Uni Prot Eric Jain Swiss Institute of Bioinformatics

Uni. Prot Eric Jain Swiss Institute of Bioinformatics, Geneva W 3 C Workshop on Semantic Web for Life Sciences, October 2004

What is it?
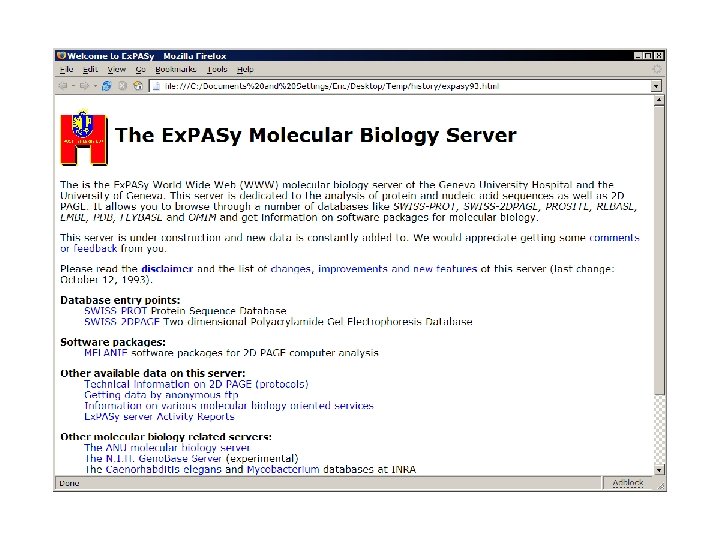


ID AC DT DT DT DE GN OS OC OC OX RN RP RC RX RA RT RT RL CC CC CC DR DR DR KW KW FT FT FT SQ // ATPB_CANFA STANDARD; PRT; 19 AA. P 99504; 15 -JUL-1998 (Rel. 36, Created) 15 -JUL-1998 (Rel. 36, Last sequence update) 05 -JUL-2004 (Rel. 44, Last annotation update) ATP synthase beta chain, mitochondrial (EC 3. 6. 3. 14) (Fragment). Name=ATP 5 B; Canis familiaris (Dog). Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi; Mammalia; Eutheria; Carnivora; Fissipedia; Canidae; Canis. NCBI_Tax. ID=9615; [1] SEQUENCE. TISSUE=Heart; MEDLINE=98163340; Pub. Med=9504812; Dunn M. J. , Corbett J. M. , Wheeler C. H. ; "HSC-2 DPAGE and the two-dimensional gel electrophoresis database of dog heart proteins. "; Electrophoresis 18: 2795 -2802(1997). -!- FUNCTION: Produces ATP from ADP in the presence of a proton gradient across the membrane. The beta chain is the catalytic subunit. -!- CATALYTIC ACTIVITY: ATP + H(2)O + H(+)(In) = ADP + phosphate + H(+)(Out). -!- SUBUNIT: F-type ATPases have 2 components, CF(1) - the catalytic core - and CF(0) - the membrane proton channel. CF(1) has five subunits: alpha(3), beta(3), gamma(1), delta(1), epsilon(1). CF(0) has three main subunits: a, b and c. -!- SUBCELLULAR LOCATION: Mitochondrial. -!- SIMILARITY: Belongs to the ATPase alpha/beta chains family. HSC-2 DPAGE; P 99504; DOG. Inter. Pro; IPR 000194; ATPase_a/bcentre. PROSITE; PS 00152; ATPASE_ALPHA_BETA; PARTIAL. ATP synthesis; ATP-binding; CF(1); Direct protein sequencing; Hydrogen ion transport; Hydrolase; Mitochondrion. UNSURE 8 8 UNSURE 17 19 NON_TER 19 19 SEQUENCE 19 AA; 1871 MW; BB 9 C 163 FDC 60 BB 42 CRC 64; ATQTSPSPKG AAAXXXRVV


What have we done so far?
![[DIR] Parent Directory 19 -Jul-2004 13: 02 [ [ [ ] ] ] 11 [DIR] Parent Directory 19 -Jul-2004 13: 02 [ [ [ ] ] ] 11](http://slidetodoc.com/presentation_image_h/17d0d5db270d2885f36642eb71ca4b86/image-8.jpg)
[DIR] Parent Directory 19 -Jul-2004 13: 02 [ [ [ ] ] ] 11 -Oct-2004 13 -Sep-2004 19 -Oct-2004 11 -Oct-2004 [ [ [ cellular-components. rdf databases. rdf. gz datasets. rdf enzymes. rdf. gz go. rdf. gz - 19: 15 11: 34 16: 32 19: 15 5 k 45 k 6 k 4 k 309 k 839 k ] intact. rdf. gz ] keywords. rdf. gz ] ontology. owl 11 -Oct-2004 19: 15 19 -Oct-2004 18: 27 636 k 96 k 77 k [ ] taxonomy. rdf. gz 11 -Oct-2004 19: 15 4. 0 M [ [ [ ] uniparc. rdf. gz ] uniprot. rdf. gz ] uniref. rdf. gz 13 -Oct-2004 10: 54 11 -Oct-2004 19: 39 01 -Oct-2004 12: 56 762 M 768 M 52. 2 M
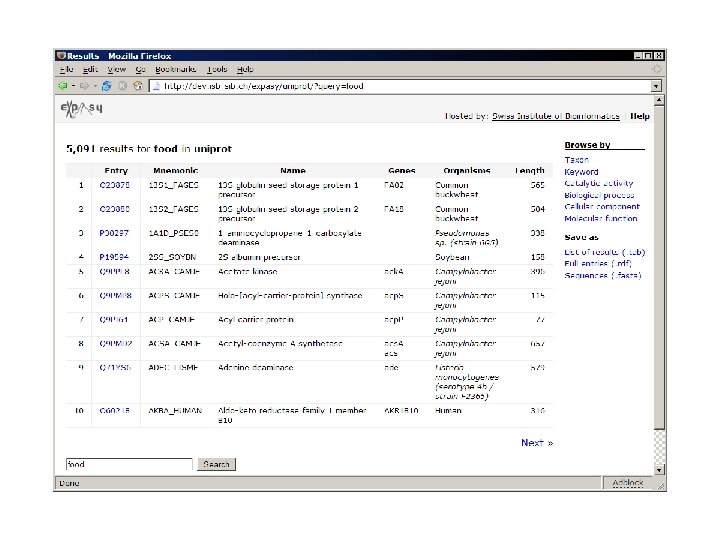
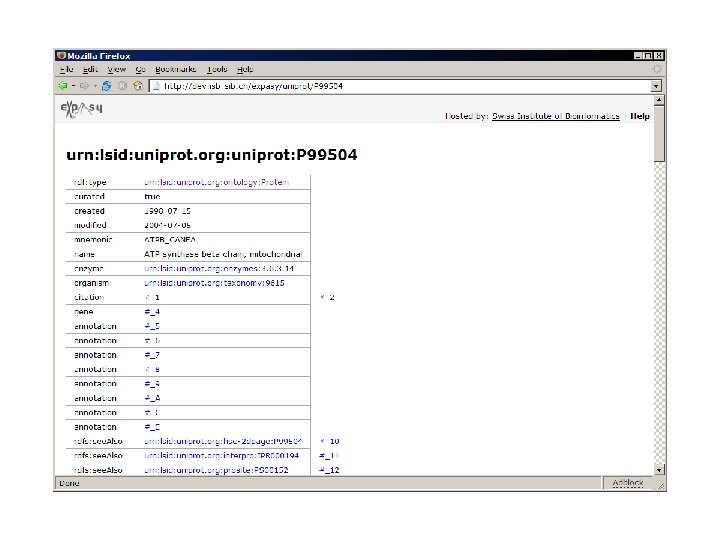
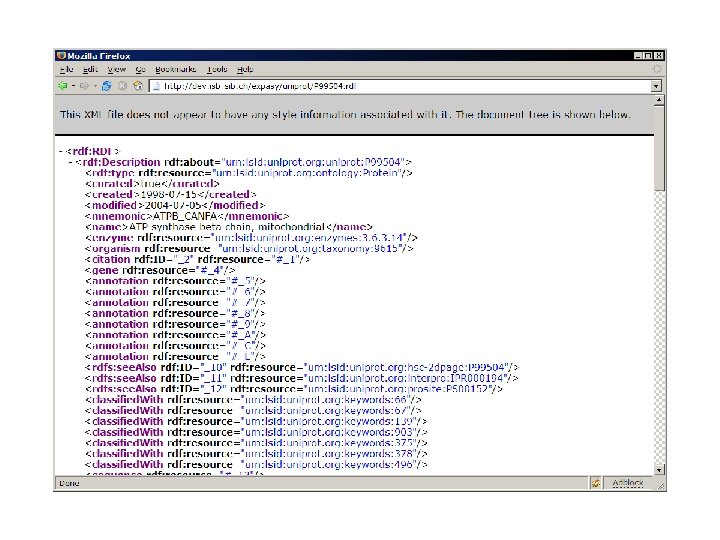
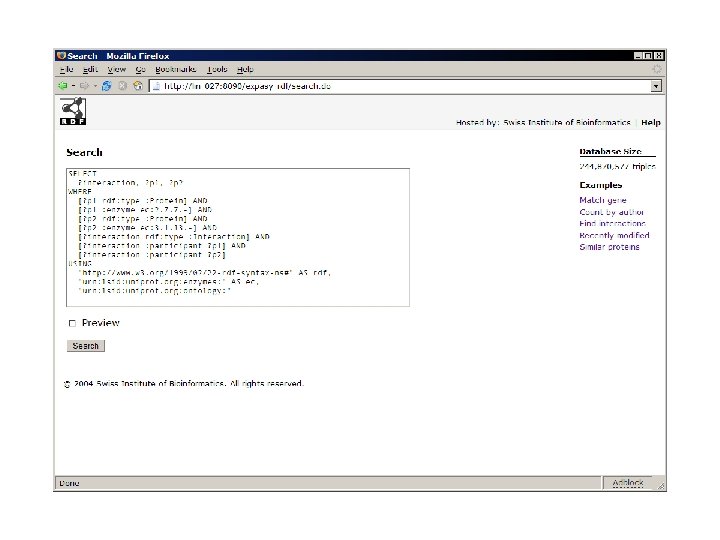

use Expasy: : RDF; my $parser = Expasy: : RDF: : Parser->new('P 12345. rdf'); while (my $protein = $parser->next) { my $id = $protein->id; my $mass = $protein->sequence->mass; print "Mass of $id is $mass. n"; print $_->type, ': ', $_->comment, "n"; foreach ($protein->annotation) } $parser->close;

Issues


XML Syntax <? xml version="1. 0" encoding="UTF-8"? > <rdf: RDF xmlns: rdfs="http: //www. w 3. org/2000/01/rdf-schema#" xmlns: rdf="http: //www. w 3. org/1999/02/22 -rdf-syntax-ns#" xmlns="urn: lsid: uniprot. org: ontology: " xmlns: owl="http: //www. w 3. org/2002/07/owl#" > <rdf: Description rdf: about="urn: lsid: uniprot. org: taxonomy: 9606"> <rdf: type rdf: resource="urn: lsid: uniprot. org: ontology: Taxon"/> <mnemonic>HUMAN</mnemonic> <scientific. Name>Homo sapiens</scientific. Name> <common. Name>Human</common. Name> <rdfs: sub. Class. Of rdf: resource="urn: lsid: uniprot. org: taxonomy: 9605"/> </rdf: Description> </rdf: RDF>

Triples, Quads and Quints ● What is the source of a triple? ● Compact reification.

Web Services ● Overkill for providing programmatic access to resources. ● Often impractical for performance reasons.

Life Science Identifiers ● Need special resolver. ● Resolution tied to retrieval. ● Explicit version numbers. ● Not widely used.

Embedded References uniprot. rdf <rdf: Description rdf: about="#_2 F 9 A"> <rdf: type rdf: resource="urn: lsid: uniprot. org: ontology: Caution_Annotation"/> <rdfs: comment>In mouse, 5 genes homologous to human CD 209/DC-SIGN and CD 209 L/DC-SIGNR have been identified. Mouse CD 209 A product was named DCSIGN by {citation 1} because of its similar expression pattern and chromosomal location in juxtaposition to CD 23, but despite of the low sequence similarity. </rdfs: comment> <citation rdf: resource="#_2 F 8 A"/> </rdf: Description> cyc. rdf <owl: Class rdf: ID="Antigen"> <rdfs: comment>The collection of substances that can stimulate immune response. For example, bacteria [#$Bacterium], #$Viruses, proteins [#$Protein. Molecule] can serve as #$Antigens. </rdfs: comment> </owl: Class>

Summary

People will adopt the technology if it provides immediate benefits and is simple to use.

Credits

<? xml version="1. 0" encoding="UTF-8"? > <rdf: RDF xmlns: rdf="http: //www. w 3. org/1999/02/22 -rdf-syntax-ns#" xmlns: foaf="http: //xmlns. com/foaf/0. 1/" > <foaf: Project> <foaf: name>Uni. Prot</foaf: name> <foaf: homepage rdf: resource="http: //uniprot. org/"/> <foaf: funded. By> <foaf: Organization> <foaf: name>National Institutes of Health</foaf: name> <foaf: homepage rdf: resource="http: //www. nih. gov/"/> </foaf: Organization> </foaf: funded. By> </foaf: Project> <foaf: Organization> <foaf: name>Swiss Institute of Bioinformatics</foaf: name> <foaf: nick>SIB</foaf: nick> <foaf: homepage rdf: resource="http: //www. isb-sib. ch/"/> </foaf: Organization> <foaf: name>European Bioinformatics Institute</foaf: name> <foaf: nick>EBI</foaf: nick> <foaf: homepage rdf: resource="http: //www. ebi. ac. uk/"/> </foaf: Organization> <foaf: name>Georgetown University</foaf: name> <foaf: homepage rdf: resource="http: //www. georgetown. edu/"/> </foaf: Organization>. . .
- Slides: 24