Tutorial Bioinformatics Resources BIOTRAC 25 Proteomics Principles and

Tutorial: Bioinformatics Resources BIO-TRAC 25 (Proteomics: Principles and Methods) October 10, 2003 NIH, Bethesda, MD Zhang-Zhi Hu, M. D. Senior Bioinformatics Scientist, Protein Information Resource National Biomedical Research Foundation, GUMC

What is Bioinformatics? Bioinformatics is the application of information technology to the analysis, organization and distribution of biological data in order to answer complex biological questions. NIH Biomedical Information Science and Technology Initiative (BISTI) Working Definition (2002) - Research, development, or application of computational tools and approaches for expanding the use of biological, medical, behavioral or health data, including those to acquire, store, organize, archive, analyze, or visualize such data. 2

Bioinformatics Resources The Molecular Biology Database Collection: An Online Compilation of Relevant Database Resources l 2003 update: http: //www 3. oup. co. uk/nar/database/ l Nucleic Acids Research Database Issues (January Annually) (2003 - http: //nar. oupjournals. org/content/vol 31/issue 1/) DBcat: A Catalog of > 500 Biological Databases l http: //www. infobiogen. fr/services/dbcat/ 3

Molecular Biology Database Collection (http: //nar. oupjournals. org/cgi/content/full/31/1/1#GKG 120 TB 1) 4

The Molecular Biology Database Collection: 2003 update (Baxevanis, A. D. ) -- An online resource of 386 key databases of 18 categories Major sequence repositories Comparative Genomics Gene Expression Gene Identification and Structure Genetic and Physical Maps Genomic Databases Intermolecular Interactions Metabolic Pathways and Cellular Regulation Mutation Databases Pathology Protein Sequence Motifs Proteome Resources Retrieval Systems and Database Structure RNA Sequences Structure Transgenics Varied Biomedical Content 5

Overview Protein Sequence Analysis I. III. Sequence Similarity Search and Alignment Family Classification Methods Structure Prediction Methods Molecular Biology Databases IV. Protein Family Databases V. Database of Protein Functions VI. Databases of Protein Structures Proteomic Resources VII. 2 D-gel databases VIII. Proteomic analyses 6

I. Sequence Similarity Search Find a protein sequence: text search Based on Pair-Wise Comparisons l BLOSUM scoring matrix l PAM scoring matrix Dynamic Programming Algorithms l Global Similarity: Needleman-Wunsch (GAP/Best. Fit) l Local Similarity: Smith-Waterman (SSEARCH) Heuristic Algorithms (Sequence Database Searching) l FASTA: Based on K-Tuples (2 -Amino Acid) l BLAST: Triples of Conserved Amino Acids l Gapped-BLAST: Allow Gaps in Segment Pairs (NREF) l PHI-BLAST: Pattern-Hit Initiated Search (NCBI) l PSI-BLAST: Iterative Search (NCBI) 7
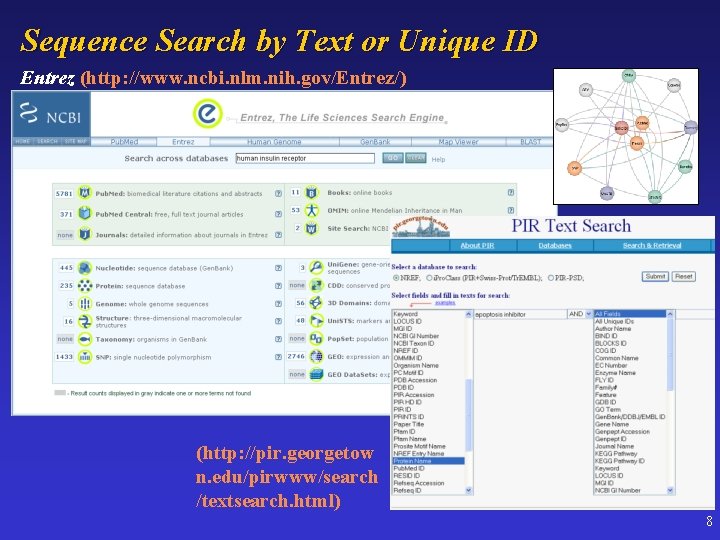
Sequence Search by Text or Unique ID Entrez (http: //www. ncbi. nlm. nih. gov/Entrez/) (http: //pir. georgetow n. edu/pirwww/search /textsearch. html) 8
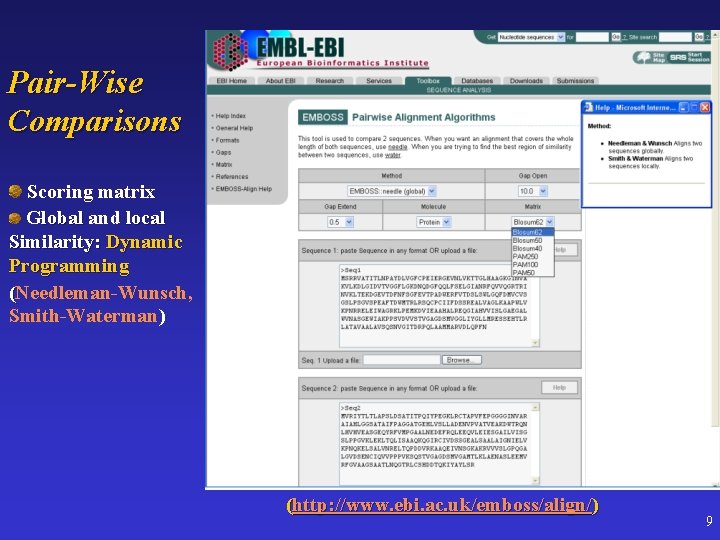
Pair-Wise Comparisons Scoring matrix Global and local Similarity: Dynamic Programming (Needleman-Wunsch, Smith-Waterman) (http: //www. ebi. ac. uk/emboss/align/) 9
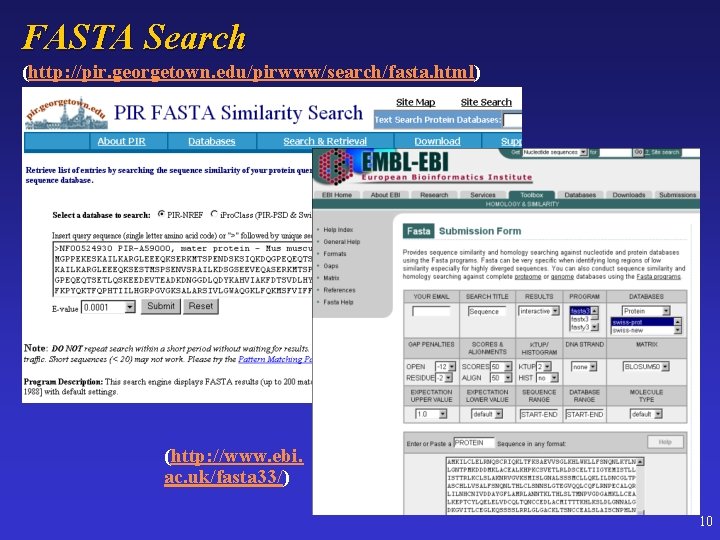
FASTA Search (http: //pir. georgetown. edu/pirwww/search/fasta. html) (http: //www. ebi. ac. uk/fasta 33/) 10
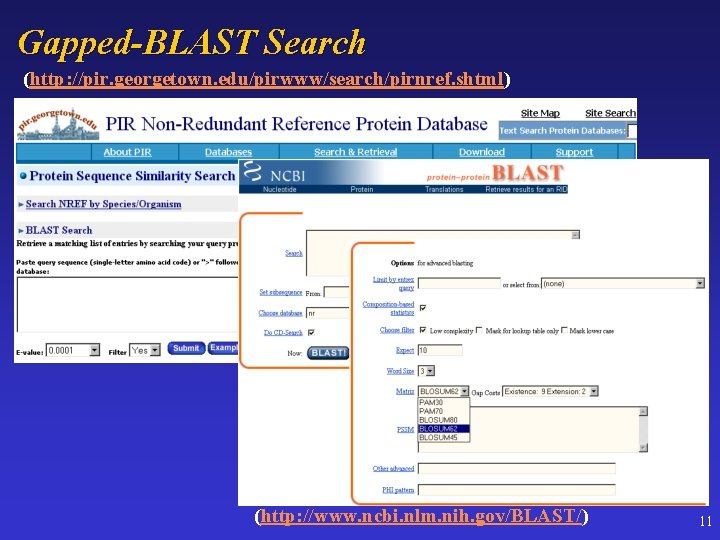
Gapped-BLAST Search (http: //pir. georgetown. edu/pirwww/search/pirnref. shtml) (http: //www. ncbi. nlm. nih. gov/BLAST/) 11

A BLAST Result
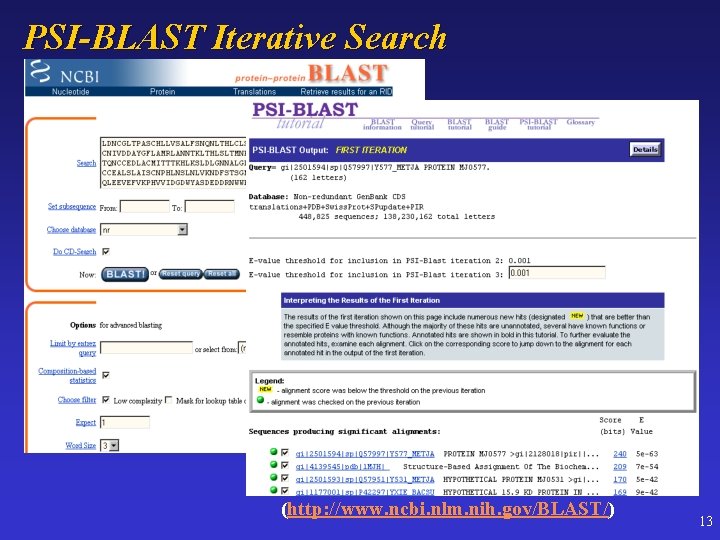
PSI-BLAST Iterative Search (http: //www. ncbi. nlm. nih. gov/BLAST/) 13
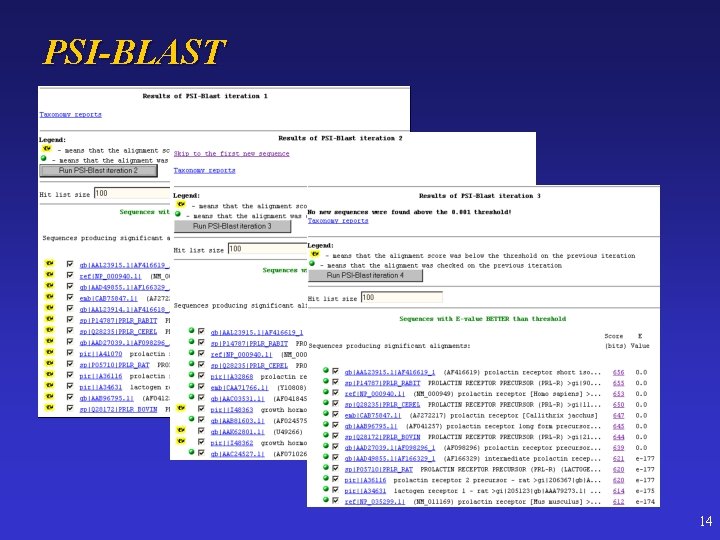
PSI-BLAST 14

II. Family Classification Methods Multiple Sequence Alignment and Phylogenetic Analysis l Clustal. W Multiple Sequence Alignment l Alignment Editor & Phylogenetic Trees Searches Based on Family Information l PROSITE Pattern Search l Motif and Profile Search l Hidden Markov Model (HMMs) 15

Multiple Sequence Alignment Clustal. W (http: //pir. georgetown. edu/pirwww/search/multaln. html) 16
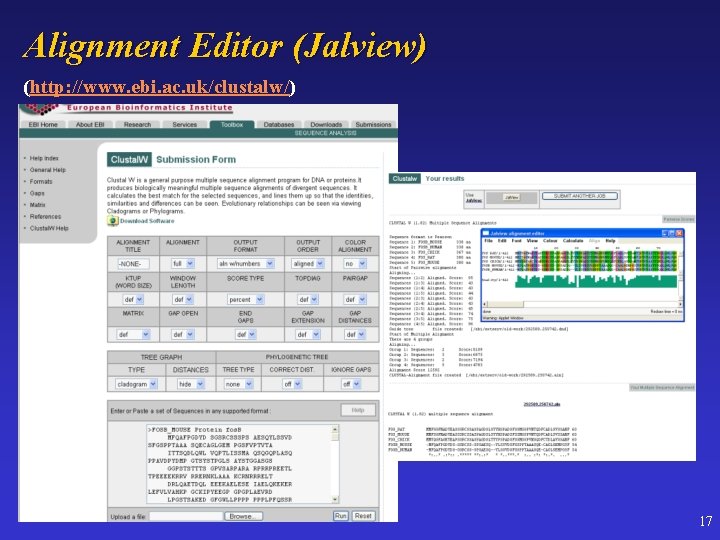
Alignment Editor (Jalview) (http: //www. ebi. ac. uk/clustalw/) 17
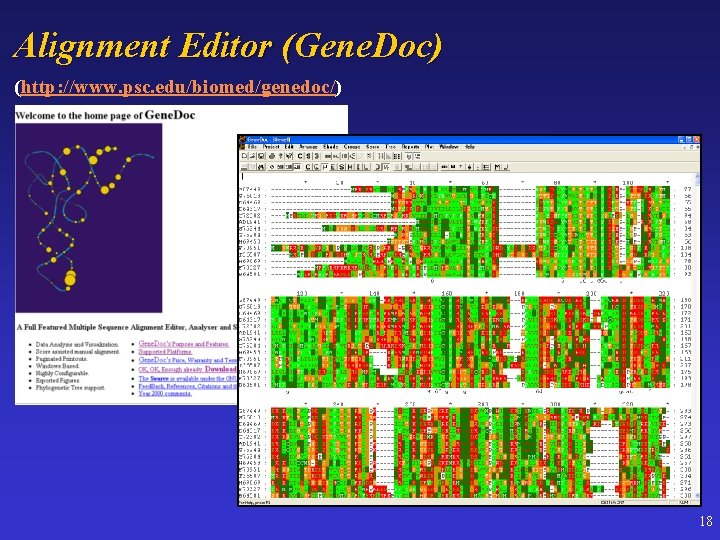
Alignment Editor (Gene. Doc) (http: //www. psc. edu/biomed/genedoc/) 18
Phylogenetic Analysis Tree Programs: (http: //evolution. genetics. washington. edu/phylip. html) Tree Searches: (http: //pauling. mbu. iisc. ernet. in/~pali/index. html) 19
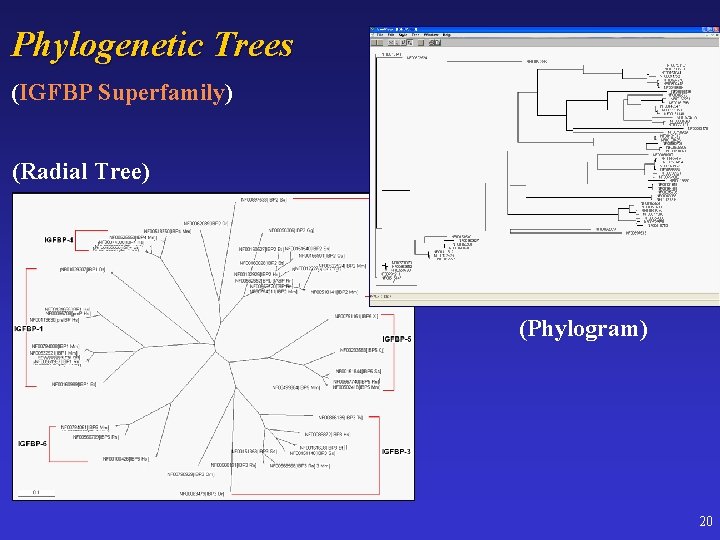
Phylogenetic Trees (IGFBP Superfamily) (Radial Tree) (Phylogram) 20
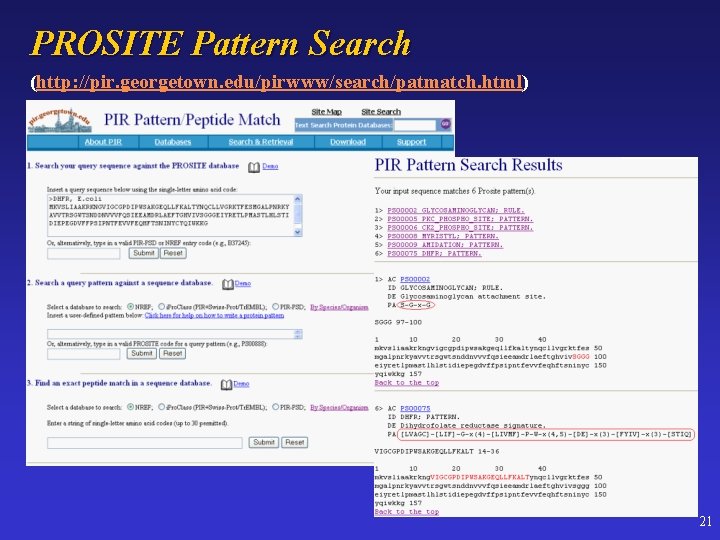
PROSITE Pattern Search (http: //pir. georgetown. edu/pirwww/search/patmatch. html) 21
Profile Search (http: //bmerc-www. bu. edu/bioinformatics/profile_request. html) 22
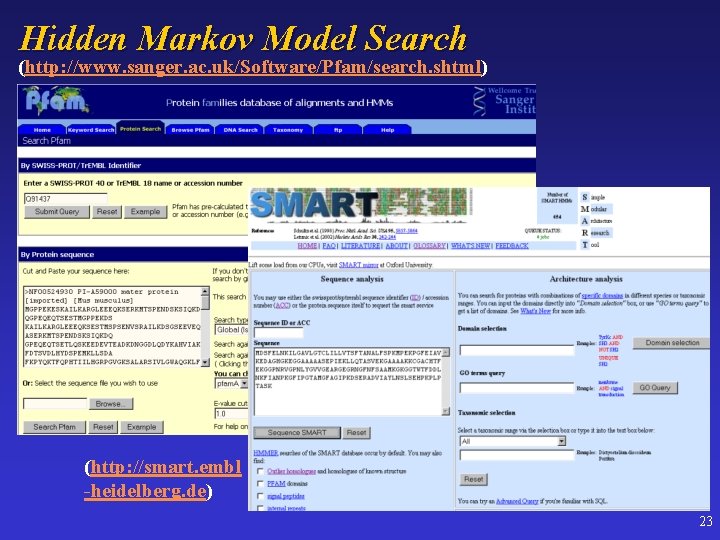
Hidden Markov Model Search (http: //www. sanger. ac. uk/Software/Pfam/search. shtml) (http: //smart. embl -heidelberg. de) 23

III. Structural Prediction Methods Signal Peptide: SIGFIND, Signal. P Transmembrane Helix: TMHMM, TMAP 2 D Prediction (a-helix, b-sheet, Coiled-coils): PHD, JPred 3 D Modeling: Homology Modeling (Modeller, SWISSMODEL), Threading, Ab-initio Prediction 24
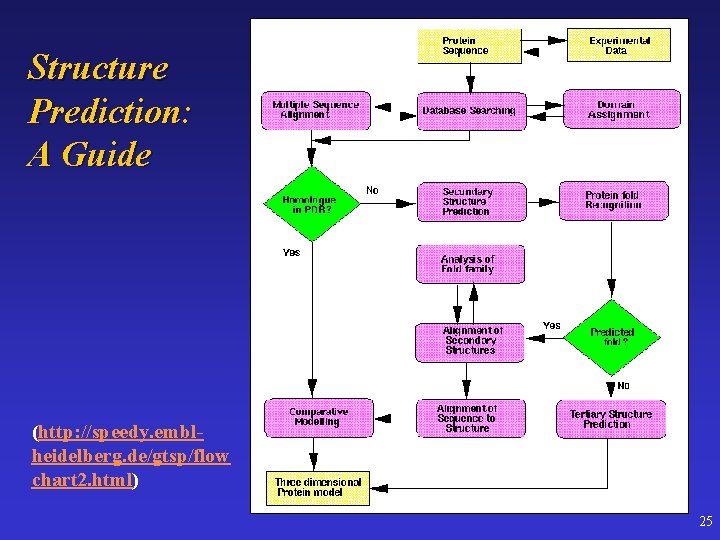
Structure Prediction: A Guide (http: //speedy. emblheidelberg. de/gtsp/flow chart 2. html) 25

Protein Prediction Server (http: //www. cbs. dtu. dk/services/) 26
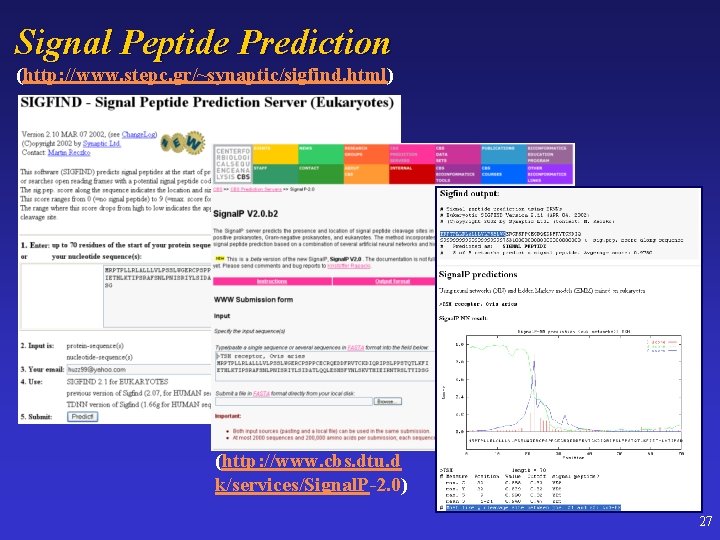
Signal Peptide Prediction (http: //www. stepc. gr/~synaptic/sigfind. html) (http: //www. cbs. dtu. d k/services/Signal. P-2. 0) 27
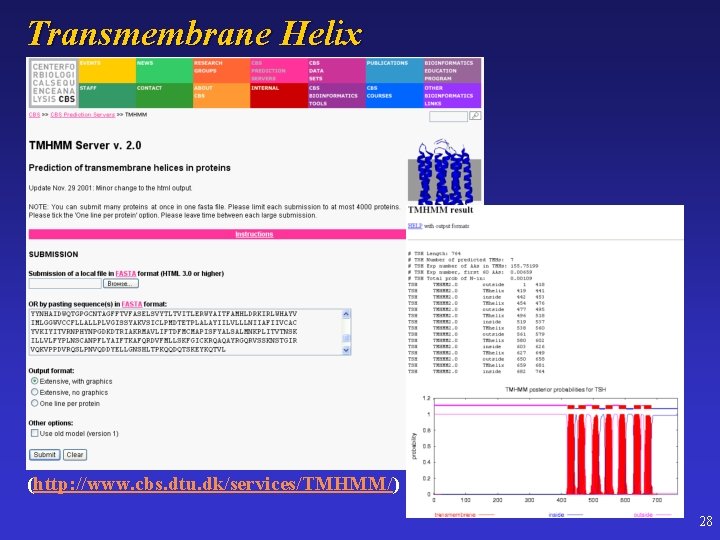
Transmembrane Helix (http: //www. cbs. dtu. dk/services/TMHMM/) 28
Protein Structure Prediction (http: //cmgm. stanford. edu/WWW/www_predict. html) (http: //restools. sdsc. edu/ biotools/biotools 9. html) 29
Structure Prediction Server (http: //cubic. bioc. columbia. edu/predictprotein/) (http: //www. compbio. dun dee. ac. uk/WWW_Servers/ JPred/jpred. html) 30

3 D-Modelling (http: //www. salilab. org/modeller. html) (http: //www. expasy. ch/swissmod/SWISS -MODEL. html) 31

IV. Protein Family Databases Whole Proteins l PIR: Superfamilies and Families l COG (Clusters of Orthologous Groups) of Complete Genomes l Proto. Net: Automated Hierarchical Classification of Proteins Protein Domains l Pfam: Alignments and HMM Models of Protein Domains l SMART: Protein Domain Families Protein Motifs l PROSITE: Protein Patterns and Profiles l BLOCKS: Protein Sequence Motifs and Alignments l PRINTS: Protein Sequence Motifs and Signatures Integrated Family Databases l i. Pro. Class: Superfamilies/Families, Domains, Motifs, Rich Links l Inter. Pro: Integrate Pfam, PRINTS, PROSITES, Pro. Dom, SMART 32
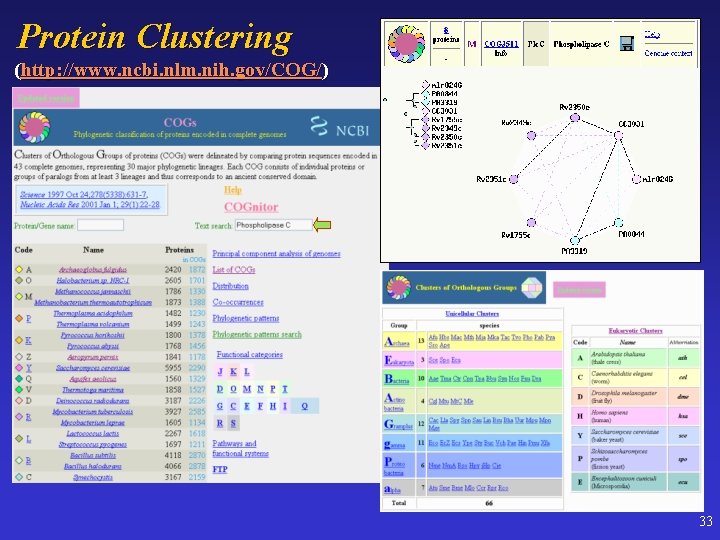
Protein Clustering (http: //www. ncbi. nlm. nih. gov/COG/) 33
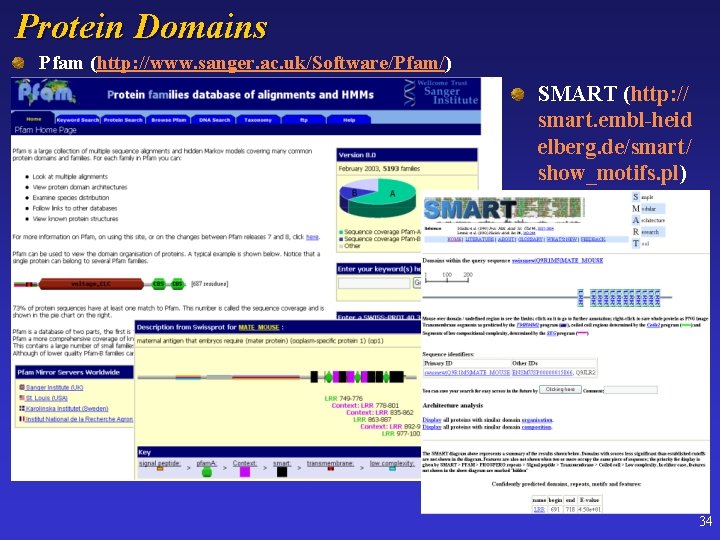
Protein Domains Pfam (http: //www. sanger. ac. uk/Software/Pfam/) SMART (http: // smart. embl-heid elberg. de/smart/ show_motifs. pl) 34
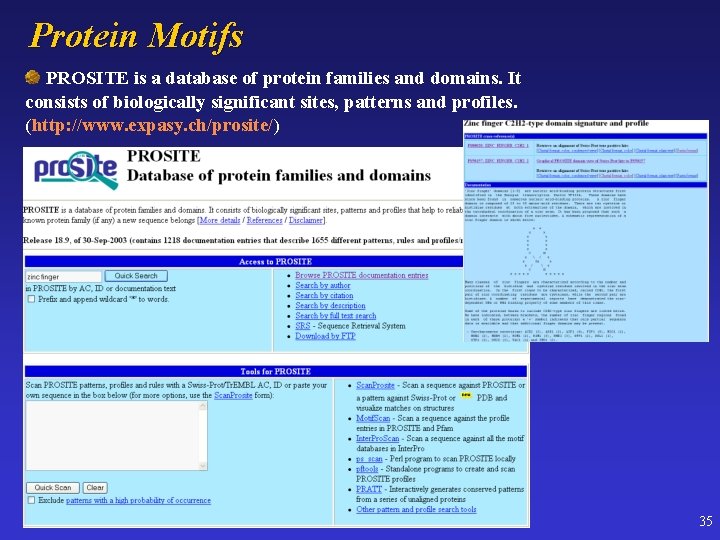
Protein Motifs PROSITE is a database of protein families and domains. It consists of biologically significant sites, patterns and profiles. (http: //www. expasy. ch/prosite/) 35
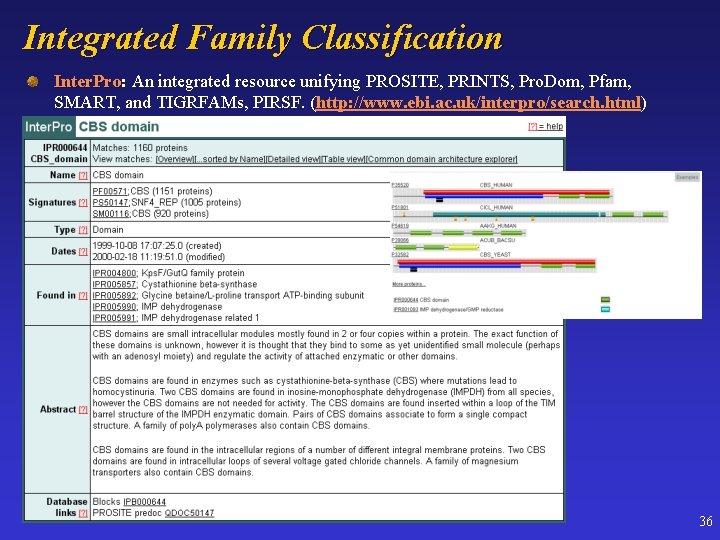
Integrated Family Classification Inter. Pro: Inter. Pro An integrated resource unifying PROSITE, PRINTS, Pro. Dom, Pfam, SMART, and TIGRFAMs, PIRSF. (http: //www. ebi. ac. uk/interpro/search. html) 36

V. Databases of Protein Functions Metabolic Pathways, Enzymes, and Compounds l l l l l Enzyme Classification: Classification and Nomenclature of Enzyme-Catalysed Reactions (EC-IUBMB) KEGG (Kyoto Encyclopedia of Genes and Genomes): Metabolic Pathways LIGAND (at KEGG): Chemical Compounds, Reactions and Enzymes Eco. Cyc: Encyclopedia of E. coli Genes and Metabolism Meta. Cyc: Metabolic Encyclopedia (Metabolic Pathways) WIT: Functional Curation and Metabolic Models BRENDA: Enzyme Database UM-BBD: Microbial Biocatalytic Reactions and Biodegradation Pathways Klotho: Collection and Categorization of Biological Compounds Cellular Regulation and Gene Networks l Epo. DB: Genes Expressed during Human Erythropoiesis BIND: Descriptions of interactions, molecular complexes and pathways l DIP: Catalogs experimentally determined interactions between proteins l Regulon. DB: Escherichia coli Pathways and Regulation l 37
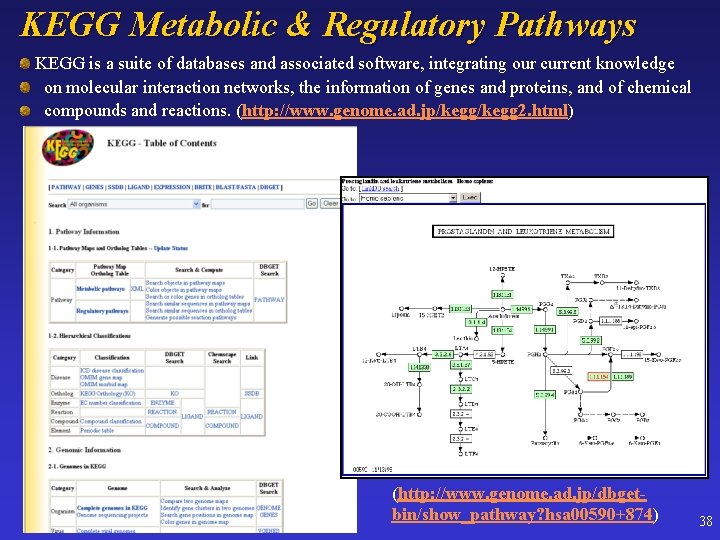
KEGG Metabolic & Regulatory Pathways KEGG is a suite of databases and associated software, integrating our current knowledge on molecular interaction networks, the information of genes and proteins, and of chemical compounds and reactions. (http: //www. genome. ad. jp/kegg 2. html) (http: //www. genome. ad. jp/dbgetbin/show_pathway? hsa 00590+874) 38

Bio. Cyc (Eco. Cyc/Meta. Cyc Metabolic Pathways) The Bio. Cyc Knowledge Library is a collection of Pathway/Genome Databases (http: //biocyc. org/) 39

Protein-Protein Interactions: DIP (http: //dip. doe-mbi. ucla. edu/) 40

Protein-Protein Interaction: BIND (http: //www. bind. ca/) 41

Bio. Carta Cellular Pathways (http: //www. biocarta. com/index. asp) 42

VI. Databases of Protein Structures Protein Structure and Classification l l PDB: Structure Determined by X-ray Crystallography and NMR CATH: Hierarchical Classification of Protein Domain Structures SCOP: Familial and Structural Protein Relationships FSSP: Protein Fold Family Database Protein Sequence-Structure Relationship l l l PIR-NRL 3 D: Protein Sequence-Structure Database PIR-RESID: Protein Structure/Post-Translational Modifications HSSP: Families and Alignments of Structurally-Conserved Regions 43
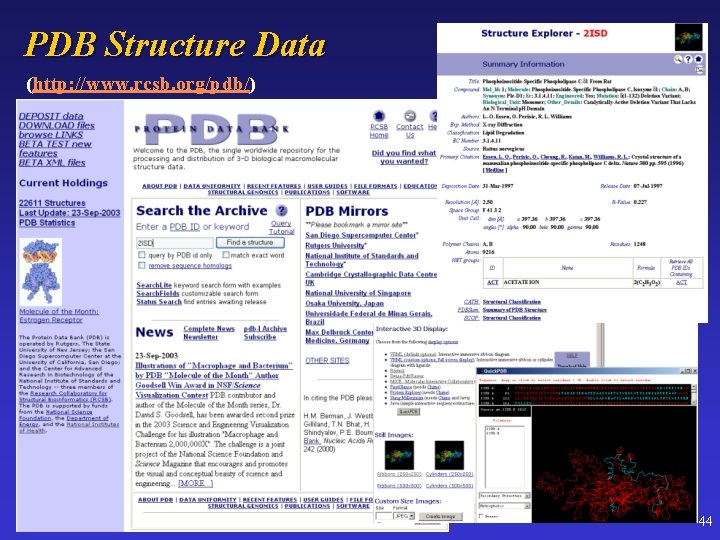
PDB Structure Data (http: //www. rcsb. org/pdb/) 44
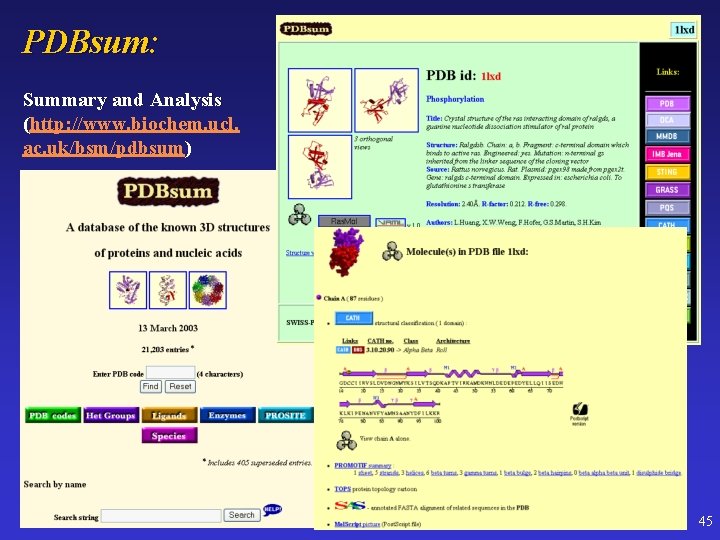
PDBsum: Summary and Analysis (http: //www. biochem. ucl. ac. uk/bsm/pdbsum) 45
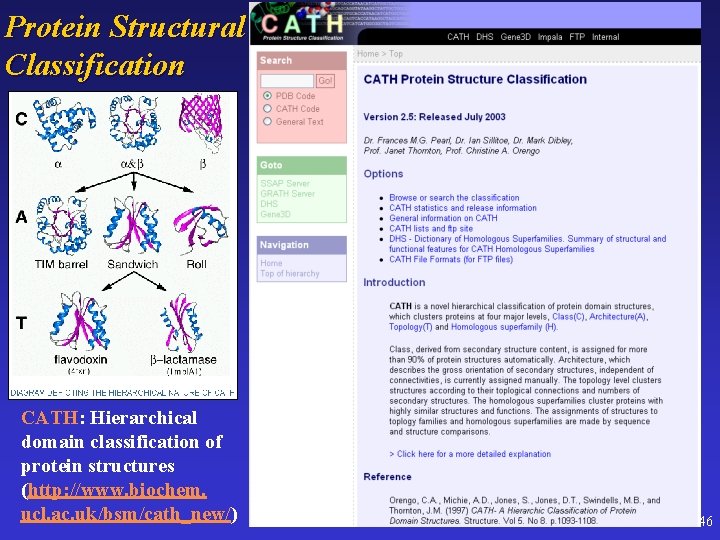
Protein Structural Classification CATH: Hierarchical domain classification of protein structures (http: //www. biochem. ucl. ac. uk/bsm/cath_new/) 46
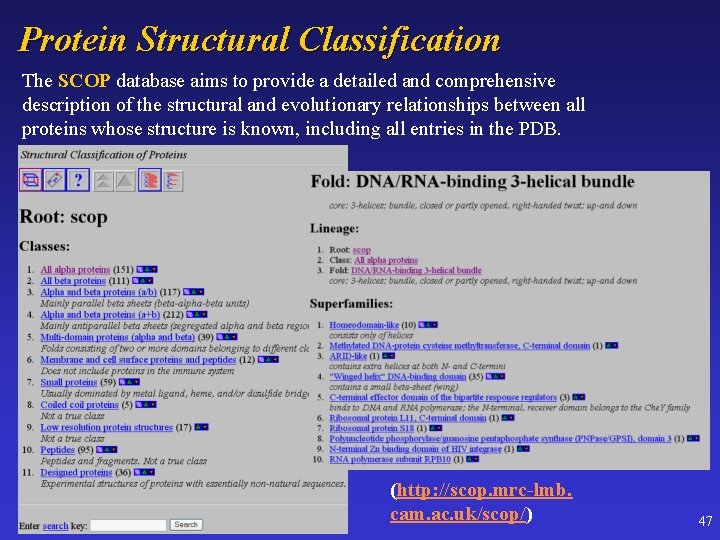
Protein Structural Classification The SCOP database aims to provide a detailed and comprehensive description of the structural and evolutionary relationships between all proteins whose structure is known, including all entries in the PDB. (http: //scop. mrc-lmb. cam. ac. uk/scop/) 47

VII. Proteomic Resources GELBANK (http: //gelbank. anl. gov): 2 D-gel patterns from completed genomes; SWISS-2 DPAGE (http: //www. expasy. org/ch 2 d/) PEP: Predictions for Entire Proteomes: (http: //cubic. bioc. columbia. edu/ pep/): Summarized analyses of protein sequences Proteome Bio. Knowledge Library: (http: //www. proteome. com): Detailed information on human, mouse and rat proteomes Proteome Analysis Database (http: //www. ebi. ac. uk/proteome/): Online application of Inter. Pro and Clu. STr for the functional classification of proteins in whole genomes Expression Profiling databases: GNF (http: //expression. gnf. org/cgibin/index. cgi, human and mouse transcriptome), SMD (http: //genomewww 5. stanford. edu/Micro. Array/SMD/, Stanford microarray data analysis), EBI Microarray Informatics (http: //www. ebi. ac. uk/microarray/ index. html , managing, storing and analyzing microarray data) 48
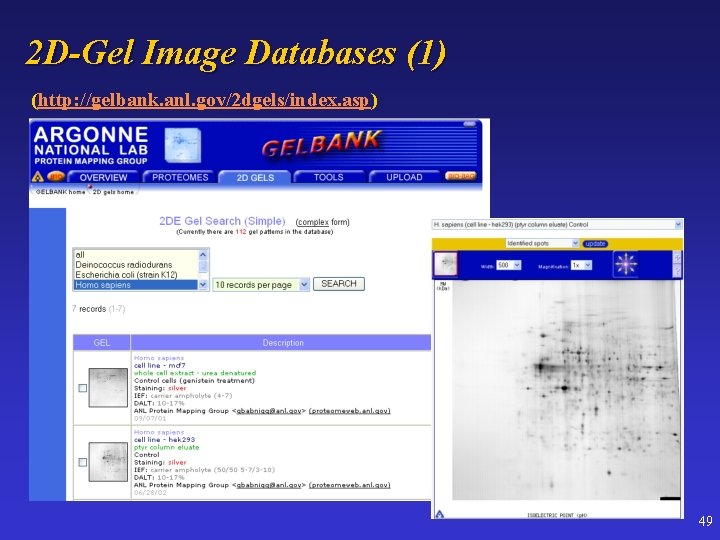
2 D-Gel Image Databases (1) (http: //gelbank. anl. gov/2 dgels/index. asp) 49
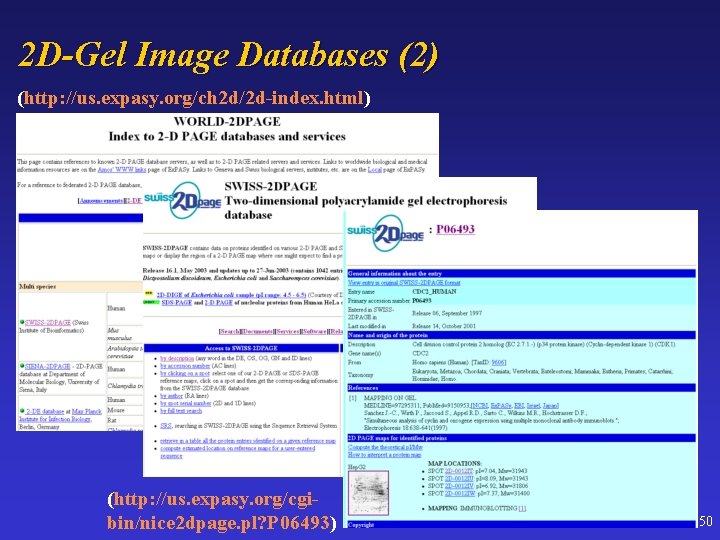
2 D-Gel Image Databases (2) (http: //us. expasy. org/ch 2 d/2 d-index. html) (http: //us. expasy. org/cgibin/nice 2 dpage. pl? P 06493) 50

VIII. Proteome Analysis (http: //www. ebi. ac. uk/proteome) 51
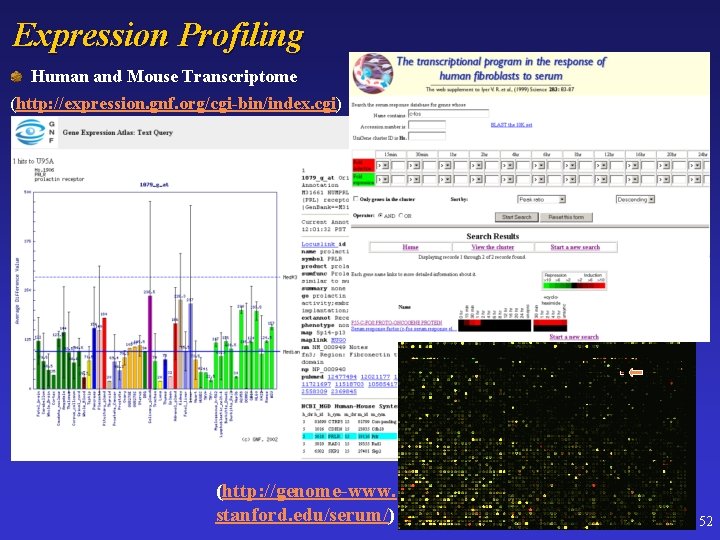
Expression Profiling Human and Mouse Transcriptome (http: //expression. gnf. org/cgi-bin/index. cgi) (http: //genome-www. stanford. edu/serum/) 52

Lab: Visit selected websites and analyze some protein sequences of your own choices. - List of Bioinformatics Resources of this tutorial available: http: //pir. georgetown. edu/~huz/bioinfo_resource. html Try some of the following sequences for analysis: 1) well characterized proteins: PIR: A 26366(CYP 17), JS 0747(Sp 1) 2) less characterized proteins: PIR: A 59000(MATER) Tr. EMBL: Q 9 QY 16(GRTH) 3) hypothetical protein: PIR: T 12515, T 00338 , T 47130 SWISS-PROT: Q 9 BWT 7 53
- Slides: 53