Types of Polymorphisms I Proteinenzyme polymorphisms Blood groups

Types of Polymorphisms I. Protein/enzyme polymorphisms Blood groups II. DNA Polymorphisms 1. Single Nucleotide Polymorphisms (SNP) 2. Tandem Repeat Polymorphisms • Microsattelites, Short Sequence Repeats (SSR) • Variable number of tandem repeats (VNTR) 3. Structural Variants • Insertion/Deletion/Inversion/Duplication/Translocation • Copy Number Variants (CNV)
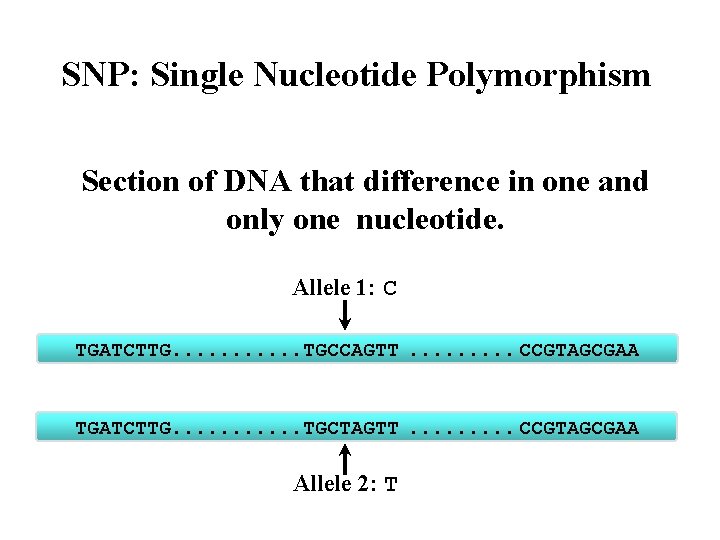
SNP: Single Nucleotide Polymorphism Section of DNA that difference in one and only one nucleotide. Allele 1: C TGATCTTG. . . TGCCAGTT . . CCGTAGCGAA TGATCTTG. . . TGCTAGTT . . CCGTAGCGAA Allele 2: T

Tandem Repeat Polymorphisms: A nucleotide sequence is repeated over and over again and the polymorphism is in the number of times it is repeated.
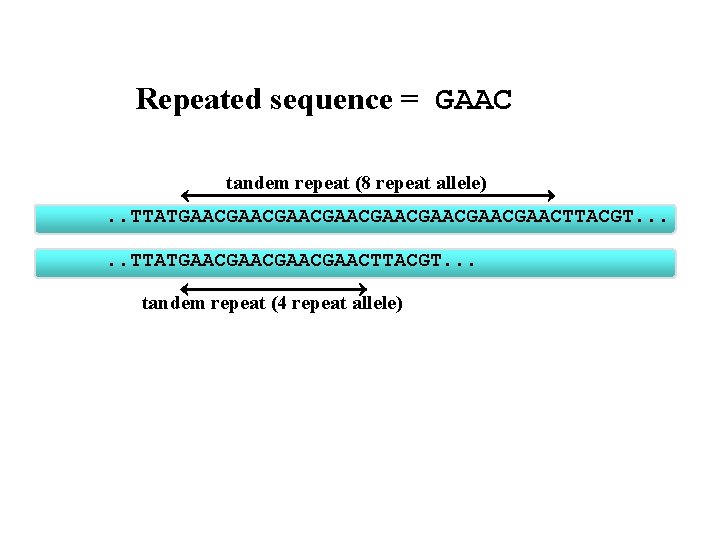
Repeated sequence = GAAC tandem repeat (8 repeat allele). . TTATGAACGAACGAACGAACTTACGT. . . TTATGAACGAACTTACGT. . . tandem repeat (4 repeat allele)
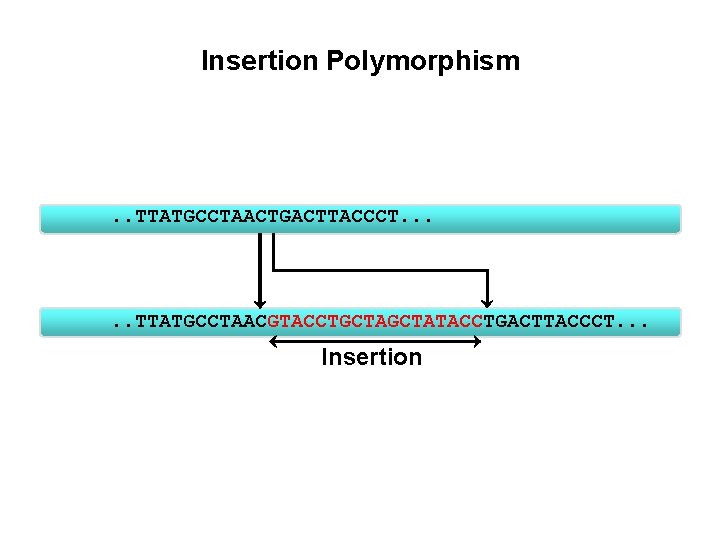
Insertion Polymorphism . . TTATGCCTAACTGACTTACCCT. . . TTATGCCTAACGTACCTGCTATACCTGACTTACCCT. . . Insertion
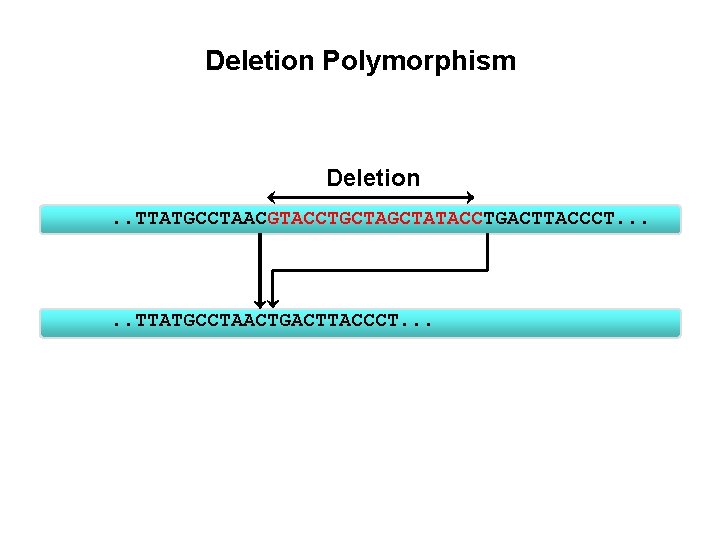
Deletion Polymorphism Deletion. . TTATGCCTAACGTACCTGCTATACCTGACTTACCCT. . . TTATGCCTAACTGACTTACCCT. . .
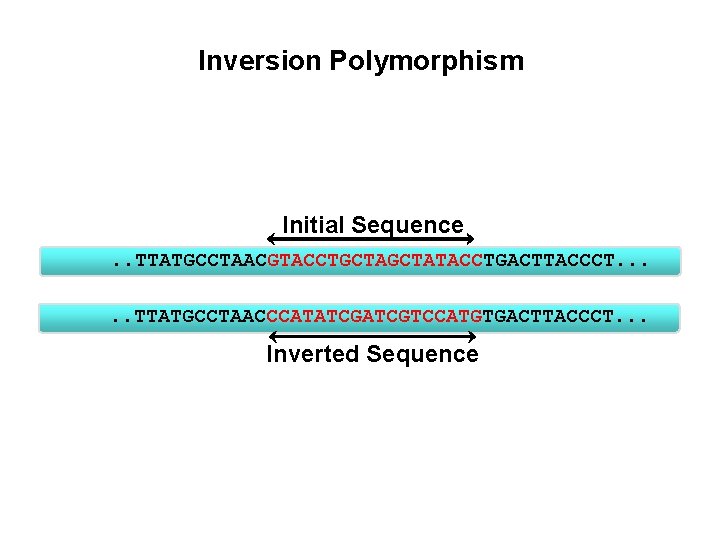
Inversion Polymorphism Initial Sequence. . TTATGCCTAACGTACCTGCTATACCTGACTTACCCT. . . TTATGCCTAACCCATATCGTCCATGTGACTTACCCT. . . Inverted Sequence
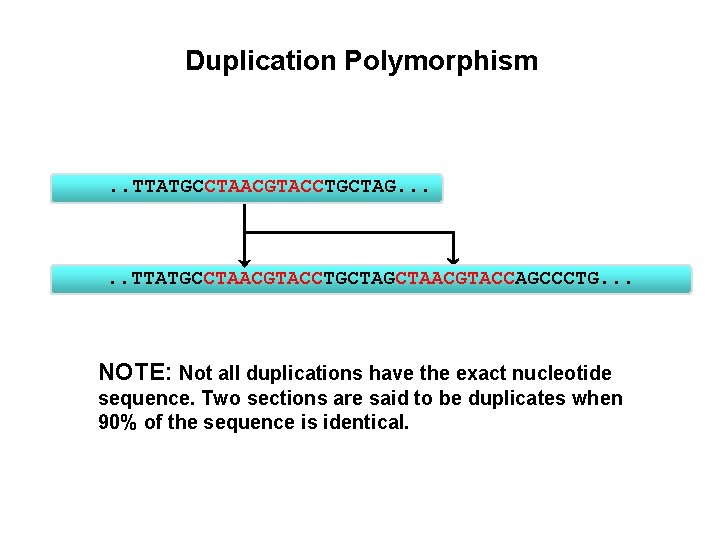
Duplication Polymorphism . . TTATGCCTAACGTACCTGCTAG. . . TTATGCCTAACGTACCTGCTAACGTACCAGCCCTG. . . NOTE: Not all duplications have the exact nucleotide sequence. Two sections are said to be duplicates when 90% of the sequence is identical.
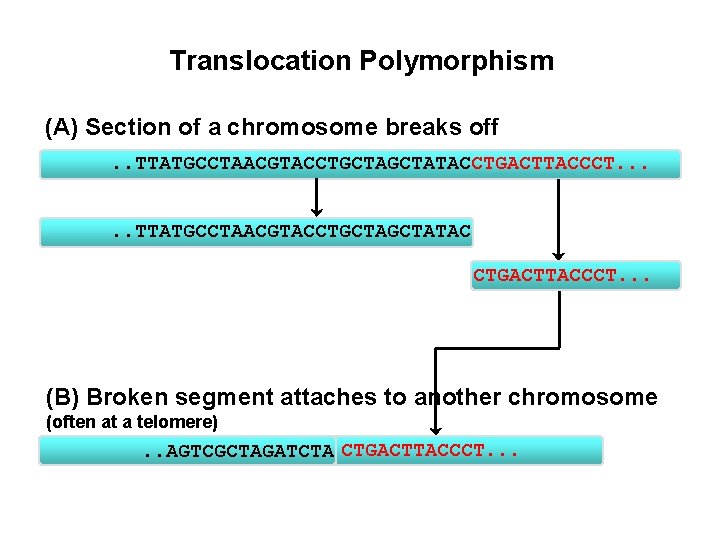
Translocation Polymorphism (A) Section of a chromosome breaks off. . TTATGCCTAACGTACCTGCTATACCTGACTTACCCT. . . TTATGCCTAACGTACCTGCTATAC CTGACTTACCCT. . . (B) Broken segment attaches to another chromosome (often at a telomere) . . AGTCGCTAGATCTA CTGACTTACCCT. . .

Copy Number Variant (CNV) (1) Somewhat long (> 1 kb) section of DNA that is repeated throughout the genome with variable copy numbers. (2) Repeats do not have to be in tandem. (3) Includes long insertions, deletions, and duplications.
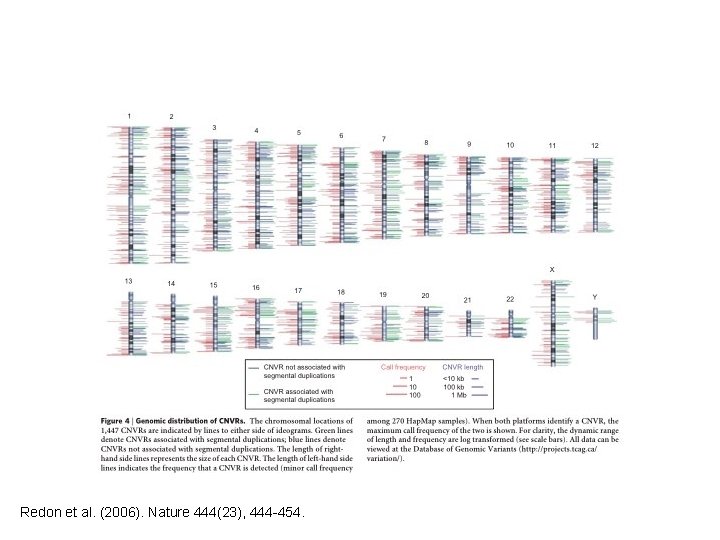
Redon et al. (2006). Nature 444(23), 444 -454.

Tools in Molecular Genetics: 1. Electrophoresis 2. Probes 3. Polymerase Chain Reaction 4. Restriction Enzyme 5. Dideoxy Nucleotides 6. DNA Arrays (Gene Chips)
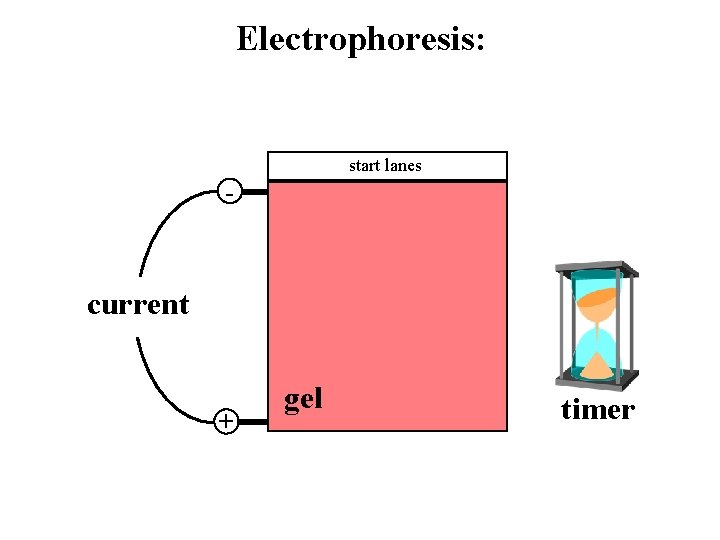
Electrophoresis: start lanes - current + gel timer
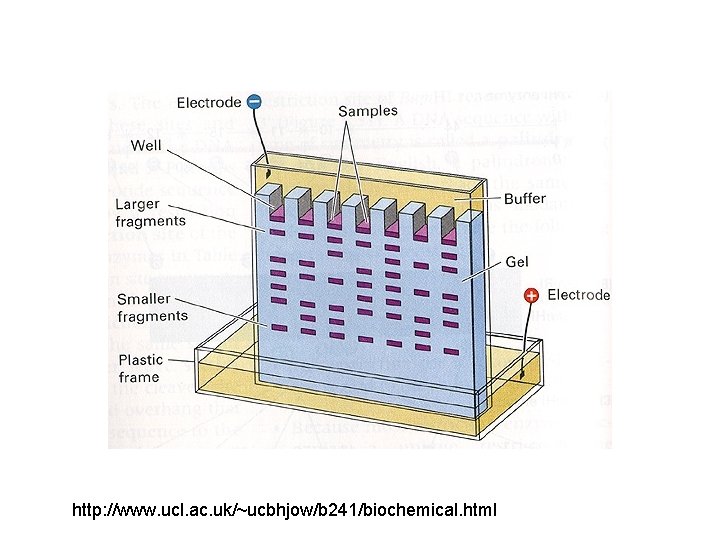
http: //www. ucl. ac. uk/~ucbhjow/b 241/biochemical. html
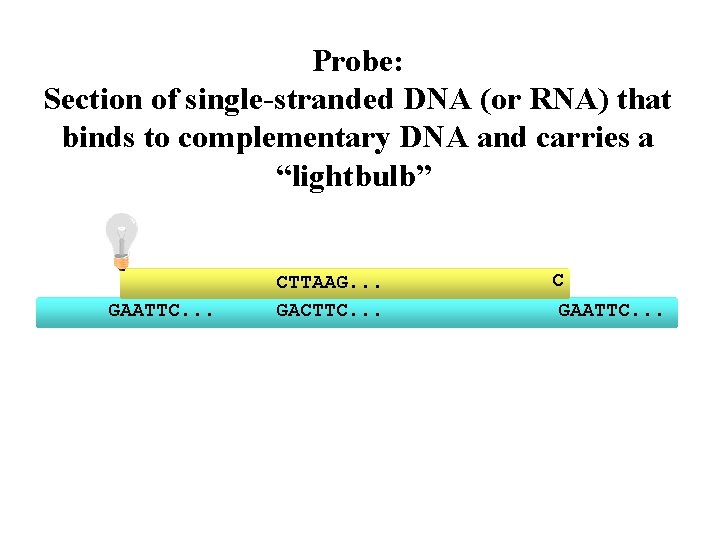
Probe: Section of single-stranded DNA (or RNA) that binds to complementary DNA and carries a “lightbulb” TTAAG. . . GAATTC. . . CTTAAG. . . GACTTC. . . C GAATTC. . .

PCR: Polymerase Chain Reaction Purpose = Make a lot of copies of a desired piece of DNA (i. e. , “amplify” the DNA)
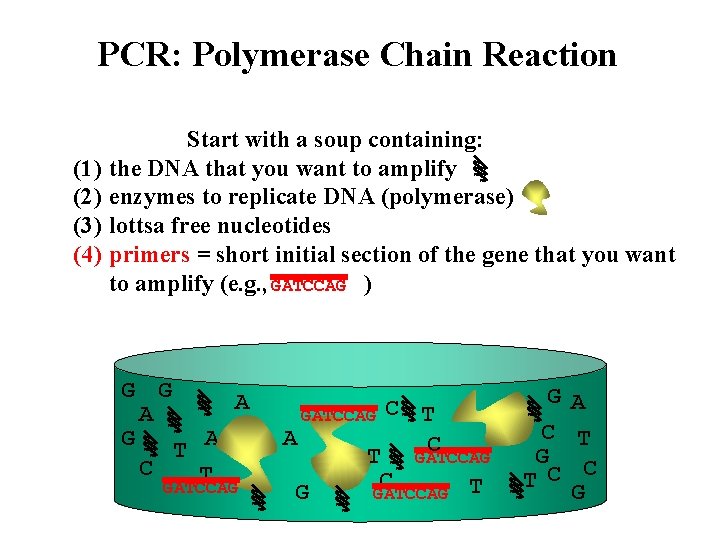
PCR: Polymerase Chain Reaction (1) (2) (3) (4) Start with a soup containing: the DNA that you want to amplify enzymes to replicate DNA (polymerase) lottsa free nucleotides primers = short initial section of the gene that you want to amplify (e. g. , GATCCAG ) G G A A G T A C T GATCCAG C T G GATCCAG G C G T C A T C G

PCR: Polymerase Chain Reaction Procedure: 1. Heat the mixture. Just before the boiling point of water, the DNA will become single-stranded. 2. Cool the mixture. As the mixture cools, the primer will bind to the DNA and the polymerase will synthesize a new strand for each strand of DNA. 3. Repeat steps 1 and 2 until a sufficient amount of the desired gene is available for analysis
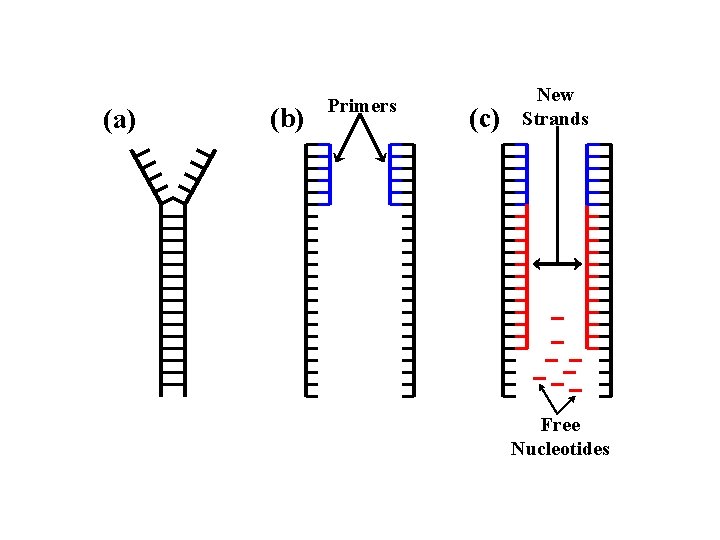
(a) (b) Primers (c) New Strands Free Nucleotides
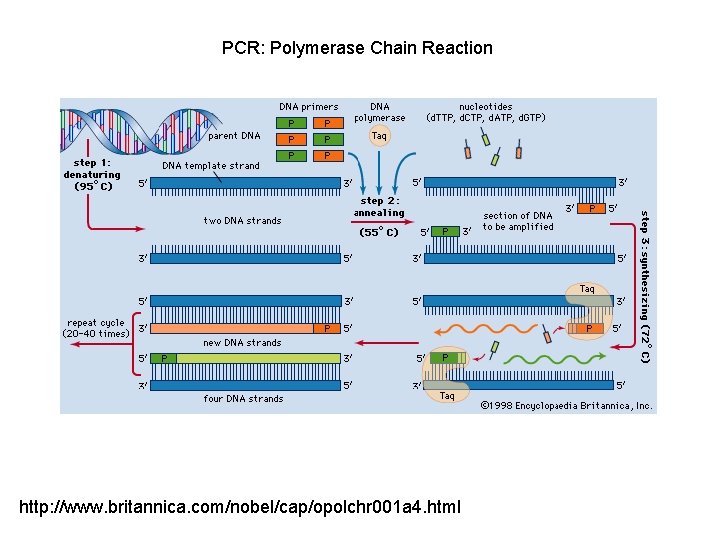
PCR: Polymerase Chain Reaction http: //www. britannica. com/nobel/cap/opolchr 001 a 4. html

RFLP: Restriction Fragment Length Polymorphism based on whether or not a restriction enzyme cuts a section of DNA. Usually two alleles: (1) the restriction enzyme does cut in the middle of the genes; and (2) the restriction enzyme does not cut in the middle of the gene. restriction enzyme = enzyme that recognizes a specific nucleotide sequence and cuts the DNA at that sequence.
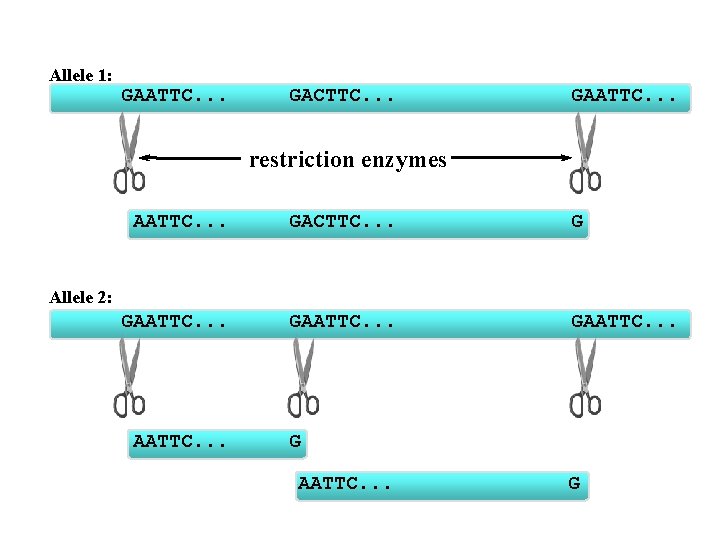
Allele 1: GAATTC. . . GACTTC. . . GAATTC. . . restriction enzymes AATTC. . . GACTTC. . . G GAATTC. . . Allele 2: AATTC. . . G
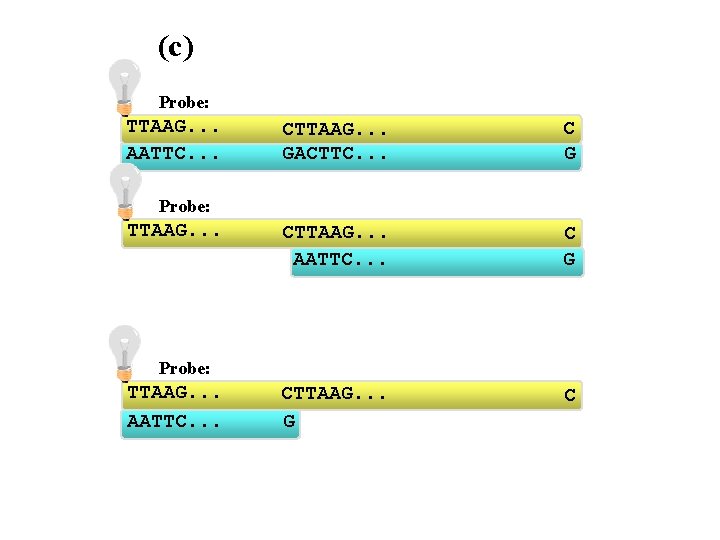
(c) Probe: TTAAG. . . AATTC. . . CTTAAG. . . GACTTC. . . C G CTTAAG. . . AATTC. . . C G CTTAAG. . . G C Probe: TTAAG. . . AATTC. . .
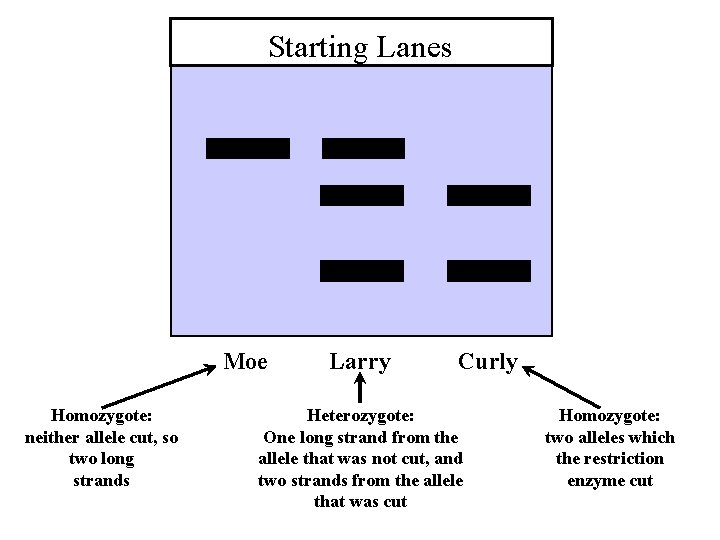
Starting Lanes Moe Homozygote: neither allele cut, so two long strands Larry Curly Heterozygote: One long strand from the allele that was not cut, and two strands from the allele that was cut Homozygote: two alleles which the restriction enzyme cut

The “Olden” Days

Current Technology 1. PCR that includes dideoxy nucleotides with ordinary nucleotides. 2. Have a laser scan the fragments from electrophoresis. dideoxy nucleotides = “color coded” nucleotides (e. g. , A, T, C, G) that stop the synthesis of a new DNA chain when they are inserted into the chain.
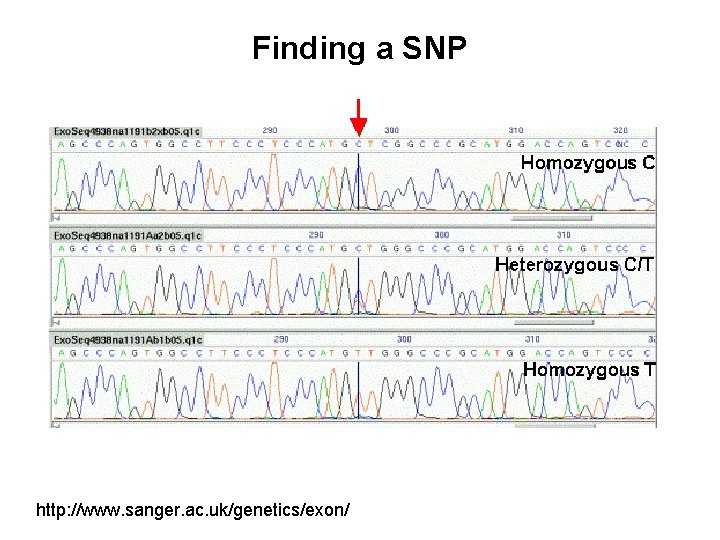
Finding a SNP http: //www. sanger. ac. uk/genetics/exon/

SNP Genotyping Machine (7000 genotypes per day) http: //www. wtcrf. ed. ac. uk/genetics/images/snp 1. jpg Wellcome Trust Clinical Research Facilities

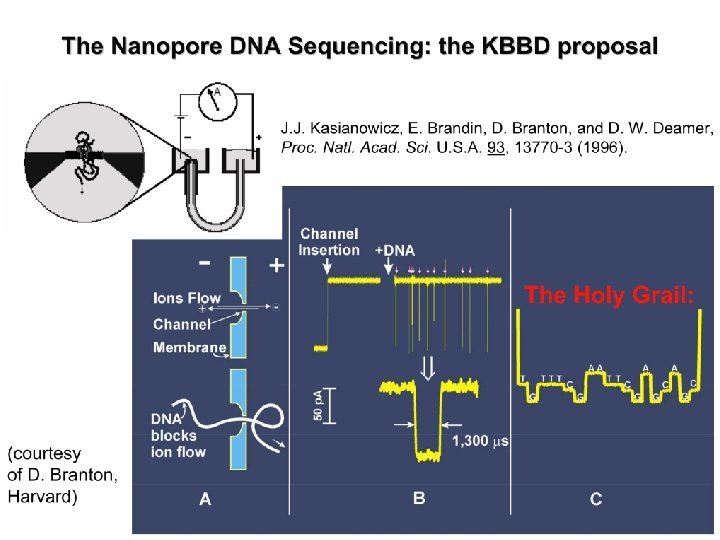
Potential New Method: Nanopores

How the Human Genome was Sequenced: (See Text)
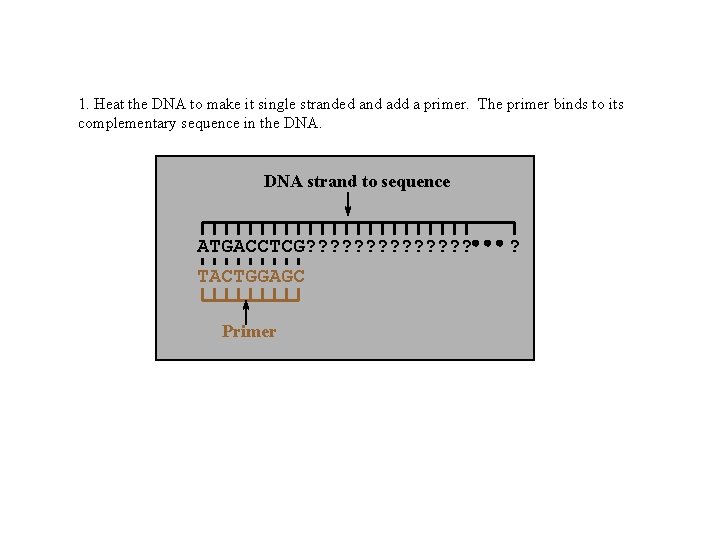
1. Heat the DNA to make it single stranded and add a primer. The primer binds to its complementary sequence in the DNA strand to sequence ATGACCTCG? ? ? ? TACTGGAGC Primer ?
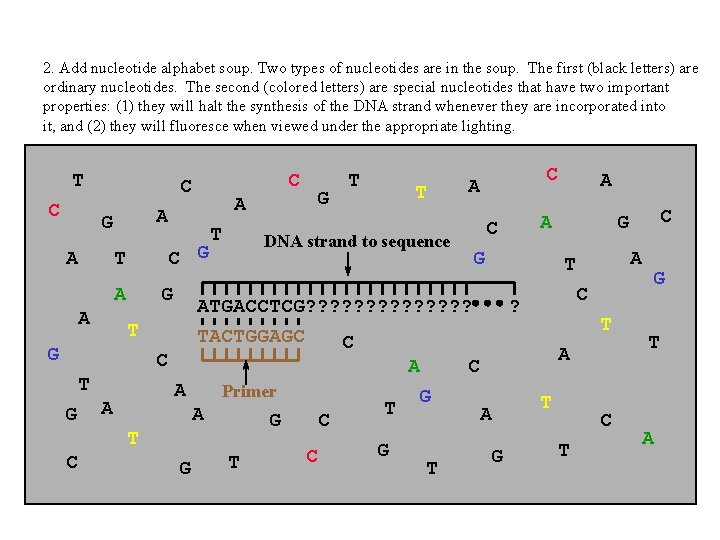
2. Add nucleotide alphabet soup. Two types of nucleotides are in the soup. The first (black letters) are ordinary nucleotides. The second (colored letters) are special nucleotides that have two important properties: (1) they will halt the synthesis of the DNA strand whenever they are incorporated into it, and (2) they will fluoresce when viewed under the appropriate lighting. T C A C G T A A G G C T A A T T C A Primer A G G T C C T G A G T A A C DNA strand to sequence T C G T G A C ? T A C A G C G T ATGACCTCG? ? ? ? TACTGGAGC C T G G C C T G T A
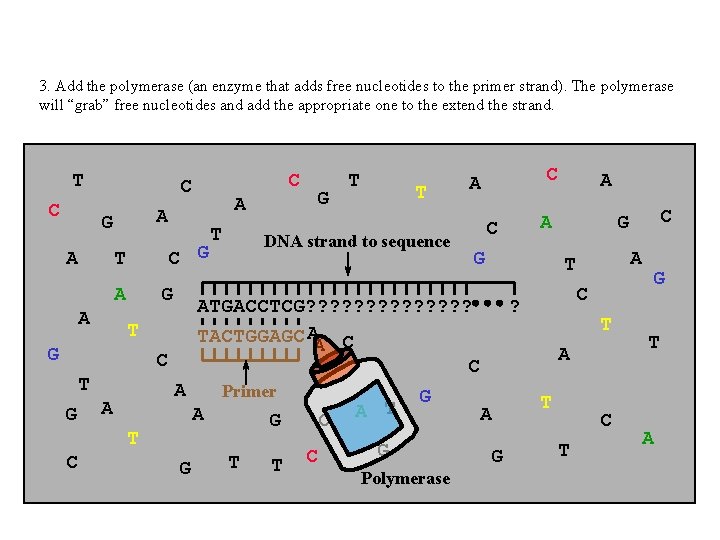
3. Add the polymerase (an enzyme that adds free nucleotides to the primer strand). The polymerase will “grab” free nucleotides and add the appropriate one to the extend the strand. T C A C G T A A G G C T Primer A G G T T C C A T G G Polymerase A A C DNA strand to sequence T C A G A C ? T A A G C G T C A A T G T ATGACCTCG? ? ? ? TACTGGAGC AA C T G G C C T G T A
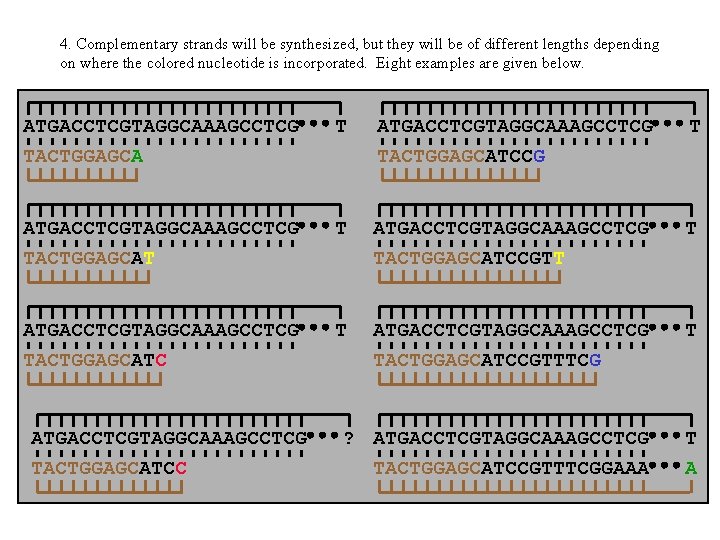
4. Complementary strands will be synthesized, but they will be of different lengths depending on where the colored nucleotide is incorporated. Eight examples are given below. ATGACCTCGTAGGCAAAGCCTCG TACTGGAGCA T ATGACCTCGTAGGCAAAGCCTCG TACTGGAGCATCCGTT T ATGACCTCGTAGGCAAAGCCTCG TACTGGAGCATCCGTTTCG T ? ATGACCTCGTAGGCAAAGCCTCG TACTGGAGCATCCGTTTCGGAAA T A ATGACCTCGTAGGCAAAGCCTCG TACTGGAGCATCC
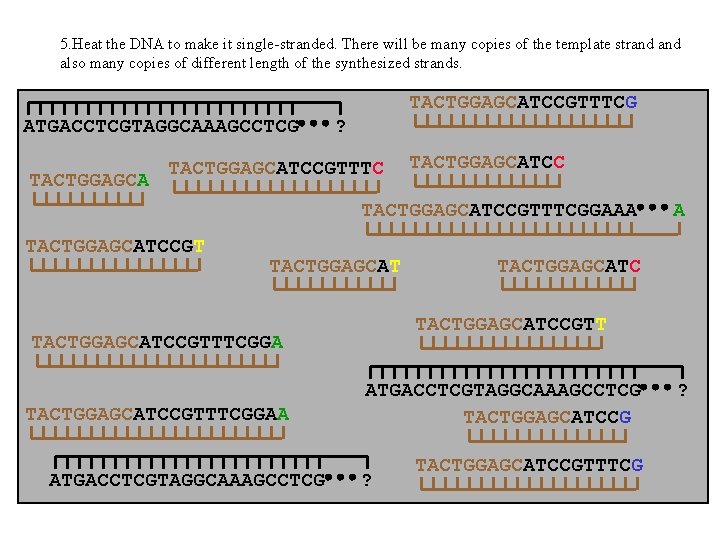
5. Heat the DNA to make it single-stranded. There will be many copies of the template strand also many copies of different length of the synthesized strands. TACTGGAGCATCCGTTTCG ATGACCTCGTAGGCAAAGCCTCG TACTGGAGCA ? TACTGGAGCATCCGTTTCGGAAA TACTGGAGCATCCGT TACTGGAGCAT ATGACCTCGTAGGCAAAGCCTCG TACTGGAGCATCCGTTTCGGAA A ATGACCTCGTAGGCAAAGCCTCG TACTGGAGCATCCG ? TACTGGAGCATCCGTTTCG ?

6. Use electrophoresis to separate the strands according to size. TACTGGAGCATCCGTTTCGGAAA TACTGGAGCATCCGTTTCGG TACTGGAGCATCCGTTTC TACTGGAGCATCCGTTT TACTGGAGCATCCGT TACTGGAGCATCCG TACTGGAGCATCC TACTGGAGCAT TACTGGAGCA A

7. Viewing the gel under a special light allows the colored nucleotides to fluoresce. This lights up the band. The color-coding permits the DNA sequence to be read. TACTGGAGCATCCGTTTCGGAAA TACTGGAGCATCCGTTTCGG TACTGGAGCATCCGTTTC TACTGGAGCATCCGTTT TACTGGAGCATCCGT TACTGGAGCATCCG TACTGGAGCATCC TACTGGAGCAT TACTGGAGCA A
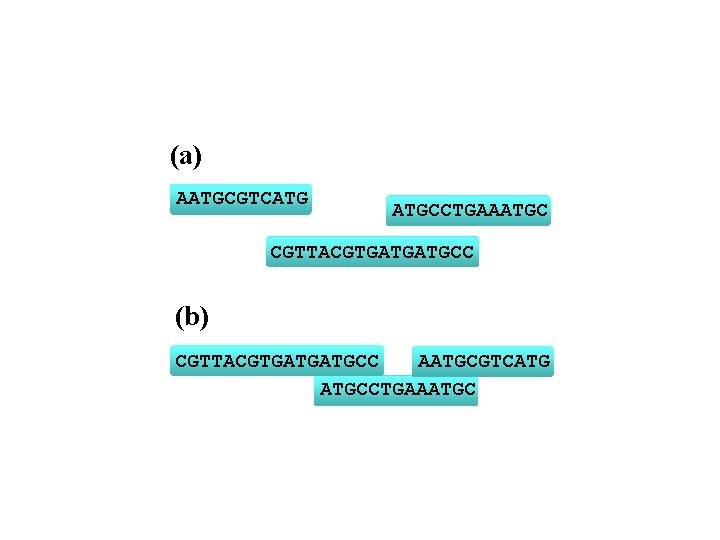
(a) AATGCGTCATG ATGCCTGAAATGC CGTTACGTGATGATGCC (b) AATGCGTCATG CGTTACGTGATGATGCCTGAAATGC
- Slides: 39