Tutorial on GRID Computing EMBnet Conference 2008 Tutorial

Tutorial on "GRID Computing“ EMBnet Conference 2008 Tutorial on GRID computing Giorgio Maggi INFN and Politecnico di Bari CNR - ITB http: //www. libi. it

LIBI • International Laboratory of Bioinformatics – A Virtual Laboratory for composing and executing complex experiments in Bioinformatics – Project Coordinator : Prof. Cecilia Saccone – Project funded by the MIUR (Italian Ministry for Education, University and Research), F. I. R. B. 2003, under grant RBLA 039 M 7 M. Tutorial on "GRID Computing“, EMBnet Conference 2008, 17 September 2008

LIBI partners • Technological RU’s (provide the platform): § University of Salento – SPACI Consortium, Lecce. • Prof. Giovanni Aloisio § INFN – Sections of Bari, Catania and Padova and CNAF-Bologna • Dr. Mirco Mazzucato § IBM Italia S. p. A - IBM Innovation Lab, Bari • Dr. Luigi Di Pace § CINECA – Bologna • Dr. Elda Rossi • Scientific RU’s (use the platform): § CNR – Institute for Biomedical Technologies, Bari • Prof. Cecilia Saccone, project coordinator § University of Bologna – Biocomputing Group, Bologna • Prof. Rita Casadio § University of Milano • Prof. Graziano Pesole § Centro di Biomedicina Molecolare – Trieste • Prof. Claudio Schneider Tutorial on "GRID Computing“, EMBnet Conference 2008, 17 September 2008

The LIBI architecture • a) Computing Virtualization layer: – a heterogeneous computing environment able to integrate and virtualize the access to different production Grids (g. Lite, Unicore and Globus) and HPC resources; • b) Databases & Data Virtualization layer: – a suite of bioinformatics databases and resources, integrated in a federated approach, including private and public data. Data are integrated by many services and tools and are accessible from the virtualization layer; • d) Bioinformatics Applications layer: – a number of specialized bioinformatics applications (Case Studies), targeting scientific problems, built by mashing-up services, tools, data and computing resources. Tutorial on "GRID Computing“, EMBnet Conference 2008, 17 September 2008
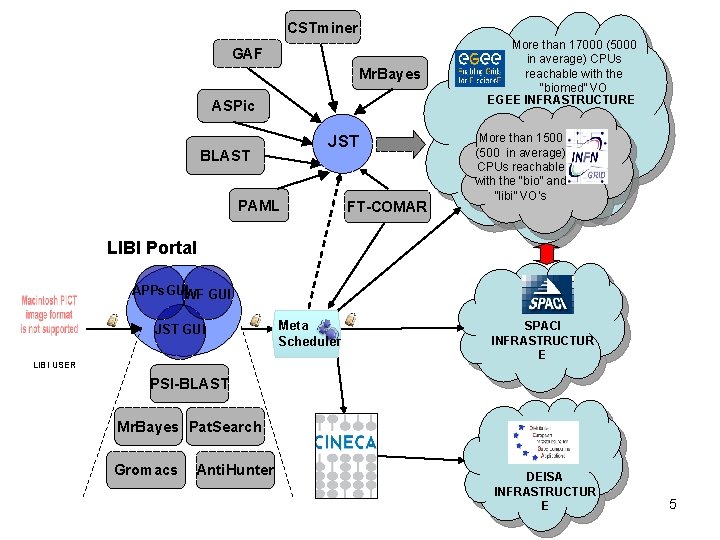
CSTminer GAF Mr. Bayes ASPic JST BLAST PAML FT-COMAR More than 17000 (5000 in average) CPUs reachable with the “biomed” VO EGEE INFRASTRUCTURE More than 1500 (500 in average) CPUs reachable with the “bio” and “libi” VO’s LIBI Portal APPs. GUIWF GUI JST GUI LIBI USER Meta Scheduler SPACI INFRASTRUCTUR E PSI-BLAST Mr. Bayes Pat. Search Gromacs Anti. Hunter DEISA INFRASTRUCTUR E 5
6
- Slides: 6