Tumour karyotype Spectral karyotyping showing chromosomal aberrations in
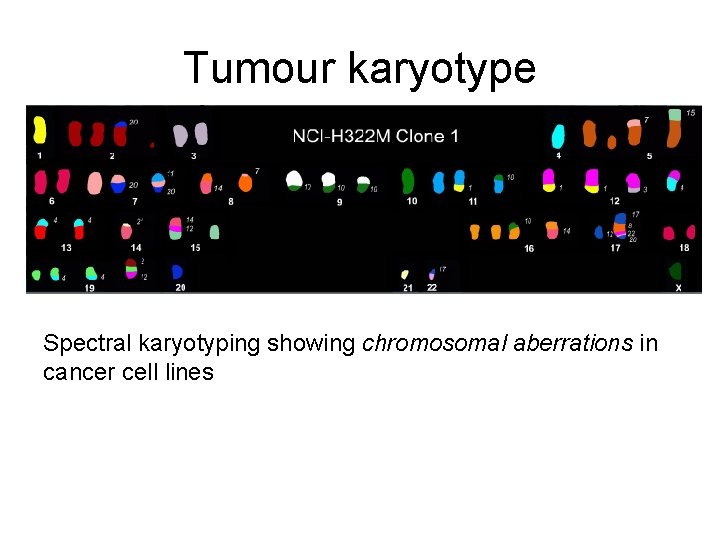
Tumour karyotype Spectral karyotyping showing chromosomal aberrations in cancer cell lines

Chromosomal aberrations • Segments of DNA that get duplicated – Gains • Segments of DNA that get deleted – Losses • Chromosomal aberrations are being investigated as diagnostic indicators of cancer and other diseases – Better diagnosis of disease – Potentially reveals biomolecular mechanisms of disease • This research is done using: – array comparative genomic hybridization (a. CGH) – Measures DNA copy number changes
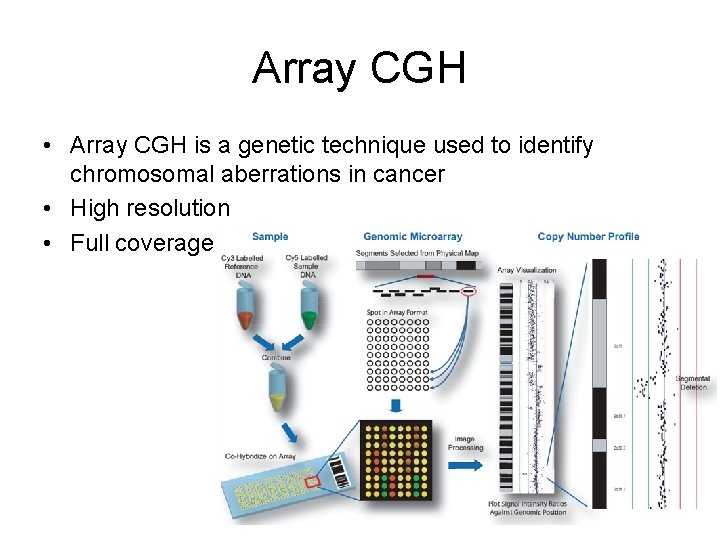
Array CGH • Array CGH is a genetic technique used to identify chromosomal aberrations in cancer • High resolution • Full coverage

Array CGH data • Measures log 2 ratios of normal vs sample for prespecified segments of the genome called clones – Theoretical log 2 ratios: • 1 copy gain (duplication) = log(3/2) = 0. 58 • Neutral = log (2/2) = 0 • 1 copy loss (deletion) = log(1/2) = -1 • Measurement based on detection of hybridization level to probes on an array • ~30, 000 measurements per sample
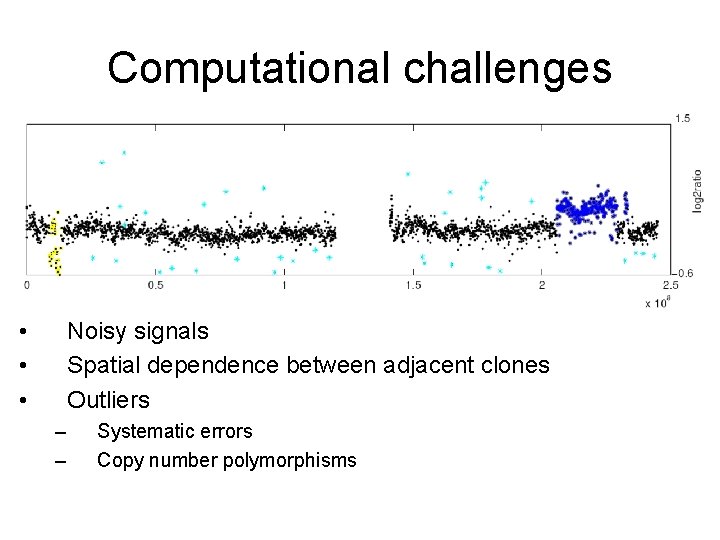
Computational challenges • • • Noisy signals Spatial dependence between adjacent clones Outliers – – Systematic errors Copy number polymorphisms

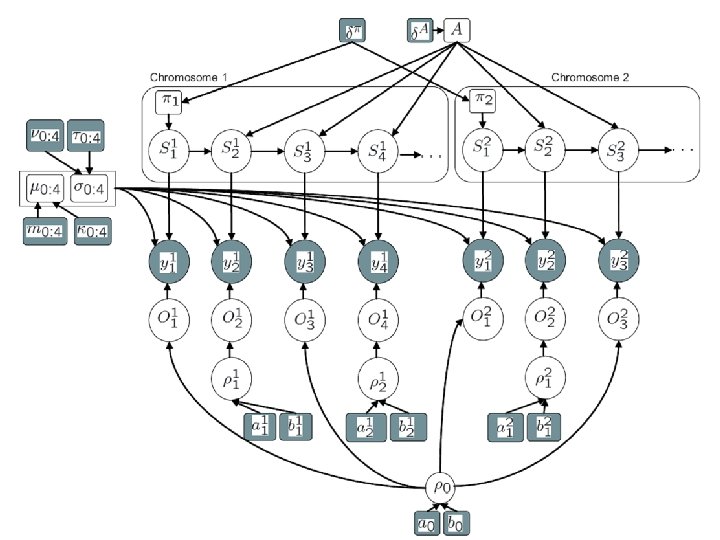
Our approach • Use a “supervised” hidden Markov model (HMM) to model spatial dependency between clones – States are loss, neutral, gain-one, gain-many – Infer the unobserved state sequence from the data using a standard efficient algorithm called forwards-backwards – This part of the model is standard v Use a Gaussian mixture model to model the outliers separately from the inliers • Inliers have spatial dependence • Outliers do not v Use prior knowledge about locations of CNPs to ‘inform’ the model about possible locations of outliers • Several published lists of CNPs are available • Internally generated list more comprehensive v Pool data across chromosomes to gain “statistical strength”

Test data • Mantle cell lymphoma cell lines with “ground truth” • 123 losses and 72 gains covering approximately 1% of the human genome • Compare results to state-of-the-art algorithms
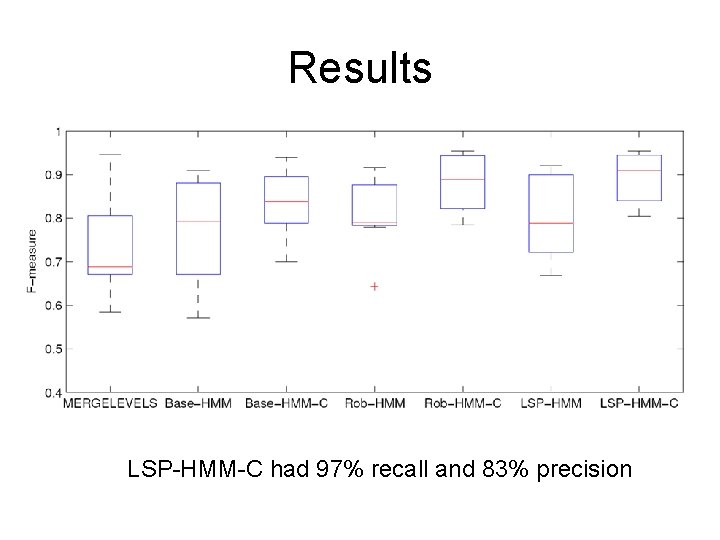
Results LSP-HMM-C had 97% recall and 83% precision
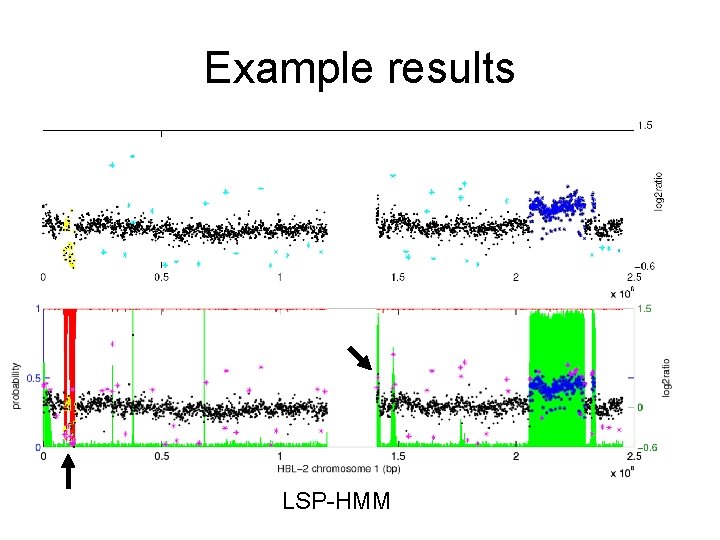
Example results LSP-HMM
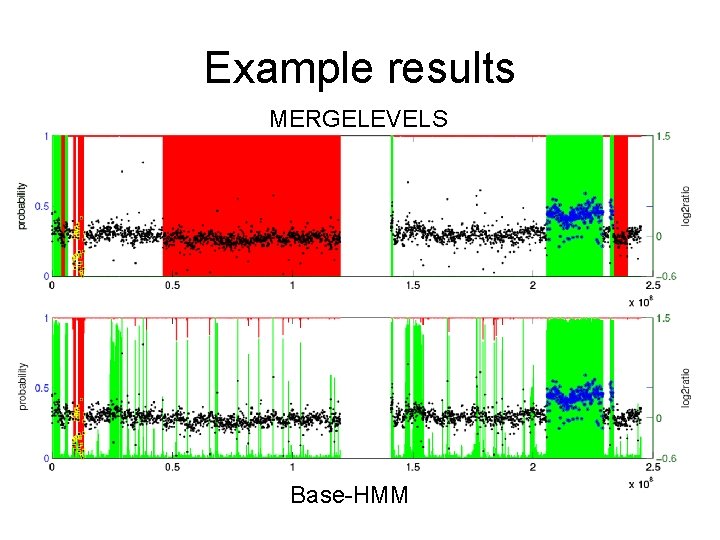
Example results MERGELEVELS Base-HMM

Conclusions • HMM framework superior to Merge. Levels • Adding robustness further improves performance over the ‘Base-HMM’ • Adding LSP information improves performance marginally over robust HMM, but importantly does not make results worse – Motivation for using more comprehensive lists to improve results – Some false positives – are they real? • Pooling across chromosomes results in big gains – Data more easily overwhelms incorrect priors – Makes the algorithm less sensitive to parameter settings
- Slides: 12