Tutorials for protein data bank and swiss PDB

Tutorials for protein data bank and swiss PDB viewer 2010/04/19 Prof. Jinn-Moon Yang Yen-Fu Chen and Kai-Cheng Hsu http: //gemdock. life. nctu. edu. tw/dock/download/20100419_spv. ppt http: //gemdock. life. nctu. edu. tw/dock/download/20100419_spv. pdf http: //ppt. cc/cnq. T (ppt) http: //ppt. cc/WWxd (pdf)

Contents Introduction of protein structures Using thymidine kinase as an example Download and install Tutorial Download protein structures from PDB Basic Operation Advance Operation Resources for tutorial http: //www. youtube. com/watch? v=y. FE 3 CAHNk. Zg&feat ure=related
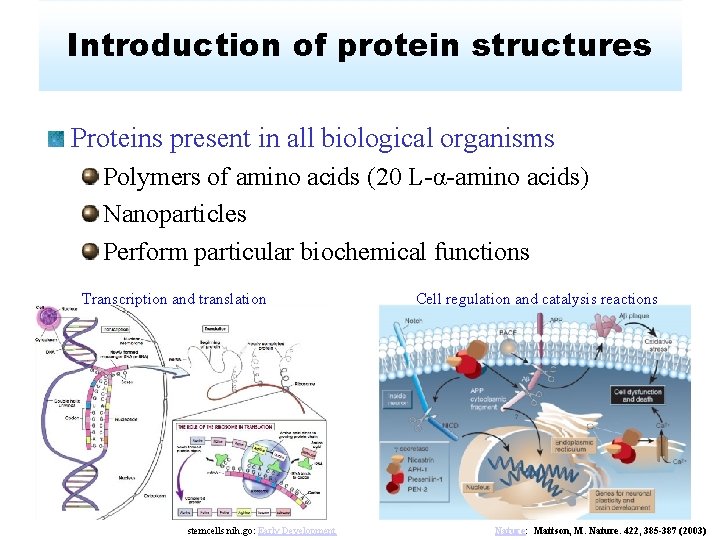
Introduction of protein structures Proteins present in all biological organisms Polymers of amino acids (20 L-α-amino acids) Nanoparticles Perform particular biochemical functions Transcription and translation stemcells. nih. go: Early Development Cell regulation and catalysis reactions Nature: Mattson, M. Nature. 422, 385 -387 (2003)
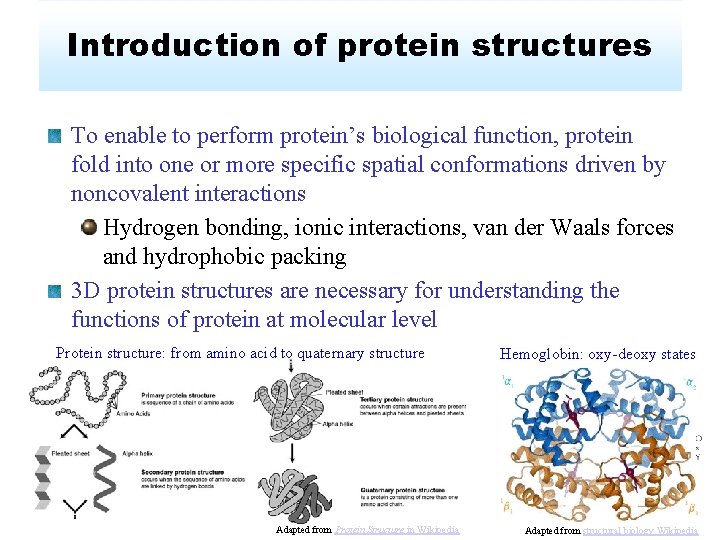
Introduction of protein structures To enable to perform protein’s biological function, protein fold into one or more specific spatial conformations driven by noncovalent interactions Hydrogen bonding, ionic interactions, van der Waals forces and hydrophobic packing 3 D protein structures are necessary for understanding the functions of protein at molecular level Protein structure: from amino acid to quaternary structure Adapted from Protein Structure in Wikipedia Hemoglobin: oxy-deoxy states Adapted from structural biology Wikipedia
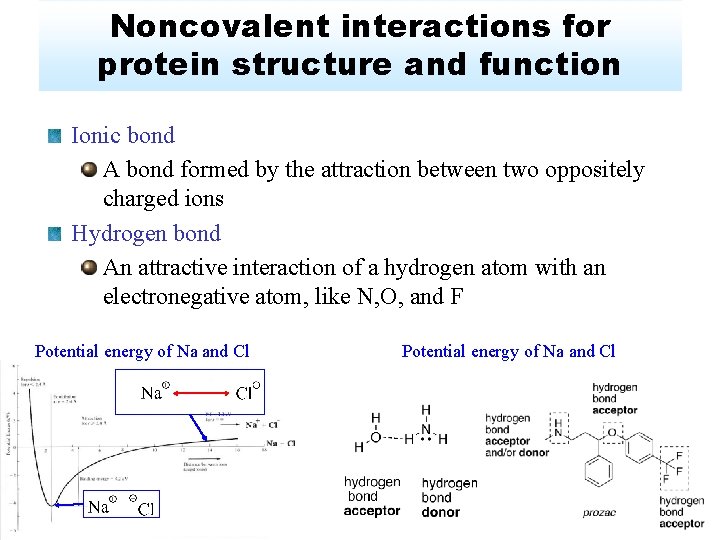
Noncovalent interactions for protein structure and function Ionic bond A bond formed by the attraction between two oppositely charged ions Hydrogen bond An attractive interaction of a hydrogen atom with an electronegative atom, like N, O, and F Potential energy of Na and Cl

Noncovalent interactions for protein structure and function van der Waals force Attractive or repulsive force between molecules Hydrophobic interaction The physical property of a molecule (known as a hydrophobe) that is repelled from a mass of water An example of van der Waals force: Gecko climbs on the glass An example of hydrophobic interaction: Water drops on hydrophobic surface

Protein structure database: Protein data bank (PDB) Techniques for determining atomic structures X-ray crystallography, NMR spectroscopy and electron microscopy PDB contains information about experimentallydetermined structures of biological marcomoleculeas (proteins, and DNA/RNA) Proteins (1 kim) X-ray NMR DNAs/RNAs (2 k 7 e) EM http: //www. pdb. org/ Biological complexes (1 zrc)

Search protein structures in PDB provides search by protein name, ligand, or structrue related keywords Search example: thymidine kinase (TK) • Function: DNA synthesis • Therapeutic: Anticancer and antivirus drug target

Example: X-ray structures of virus’ thymidine kinase with substrates/inhibitors Protein name Source spices Experimental method has ligands
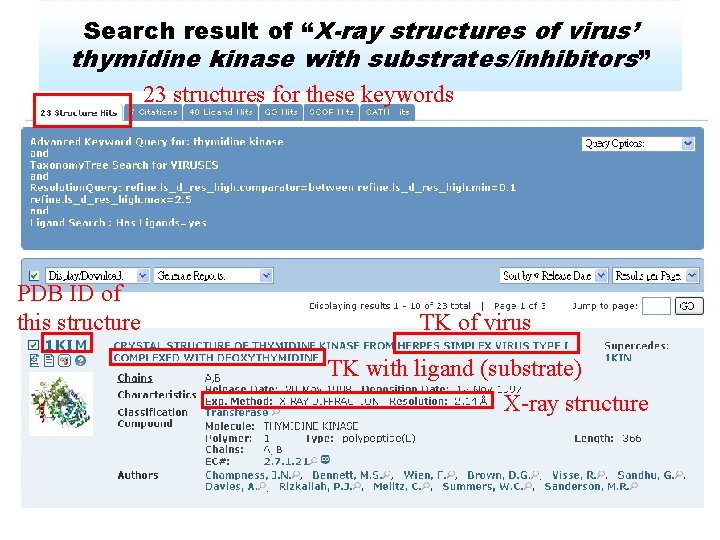
Search result of “X-ray structures of virus’ thymidine kinase with substrates/inhibitors” 23 structures for these keywords PDB ID of this structure TK of virus TK with ligand (substrate) X-ray structure

Structure and related data (1 kim) The title of this structure The citation of this structure Related data of this structure Visualization of biological assembly
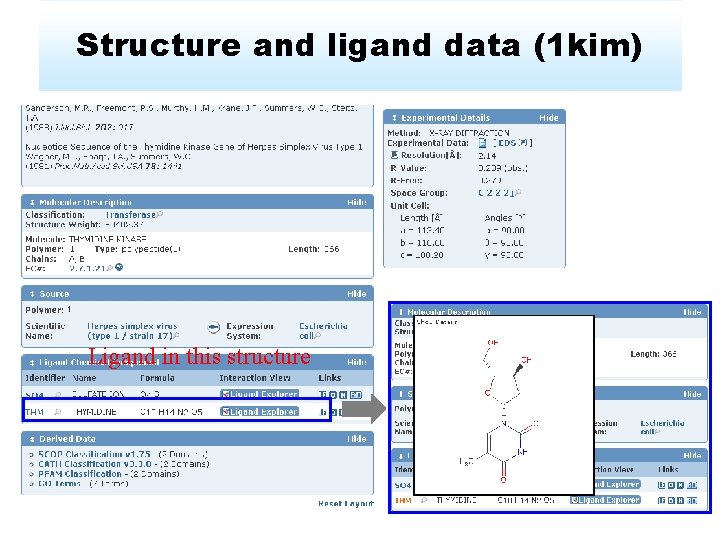
Structure and ligand data (1 kim) Ligand in this structure
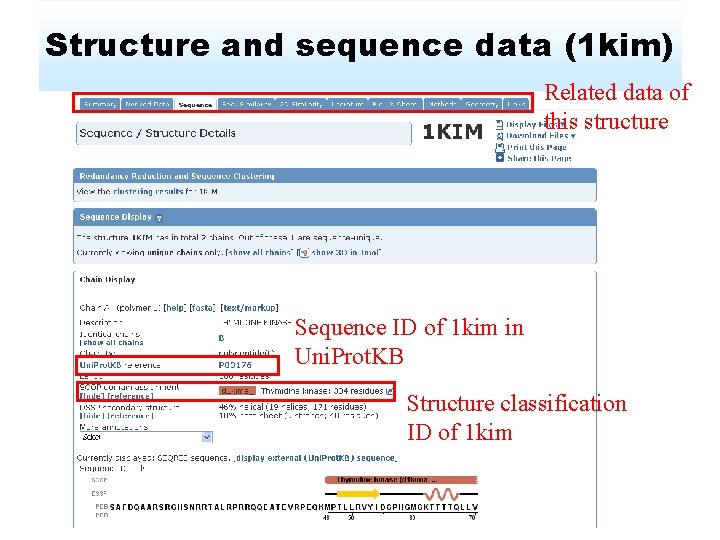
Structure and sequence data (1 kim) Related data of this structure Sequence ID of 1 kim in Uni. Prot. KB Structure classification ID of 1 kim

Advance inspection for protein structure: download structure from PDB 1. Save the data on your PC 2. Open the file on a structure viewer program (swiss PDBviewer, pymol, and etc. )
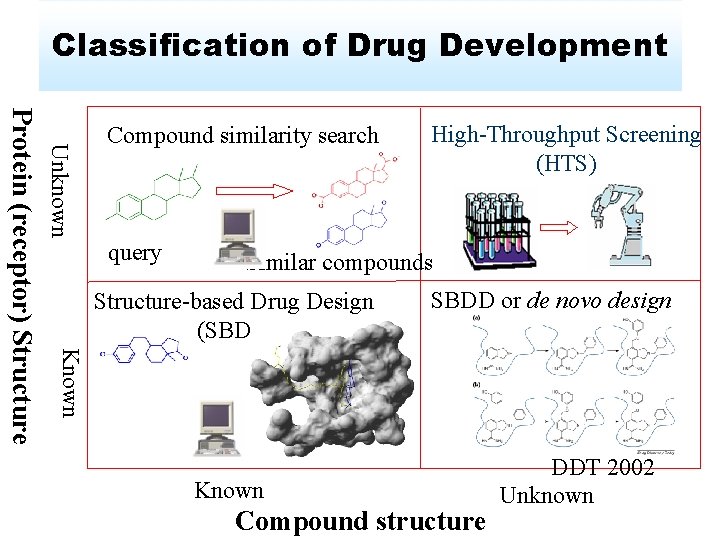
Classification of Drug Development Unknown query High-Throughput Screening (HTS) Similar compounds Structure-based Drug Design (SBDD) SBDD or de novo design Known Protein (receptor) Structure Compound similarity search Known Compound structure DDT 2002 Unknown
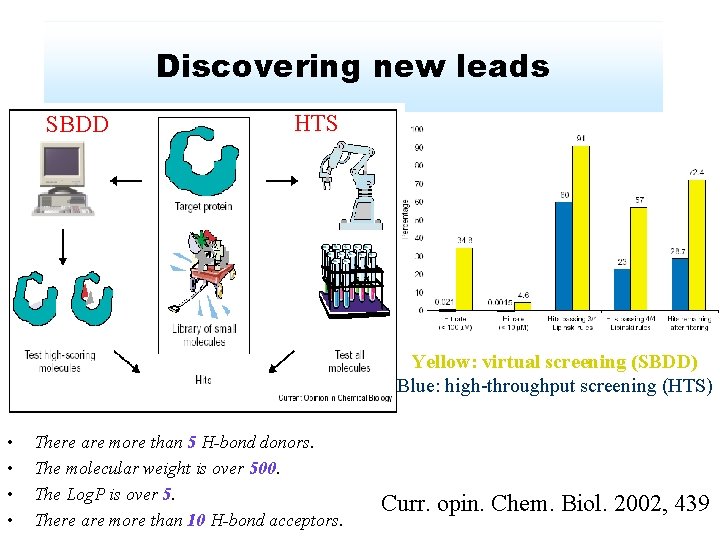
Discovering new leads SBDD HTS Yellow: virtual screening (SBDD) Blue: high-throughput screening (HTS) • • There are more than 5 H-bond donors. The molecular weight is over 500. The Log. P is over 5. There are more than 10 H-bond acceptors. Curr. opin. Chem. Biol. 2002, 439
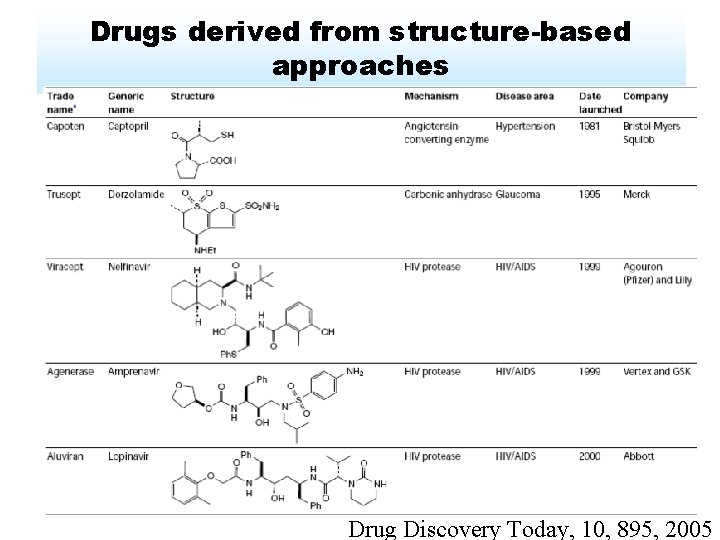
Drugs derived from structure-based approaches Drug Discovery Today, 10, 895, 2005
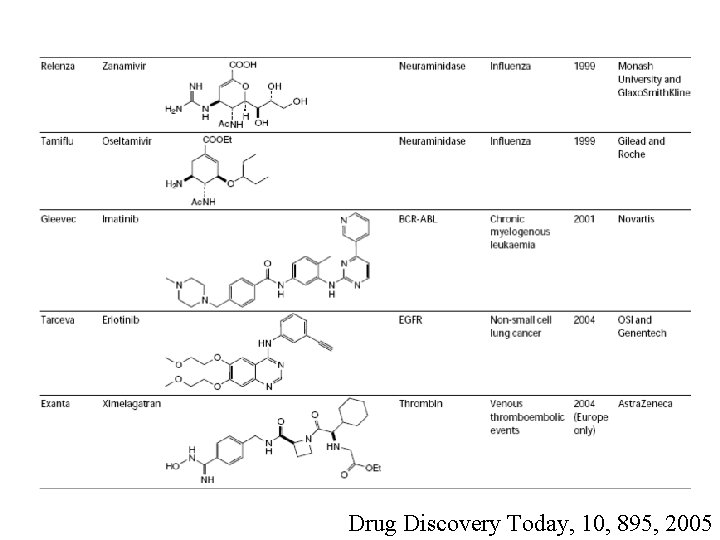
Drug Discovery Today, 10, 895, 2005

Tutorial for Swiss PDB viewer

Download and install Download Swiss Pdb. Viewer http: //spdbv. vital-it. ch/download. html Download user guide http: //spdbv. vital-it. ch/Swiss. Pdb. Viewer. Manualv 3. 7. pdf Tutorial video (English) http: //www. youtube. com/watch? v=n. YT 5 qwtf. New&fe ature=related http: //www. youtube. com/watch? v=y. FE 3 CAHNk. Zg
Download page
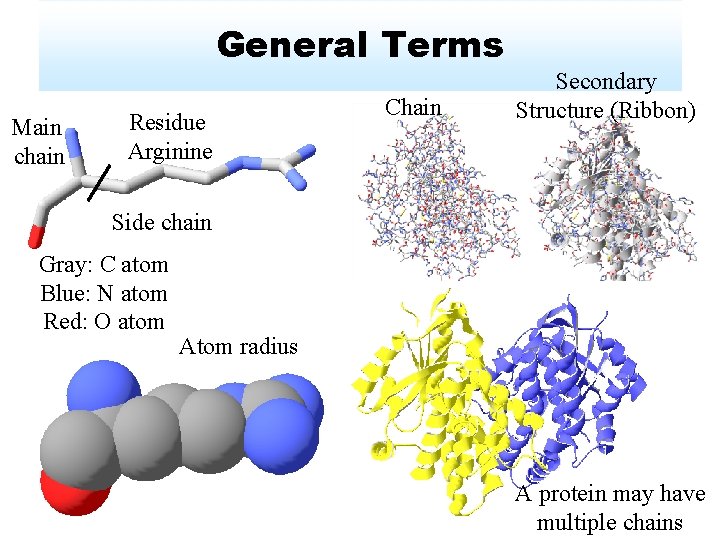
General Terms Main chain Residue Arginine Chain Secondary Structure (Ribbon) Side chain Gray: C atom Blue: N atom Red: O atom Atom radius A protein may have multiple chains
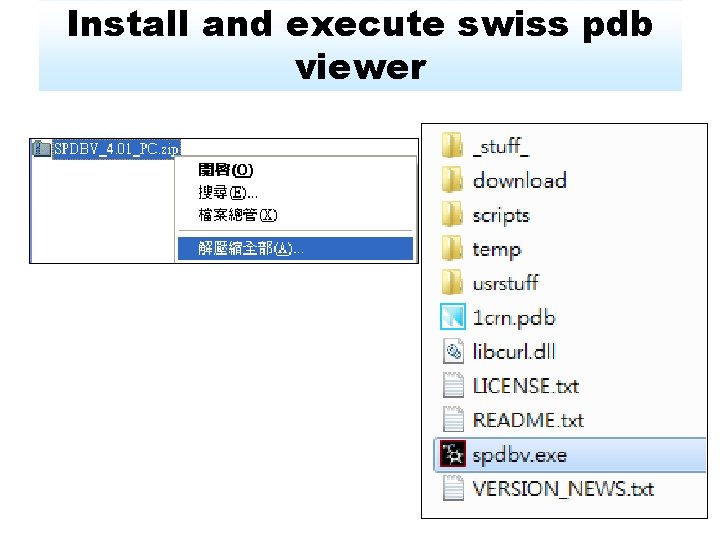
Install and execute swiss pdb viewer
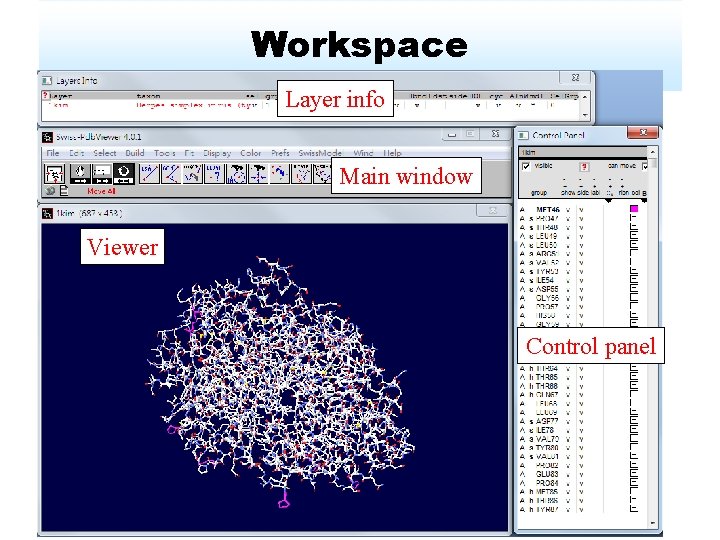
Workspace Layer info Main window Viewer Control panel
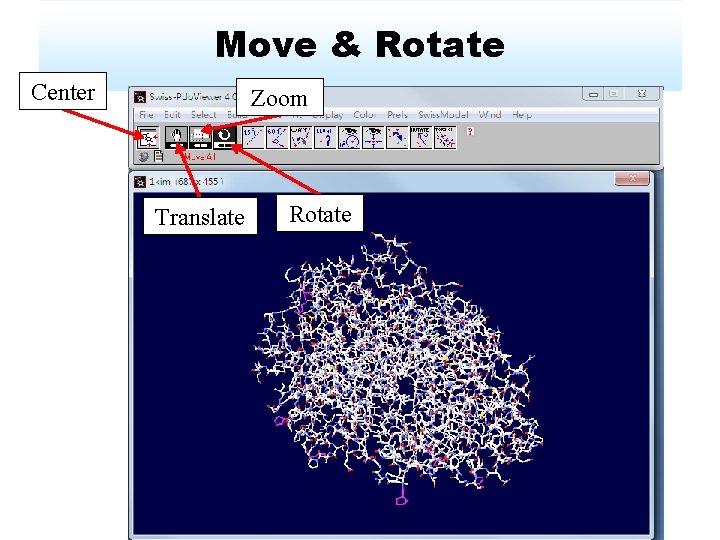
Move & Rotate Center Zoom Translate Rotate
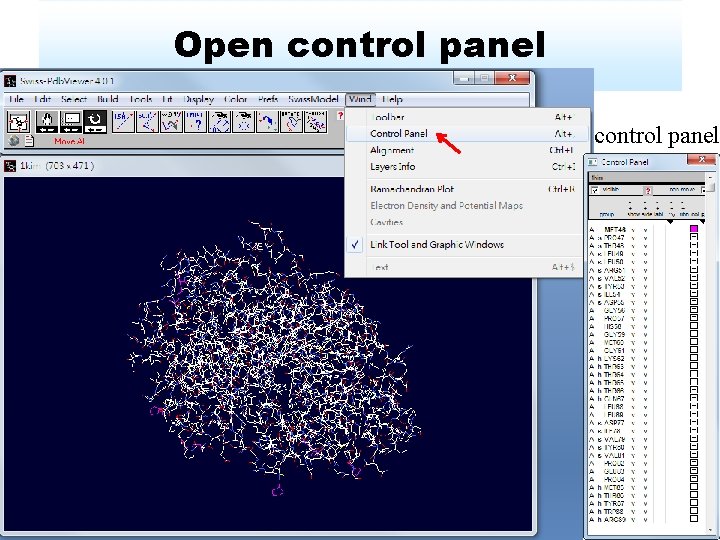
Open control panel
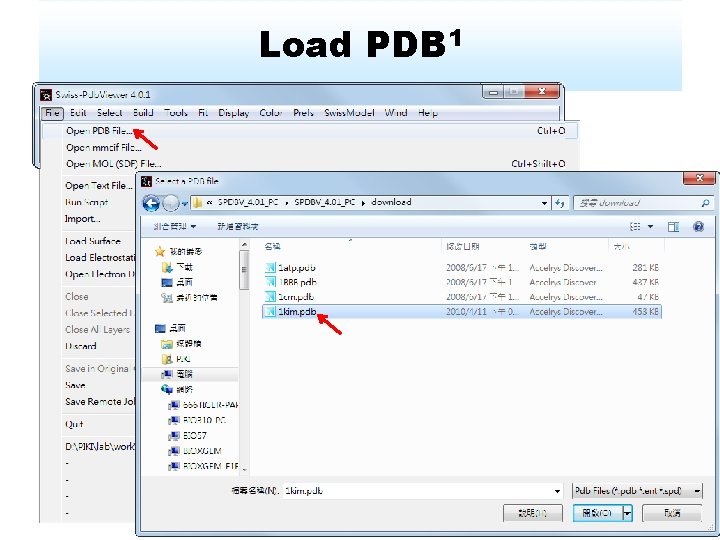
Load PDB 1
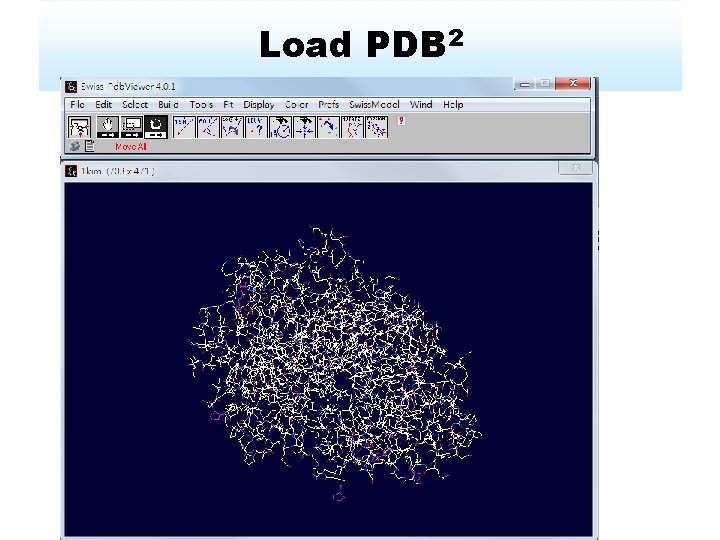
Load PDB 2
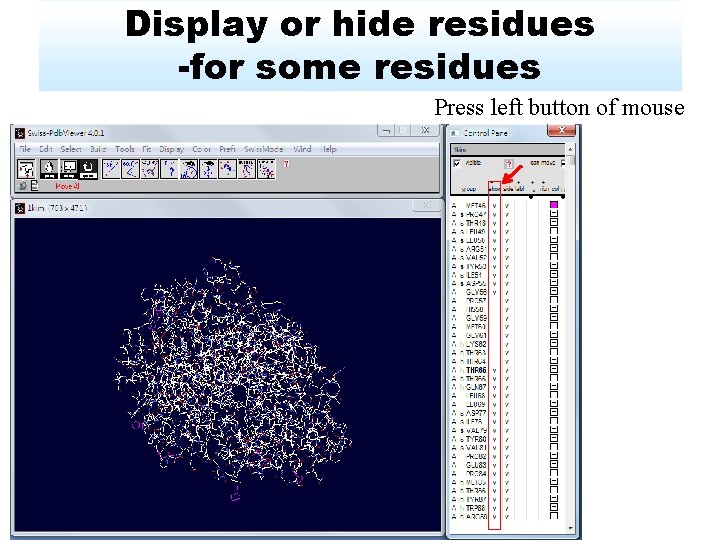
Display or hide residues -for some residues Press left button of mouse
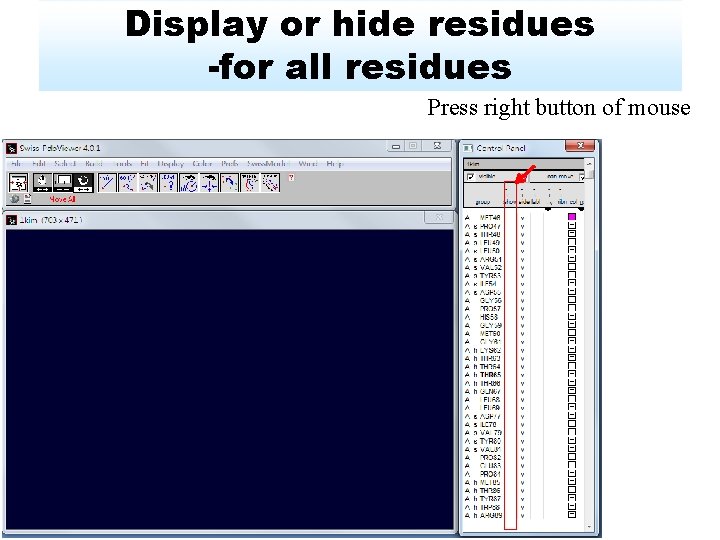
Display or hide residues -for all residues Press right button of mouse
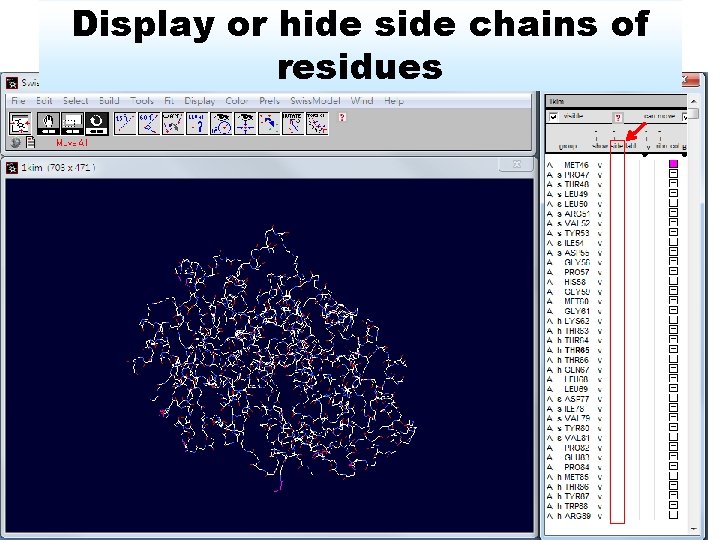
Display or hide side chains of residues
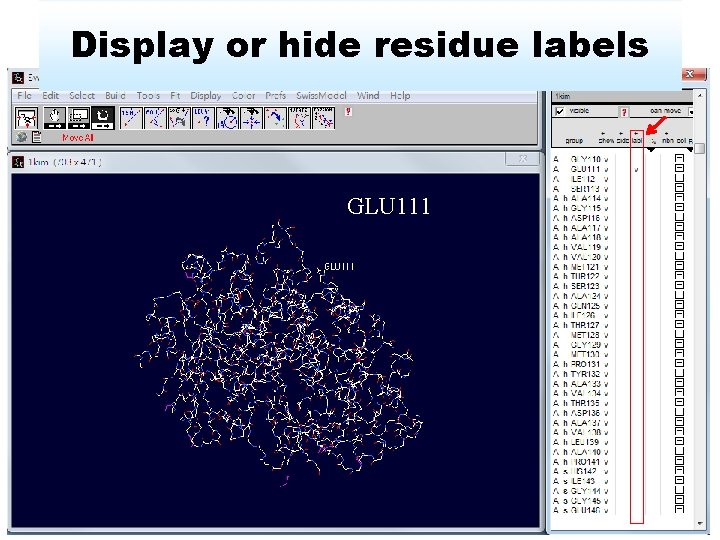
Display or hide residue labels GLU 111
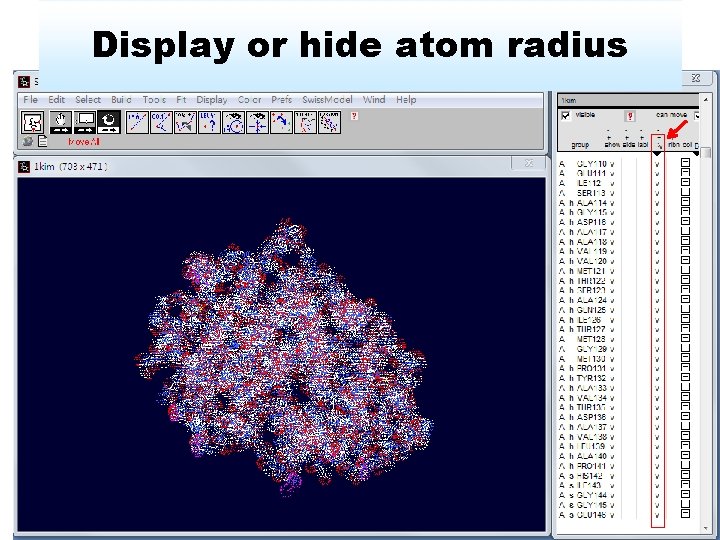
Display or hide atom radius
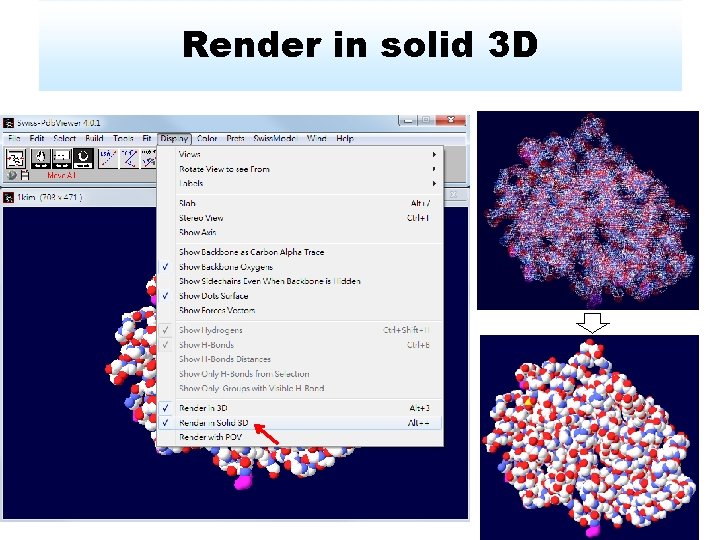
Render in solid 3 D
Show secondary structures -Display or hide ribbons
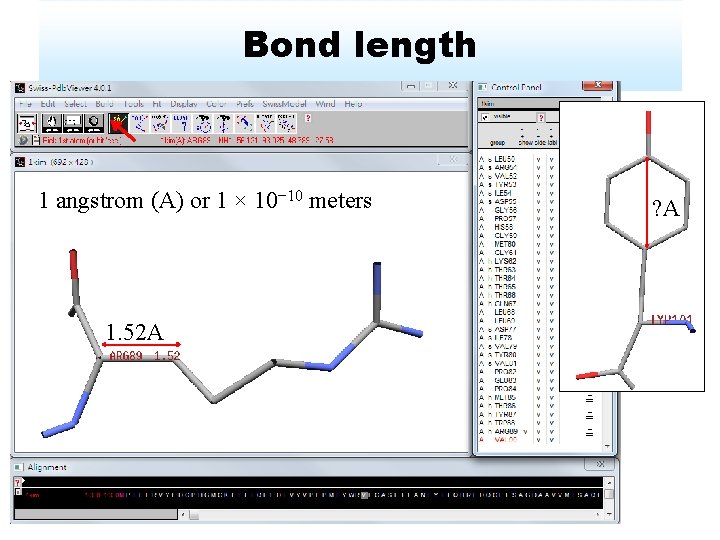
Bond length 1 angstrom (A) or 1 × 10− 10 meters 1. 52 A ? A
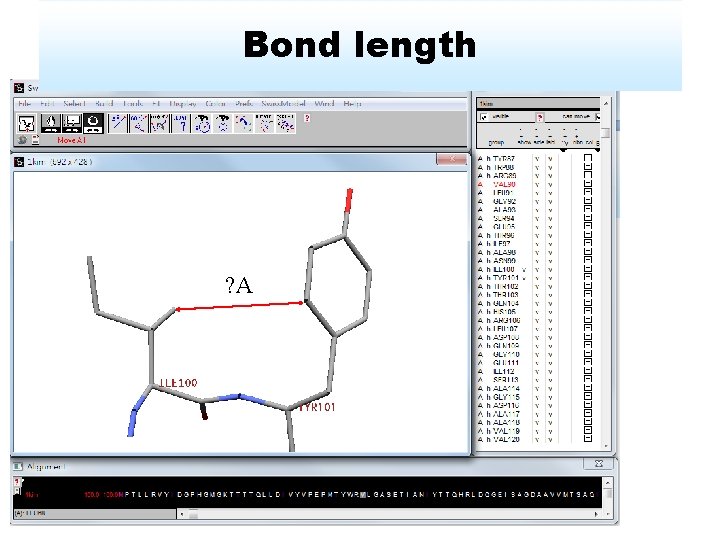
Bond length ? A
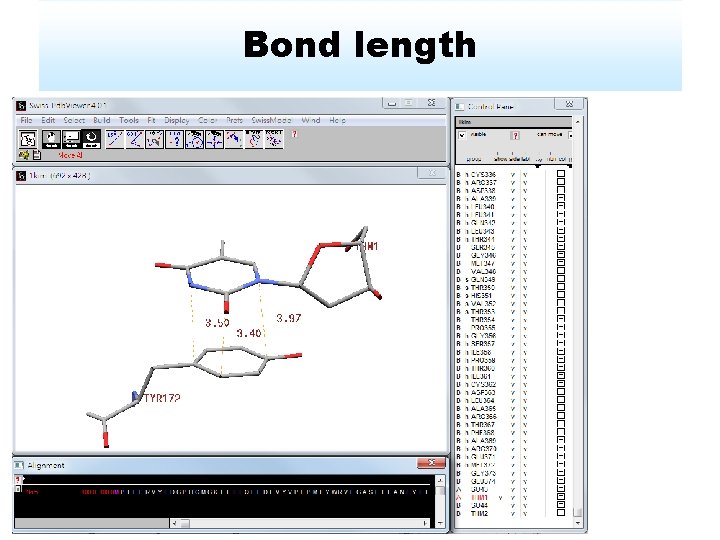
Bond length
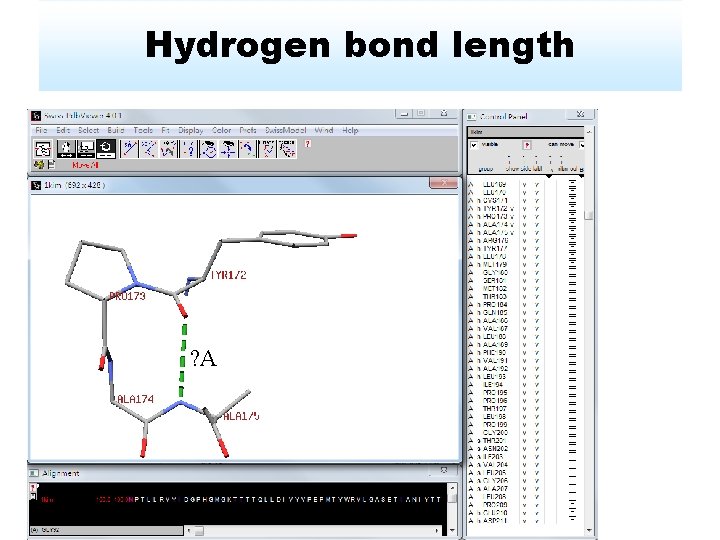
Hydrogen bond length ? A
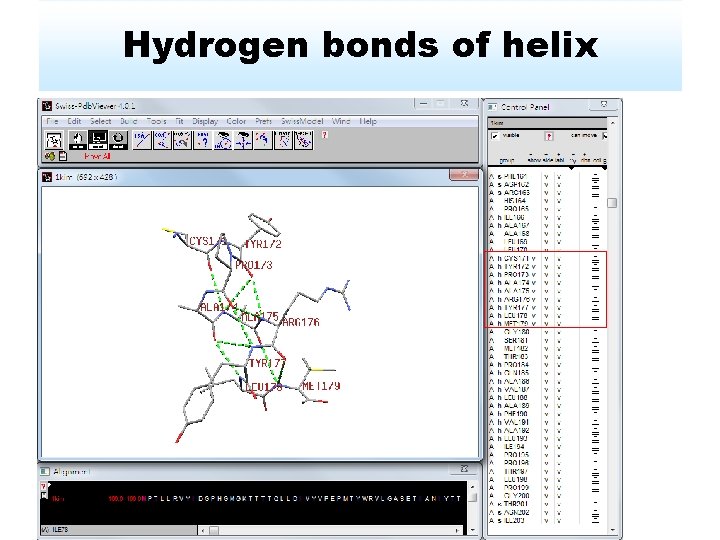
Hydrogen bonds of helix
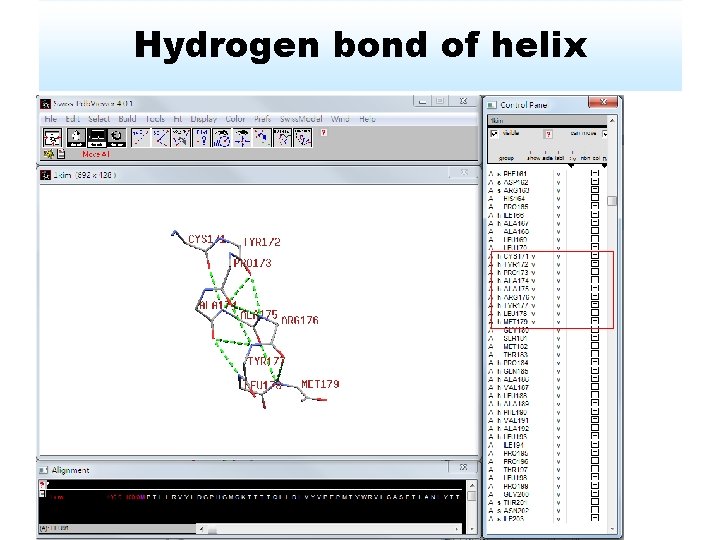
Hydrogen bond of helix
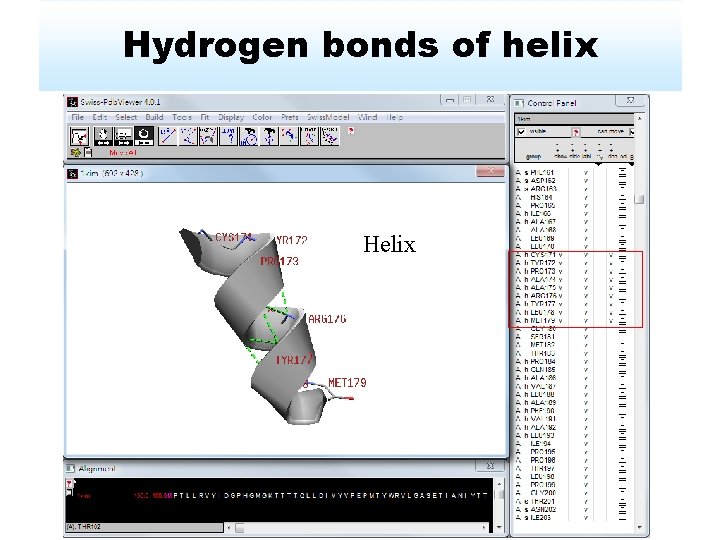
Hydrogen bonds of helix Helix

Hydrogen bonds of sheet Sheet Residue 50~55 201~207 323~328
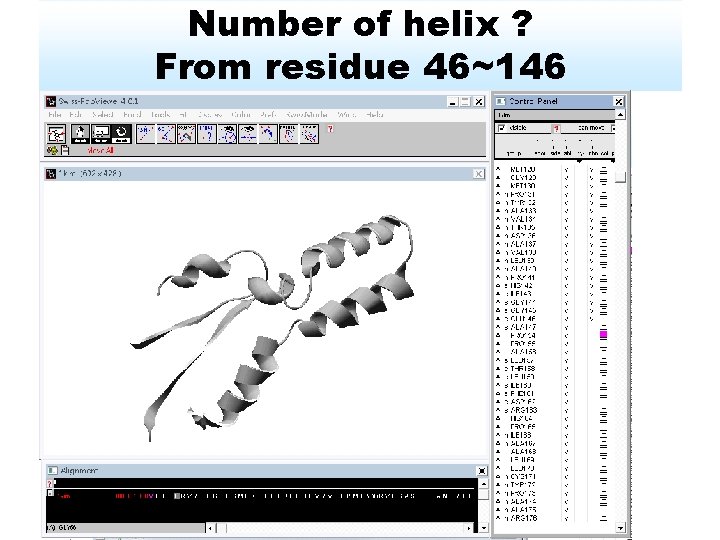
Number of helix ? From residue 46~146
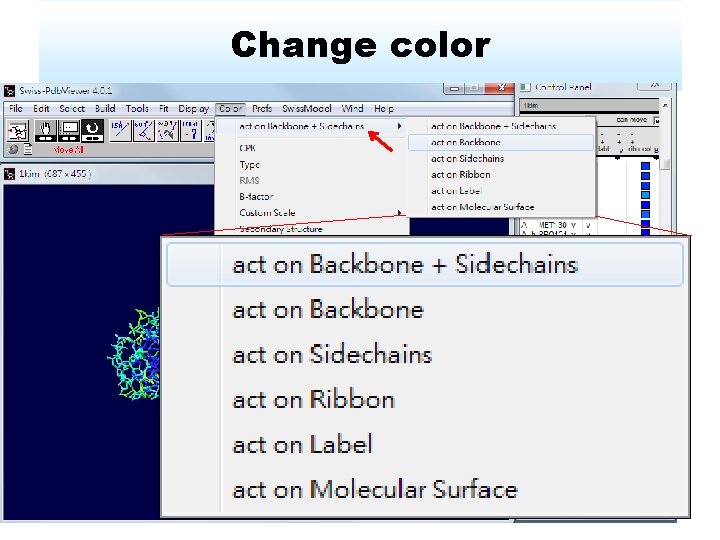
Change color
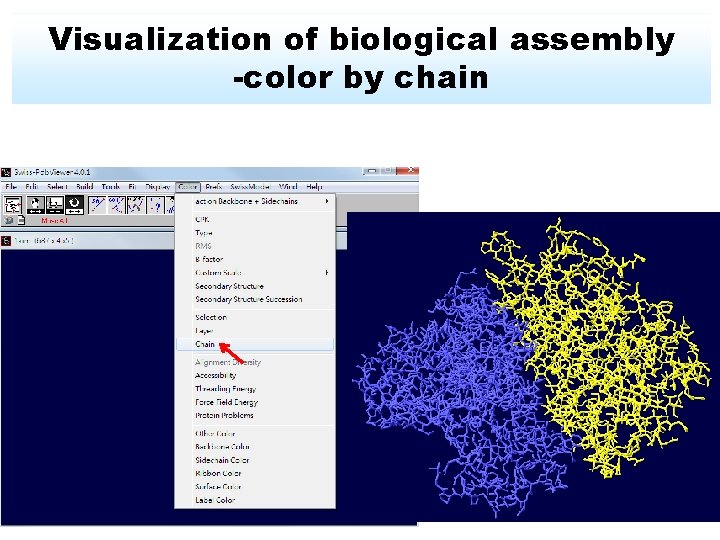
Visualization of biological assembly -color by chain

Change color by chain -act on Ribbons
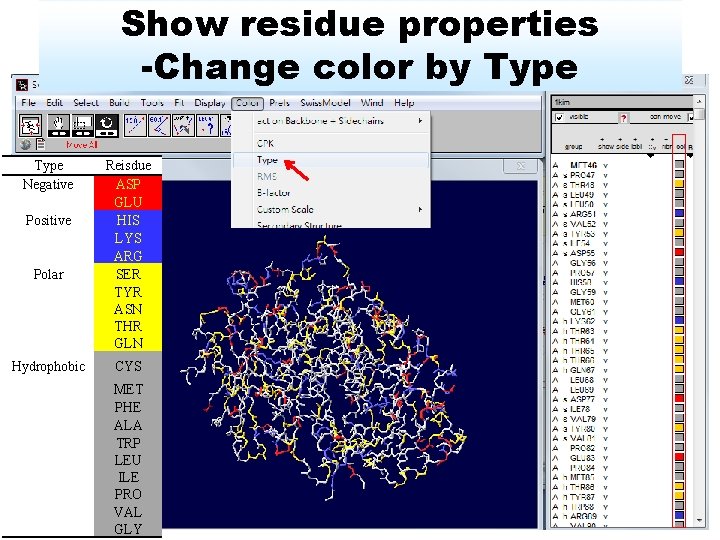
Show residue properties -Change color by Type Negative Positive Polar Reisdue ASP GLU HIS LYS ARG SER TYR ASN THR GLN Hydrophobic CYS MET PHE ALA TRP LEU ILE PRO VAL GLY
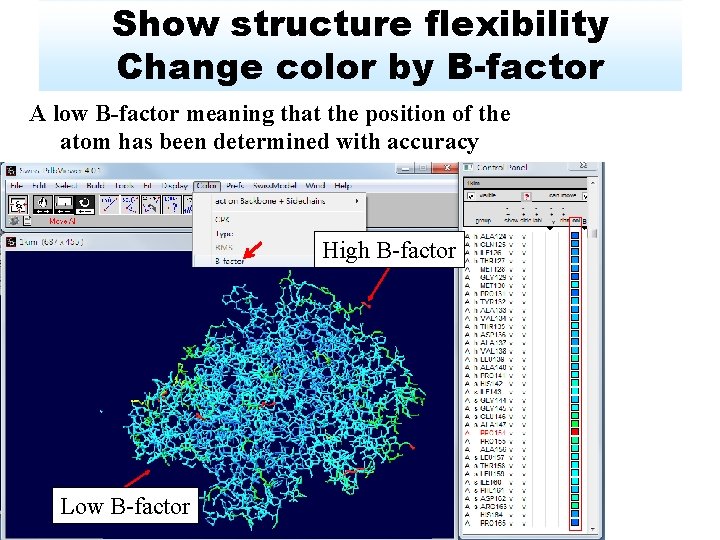
Show structure flexibility Change color by B-factor A low B-factor meaning that the position of the atom has been determined with accuracy High B-factor Low B-factor

Other color types default atom colors Root mean square between 2 molecules Secondary Structure Selected residues Relative accessibility
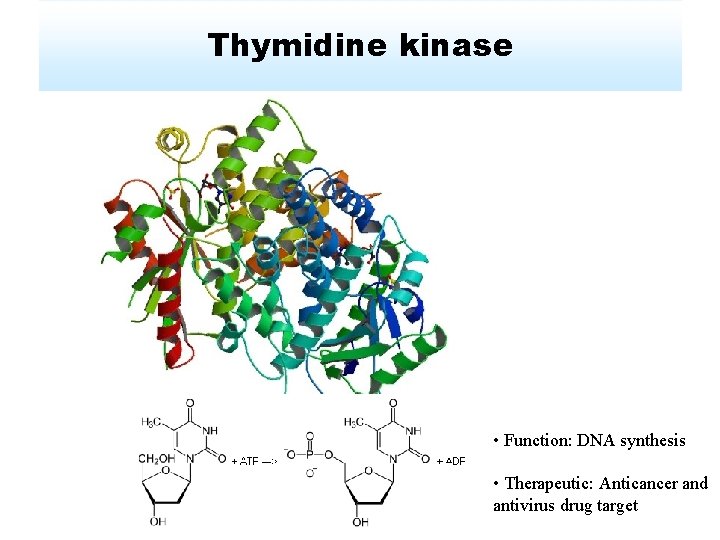
Thymidine kinase • Function: DNA synthesis • Therapeutic: Anticancer and antivirus drug target
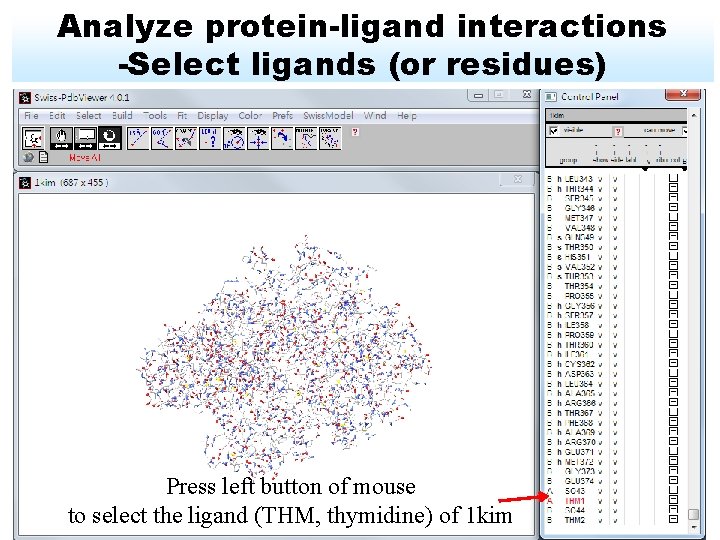
Analyze protein-ligand interactions -Select ligands (or residues) Press left button of mouse to select the ligand (THM, thymidine) of 1 kim
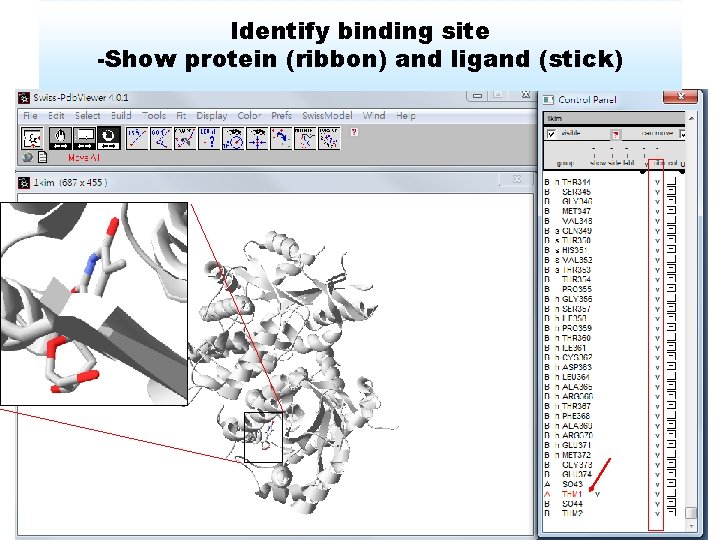
Identify binding site -Show protein (ribbon) and ligand (stick)
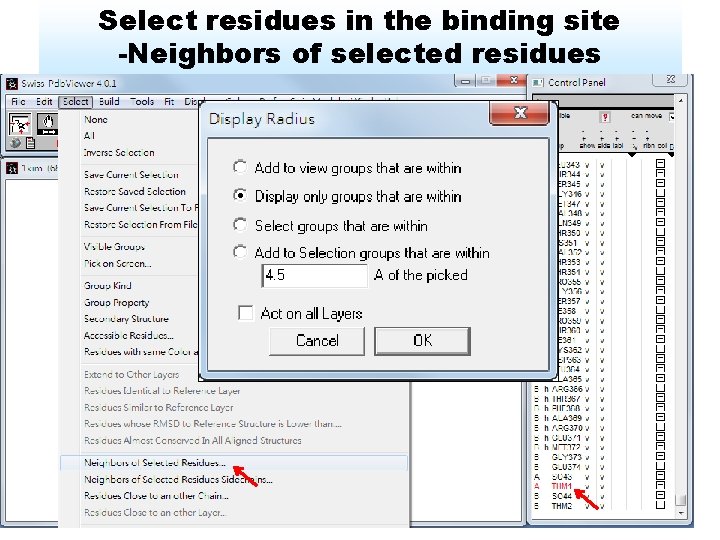
Select residues in the binding site -Neighbors of selected residues
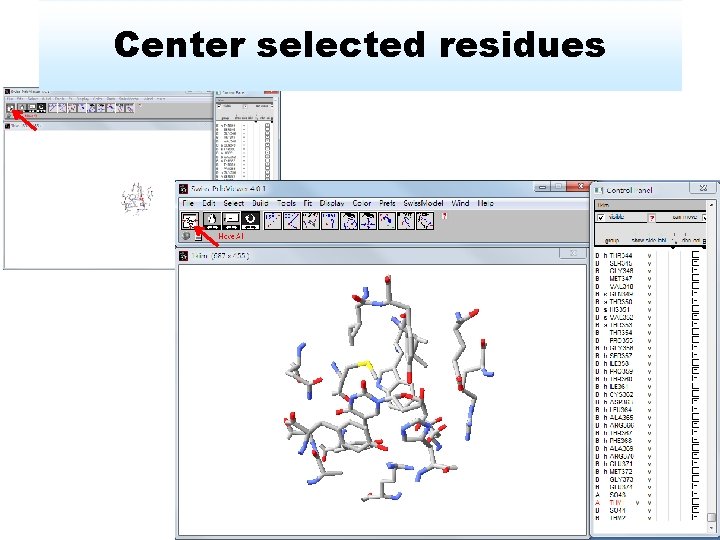
Center selected residues
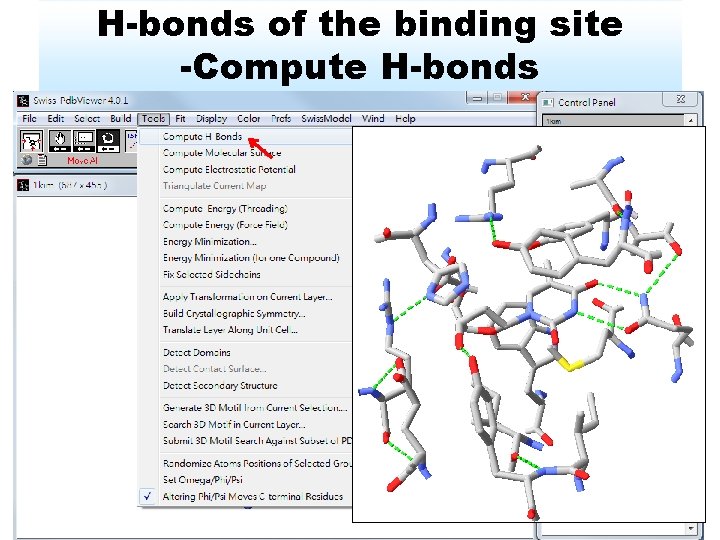
H-bonds of the binding site -Compute H-bonds
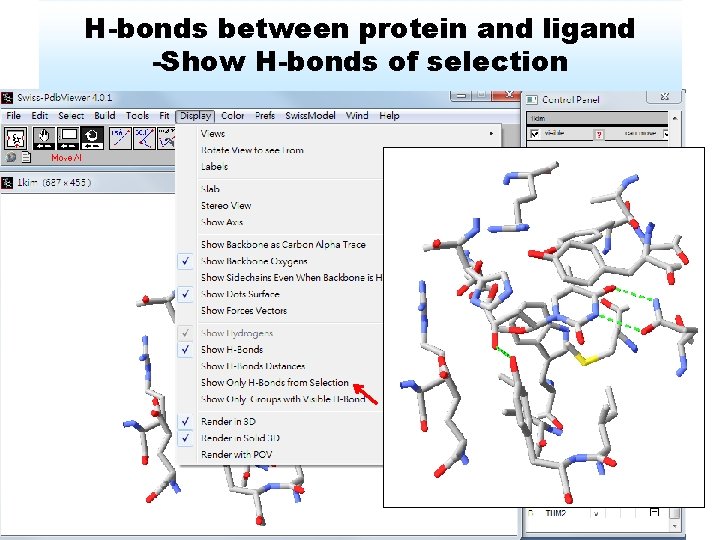
H-bonds between protein and ligand -Show H-bonds of selection
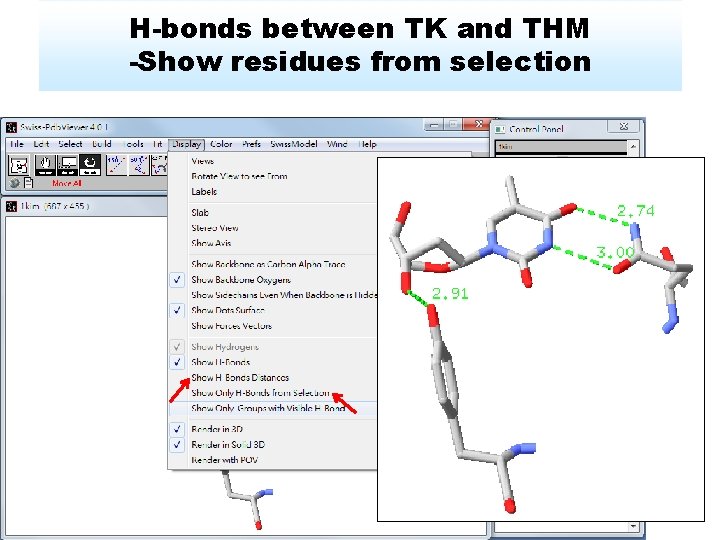
H-bonds between TK and THM -Show residues from selection

Show residue label Press right button of mouse 1. Q 125 recognize thymine moiety 2. Activity was decreased by over 90% if Q 125 mutated (Biochemistry, 2000. 39: p. 4105 -4111)
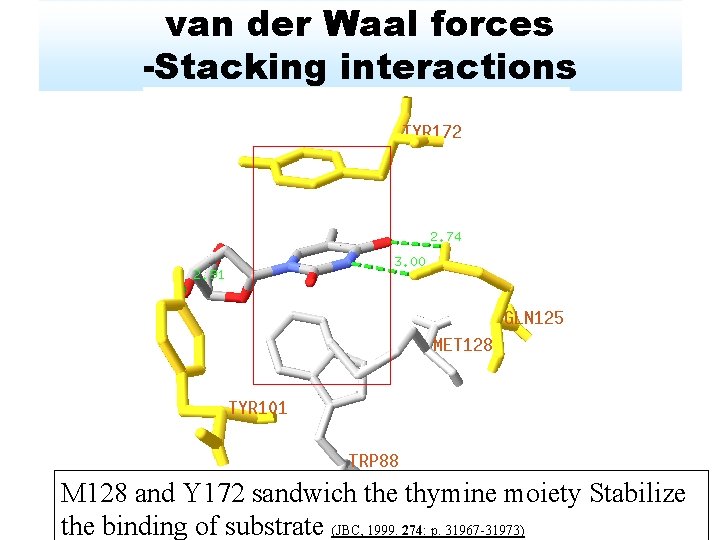
van der Waal forces -Stacking interactions M 128 and Y 172 sandwich the thymine moiety Stabilize the binding of substrate (JBC, 1999. 274: p. 31967 -31973)
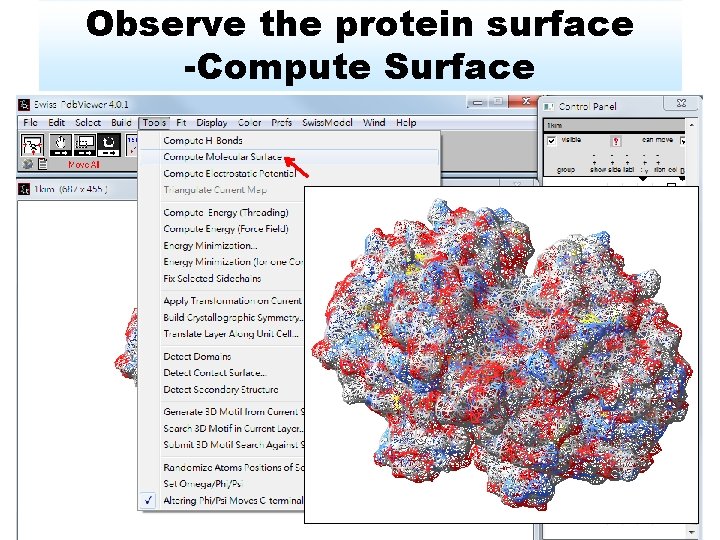
Observe the protein surface -Compute Surface
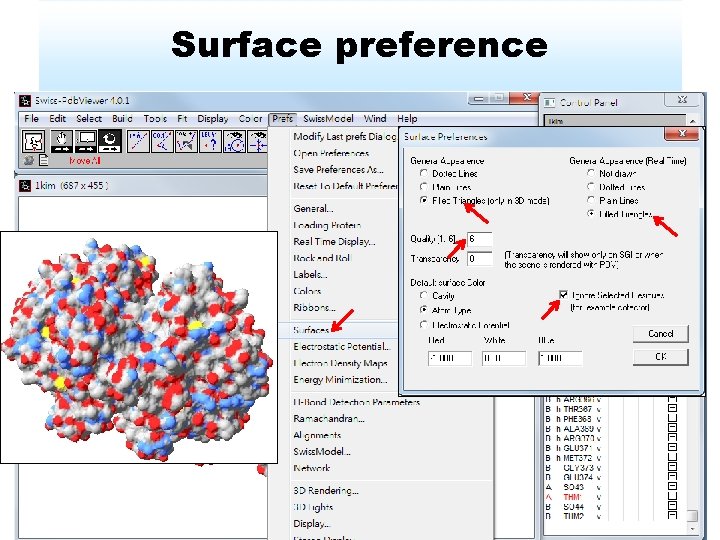
Surface preference
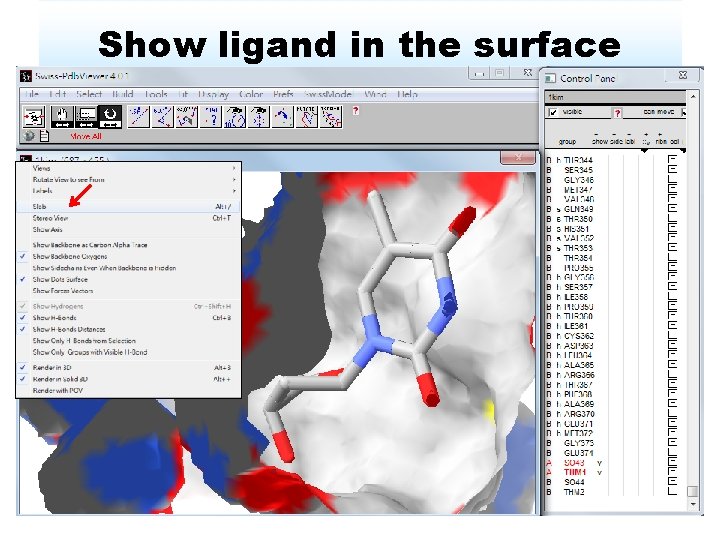
Show ligand in the surface
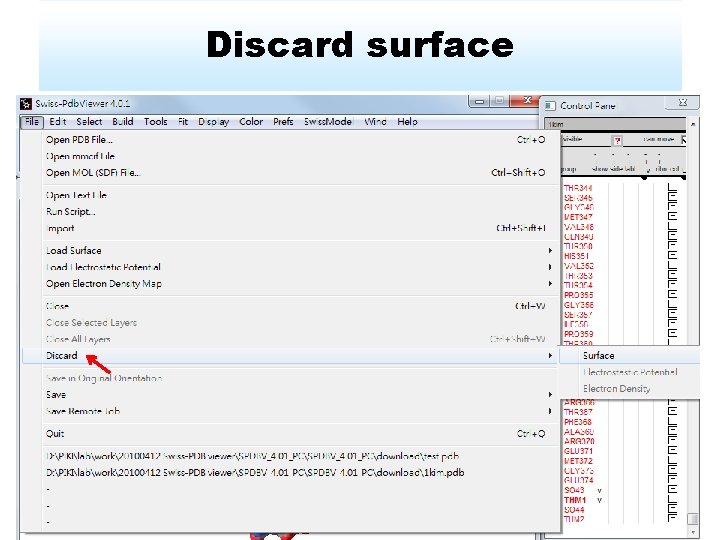
Discard surface
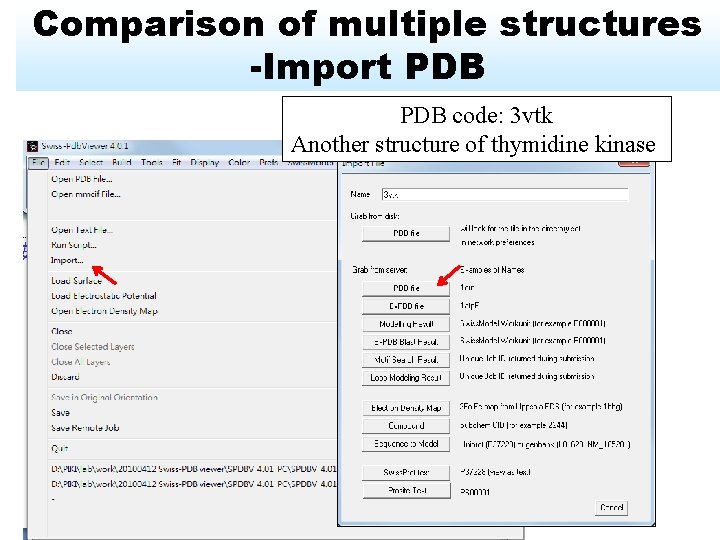
Comparison of multiple structures -Import PDB code: 3 vtk Another structure of thymidine kinase
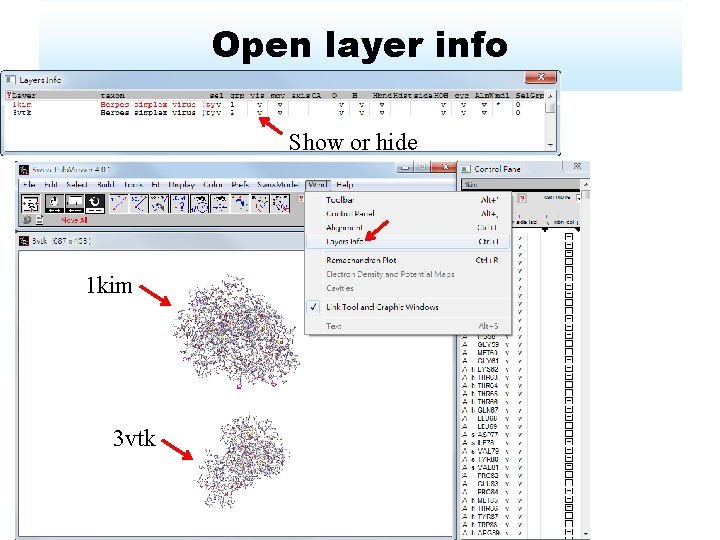
Open layer info Show or hide 1 kim 3 vtk
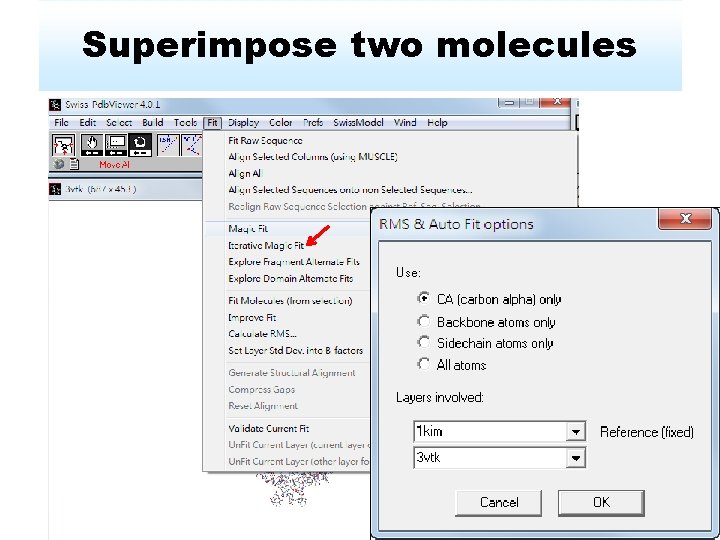
Superimpose two molecules

Results of superimposition RMS: 0. 63 Ǻ Measure the structure similarity
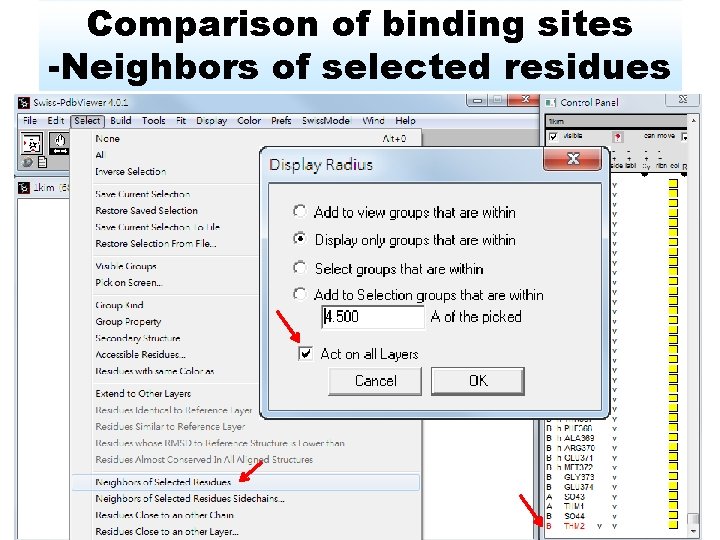
Comparison of binding sites -Neighbors of selected residues
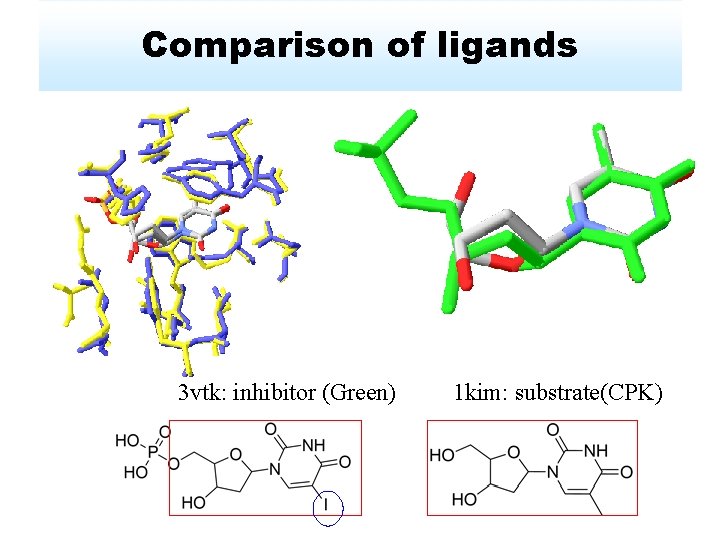
Comparison of ligands 3 vtk: inhibitor (Green) 1 kim: substrate(CPK)
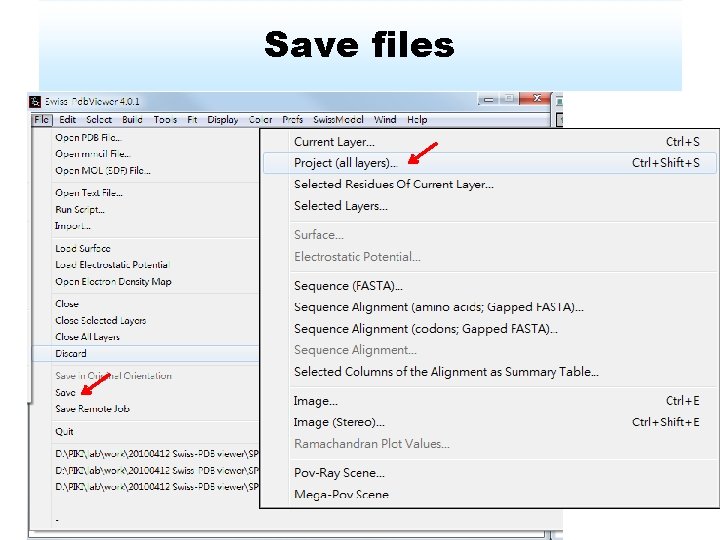
Save files

Homework Keyword 1. Capture a picture of N 1 neuraminidase (ribbon) and its ligand (stick) 2. Capture a picture of H-bonds between protein and ligand E-mail pikihsu@gmail. com Mail title: 學號姓名_HW 2
- Slides: 72