UBIQUITIN 76 Amino Acid polypeptide Highly conserved in
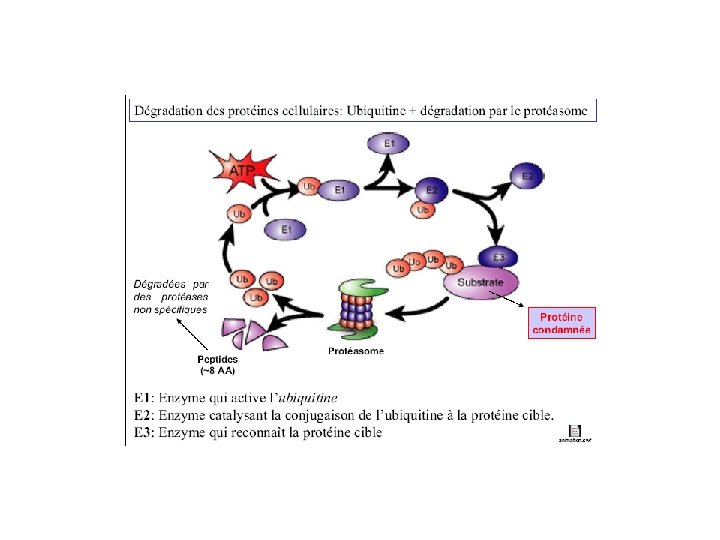

UBIQUITIN • 76 Amino Acid polypeptide • Highly conserved in evolution • 3 Amino acid differences between yeast and human homologues • C-Terminal Gly residue is activated via an ATP to form a thiol ester

UBIQUITIN
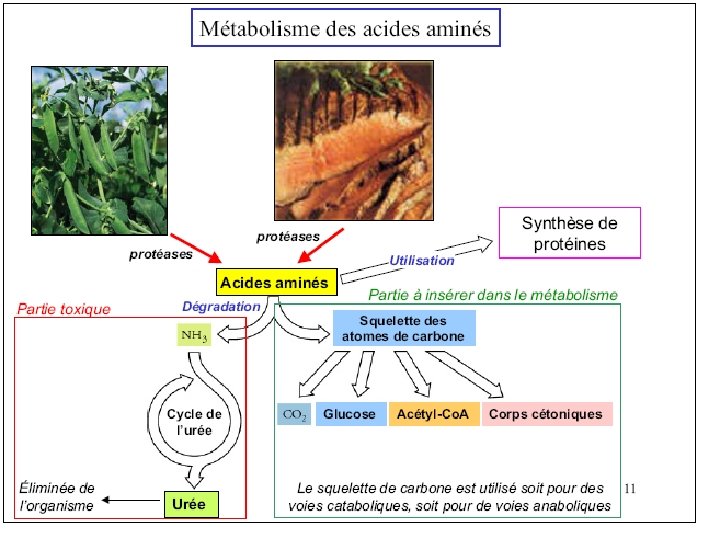

Amino Acid Metabolism • Metabolism of the 20 common amino acids is considered from the origins and fates of their: (1) Nitrogen atoms (2) Carbon skeletons • For mammals: Essential amino acids must be obtained from diet Nonessential amino acids - can be synthesized

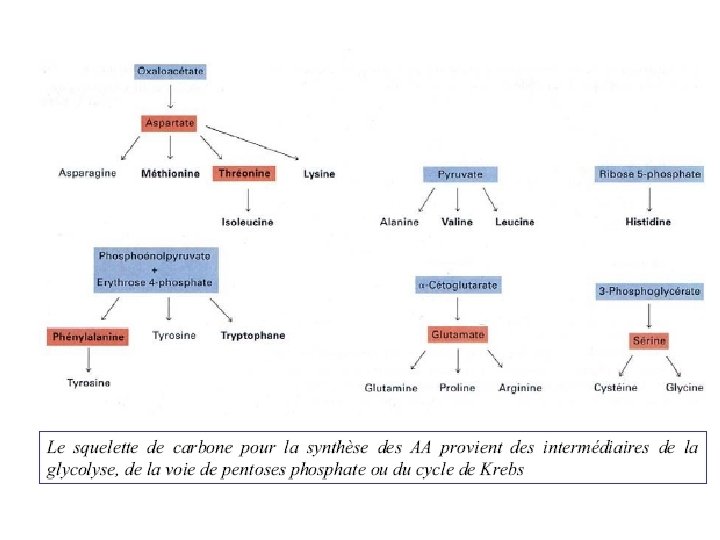
Non-Essential Amino Acids in Humans • Not required in diet • Can be formed from a-keto acids by transamination and subsequent reactions • Alanine • Asparagine • Aspartate • Glutamine • Glycine • Proline • Serine • Cysteine (from Met*) • Tyrosine (from Phe*) * Essential amino acids

Essential Amino Acids in Humans • Required in diet • Humans incapable of forming requisite carbon skeleton • Arginine* • Histidine* • Isoleucine • Leucine • Valine • Lysine • Methionine • Threonine • Phenylalanine • Tryptophan * Essential in children, not in adults

. Fournisseur de peu d’énergie (10%). AA se jettent dans le cycle de Krebbs . AA pour synthèse de protéines. Catabolisme des aa absent chez les plantes
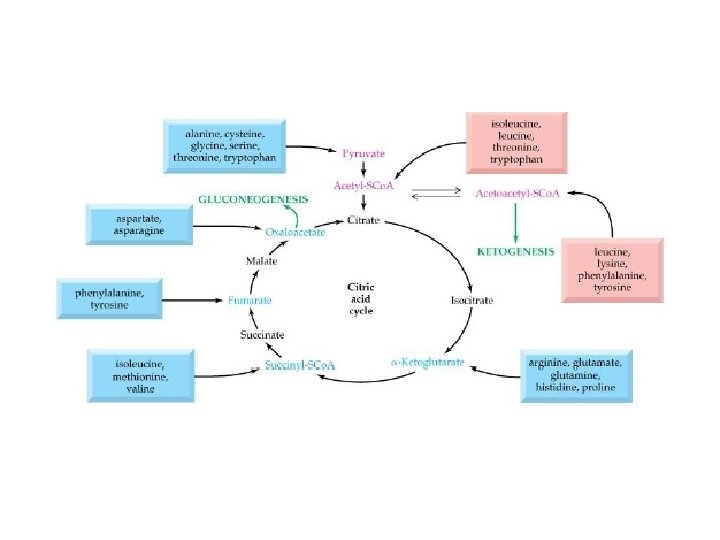

Catabolisme
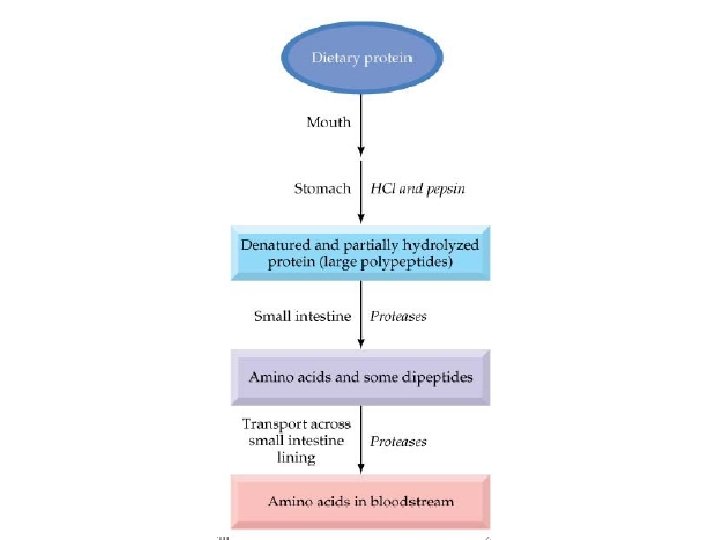
• Whole proteins are not absorbed – Too large to pass through cell membranes intact – Some combined with carbohydrates (glycoproteins) • Digestive enzymes – Break peptide bonds • Secreted as inactive pre-enzymes – Prevents self-digestion

• Initiated in stomach – HCl from parietal cells • Stomach p. H 1. 6 to 3. 2 • Denatures 40, 30, and 20 structures – Pepsinogen from chief cells Pepsinogen HCl Pepsin • Cleaves at phenylalanine, tyrosine, tryptophan Aromatic amino acids • Protein leaves stomach as mix of insoluble protein, peptides and amino acids
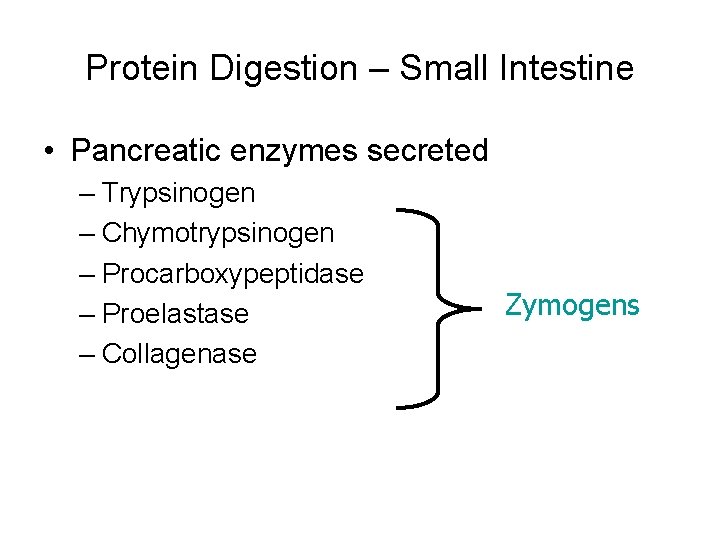
Protein Digestion – Small Intestine • Pancreatic enzymes secreted – Trypsinogen – Chymotrypsinogen – Procarboxypeptidase – Proelastase – Collagenase Zymogens

– Small Intestine • Zymogens must be converted to active form – Trypsinogen Enteropeptidase/Trypsin • Endopeptidase – Cleaves on carbonyl side of Lys & Arg – Chymotrypsinogen Trypsin Chymotrypsin • Endopeptidase – Cleaves carboxy terminal Phe, Tyr and Trp – Procarboxypeptidase Trypsin Carboxypeptidase • Exopeptidase – Removes carboxy terminal residues Endo dans Exo à partir des extrémités
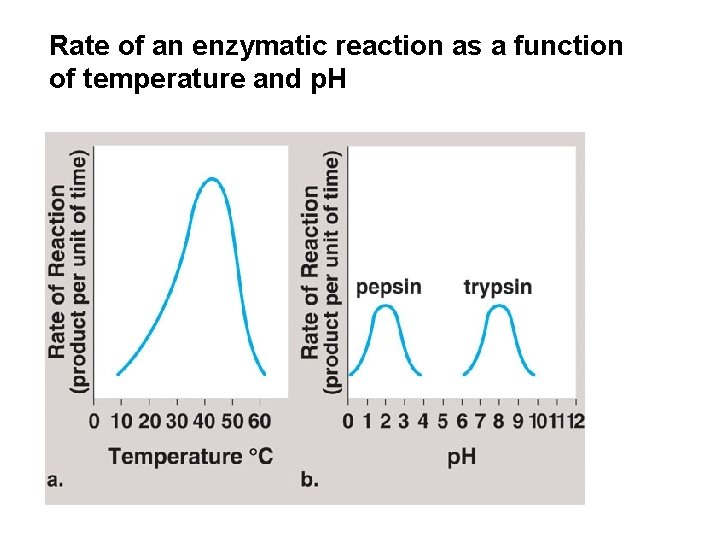
Rate of an enzymatic reaction as a function of temperature and p. H
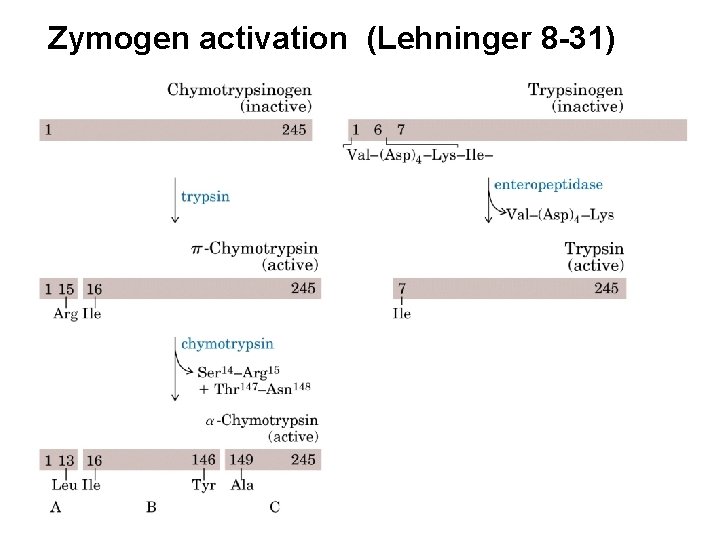
Zymogen activation (Lehninger 8 -31)
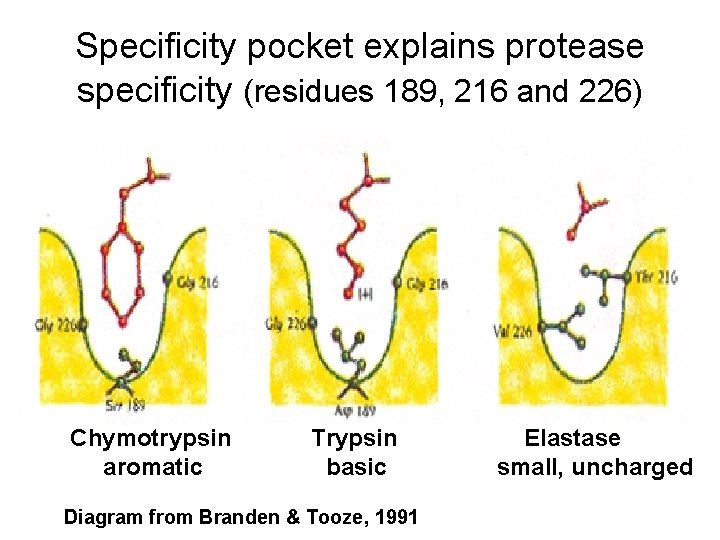
Specificity pocket explains protease specificity (residues 189, 216 and 226) Chymotrypsin aromatic Trypsin basic Diagram from Branden & Tooze, 1991 Elastase small, uncharged

Protein Digestion • Small intestine (brush border) – Aminopeptidases • Cleave at N-terminal AA – Dipeptidases • Cleave dipeptides – (Enterokinase or enteropeptidase) • Trypsinogen trypsin • Trypsin then activates all the other enzymes

Peptide Absorption • Form in which the majority of protein is absorbed • More rapid than absorption of free amino acids • Coupled to H gradient – Na/H exchange maintains gradient • Metabolized into free amino acids in enterocyte • Only free amino acids absorbed into blood

• AA exogène >>>>>foie • AA endogène • Ammoniaque: Toxique Rare organisme qui transforment N 2 en ammonique

The Nitrogen Cycle and Nitrogen Fixation • Nitrogen is needed for amino acids, nucleotides • Atmospheric N 2 is the ultimate source of biological nitrogen • Nitrogen fixation: a few bacteria possess nitrogenase which can reduce N 2 to ammonia • Nitrogen is recycled in nature through the nitrogen cycle

The Nitrogen cycle


Nitrogenase • An enzyme present in Rhizobium bacteria that live in root nodules of leguminous plants • Some free-living soil and aquatic bacteria also possess nitrogenase • Nitrogenase reaction: N 2 + 8 H+ + 8 e- + 16 ATP 2 NH 3 + H 2 + 16 ADP + 16 Pi
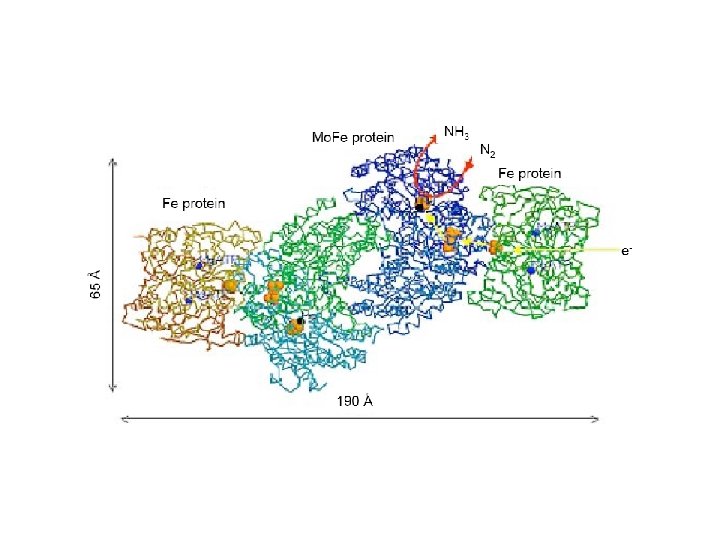

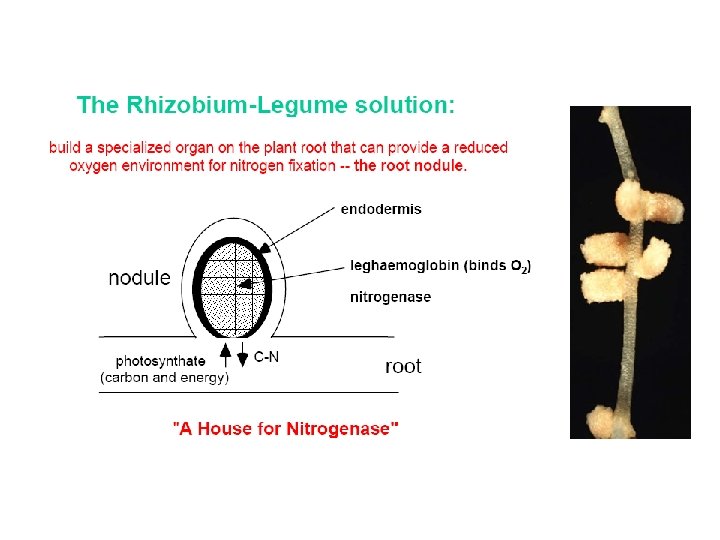

Assimilation of Ammonia • Ammonia generated from N 2 is assimilated into low molecular weight metabolites such as glutamate or glutamine • At p. H 7 ammonium ion predominates (NH 4+)

Ammonia Is Incorporated into Glutamate • Reductive amination of a-ketoglutarate by glutamate dehydrogenase occurs in plants, animals and microorganisms • In mammals & plants, located in mitochondria.

Glutamine Is a Nitrogen Carrier in Many Biosynthetic Reactions • A second important route in assimilation of ammonia is via glutamine synthetase

Glutamate synthase transfers a nitrogen to a-ketoglutarate Prokaryotes & plants

Alternate amino acid production in prokaryotes

How some enzymes transfer ammonia from glutamine • CP synthetase has 3 active sites connected by a tunnel running through the interior • Protects intermediates from being degraded by water
Carbamoyl phosphate synthase backbone structure • Tunnel connecting active sites (blue wire)

C. Regulation of Glutamine Synthetase in E. coli • Glutamine synthetase (GS) plays a critical role in nitrogen metabolism • E. coli enzyme regulated by: (1) Cumulative feedback inhibition (9 allosteric inhibitors with additive effects) (2) Covalent modification (3) Regulation of enzyme synthesis

Allosteric inhibition of GS in E. coli

Regulation of E. coli GS by covalent modification

Regulation of mammalian GS • Regulation not as extensive as in microorganisms • No covalent regulation • Allosteric inhibitors: glycine, serine, alanine, and carbamoyl phosphate • Allosteric activator: a-ketoglutarate

Transamination Reactions • Transfer of an amino group from an a-amino acid to an a-keto acid • In amino acid biosynthesis, the amino group of glutamate is transferred to various a-keto acids generating a-amino acids • In amino acid catabolism, transamination reactions generate glutamate or aspartate

Transamination reactions

Transaminases (aminotransferase s) catalyze the reversible reaction at right. There are multiple transaminase enzymes which vary in substrate specificity. Some show preference for particular amino acids or classes of amino acids as amino group donors, and/or for particular a-keto acid acceptors.
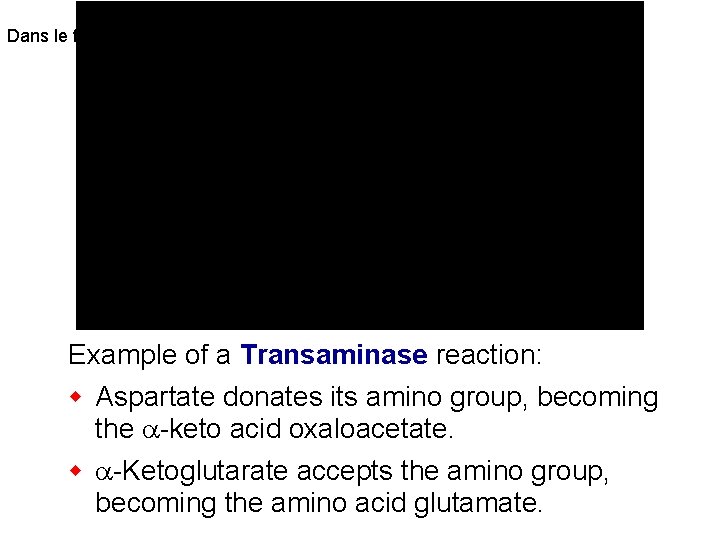
Dans le foie Example of a Transaminase reaction: w Aspartate donates its amino group, becoming the a-keto acid oxaloacetate. w a-Ketoglutarate accepts the amino group, becoming the amino acid glutamate.

Transaminases equilibrate amino groups among available a-keto acids. This permits synthesis of non-essential amino acids, using amino groups from other amino acids & carbon skeletons synthesized in a cell. Thus a balance of different amino acids is maintained, as proteins of varied amino acid contents are synthesized. Although the amino N of one amino acid can be used to synthesize another amino acid, N must be obtained in the diet as amino acids (proteins).

The prosthetic group of Transaminase is pyridoxal phosphate (PLP), a derivative of vitamin B 6.

In the resting state, the aldehyde group of pyridoxal phosphate is in a Schiff base linkage to the e-amino group of an enzyme lysine sidechain.

The a-amino group of a substrate amino acid displaces the enzyme lysine, to form a Schiff base linkage to PLP. The active site lysine extracts H+, promoting tautomerization, followed by reprotonation & hydrolysis.

What was an amino acid leaves as an aketo acid. The amino group remains on what is now pyridoxamine phosphate (PMP). A different a-keto acid reacts with PMP and the process reverses, to complete the reaction.

Several other enzymes that catalyze metabolism or synthesis of amino acids also utilize PLP as prosthetic group, and have mechanisms involving a Schiff base linkage of the amino group to PLP.

In addition to equilibrating amino groups among available a-keto acids, transaminases funnel amino groups from excess dietary amino acids to those amino acids (e. g. , glutamate) that can be deaminated. Carbon skeletons of deaminated amino acids can be catabolized for energy, or used to synthesize glucose or fatty acids for energy storage. Only a few amino acids are deaminated directly.

Glutamate Dehydrogenase catalyzes a major reaction that effects net removal of N from the amino acid pool. It is one of the few enzymes that can use NAD+ or NADP+ as e- acceptor. Oxidation at the a-carbon is followed by hydrolysis, releasing NH 4+.

Summarized above: The role of transaminases in funneling amino N to glutamate, which is deaminated via Glutamate Dehydrogenase, producing NH 4+.

Some other pathways for deamination of amino acids: 1. Serine Dehydratase catalyzes: serine pyruvate + NH 4+

2. Peroxisomal L- and D-amino acid oxidases catalyze: amino acid + FAD + H 2 O a-keto acid + NH 4+ + FADH 2 + O 2 FAD + H 2 O 2 Catalase catalyzes: 2 H 2 O 2 2 H 2 O + O 2 Postulated mechanisms for toxicity of high [ammonia]: 1. High [NH 3] would drive Glutamine Synthase: glutamate + ATP + NH 3 glutamine + ADP + Pi This would deplete glutamate – a neurotransmitter & precursor for synthesis of the neurotransmitter GABA. 2. Depletion of glutamate & high ammonia level would drive Glutamate Dehydrogenase reaction to reverse: glutamate + NAD(P)+ a-ketoglutarate + NAD(P)H + NH 4+ The resulting depletion of a-ketoglutarate, an essential Krebs Cycle intermediate, could impair energy metabolism in the brain.
Most terrestrial land animals convert excess nitrogen to urea, prior to excreting it. Urea is less toxic than ammonia. The Urea Cycle occurs mainly in liver. The 2 nitrogen atoms of urea enter the Urea Cycle as NH 3 (produced mainly via Glutamate Dehydrogenase) and as the amino N of aspartate. The NH 3 and HCO 3 - (carbonyl C) that will be part of urea are incorporated first into carbamoyl phosphate.
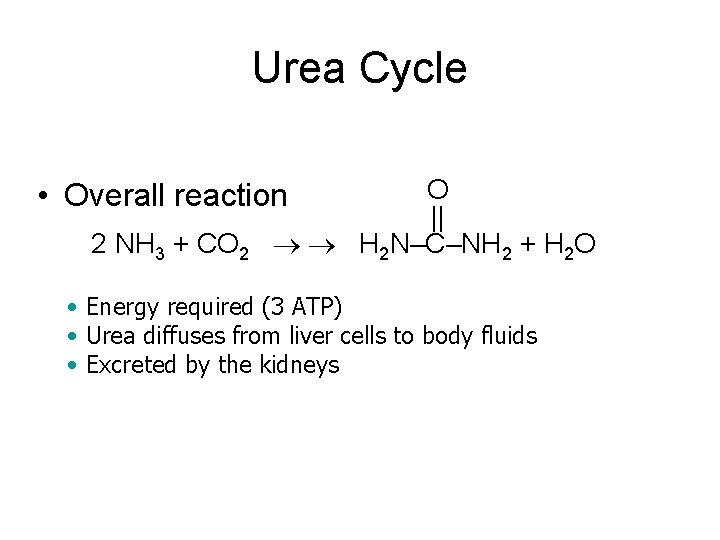
Urea Cycle O || 2 NH 3 + CO 2 H 2 N–C–NH 2 + H 2 O • Overall reaction • Energy required (3 ATP) • Urea diffuses from liver cells to body fluids • Excreted by the kidneys

Carbamoyl Phosphate Synthase (Type I) catalyzes a 3 -step reaction, with carbonyl phosphate and carbamate intermediates. Ammonia is the N input. The reaction, which involves cleavage of 2 ~P bonds of ATP, is essentially irreversible.

Urea Cycle Enzymes in mitochondria: 1. Ornithine Trans carbamylase Enzymes in cytosol: 2. Arginino Succinate Synthase 3. Arginino succinase 4. Arginase.
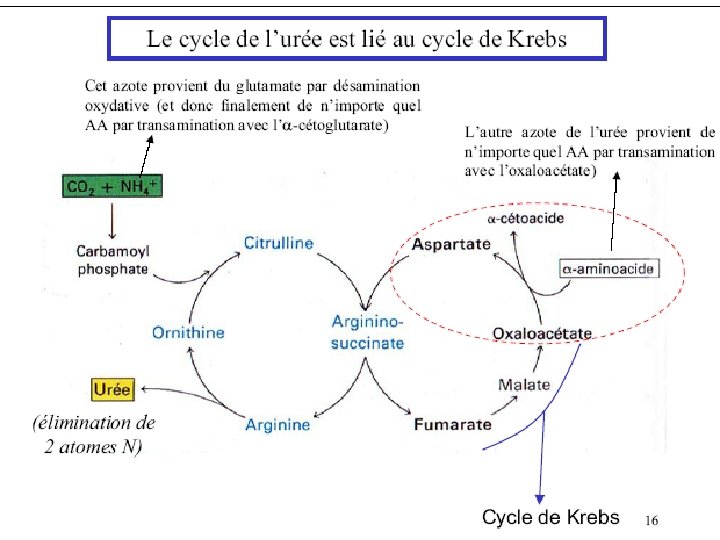
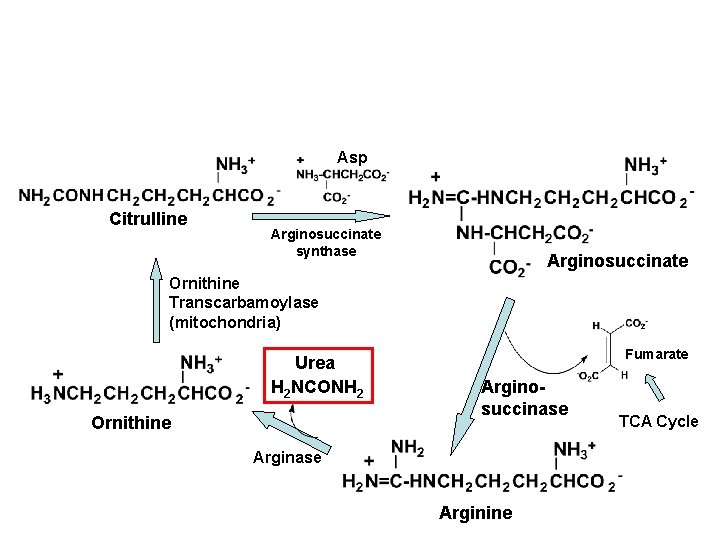
Asp Citrulline Arginosuccinate synthase Arginosuccinate Ornithine Transcarbamoylase (mitochondria) Urea H 2 NCONH 2 Ornithine Fumarate Arginosuccinase Arginine TCA Cycle

The complete Urea Cycle is significantly only in liver. However some enzymes of the pathway are in other cells and tissues where they generate arginine & ornithine, which are precursors for other important molecules. E. g. , Argininosuccinate Synthase, which catalyzes synthesis of the precursor to arginine, is in most tissues. Mitochondrial Arginase II, distinct from the cytosolic Urea Cycle Arginase, cleaves arginine to yield ornithine.

For each cycle, citrulline must leave the mitochondria, and ornithine must enter the mitochondrial matrix. An ornithine/citrulline transporter in the inner mitochondrial membrane facilitates transmembrane fluxes of citrulline & ornithine.

Arginine is a constituent of proteins, & is precursor for synthesis of creatine and nitric oxide (·NO). ·NO is a short-lived signal molecule with diverse roles in different cell types: regulating smooth muscle contraction, gene transcription, metabolism, & neurotransmission. Many effects of ·NO arise from its activation of a soluble cytosolic Guanylate Cyclase that synthesizes cyclic-GMP.
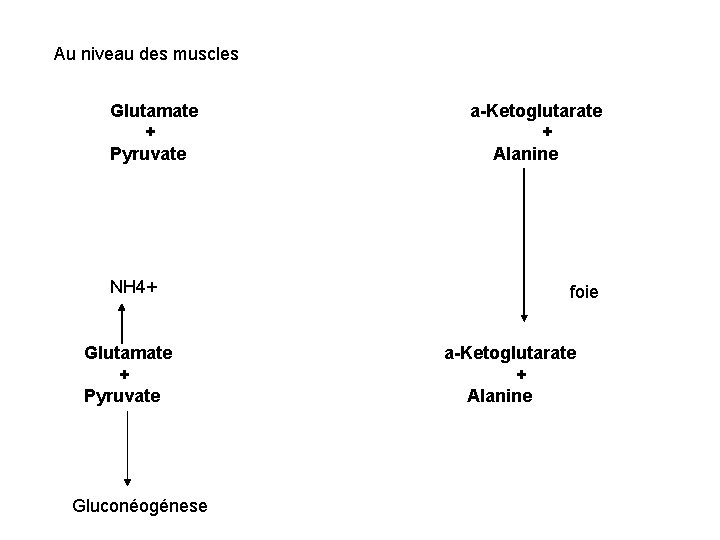
Au niveau des muscles Glutamate + Pyruvate NH 4+ Glutamate + Pyruvate Gluconéogénese a-Ketoglutarate + Alanine foie a-Ketoglutarate + Alanine
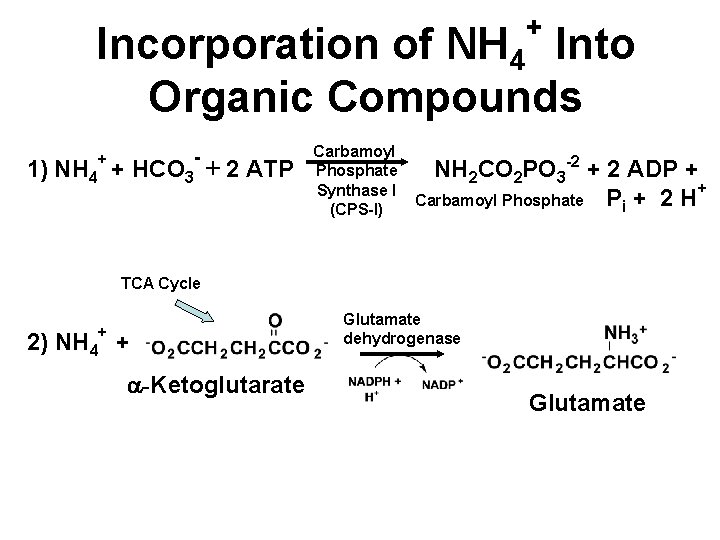
+ Incorporation of NH 4 Into Organic Compounds - + 1) NH 4 + HCO 3 + 2 ATP Carbamoyl Phosphate Synthase I (CPS-I) NH 2 CO 2 PO 3 -2 + 2 ADP + + Carbamoyl Phosphate Pi + 2 H TCA Cycle + 2) NH 4 + a-Ketoglutarate Glutamate dehydrogenase Glutamate
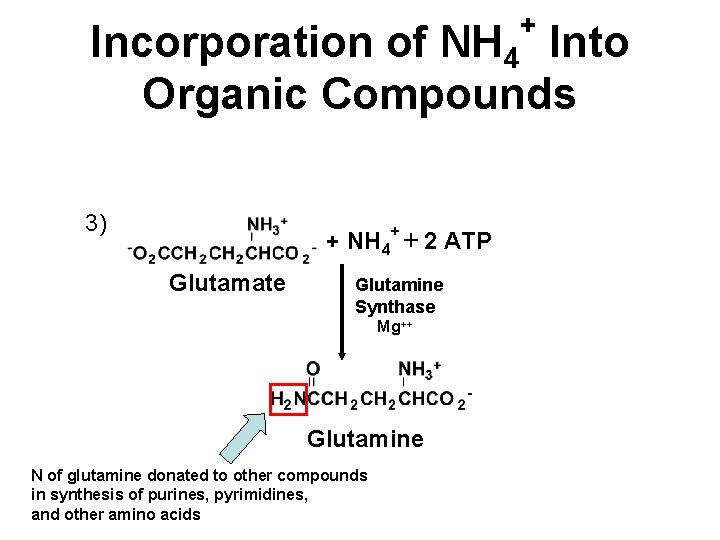
+ Incorporation of NH 4 Into Organic Compounds 3) + NH 4+ + 2 ATP Glutamate Glutamine Synthase Mg++ Glutamine N of glutamine donated to other compounds in synthesis of purines, pyrimidines, and other amino acids
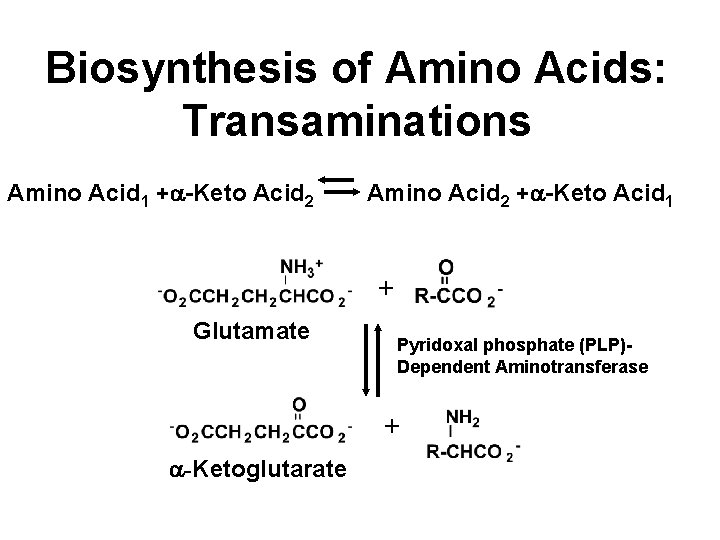
Biosynthesis of Amino Acids: Transaminations Amino Acid 1 +a-Keto Acid 2 Amino Acid 2 +a-Keto Acid 1 + Glutamate Pyridoxal phosphate (PLP)Dependent Aminotransferase + a-Ketoglutarate

Transaminations: Role of PLP Tautomerization
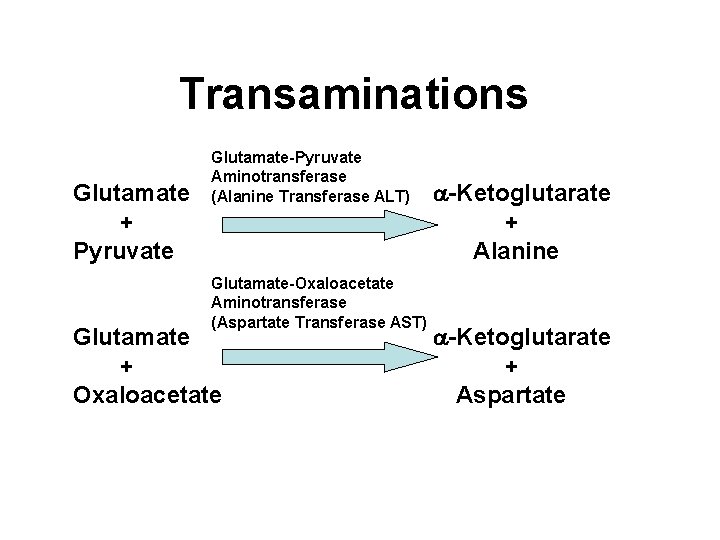
Transaminations Glutamate + Pyruvate Glutamate-Pyruvate Aminotransferase (Alanine Transferase ALT) Glutamate-Oxaloacetate Aminotransferase (Aspartate Transferase AST) Glutamate + Oxaloacetate a-Ketoglutarate + Alanine a-Ketoglutarate + Aspartate

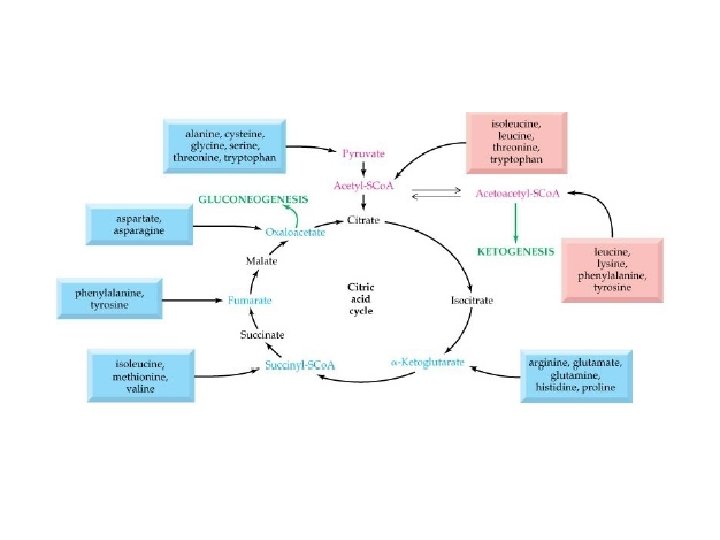
Ketogenic Amino Acids • Metabolized to acetyl Co. A or acetoacetate • Isoleucine • Leucine • Threonine • Tryptophan • Lysine • Phenylalanine • Tyrosine
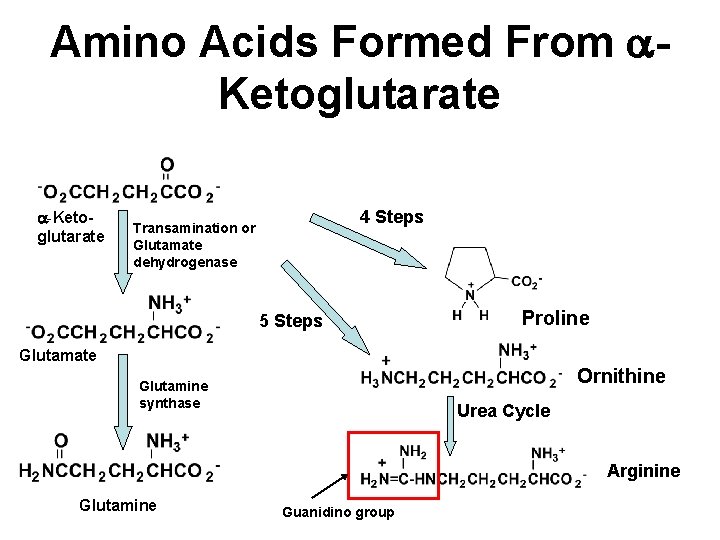
Amino Acids Formed From a. Ketoglutarate a-Ketoglutarate 4 Steps Transamination or Glutamate dehydrogenase 5 Steps Proline Glutamate Ornithine Glutamine synthase Urea Cycle Arginine Glutamine Guanidino group

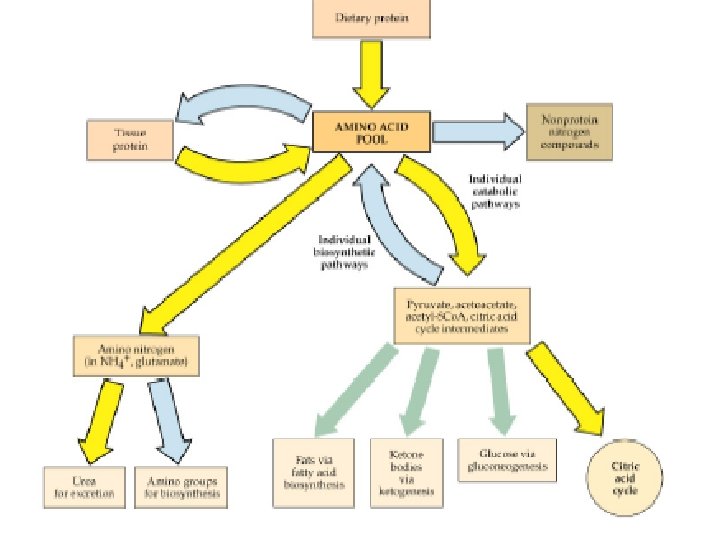
Use of Keto Acids for Energy • Keto acids can – Enter the TCA cycle and be broken down to CO 2 and H 2 O with release of energy – Be used for gluconeogenesis • Some, not all amino acids • In liver (and kidney) – Lipogenesis – Ketogenesis • Ketone bodies (acetoacetate, acetyl-Co. A) • Used as energy source in various tissues

Disposal of NH 3 • NH 3 is very toxic and must be detoxified and excreted from the body – Fish: elimination au niveau des branchies NH 3 Organismes ammnoteliques – Mammals: Urea plus soluble et moins toxique NH 3 Organismes ureoteliques – Birds: Uric acid Organismes ureotiliques • Synthesis of uric acid – Same pathway as for purines • Synthesis of urea—the urea cycle – Detoxifies NH 3 to urea – Synthesizes arginine
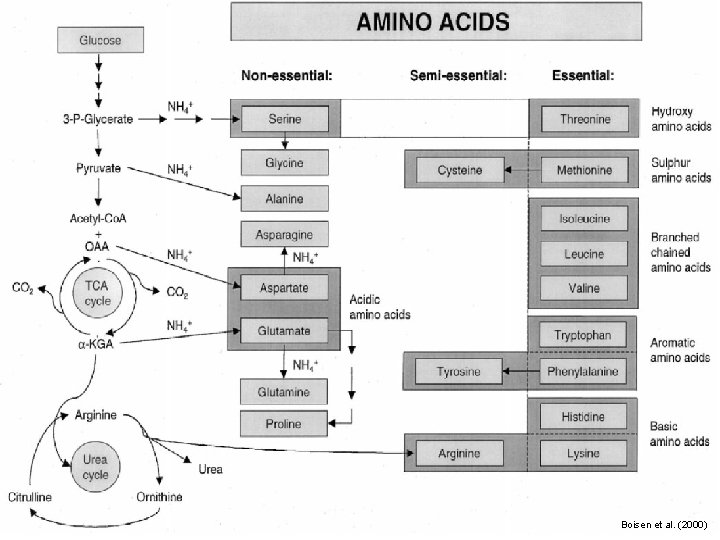
Boisen et al. (2000)

Photosynthesis Light energy is captured and stored as chemical potential energy in the covalent bonds of carbohydrate molecules. The overall chemical reaction for photosynthesis is: 6 CO 2 + 6 H 2 O + light energy C 6 H 12 O 6 + 6 O 2 Photosynthesis occurs in plants, algae, and certain types of bacteria, all of which are autotrophs (literally, “self-feeders”)

Chloroplasts are the sites of photosynthesis in plants. Evidence that chloroplasts split water molecules enabled researchers to track atoms through photosynthesis.
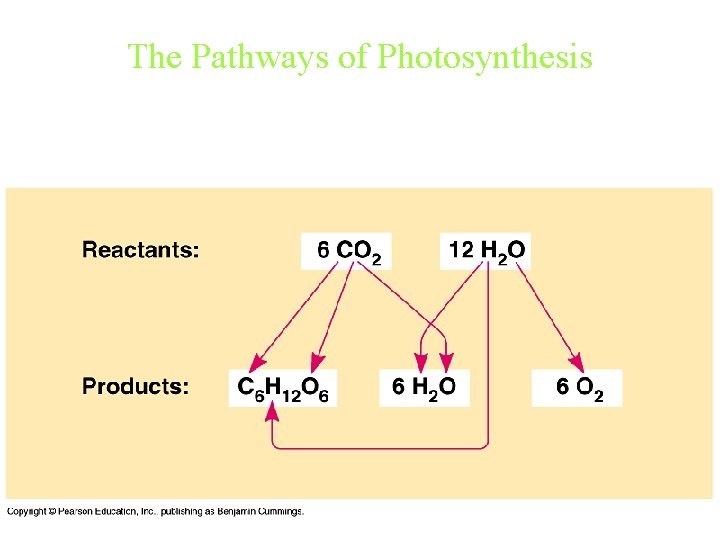
The Pathways of Photosynthesis
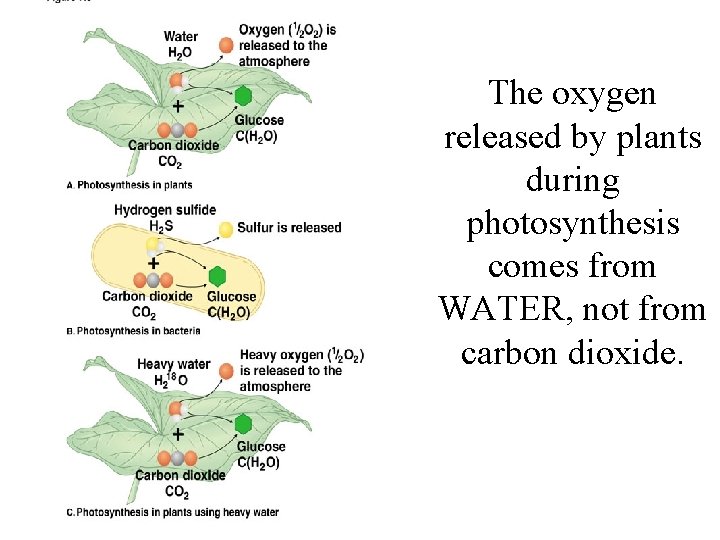
The oxygen released by plants during photosynthesis comes from WATER, not from carbon dioxide.
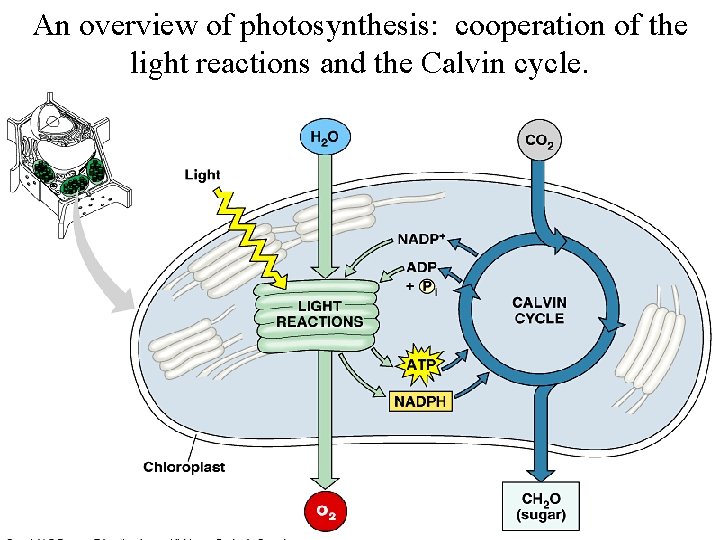
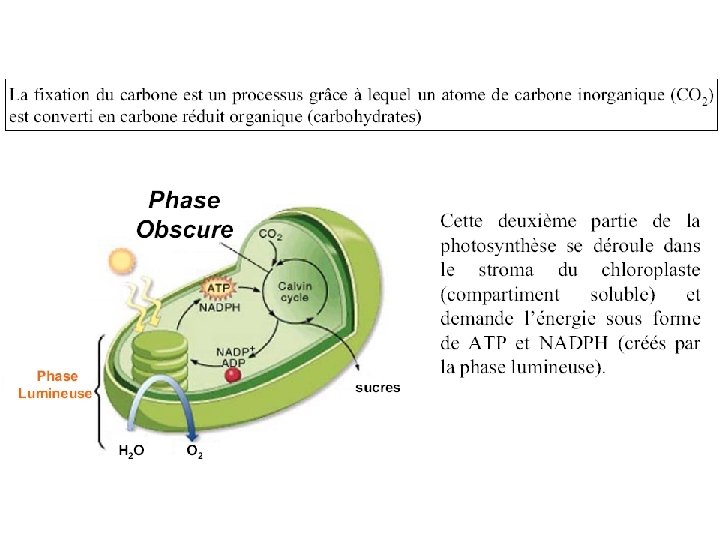
An overview of photosynthesis: cooperation of the light reactions and the Calvin cycle.
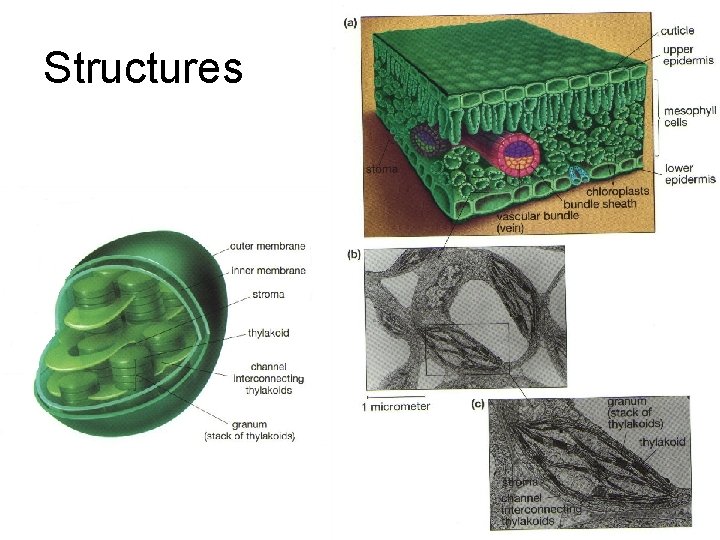
Structures
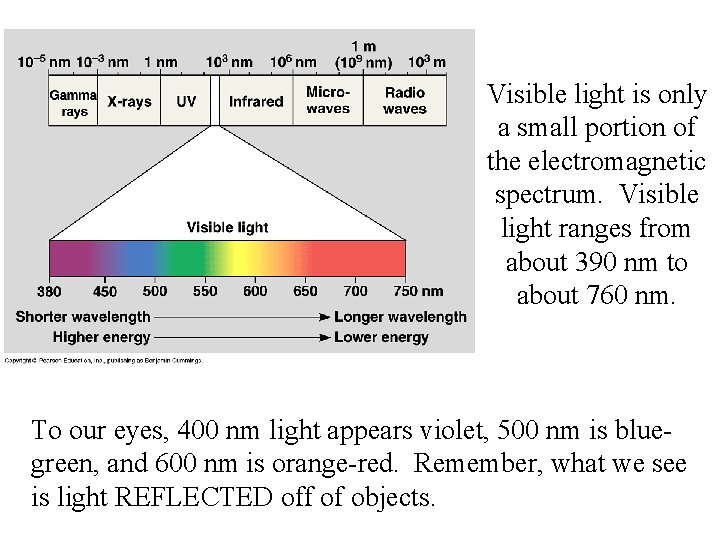
Visible light is only a small portion of the electromagnetic spectrum. Visible light ranges from about 390 nm to about 760 nm. To our eyes, 400 nm light appears violet, 500 nm is bluegreen, and 600 nm is orange-red. Remember, what we see is light REFLECTED off of objects.
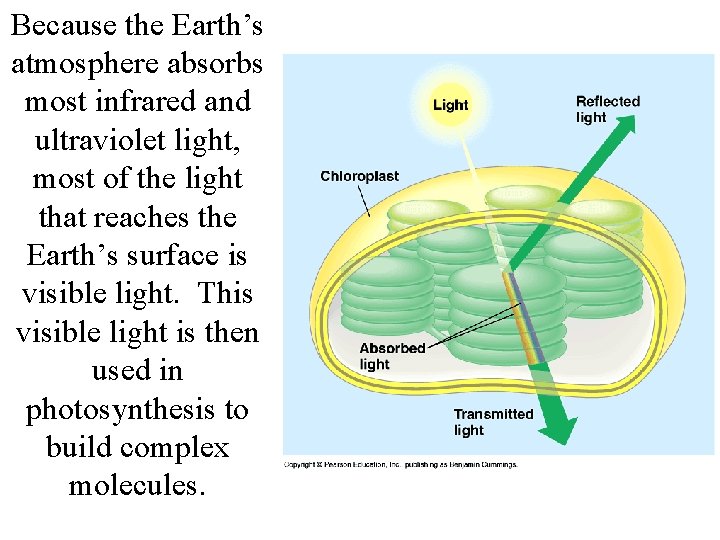
Because the Earth’s atmosphere absorbs most infrared and ultraviolet light, most of the light that reaches the Earth’s surface is visible light. This visible light is then used in photosynthesis to build complex molecules.

Mechanisms of Photosynthesis Substances needed: Photons … sunlight CO 2 … atmosphere H 2 O … soil, rain Chlorophyll … chloroplasts Light-dependent reactions occur in two steps: 1. Conversion of solar energy (photons) to chemical energy (high energy ATP) phosphorylation 2. ATP is used to join CO 2 and H 2 O PGAL Glucose

Light-dependent reactions • Are carried out by molecules in the thylakoid membranes. • Convert light energy to the chemical energy of ATP and NADPH • Split H 2 O and release O 2 atmosphere. • Involves two PHOTOSYSTEMS: • Photosystem I absorbs light at 700 nm and is called P 700 (P stands for “pigment”. ) • Photosystem II absorbs light at 680 nm and is called P 680
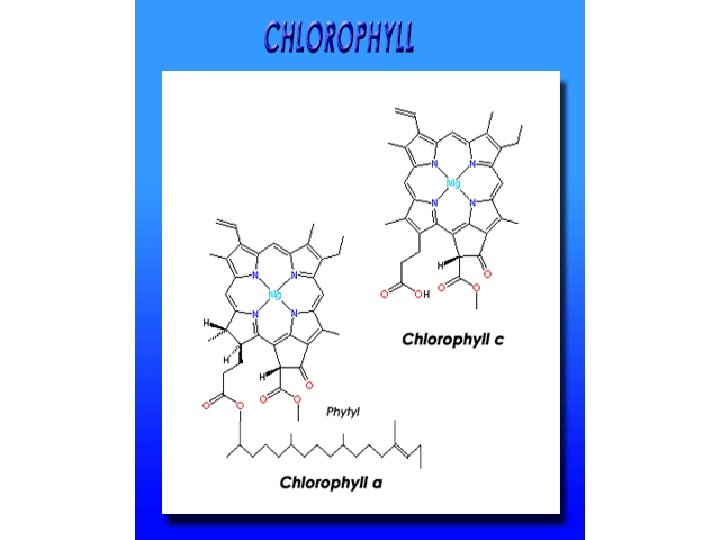
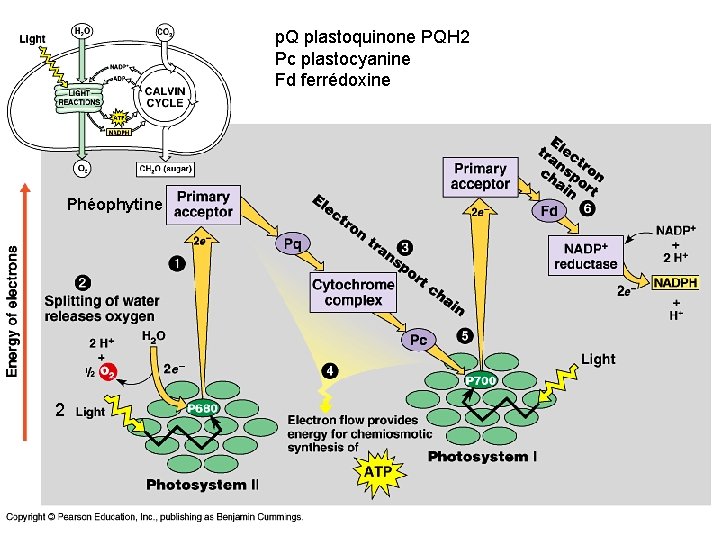
p. Q plastoquinone PQH 2 Pc plastocyanine Fd ferrédoxine Phéophytine 2
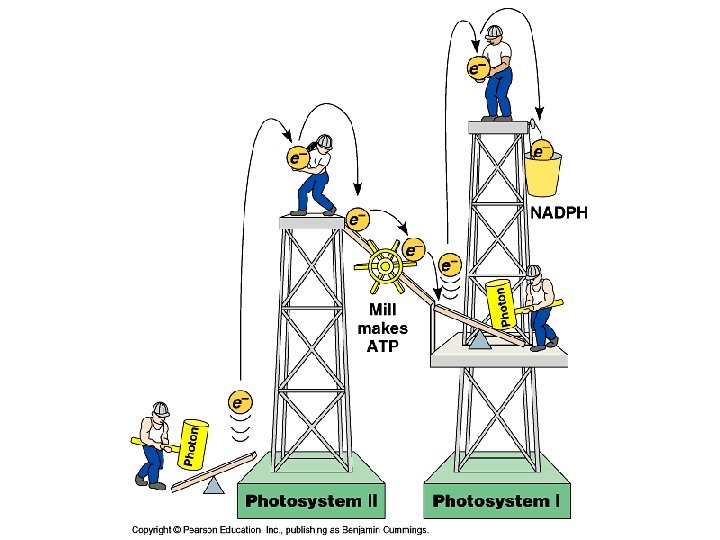
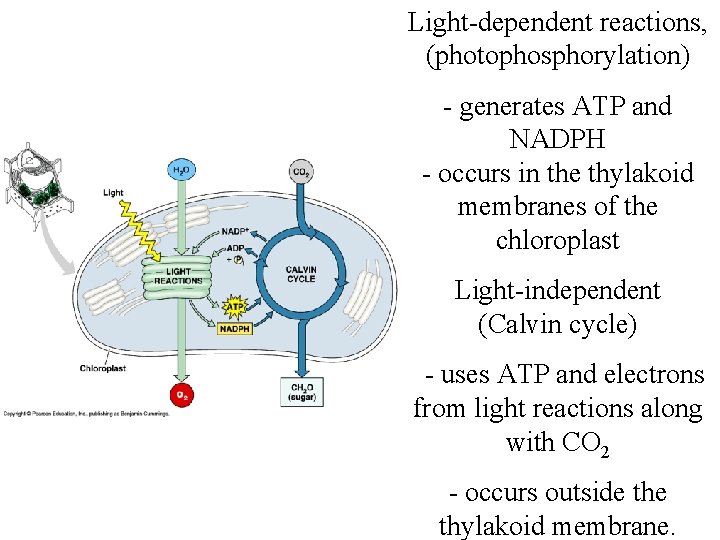
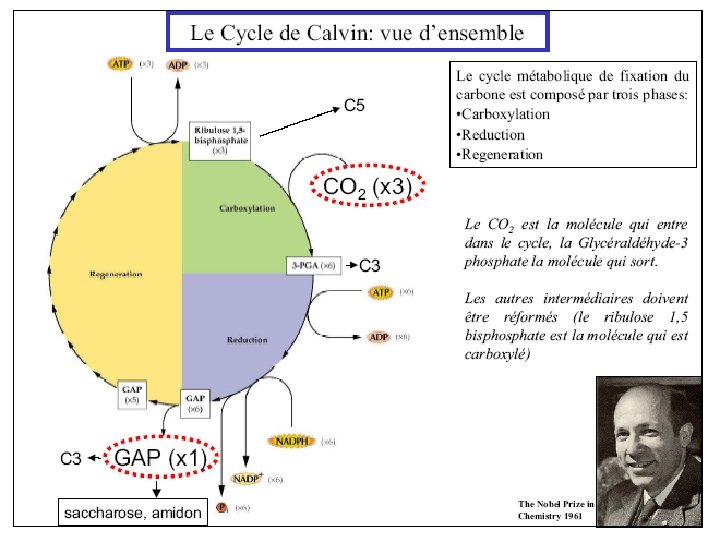
Light-dependent reactions, (photophosphorylation) - generates ATP and NADPH - occurs in the thylakoid membranes of the chloroplast Light-independent (Calvin cycle) - uses ATP and electrons from light reactions along with CO 2 - occurs outside thylakoid membrane.
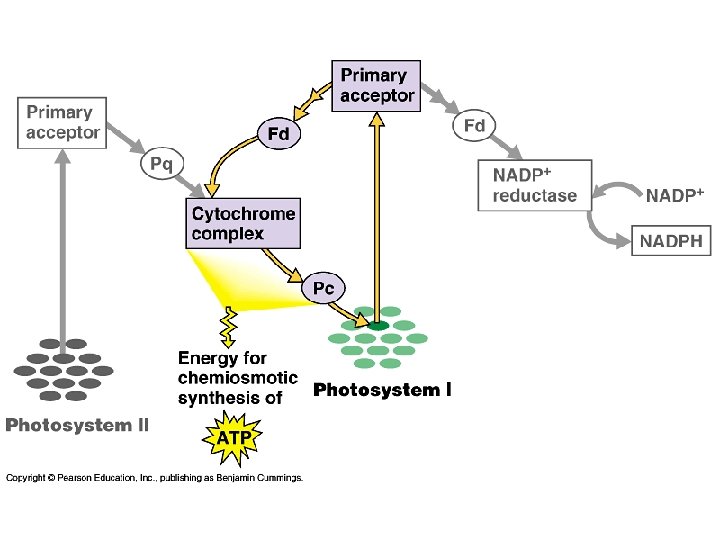
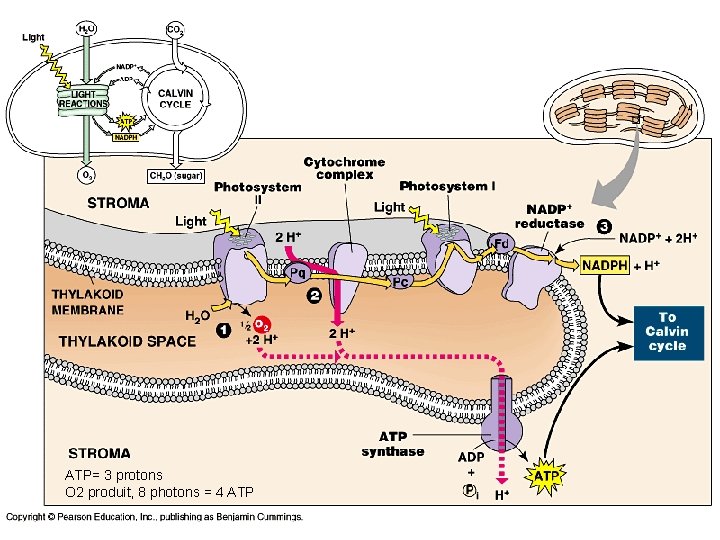
ATP= 3 protons O 2 produit, 8 photons = 4 ATP
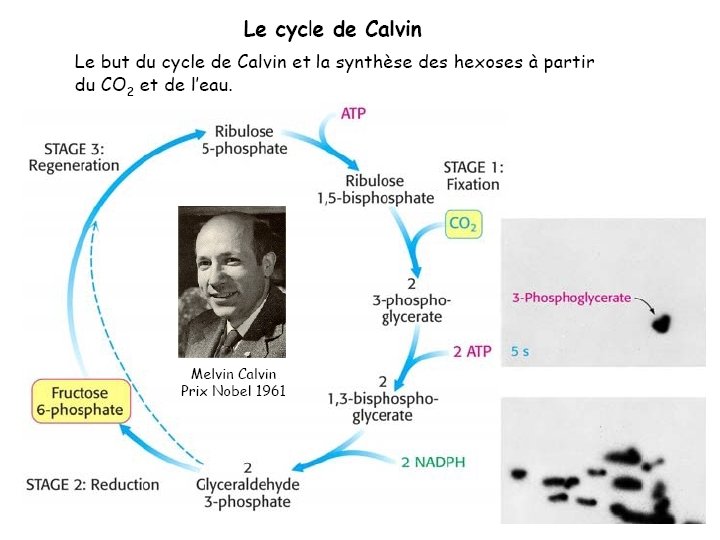
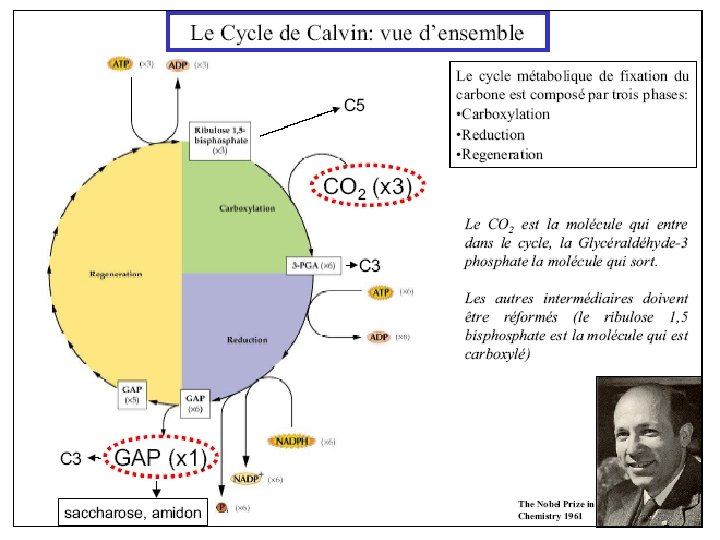


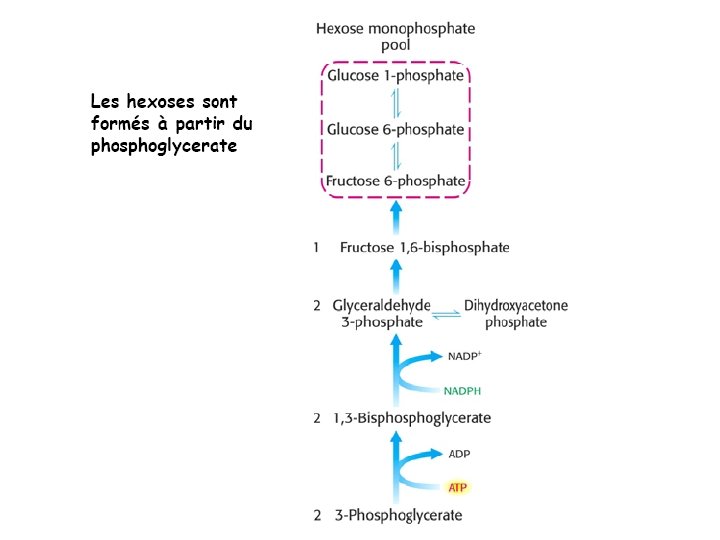
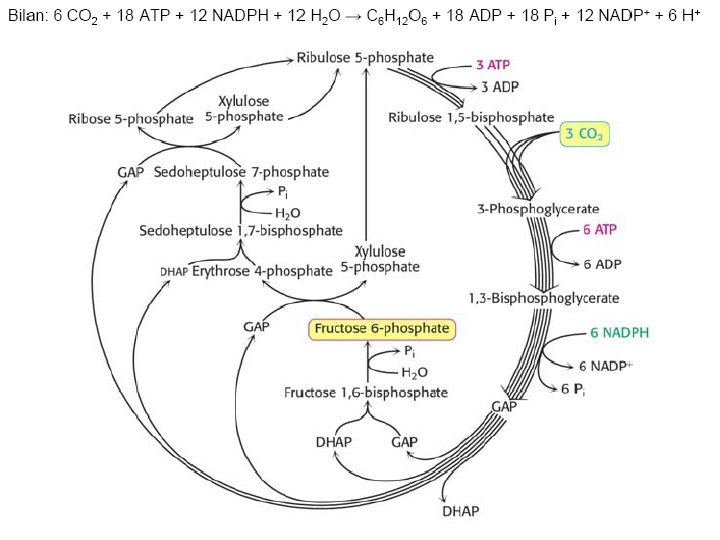
Ru. BP Carboxylase - postulated mechanism: Extraction of H+ from C 3 of ribulose-1, 5 -bisphosphate promotes formation of an enediolate intermediate. Nucleophilic attack on CO 2 leads to formation of a b-keto acid intermediate, that reacts with water and cleaves to form 2 molecules of 3 phosphoglycerate.
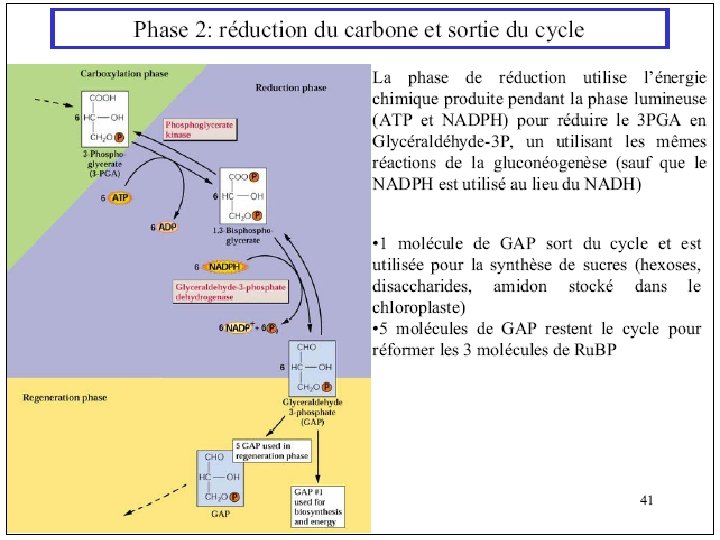

- Slides: 111