UM BDSI June 21 2019 Early years Born

UM BDSI June 21, 2019

Early years • Born May 16, 1956 in Eugene, Oregon • Father George: printer, small business owner • Mother Chari: homemaker, volunteer • Parents active in community (Y, Active Club Rotary, Blood Bank, Republican Party, …) • Siblings: Richard (1958 -88), Barb (1961 -present) ,

College • U Oregon Honors College, Math, 1973 -77 • Broad liberal arts education: lots of mostly pure math, physical sciences, but also History of Ideas, Europe since 1789, Novels of Kurt Vonnegut, … • Plan A: Math major obvious choice, but by junior year of college: what will I do after I finish college? ? • Plan B: Law school at Stanford, UC Berkeley, Oregon? • Two critical pieces of advice (10 minute conversations): • apply for Fulbright scholarship (Ed Diller) • study quantitative biology (Bill Bradshaw) Mike: I haven't had biology since 7 th grade. Bill: Who cares?

Postgraduate year in Germany 1977 -78 • Goethe Institute in Blaubeuren • Albert-Ludwigs-Universitӓt Freiburg • Analysis, topology, American history, … • No real evaluations, lots of travel • Pan C: decided who care's? • GREs in Heidelberg, October 1977 • Read Keeton: "Biological Science" and discovered genetics (and ecology, epidemiology, …) • Considered UCLA and U Washington; chose UCLA

Graduate school at UCLA 1978 -1983 • Biomathematics: see also R Little, J Taylor, T Johnson • Advisors: Ken Lange and Anne Spence • Academic siblings: Neil Risch, Dan Weeks • Academic grandfather: Robert Elston • Key event: Henry Tuckwell visited and started a journal club, said I should give the first talk; yikes!

Why statistical genetics? • First term at UCLA took computational statistics with Bob Jennrich; needed project • Ken had just gotten first NIH grant; offered to pay me to write first version of MENDEL • Good project, added to $325/month NIH traineeship • Liked Ken, liked project • Chose statistics to do genetics with Ken and because I liked probability and computing

Summer internship at Cedars-Sinai • Ken went to Boston on sabbatical • Anne Spence helped find me a summer job with Richard Gatti at Cedars-Sinai hospital with • Developed method to classify HLA-D genotypes (now recognized as multigene family!), wrote my firstauthor paper (with lots of help from Dick) • Played (Colossal Cave) Adventure: "You are standing at the end of a road before a small brick building. Around you is a forest. A small stream flows out of the building and down a gully…"


Graduate school not always just about learning • The most important events of my life occurred May 10 to September 21, 1980 • Henry's advice: don't get married (some advice should not be taken!) • Ken's advice: any time you go anywhere (both of you) will visit and give talks: Salt Lake City, Cincinnati, Indianapolis, Berkeley, Portland, Ann Arbor

While in graduate school at UCLA • Lots of coursework • Qualifying exams in biomathematics, genetics • RA/programmer; TA for population genetics • Learned to collaborate, write, present (better) • Platform presentations at ASHG in 1981, 1982 • 9 papers (4 first author), 7 with Ken • Postdoc offers from Mary Claire King, Jerry Rotter, Michael Conneally, Mark Skolnick

Choosing Michigan • ASHG in 1982 in Detroit • Ken arranged for Betsy and me to give talks at U-M • Came back in April 1983 for actual job interviews • Choose Michigan over NIH, postdocs • Reasons: • best sum of two jobs in the same place • great university, outstanding human genetics • strong welcome from Biostatistics and other faculty: Charlie Sing, Pat Peyser • felt at home; a place if things went well we might stay a few years (>35 years later )

Early years at Michigan • Started January 1, 1984 • Coldest winter in 50 years; -21 o. F 3 weeks after arrival • 12 th faculty member in department • First two years summer months supported by Charlie • Daily faculty lunches helped with easy integration into department, SPH, U-M • Morton Brown, Pat Peyser immediate mentors; Morton and Raya Brown made us part of their family

Early years at Michigan • Established methods research independence (without Ken) • Learned to do applied research (most people call it science): Alzheimer disease, neurofibromatosis, breast cancer, lipids, blood pressure, human mutation • Learned to teach: in first 3½ years at U-M, I taught 11 courses, 7 of them different • • • 601 (now 602): Statistical inference 666: Statistical models and numerical methods in human genetics 500/503 (now 501): Introduction to biostatistics 680: Stochastic processes 511: Statistical computer packages (OJOC) 580: Mathematical modeling in clinical research (OJOC)

Early years at Michigan: our three sons • David Foxman Boehnke, Sept 23, 1985 • Kevin Foxman Boehnke, April 23, 1987 • Richard Foxman Boehnke, July 2, 1989 • "You know Mike, three kids is a lot more work than two. " Galen Shorack, April 1989 (TIMELY mentoring is valuable)

First key methods paper on my own Basis for NIH grant HG 000376 in 1988; start of continuous methods funding for >30 years Basis for Lynn Ploughman dissertation, my first Ph. D student

Other early methods research 1986 -1994 • Power of linkage studies for complex traits (with Lynn Ploughman) • Optimal design for human family studies (5 papers) • Statistical methods for radiation hybrid mapping: • 8 methods papers (+6 science papers) • focus of Kathryn Lunetta dissertation, a paper in Heather Stringham's dissertation • 1992 paper (with Ken) won Snedecor award • Allele frequency estimation in pedigrees • Limits of resolution of genetic linkage studies

Getting started in human gene mapping GENOMICS 1, 361 -363 (1987) Linkage Analysis of von Recklinghausen Neurofibromatosis to DNA Markers on Chromosome 17 S. R. DIEHL, M. BOEHNKE, R. P. ERICKSON, A. B. BAXTER, M. A. BRUCE, J. L. LIEBERMAN, D. J. PLATT, L. M. PLOUGHMAN, K. A. SEILER, A. M. SWEET, AND F. S. COLLINS best collaborators -----“Whoever has the most toys wins” 1980 s bumper sticker Francis Collins moved to U-M in 1984, same year I did 17

CDKN 2 B/A 8 q 24 #2 8 q 24 #3 8 q 24 #4 8 q 24 #5 8 q 24 #6 Cholesterol Age related macular degeneration ATG 16 L 1 Obesity Crohn’s disease 5 p 13 Myocardial Type 1 diabetes 10 q 21 infarction Systemic lupus erythematosus IRGM QT interval Asthma NKX 2 -3 Atrial fibrilliation Restless leg syndrome IL 12 B 3 p 21 Type 2 diabetes Gallstone disease 1 q 24 Prostate cancer Multiple sclerosis NOS 1 AP PTPN 2 Breast cancer Rheumatoid arthritis IFIH 1 TCF 2 Colon cancer Glaucoma PCSK 9 CDKN 2 B/ Height CFB/C 2 A LOC 38771 IGF 2 BP 2 5 CD 25 CDKAL 1 8 q 24 IRF 5 HHEX IBD 5 IL 23 R PCSK 9 NOD 2 TCF 7 L 2 SLC 30 A 8 CTLA 4 CFH PPAR KCNJ 11 PTPN 22 Early (lack of) progress in mapping genes for complex traits 2000 2001 2002 2003 2004 2005 2006 HMGA 2 MEIS 1 GDF 5 LBXCOR 1 UQCC BTBD 9 HMPG C 3 JAZF 1 8 q 24 CDC 123 ORMDL 3 ADAMTS 4 q 25 TCF 2 9 GCKR THADA WSF 1 FTO C 12 orf 30 LOXL 1 ERBB 3 IL 7 R KIAA 0350 TRAF 1/C 5 CD 226 STAT 4 16 p 13 ABCG 8 PTPN 2 GALNT 2 SH 2 B 3 PSRC 1 FGFR 2 NCAN TNRC 9 TBL 2 MAP 3 K 1 TRIB 1 LSP 1 KCTD 10 8 q 24 ANGLPT 3 GRIN 3 A 2007 Slide courtesy of David Altshuler 18

FUSION: Finland-United States Investigation of NIDDM Genetics NHGRI, Bethesda, Francis Collins Cedars Sinai, Los Angeles, Richard Bergman KTL, Helsinki, Jaakko Tuomilehto U Michigan, Michael Boehnke Soumitra Ghosh (postdoc), Beth Hauser (student) 19

FUSION: Finland-United States Investigation of NIDDM Genetics NHGRI, Bethesda, Francis Collins Cedars Sinai, Los Angeles, Richard Bergman KTL, Helsinki, Jaakko Tuomilehto U North Carolina, Karen Mohlke U Eastern Finland, Markku Laakso U Michigan, Michael Boehnke and Laura Scott 20

FUSION study goals Identify genetic variants that predispose to type 2 diabetes (T 2 D) or are responsible for variability in T 2 D-related traits 21

FUSION ASP families for linkage analysis FUSION started in 1990 s as a T 2 D family study Sampled >5000 individuals from >800 families with ≥ 2 affected siblings Obtained extensive phenotype information Genotyped participants at ~400 microsatellite markers at CIDR 22
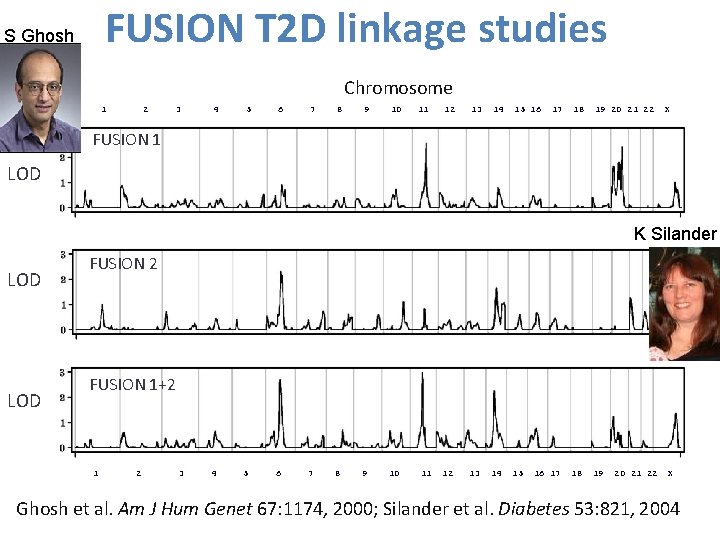
FUSION T 2 D linkage studies S Ghosh Chromosome 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 X FUSION 1 LOD K Silander LOD FUSION 2 FUSION 1+2 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 X 23 Ghosh et al. Am J Hum Genet 67: 1174, 2000; Silander et al. Diabetes 53: 821, 2004

International T 2 D Linkage Analysis Consortium Approach: combine linkage data across studies >6000 families from ~20 studies Guan et al. Human Heredity 2008 D Burns S Elbein P Froguel B Mitchell W Guan N Cox A Pluzhnikov LOD 24

“If you’re going through hell, keep going. ” Winston Churchill

The Future of Genetic Studies of Complex Human Diseases Neil Risch and Kathleen Merikangas SCIENCE VOL. 273 13 SEPTEMBER 1996 26

Genome-Wide Association Study (GWAS) • Risch and Merikangas Science 1996 • Sample individuals with and without disease (e. g. “cases” with T 2 D, “controls” without T 2 D) • Genotype individuals for 100, 000 s of single nucleotide polymorphisms (SNPs) across the genome • Test for disease-variant association • Identify variants statistically associated with disease, suggesting disease gene nearby • Anonymous reviewer: “science fiction” 27
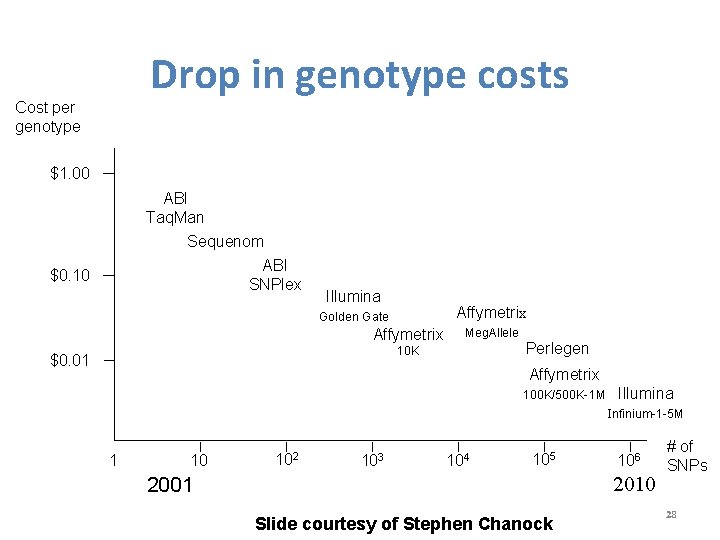
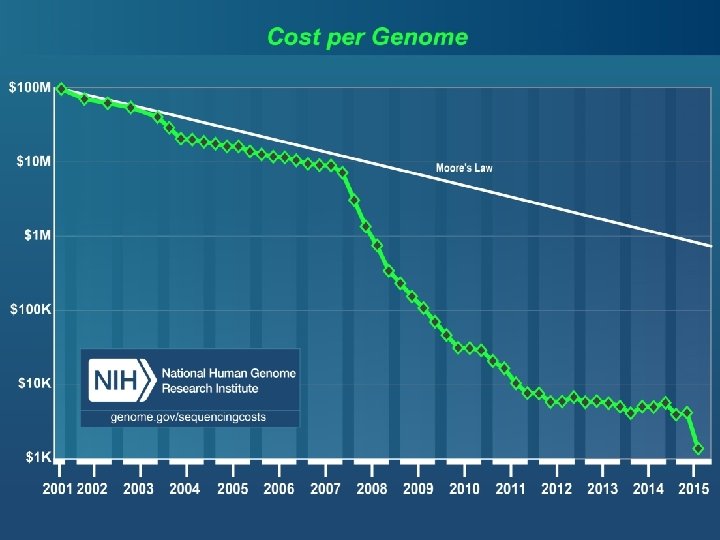
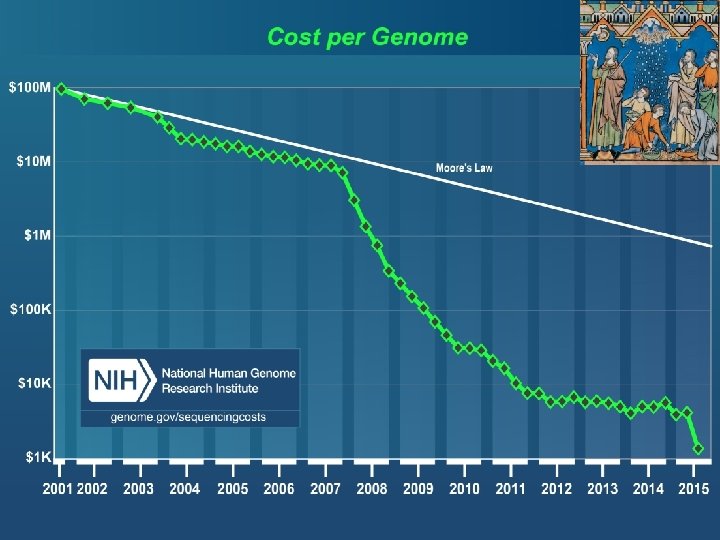
Drop in genotype costs Cost per genotype $1. 00 ABI Taq. Man Sequenom ABI SNPlex $0. 10 Illumina Affymetrix Golden Gate Affymetrix Meg. Allele Perlegen 10 K $0. 01 Affymetrix 100 K/500 K-1 M Illumina Infinium-1 -5 M 1 10 102 103 104 105 106 2010 2001 Slide courtesy of Stephen Chanock # of SNPs 28
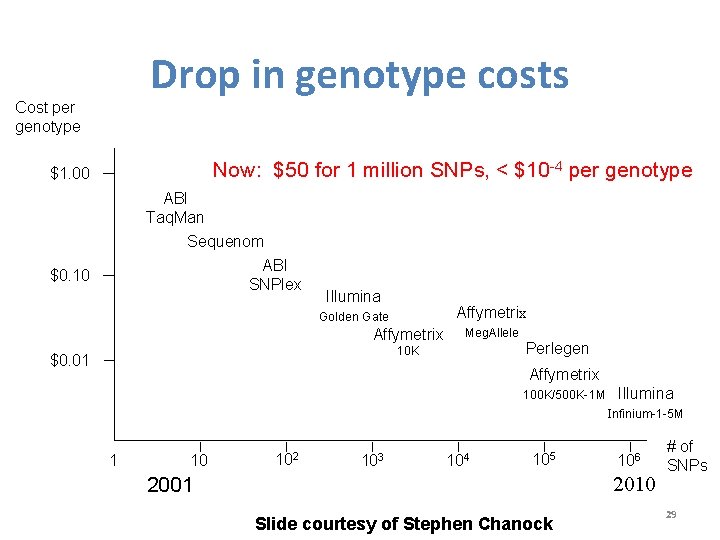
Drop in genotype costs Cost per genotype Now: $50 for 1 million SNPs, < $10 -4 per genotype $1. 00 ABI Taq. Man Sequenom ABI SNPlex $0. 10 Illumina Affymetrix Golden Gate Affymetrix Meg. Allele Perlegen 10 K $0. 01 Affymetrix 100 K/500 K-1 M Illumina Infinium-1 -5 M 1 10 102 103 104 105 106 2010 2001 Slide courtesy of Stephen Chanock # of SNPs 29

When manna falls from heaven, say thank you. And take advantage of it! Be opportunistic 30

FUSION a grant, not a contract • Grant renewal obtained in 2003 • Specific aims: fine map several T 2 D linkage peaks 31

FUSION is a grant, not a contract • Grant renewal obtained in 2003 • Specific aims: fine map several T 2 D linkage peaks • Ignored aims, instead carried out one of the first GWAS (for T 2 D) – First GWAS to use genotype imputation – First GWAS meta-analysis – Developed methods for efficient two-stage designs Laura Scott Yun Li Andrew Skol Gonçalo Abecasis 32

Joint analysis is more powerful than replication-based analysis Skol, Scott, Abecasis, Boehnke (2006) Nature Genetics 38: 209 -213 One-stage power M = 300, 000 SNPs genotyped on 1000 cases, 1000 controls Multiplicative model, prevalence 10%, GRR = 1. 4 33
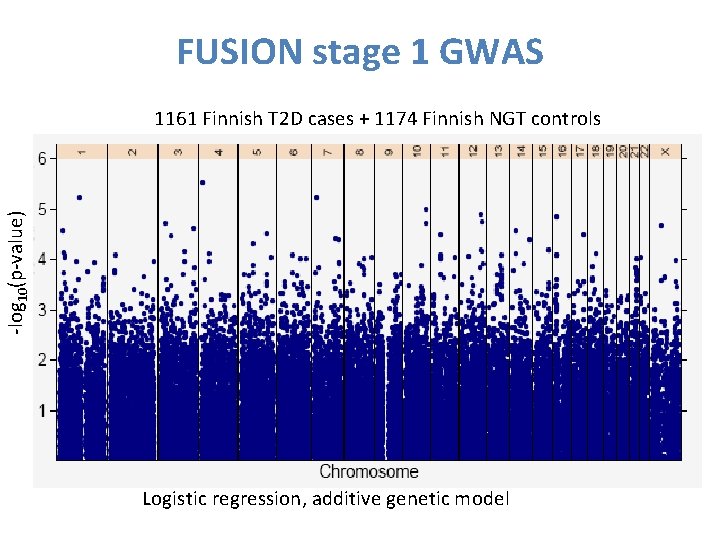
FUSION stage 1 GWAS -log 10(p-value) 1161 Finnish T 2 D cases + 1174 Finnish NGT controls Logistic regression, additive genetic model 34
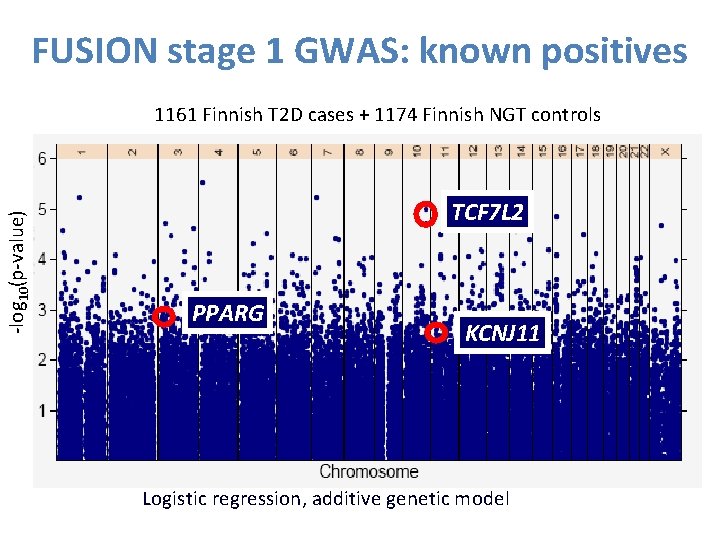
FUSION stage 1 GWAS: known positives -log 10(p-value) 1161 Finnish T 2 D cases + 1174 Finnish NGT controls TCF 7 L 2 PPARG KCNJ 11 Logistic regression, additive genetic model 35

Three-study collaboration • FUSION: Finnish cases and controls D Altshuler • Diabetes Genetic Initiative (DGI): Finnish, Swedish cases and controls • UKT 2 D: UK cases, controls M Mc. Carthy • Problem: FUSION genotyped Illumina 317 K chip; DGI, UK Affymetrix 500 K chip • Combine results across studies with different SNP sets by genotype imputation (Li, Abecasis et al. ) Y Li G Abecasis 3 6
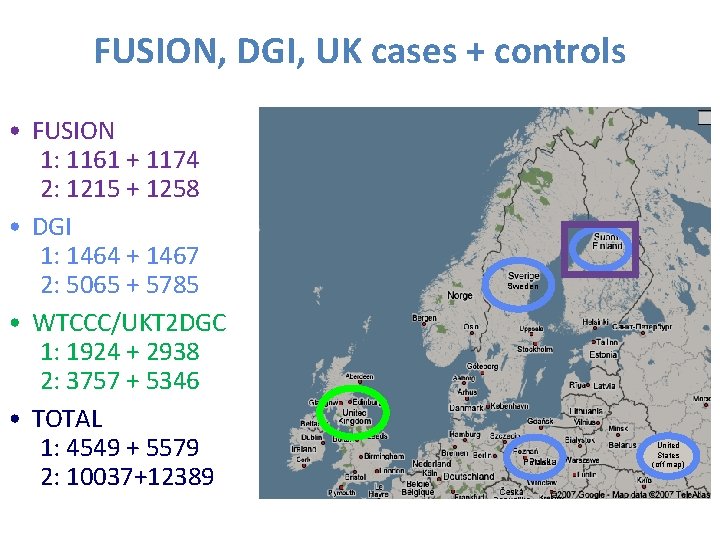
FUSION, DGI, UK cases + controls • FUSION 1: 1161 + 1174 2: 1215 + 1258 • DGI 1: 1464 + 1467 2: 5065 + 5785 • WTCCC/UKT 2 DGC 1: 1924 + 2938 2: 3757 + 5346 • TOTAL 1: 4549 + 5579 2: 10037+12389 Sweden Poland United States (off map) 37 37

E Zeggini R Saxena Science April 2007 38 38 L Scott >2800 citations to date
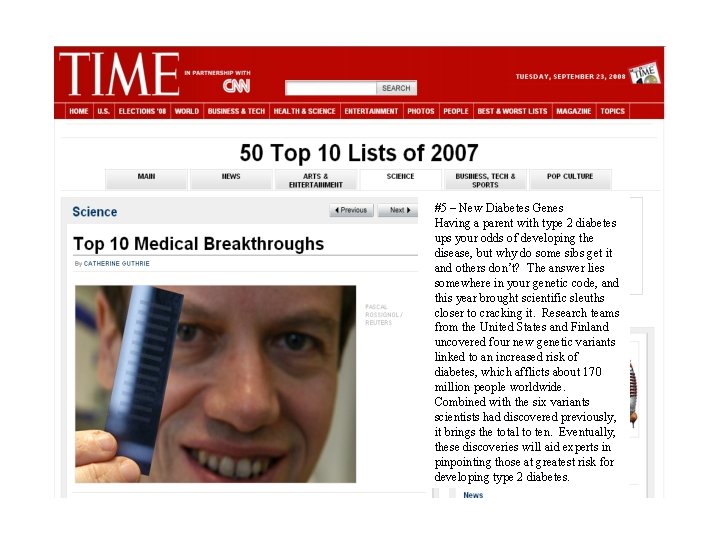
#5 – New Diabetes Genes Having a parent with type 2 diabetes ups your odds of developing the disease, but why do some sibs get it and others don’t? The answer lies somewhere in your genetic code, and this year brought scientific sleuths closer to cracking it. Research teams from the United States and Finland uncovered four new genetic variants linked to an increased risk of diabetes, which afflicts about 170 million people worldwide. Combined with the six variants scientists had discovered previously, it brings the total to ten. Eventually, these discoveries will aid experts in pinpointing those at greatest risk for developing type 2 diabetes. 39

http: //www. sciencemag. org/cgi/ content/full/318/5858/1842 40
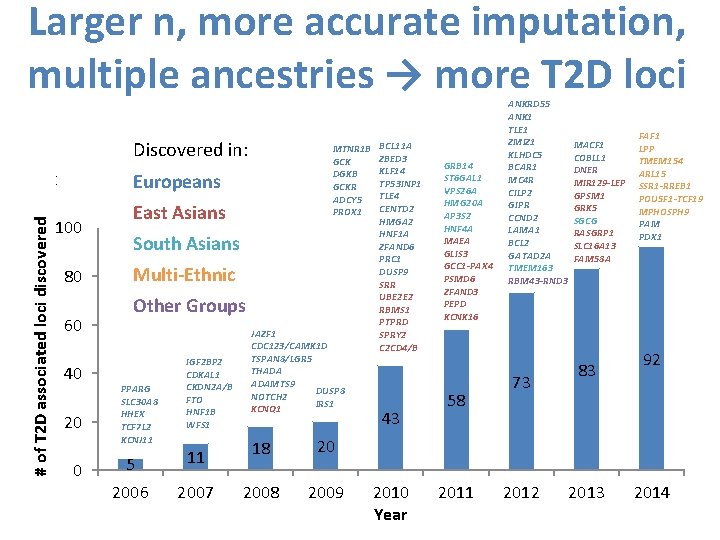
Larger n, more accurate imputation, multiple ancestries → more T 2 D loci Discovered in: # of T 2 D associated loci discovered 120 100 80 60 40 20 0 Europeans East Asians South Asians Multi-Ethnic Other Groups PPARG SLC 30 A 8 HHEX TCF 7 L 2 KCNJ 11 IGF 2 BP 2 CDKAL 1 CKDN 2 A/B FTO HNF 1 B WFS 1 MTNR 1 B BCL 11 A ZBED 3 GCK KLF 14 DGKB TP 53 INP 1 GCKR TLE 4 ADCY 5 CENTD 2 PROX 1 HMGA 2 HNF 1 A ZFAND 6 PRC 1 DUSP 9 SRR UBE 2 E 2 RBMS 1 PTPRD JAZF 1 SPRY 2 CDC 123/CAMK 1 D C 2 CD 4/B TSPAN 8/LGR 5 THADA ADAMTS 9 DUSP 8 NOTCH 2 IRS 1 KCNQ 1 43 5 11 18 20 2006 2007 2008 2009 2010 Year GRB 14 ST 6 GAL 1 VPS 26 A HMG 20 A AP 3 S 2 HNF 4 A MAEA GLIS 3 GCC 1 -PAX 4 PSMD 6 ZFAND 3 PEPD KCNK 16 58 2011 ANKRD 55 ANK 1 TLE 1 ZMIZ 1 KLHDC 5 BCAR 1 MC 4 R CILP 2 GIPR CCND 2 LAMA 1 BCL 2 GATAD 2 A TMEM 163 RBM 43 -RND 3 73 2012 MACF 1 COBLL 1 DNER MIR 129 -LEP GPSM 1 GRK 5 SGCG RASGRP 1 SLC 16 A 13 FAM 58 A 83 2013 FAF 1 LPP TMEM 154 ARL 15 SSR 1 -RREB 1 POU 5 F 1 -TCF 19 MPHOSPH 9 PAM PDX 1 92 2014
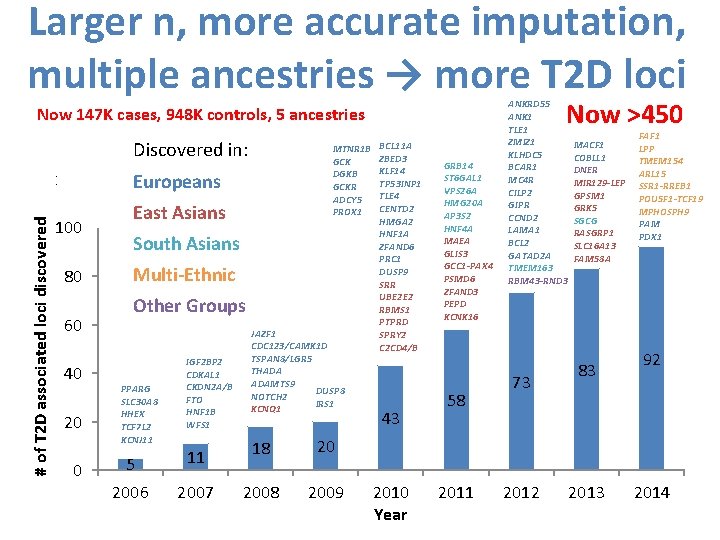
Larger n, more accurate imputation, multiple ancestries → more T 2 D loci Now >450 Now 147 K cases, 948 K controls, 5 ancestries Discovered in: # of T 2 D associated loci discovered 120 100 80 60 40 20 0 Europeans East Asians South Asians Multi-Ethnic Other Groups PPARG SLC 30 A 8 HHEX TCF 7 L 2 KCNJ 11 IGF 2 BP 2 CDKAL 1 CKDN 2 A/B FTO HNF 1 B WFS 1 MTNR 1 B BCL 11 A ZBED 3 GCK KLF 14 DGKB TP 53 INP 1 GCKR TLE 4 ADCY 5 CENTD 2 PROX 1 HMGA 2 HNF 1 A ZFAND 6 PRC 1 DUSP 9 SRR UBE 2 E 2 RBMS 1 PTPRD JAZF 1 SPRY 2 CDC 123/CAMK 1 D C 2 CD 4/B TSPAN 8/LGR 5 THADA ADAMTS 9 DUSP 8 NOTCH 2 IRS 1 KCNQ 1 43 5 11 18 20 2006 2007 2008 2009 2010 Year GRB 14 ST 6 GAL 1 VPS 26 A HMG 20 A AP 3 S 2 HNF 4 A MAEA GLIS 3 GCC 1 -PAX 4 PSMD 6 ZFAND 3 PEPD KCNK 16 58 2011 ANKRD 55 ANK 1 TLE 1 ZMIZ 1 KLHDC 5 BCAR 1 MC 4 R CILP 2 GIPR CCND 2 LAMA 1 BCL 2 GATAD 2 A TMEM 163 RBM 43 -RND 3 73 2012 MACF 1 COBLL 1 DNER MIR 129 -LEP GPSM 1 GRK 5 SGCG RASGRP 1 SLC 16 A 13 FAM 58 A 83 2013 FAF 1 LPP TMEM 154 ARL 15 SSR 1 -RREB 1 POU 5 F 1 -TCF 19 MPHOSPH 9 PAM PDX 1 92 2014
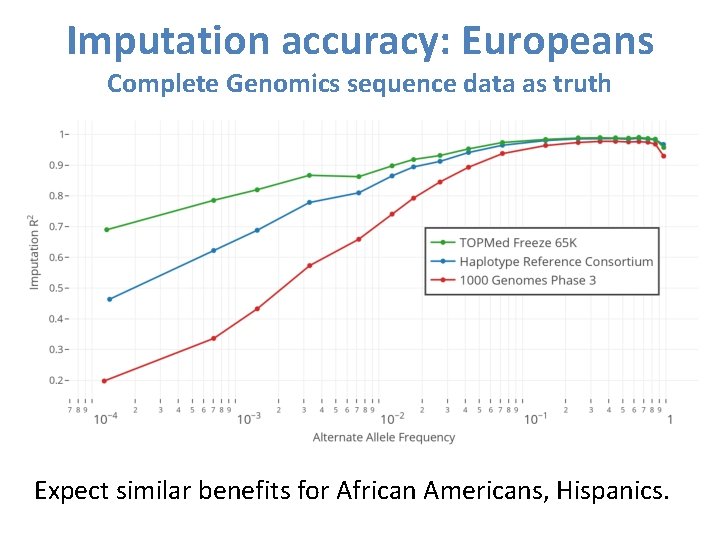
Imputation accuracy: Europeans Complete Genomics sequence data as truth Expect similar benefits for African Americans, Hispanics.

Testing for association with T 2 D-related traits • Routinely test for association with T 2 D-associated traits: glucose, insulin; BMI, WHR, height; lipids; … • Given phenotypes/genotypes, no additional experiments required • Our GWAS meta-analysis consortia (MAGIC, GIANT, Global Lipids, …) have identified >1000 loci for these traits; many papers in Nature, Nature Genetics, … 45

Next step: sequencing • Impressive successes with array-based GWAS • Now also genome and exome sequencing • Allows near-complete assay of – full allele-frequency spectrum – all classes of genetic variation • Why haven’t we been doing this all along?

T 2 D-GENES/AMP-T 2 D 45 K exomes • Published association results for 13 K exomes from 5 ancestry groups (Fuchsberger et al. Nature 2016) • Now expanded analysis to >45 K exomes as C Fuchsberger part of Accelerating Medicines Partnership project Ancestry T 2 D Cases Controls European 4, 993 7, 468 Hispanic 7, 143 7, 321 African American 2, 756 3, 660 East Asian 2, 927 3, 105 South Asian 2, 972 2, 886 Total 20, 791 24, 440 J Flannick • Testing for T 2 D association: single-variant, gene-based, gene sets (Flannick et al. Nature 2019)

Finnish exome sequencing Fin. Met. Seq: N~20 K Locke et al. Nature, in press • METSIM • Population based study of 10, 197 Kuopio men • Ages 45 -70 years at baseline • Baseline study in 2005 -2010, follow-up ongoing • FINRISK • Health surveys every five years since 1972 • Sample sizes 6, 000 -12, 000 individuals • Ages 25 -74 • Follow-up in 25 K Finns with GWAS data • Results – – 478 novel associations at 126 loci for 64 traits Many associated variants enriched in Finns Some effect sizes large E. g. AF=. 004 variant in THBSP 4 associated with 6 kg decrease in weight, 35 x enriched in Finns Kuopio K Meltz-Steinberg A Locke

Methods continue to follow science • Dealing with case-control imbalance in GWAS • Contamination detection and correction in sequencing studies • Efficient designs for genotyping-sequencing studies • Efficient multi-trait GWAS • Visualization of genetic association signals Clement Ma M Flickinger Corbin Quick Debashree Ray Ryan Welch

R Welch A Boughton

UM Genome Science Training Program • 1989: NHGRI decided to support training grants • Invited to planning meeting to discuss • Suggested program focused at interface of statistics and genomics • Funded for 5 trainees in year 1 (1990), 8 in years 2 -5 • 2009: requested 10 slots, study section said 13(!) • Funded 121 trainees to date, 51 in Biostatistics • Just submitted competing renewal for years 26 -30

A few lessons learned along the way • Life is a grant, not a contract: do what matters to you • Do work that has impact: develop methods and tools that people will use • Work hard to be lucky; be opportunistic (and grateful) when you are • Choose places (program, school, job, …) where if you do what you want to do and do it well, people will love you • When in doubt, do what you think is exciting and fun, and do it with the best (smartest, nicest) people • Anyone can be your mentor: 10 minutes, 10 years, … • Celebrate your triumphs, large and small

Summary and Comments • Real progress with T 2 D genetics: >240 loci identified, 100 s for T 2 D-related traits like glucose, insulin, body mass index • Variants of modest impact, but each can lead to better understanding of T 2 D etiology, new or better targeted therapies • Sequencing allows us to explore full range of genetic variation • Associated common variants mostly consistent across ancestries; associated rare variants often differ by ancestry • New statistical methods and computational tools key to success • Working on exciting science provides opportunity to develop statistical methods that will be used by many others 55

Acknowledgements • Michigan: L Scott, G Abecasis, T Teslovich, HM Kang, X Sim, G Jun, C Fuchsberger, A Locke, D Taliun, R Welch, H Stringham, A Jackson, T Blackwell • More FUSION/METSIM: M Laakso, K Mohlke, F Collins, J Tuomilehto, H Koistinen, R Bergman, L Bonnycastle, P Chines, M Erdos, N Narisu, A Swift, R Watanabe • CIDR: K Doheny, E Pugh • Broad/Lund: D Altshuler, L Groop, J Florez, R Saxena, B Voight, N Burtt, S Gabriel, J Flannick, A Manning, P Fontanillas, A Williams, E Banks, C Hartl • Oxford/Exeter: M Mc. Carthy, A Morris, A Hattersley, K Gaulton, P Donnelly, C Lindgren, I Prokopenko, A Mahajan • Munich: Thomas Meitinger, Tim Strom • More T 2 D-GENES: R Duggirala, J Blangero, C Hanis, N Cox, G 56 Bell • Funding from NIDDK, NHGRI, FNIH, ADA, Wellcome Trust


Louis B Pierce Rebecca B Pierce
- Slides: 58