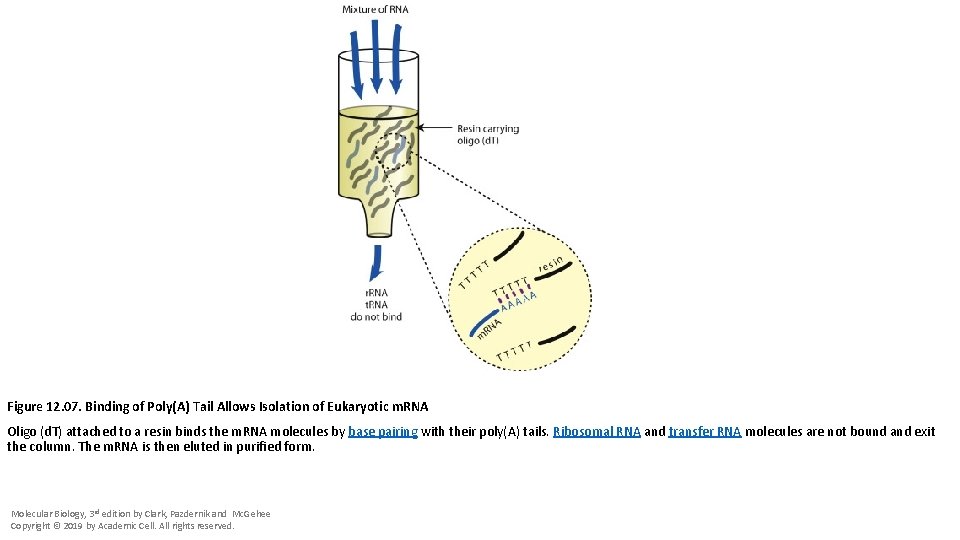
Figure 12. 07. Binding of Poly(A) Tail Allows Isolation of Eukaryotic m. RNA Oligo (d. T) attached to a resin binds the m. RNA molecules by base pairing with their poly(A) tails. Ribosomal RNA and transfer RNA molecules are not bound and exit the column. The m. RNA is then eluted in purified form. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
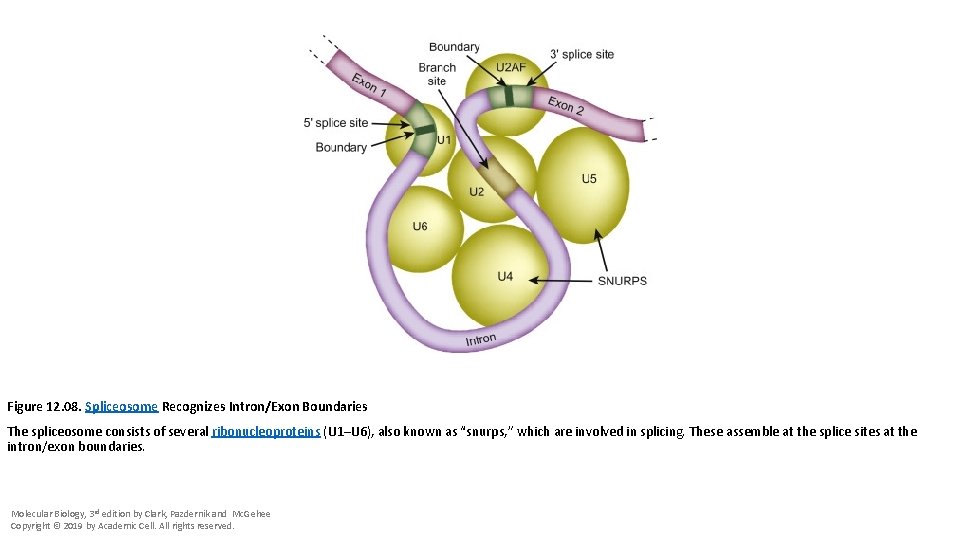
Figure 12. 08. Spliceosome Recognizes Intron/Exon Boundaries The spliceosome consists of several ribonucleoproteins (U 1–U 6), also known as “snurps, ” which are involved in splicing. These assemble at the splice sites at the intron/exon boundaries. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
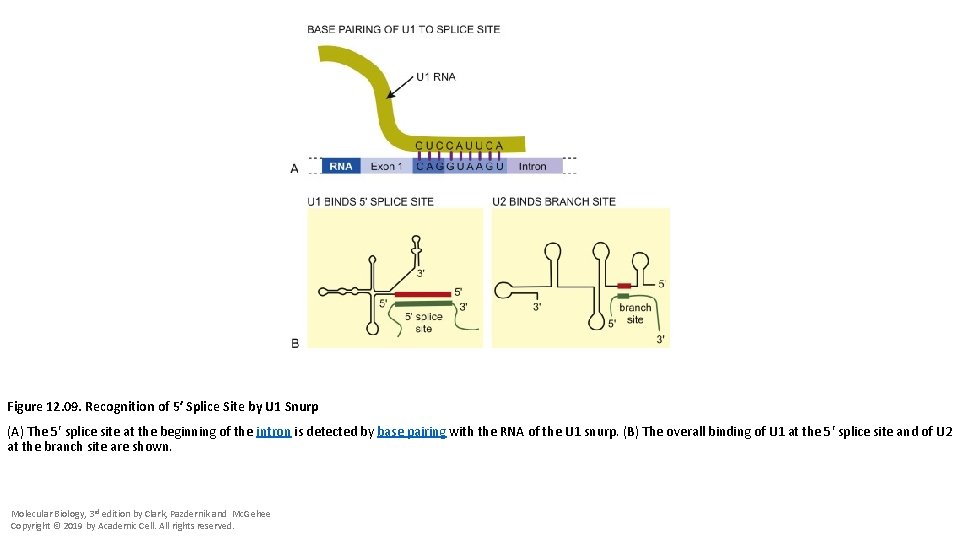
Figure 12. 09. Recognition of 5′ Splice Site by U 1 Snurp (A) The 5′ splice site at the beginning of the intron is detected by base pairing with the RNA of the U 1 snurp. (B) The overall binding of U 1 at the 5′ splice site and of U 2 at the branch site are shown. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
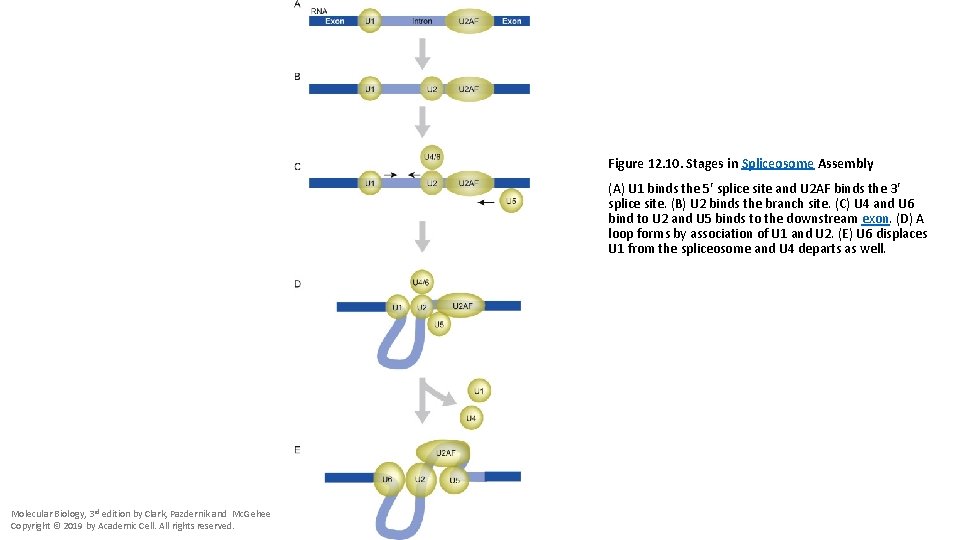
Figure 12. 10. Stages in Spliceosome Assembly (A) U 1 binds the 5′ splice site and U 2 AF binds the 3′ splice site. (B) U 2 binds the branch site. (C) U 4 and U 6 bind to U 2 and U 5 binds to the downstream exon. (D) A loop forms by association of U 1 and U 2. (E) U 6 displaces U 1 from the spliceosome and U 4 departs as well. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
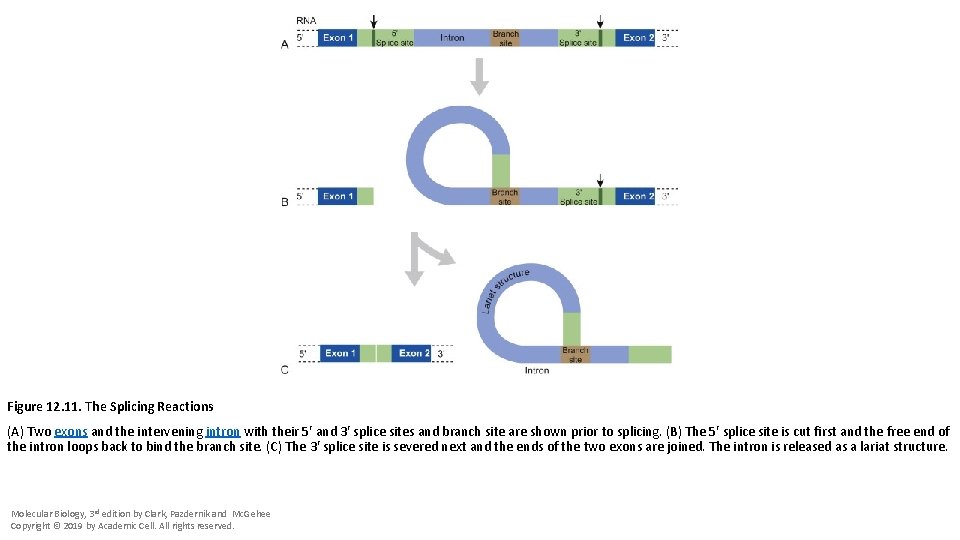
Figure 12. 11. The Splicing Reactions (A) Two exons and the intervening intron with their 5′ and 3′ splice sites and branch site are shown prior to splicing. (B) The 5′ splice site is cut first and the free end of the intron loops back to bind the branch site. (C) The 3′ splice site is severed next and the ends of the two exons are joined. The intron is released as a lariat structure. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
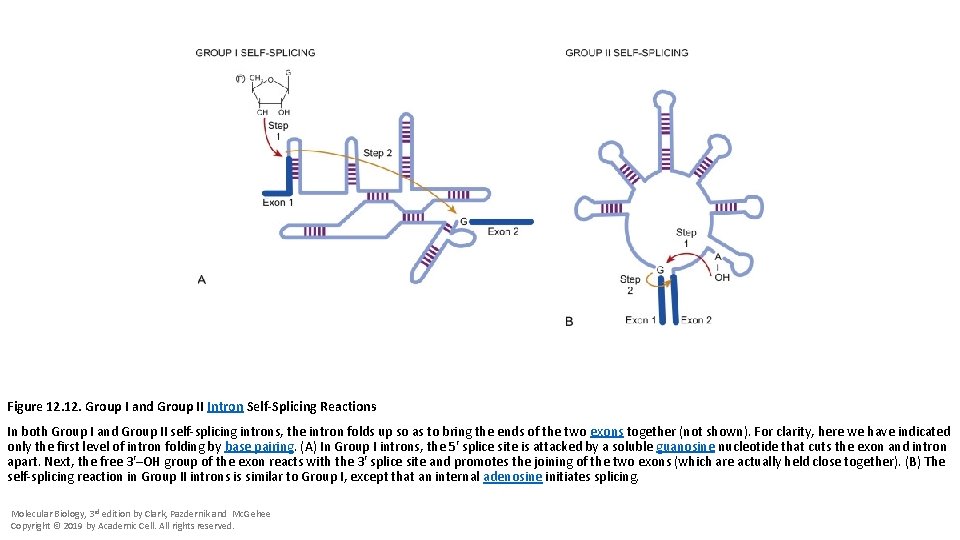
Figure 12. Group I and Group II Intron Self-Splicing Reactions In both Group I and Group II self-splicing introns, the intron folds up so as to bring the ends of the two exons together (not shown). For clarity, here we have indicated only the first level of intron folding by base pairing. (A) In Group I introns, the 5′ splice site is attacked by a soluble guanosine nucleotide that cuts the exon and intron apart. Next, the free 3′–OH group of the exon reacts with the 3′ splice site and promotes the joining of the two exons (which are actually held close together). (B) The self-splicing reaction in Group II introns is similar to Group I, except that an internal adenosine initiates splicing. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
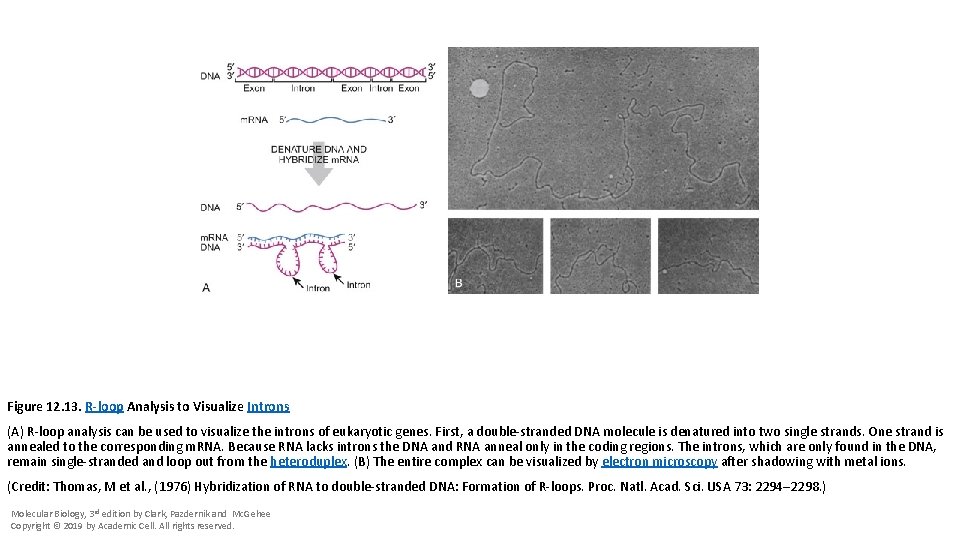
Figure 12. 13. R-loop Analysis to Visualize Introns (A) R-loop analysis can be used to visualize the introns of eukaryotic genes. First, a double-stranded DNA molecule is denatured into two single strands. One strand is annealed to the corresponding m. RNA. Because RNA lacks introns the DNA and RNA anneal only in the coding regions. The introns, which are only found in the DNA, remain single-stranded and loop out from the heteroduplex. (B) The entire complex can be visualized by electron microscopy after shadowing with metal ions. (Credit: Thomas, M et al. , (1976) Hybridization of RNA to double-stranded DNA: Formation of R-loops. Proc. Natl. Acad. Sci. USA 73: 2294– 2298. ) Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
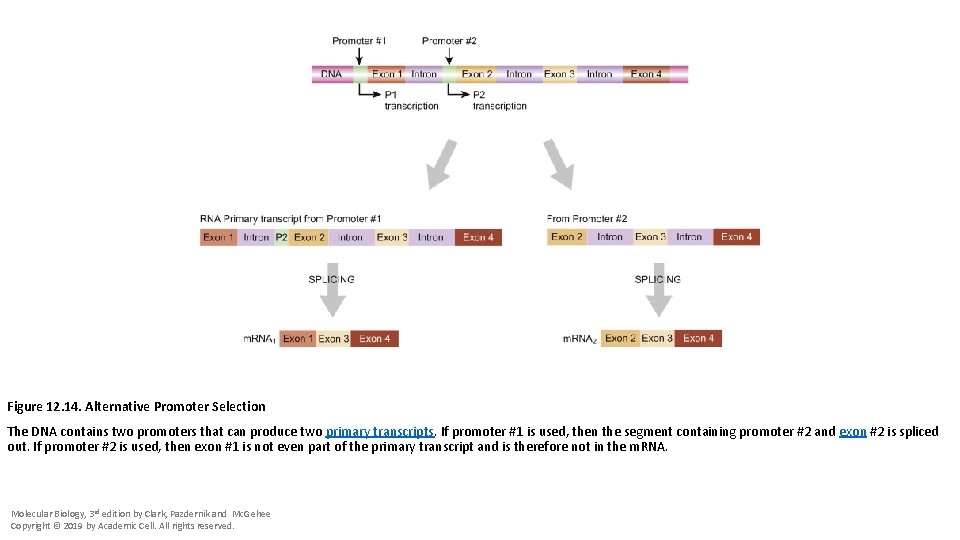
Figure 12. 14. Alternative Promoter Selection The DNA contains two promoters that can produce two primary transcripts. If promoter #1 is used, then the segment containing promoter #2 and exon #2 is spliced out. If promoter #2 is used, then exon #1 is not even part of the primary transcript and is therefore not in the m. RNA. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
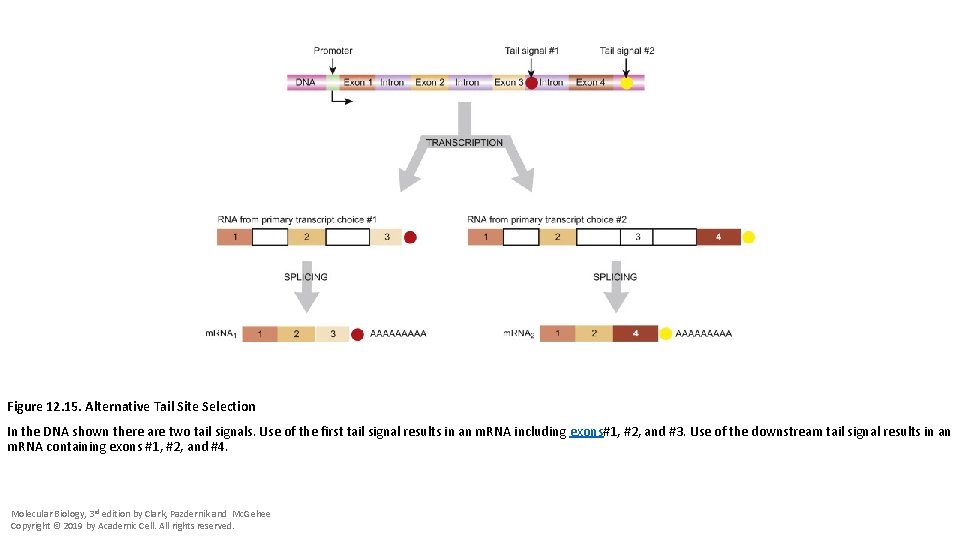
Figure 12. 15. Alternative Tail Site Selection In the DNA shown there are two tail signals. Use of the first tail signal results in an m. RNA including exons#1, #2, and #3. Use of the downstream tail signal results in an m. RNA containing exons #1, #2, and #4. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
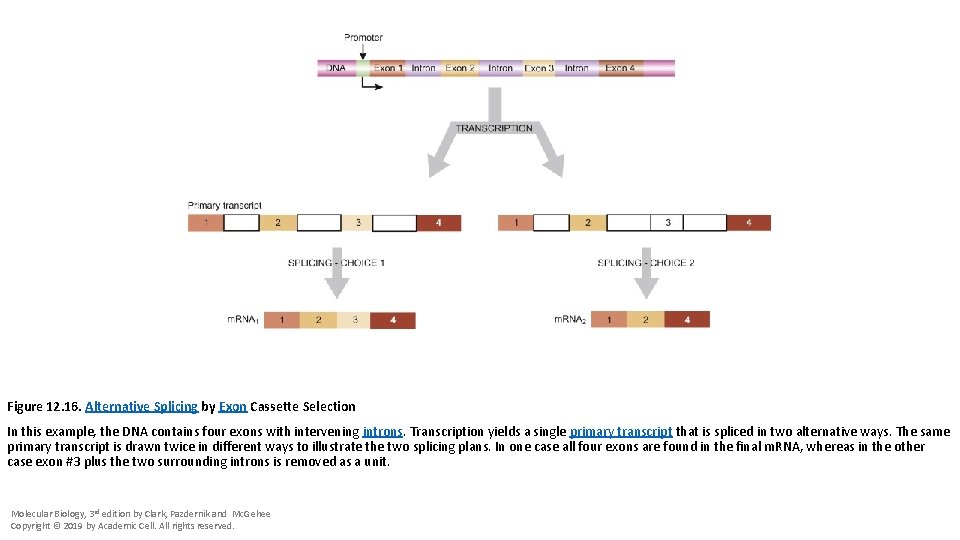
Figure 12. 16. Alternative Splicing by Exon Cassette Selection In this example, the DNA contains four exons with intervening introns. Transcription yields a single primary transcript that is spliced in two alternative ways. The same primary transcript is drawn twice in different ways to illustrate the two splicing plans. In one case all four exons are found in the final m. RNA, whereas in the other case exon #3 plus the two surrounding introns is removed as a unit. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
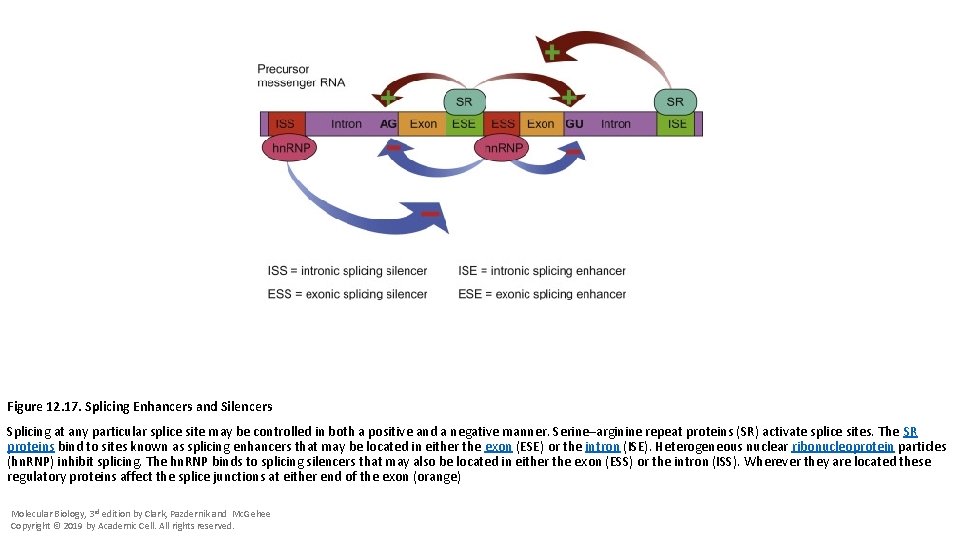
Figure 12. 17. Splicing Enhancers and Silencers Splicing at any particular splice site may be controlled in both a positive and a negative manner. Serine–arginine repeat proteins (SR) activate splice sites. The SR proteins bind to sites known as splicing enhancers that may be located in either the exon (ESE) or the intron (ISE). Heterogeneous nuclear ribonucleoprotein particles (hn. RNP) inhibit splicing. The hn. RNP binds to splicing silencers that may also be located in either the exon (ESS) or the intron (ISS). Wherever they are located these regulatory proteins affect the splice junctions at either end of the exon (orange) Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
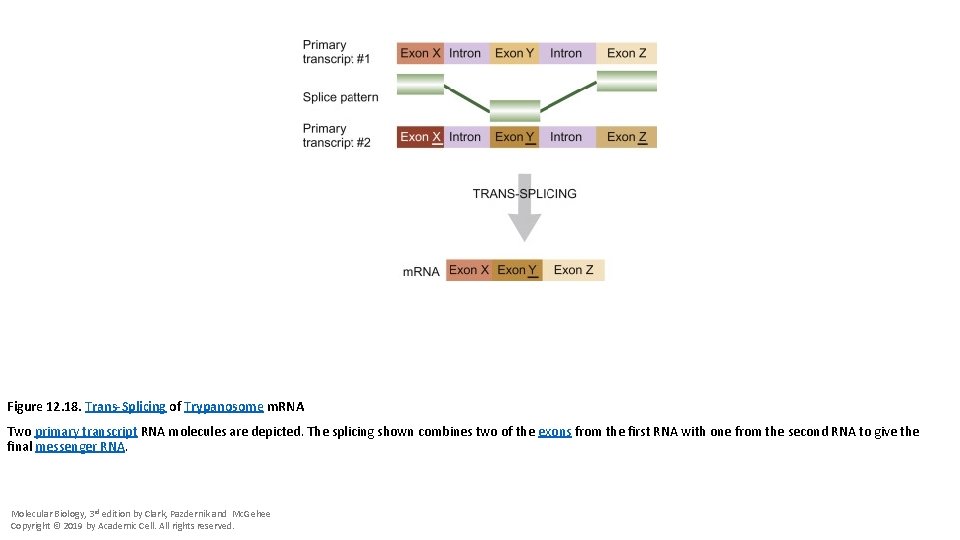
Figure 12. 18. Trans-Splicing of Trypanosome m. RNA Two primary transcript RNA molecules are depicted. The splicing shown combines two of the exons from the first RNA with one from the second RNA to give the final messenger RNA. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
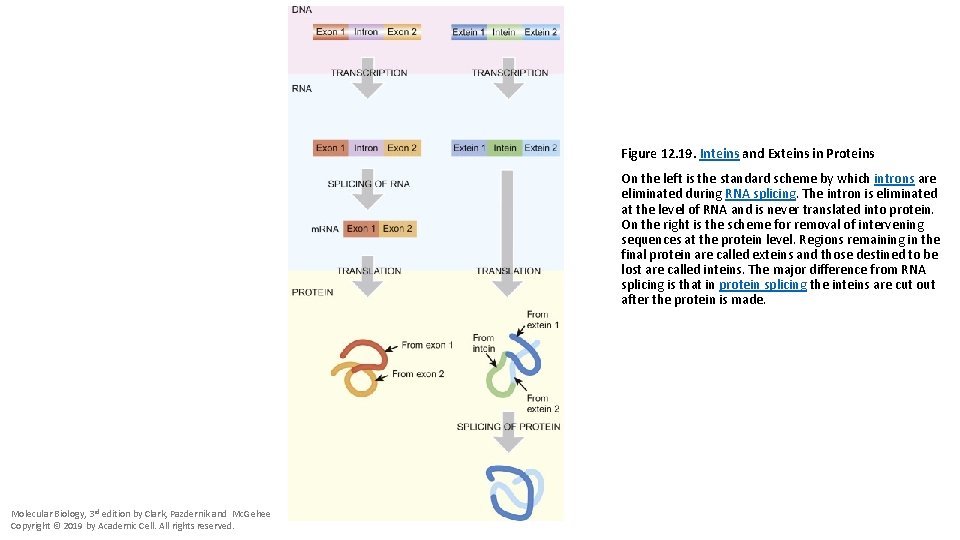
Figure 12. 19. Inteins and Exteins in Proteins On the left is the standard scheme by which introns are eliminated during RNA splicing. The intron is eliminated at the level of RNA and is never translated into protein. On the right is the scheme for removal of intervening sequences at the protein level. Regions remaining in the final protein are called exteins and those destined to be lost are called inteins. The major difference from RNA splicing is that in protein splicing the inteins are cut out after the protein is made. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
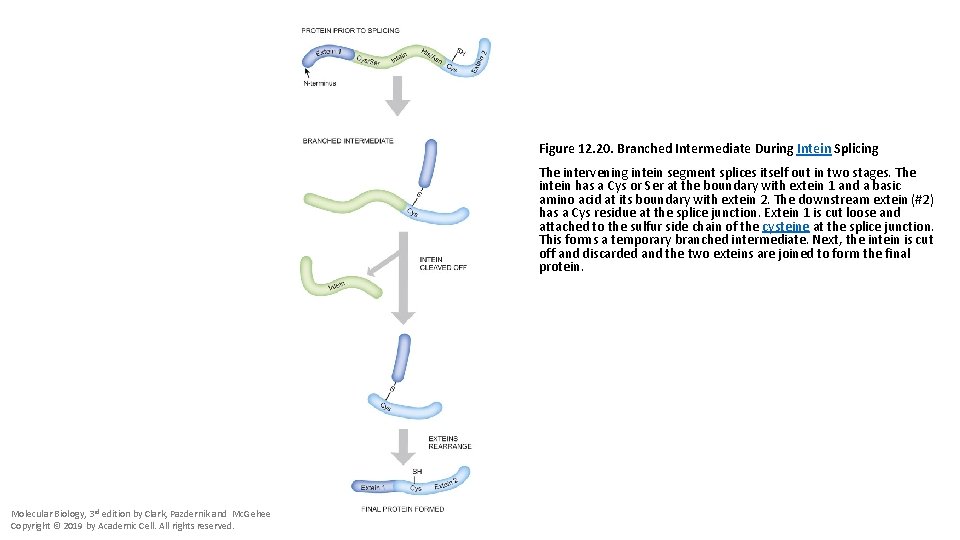
Figure 12. 20. Branched Intermediate During Intein Splicing The intervening intein segment splices itself out in two stages. The intein has a Cys or Ser at the boundary with extein 1 and a basic amino acid at its boundary with extein 2. The downstream extein (#2) has a Cys residue at the splice junction. Extein 1 is cut loose and attached to the sulfur side chain of the cysteine at the splice junction. This forms a temporary branched intermediate. Next, the intein is cut off and discarded and the two exteins are joined to form the final protein. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
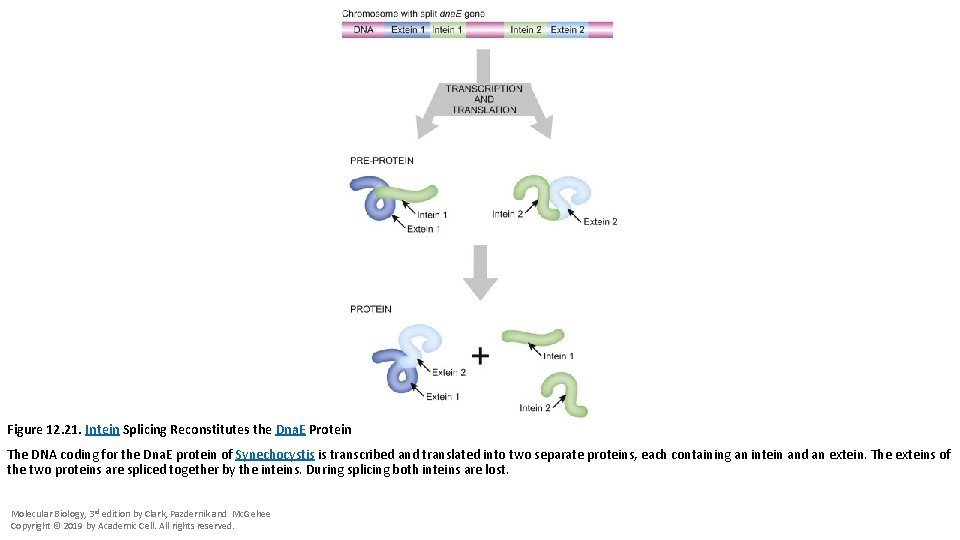
Figure 12. 21. Intein Splicing Reconstitutes the Dna. E Protein The DNA coding for the Dna. E protein of Synechocystis is transcribed and translated into two separate proteins, each containing an intein and an extein. The exteins of the two proteins are spliced together by the inteins. During splicing both inteins are lost. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
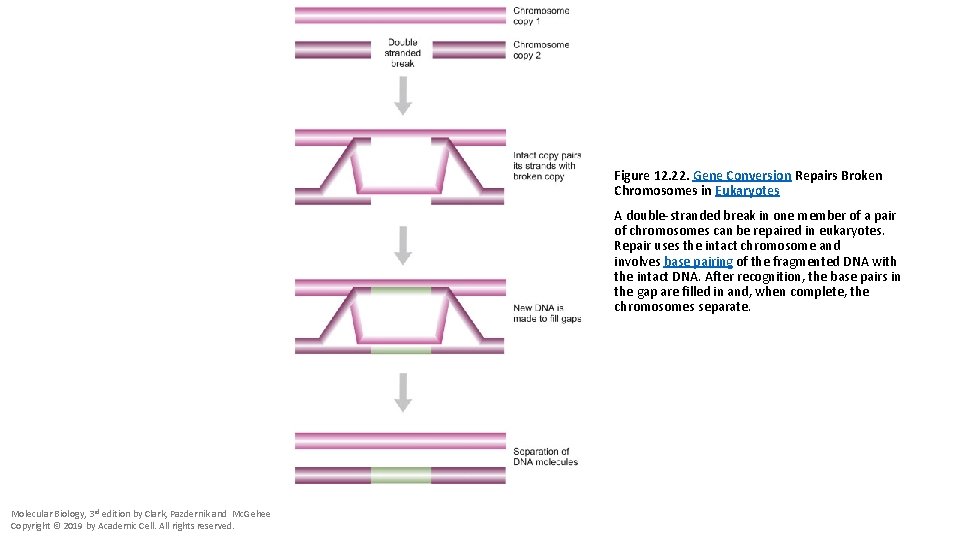
Figure 12. 22. Gene Conversion Repairs Broken Chromosomes in Eukaryotes A double-stranded break in one member of a pair of chromosomes can be repaired in eukaryotes. Repair uses the intact chromosome and involves base pairing of the fragmented DNA with the intact DNA. After recognition, the base pairs in the gap are filled in and, when complete, the chromosomes separate. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
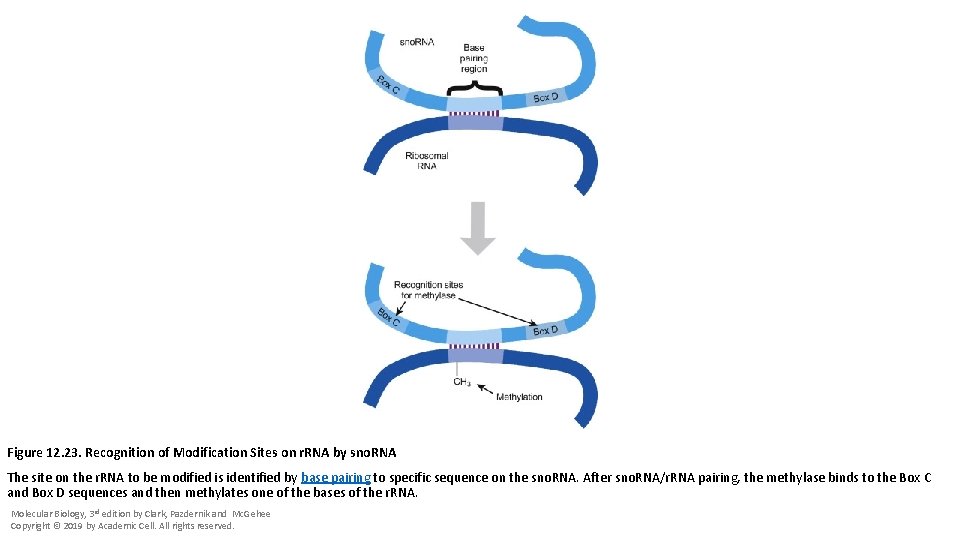
Figure 12. 23. Recognition of Modification Sites on r. RNA by sno. RNA The site on the r. RNA to be modified is identified by base pairing to specific sequence on the sno. RNA. After sno. RNA/r. RNA pairing, the methylase binds to the Box C and Box D sequences and then methylates one of the bases of the r. RNA. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
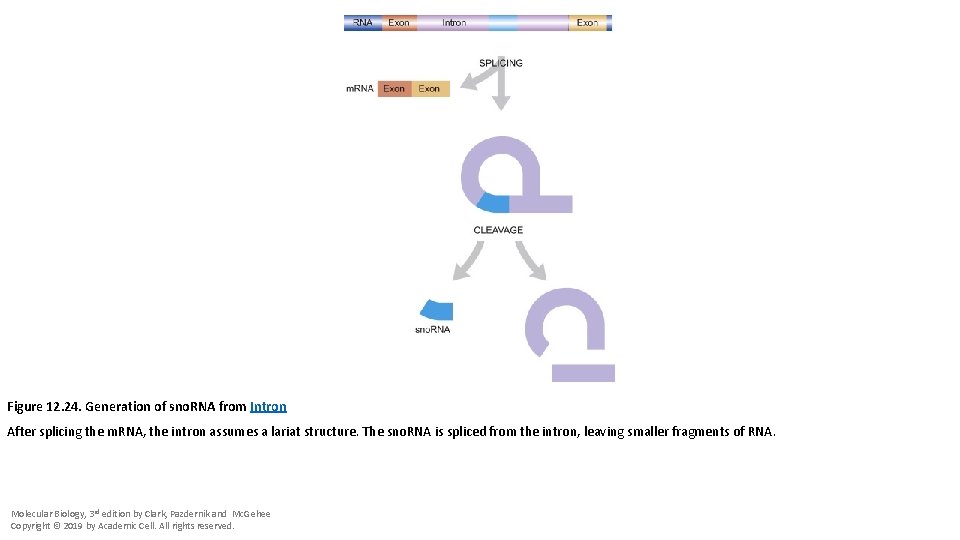
Figure 12. 24. Generation of sno. RNA from Intron After splicing the m. RNA, the intron assumes a lariat structure. The sno. RNA is spliced from the intron, leaving smaller fragments of RNA. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
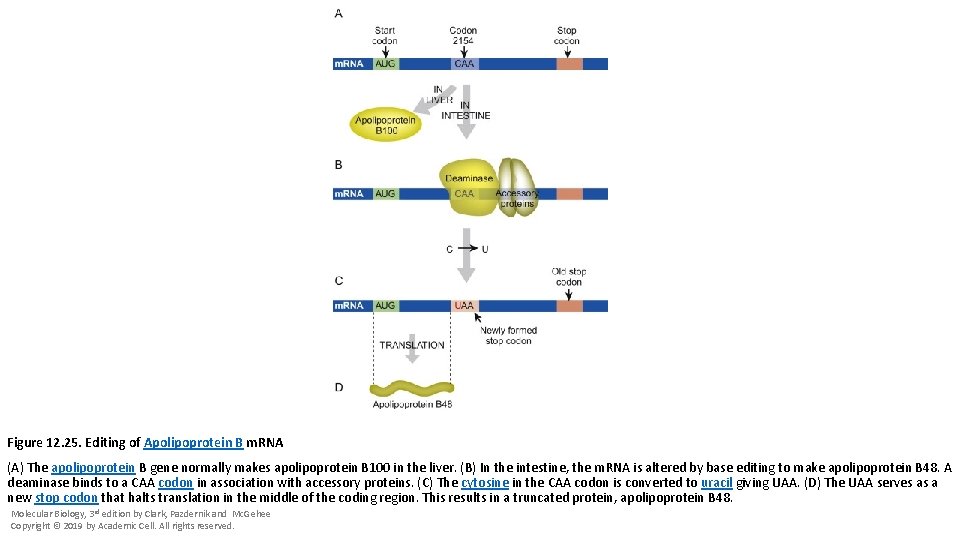
Figure 12. 25. Editing of Apolipoprotein B m. RNA (A) The apolipoprotein B gene normally makes apolipoprotein B 100 in the liver. (B) In the intestine, the m. RNA is altered by base editing to make apolipoprotein B 48. A deaminase binds to a CAA codon in association with accessory proteins. (C) The cytosine in the CAA codon is converted to uracil giving UAA. (D) The UAA serves as a new stop codon that halts translation in the middle of the coding region. This results in a truncated protein, apolipoprotein B 48. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
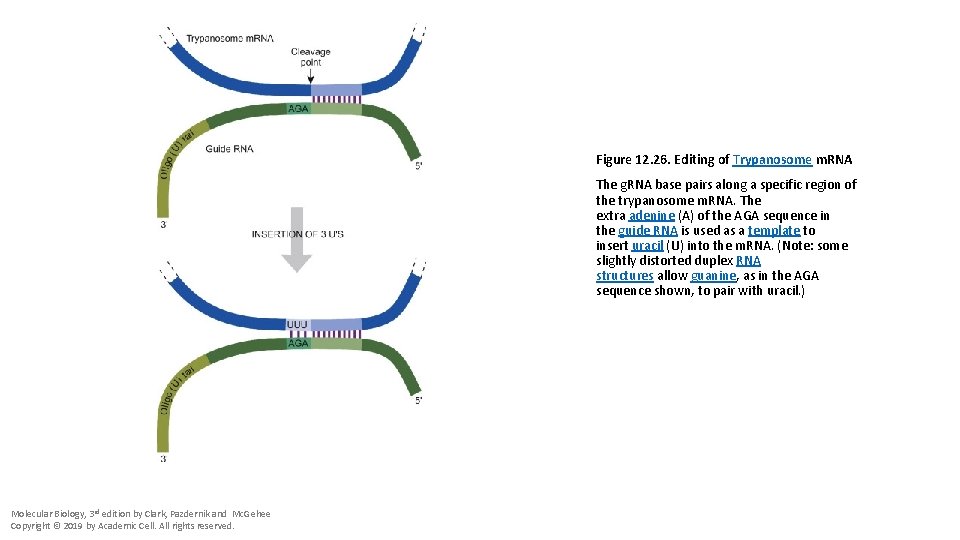
Figure 12. 26. Editing of Trypanosome m. RNA The g. RNA base pairs along a specific region of the trypanosome m. RNA. The extra adenine (A) of the AGA sequence in the guide RNA is used as a template to insert uracil (U) into the m. RNA. (Note: some slightly distorted duplex RNA structures allow guanine, as in the AGA sequence shown, to pair with uracil. ) Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
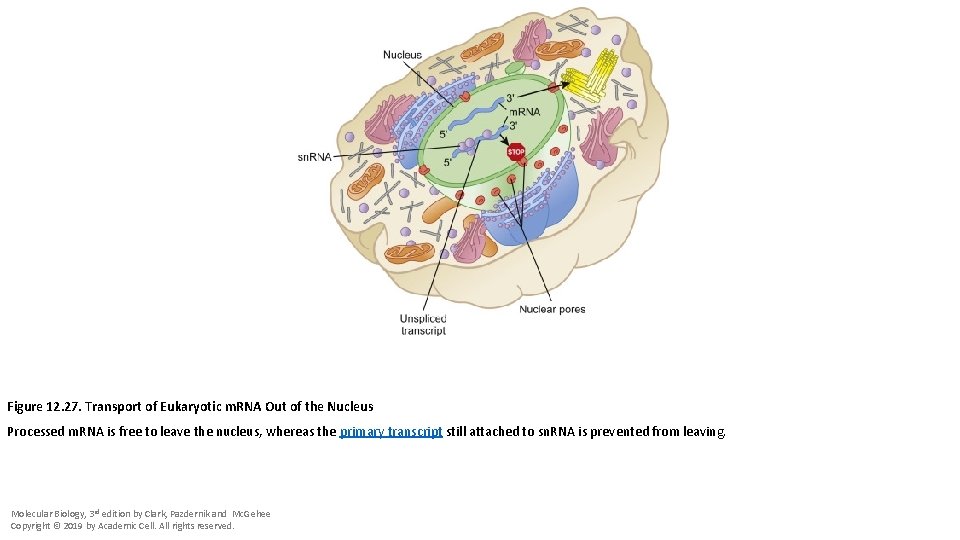
Figure 12. 27. Transport of Eukaryotic m. RNA Out of the Nucleus Processed m. RNA is free to leave the nucleus, whereas the primary transcript still attached to sn. RNA is prevented from leaving. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
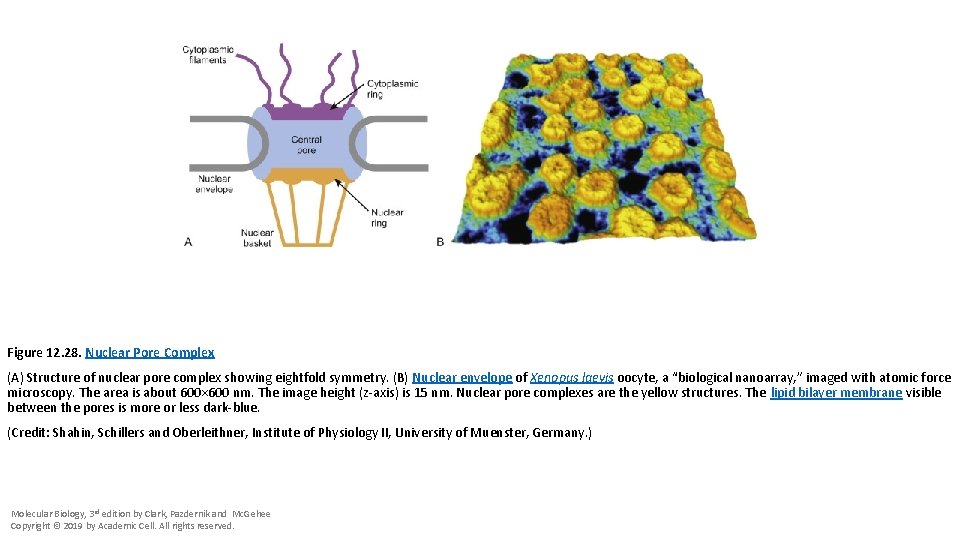
Figure 12. 28. Nuclear Pore Complex (A) Structure of nuclear pore complex showing eightfold symmetry. (B) Nuclear envelope of Xenopus laevis oocyte, a “biological nanoarray, ” imaged with atomic force microscopy. The area is about 600× 600 nm. The image height (z-axis) is 15 nm. Nuclear pore complexes are the yellow structures. The lipid bilayer membrane visible between the pores is more or less dark-blue. (Credit: Shahin, Schillers and Oberleithner, Institute of Physiology II, University of Muenster, Germany. ) Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
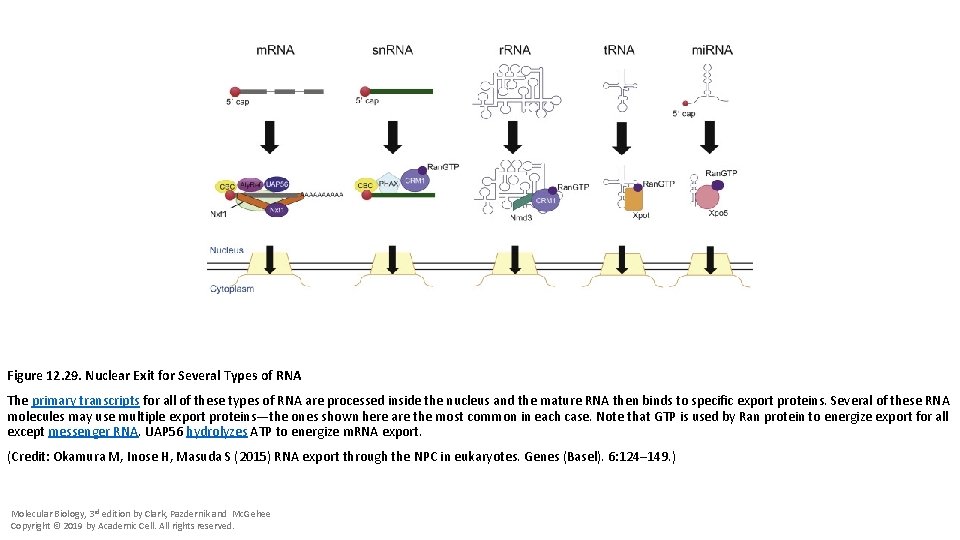
Figure 12. 29. Nuclear Exit for Several Types of RNA The primary transcripts for all of these types of RNA are processed inside the nucleus and the mature RNA then binds to specific export proteins. Several of these RNA molecules may use multiple export proteins—the ones shown here are the most common in each case. Note that GTP is used by Ran protein to energize export for all except messenger RNA. UAP 56 hydrolyzes ATP to energize m. RNA export. (Credit: Okamura M, Inose H, Masuda S (2015) RNA export through the NPC in eukaryotes. Genes (Basel). 6: 124– 149. ) Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
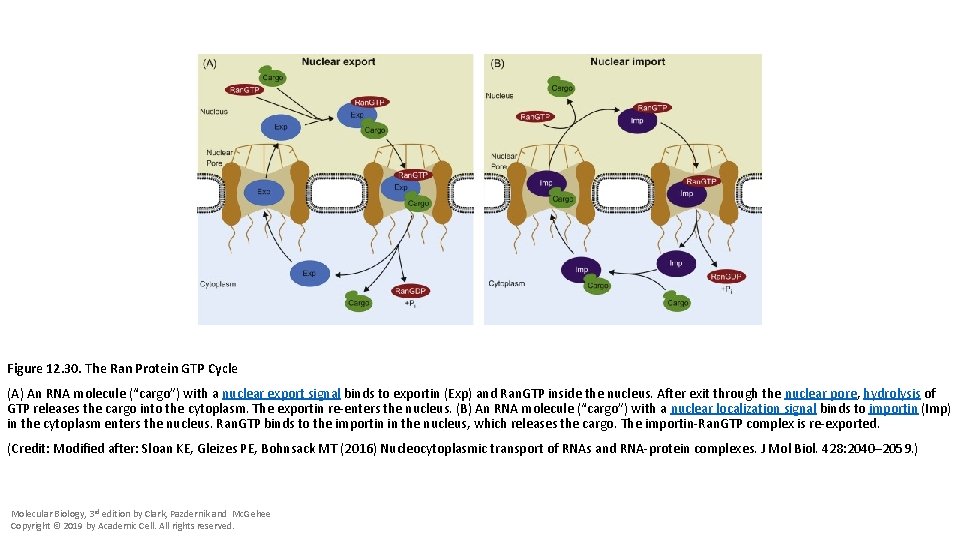
Figure 12. 30. The Ran Protein GTP Cycle (A) An RNA molecule (“cargo”) with a nuclear export signal binds to exportin (Exp) and Ran. GTP inside the nucleus. After exit through the nuclear pore, hydrolysis of GTP releases the cargo into the cytoplasm. The exportin re-enters the nucleus. (B) An RNA molecule (“cargo”) with a nuclear localization signal binds to importin (Imp) in the cytoplasm enters the nucleus. Ran. GTP binds to the importin in the nucleus, which releases the cargo. The importin-Ran. GTP complex is re-exported. (Credit: Modified after: Sloan KE, Gleizes PE, Bohnsack MT (2016) Nucleocytoplasmic transport of RNAs and RNA-protein complexes. J Mol Biol. 428: 2040– 2059. ) Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
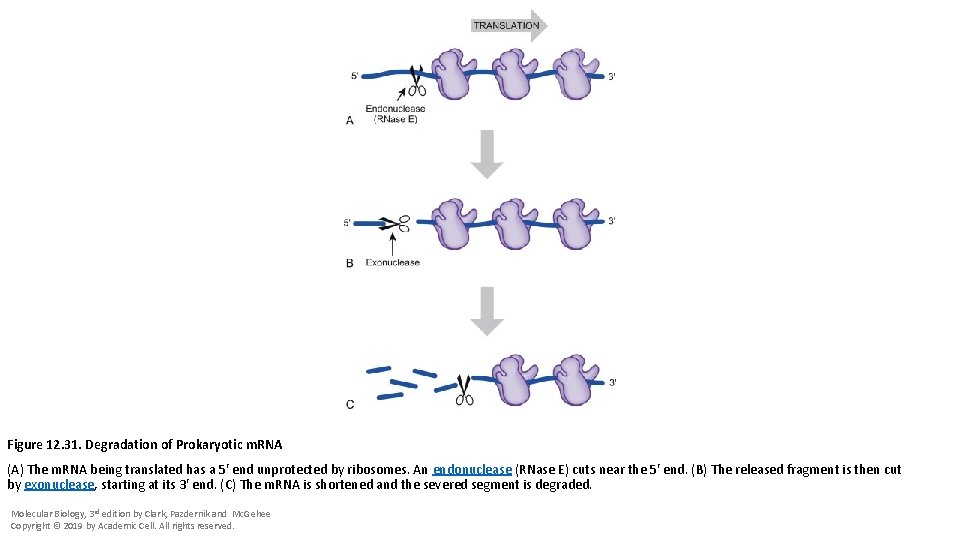
Figure 12. 31. Degradation of Prokaryotic m. RNA (A) The m. RNA being translated has a 5′ end unprotected by ribosomes. An endonuclease (RNase E) cuts near the 5′ end. (B) The released fragment is then cut by exonuclease, starting at its 3′ end. (C) The m. RNA is shortened and the severed segment is degraded. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
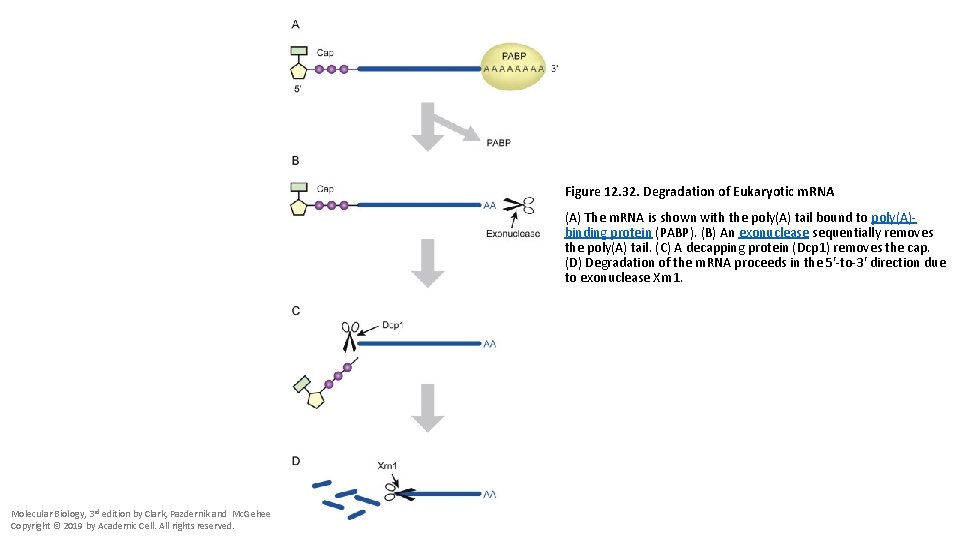
Figure 12. 32. Degradation of Eukaryotic m. RNA (A) The m. RNA is shown with the poly(A) tail bound to poly(A)binding protein (PABP). (B) An exonuclease sequentially removes the poly(A) tail. (C) A decapping protein (Dcp 1) removes the cap. (D) Degradation of the m. RNA proceeds in the 5′-to-3′ direction due to exonuclease Xrn 1. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
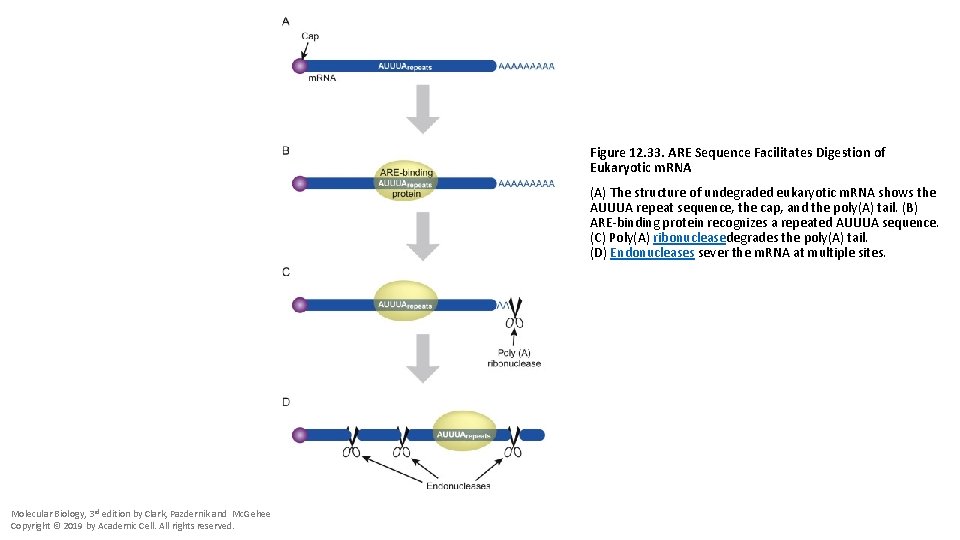
Figure 12. 33. ARE Sequence Facilitates Digestion of Eukaryotic m. RNA (A) The structure of undegraded eukaryotic m. RNA shows the AUUUA repeat sequence, the cap, and the poly(A) tail. (B) ARE-binding protein recognizes a repeated AUUUA sequence. (C) Poly(A) ribonucleasedegrades the poly(A) tail. (D) Endonucleases sever the m. RNA at multiple sites. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
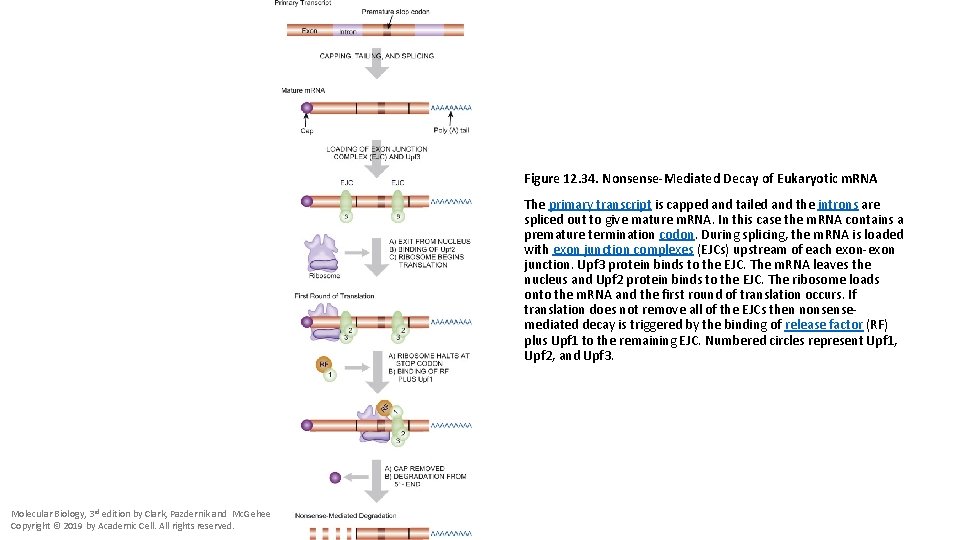
Figure 12. 34. Nonsense-Mediated Decay of Eukaryotic m. RNA The primary transcript is capped and tailed and the introns are spliced out to give mature m. RNA. In this case the m. RNA contains a premature termination codon. During splicing, the m. RNA is loaded with exon junction complexes (EJCs) upstream of each exon-exon junction. Upf 3 protein binds to the EJC. The m. RNA leaves the nucleus and Upf 2 protein binds to the EJC. The ribosome loads onto the m. RNA and the first round of translation occurs. If translation does not remove all of the EJCs then nonsensemediated decay is triggered by the binding of release factor (RF) plus Upf 1 to the remaining EJC. Numbered circles represent Upf 1, Upf 2, and Upf 3. Molecular Biology, 3 rd edition by Clark, Pazdernik and Mc. Gehee Copyright © 2019 by Academic Cell. All rights reserved.
- Slides: 35