Tumor Mutation Burder as a predictive biomarker Dr

Tumor Mutation Burder as a predictive biomarker Dr Francesco Pantano, MD Ph. D Medical Oncology Department Campus Bio Medico of Rome
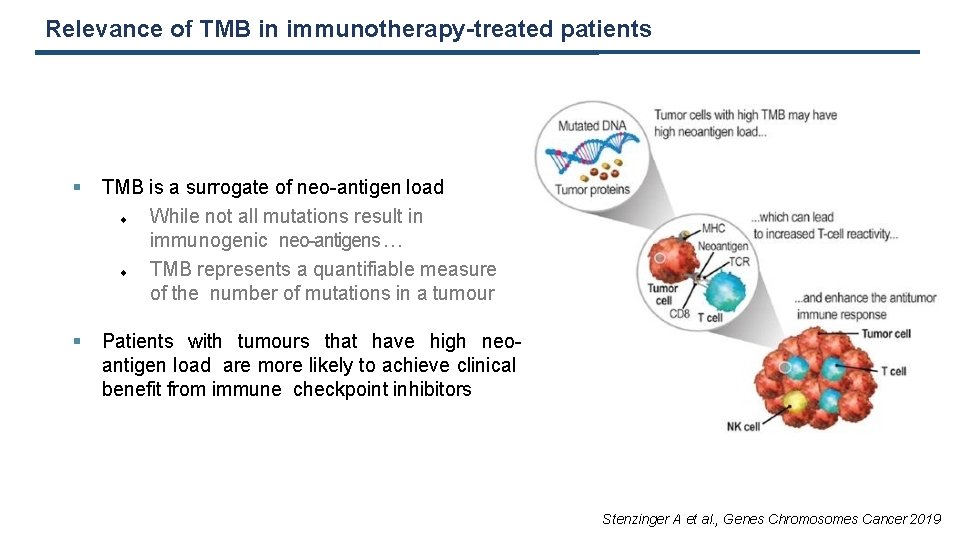
Relevance of TMB in immunotherapy-treated patients TMB is a surrogate of neo-antigen load While not all mutations result in immunogenic neo-antigens … TMB represents a quantifiable measure of the number of mutations in a tumour Patients with tumours that have high neoantigen load are more likely to achieve clinical benefit from immune checkpoint inhibitors Stenzinger A et al. , Genes Chromosomes Cancer 2019
Evolution of TMB as a biomarker Chan et al. , Annals of Oncology 2018
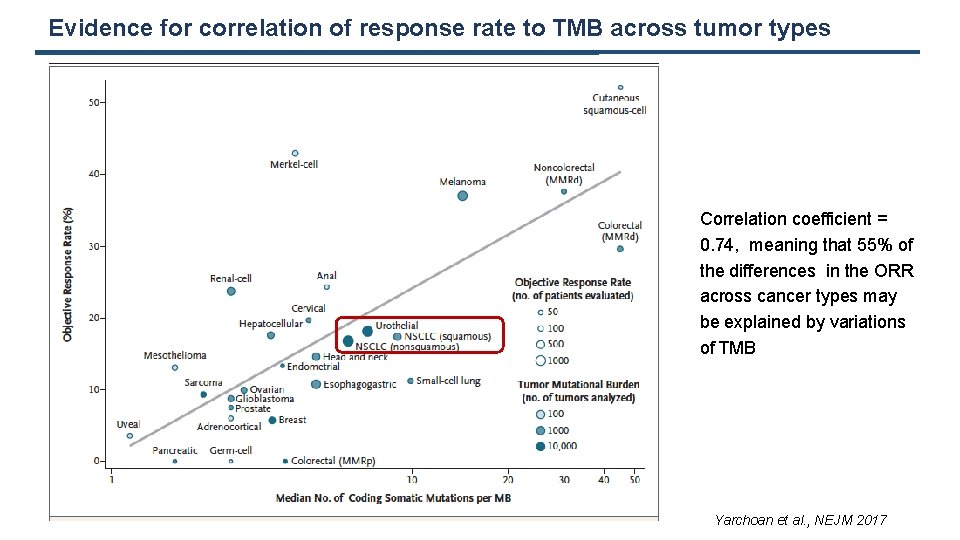
Evidence for correlation of response rate to TMB across tumor types Correlation coefficient = 0. 74, meaning that 55% of the differences in the ORR across cancer types may be explained by variations of TMB Yarchoan et al. , NEJM 2017

NSCLC is associated to a high TMB Schumacher and Schreiber, Science 2015

Data from clinical trials and cohorts Impact of TMB on response, duration of response, PFS and OS with ICIs
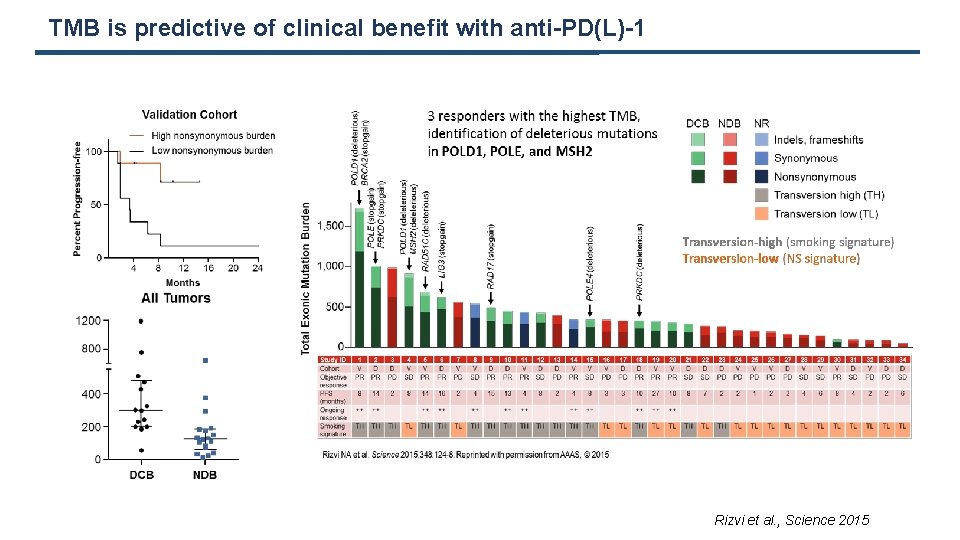
TMB is predictive of clinical benefit with anti-PD(L)-1 Rizvi et al. , Science 2015
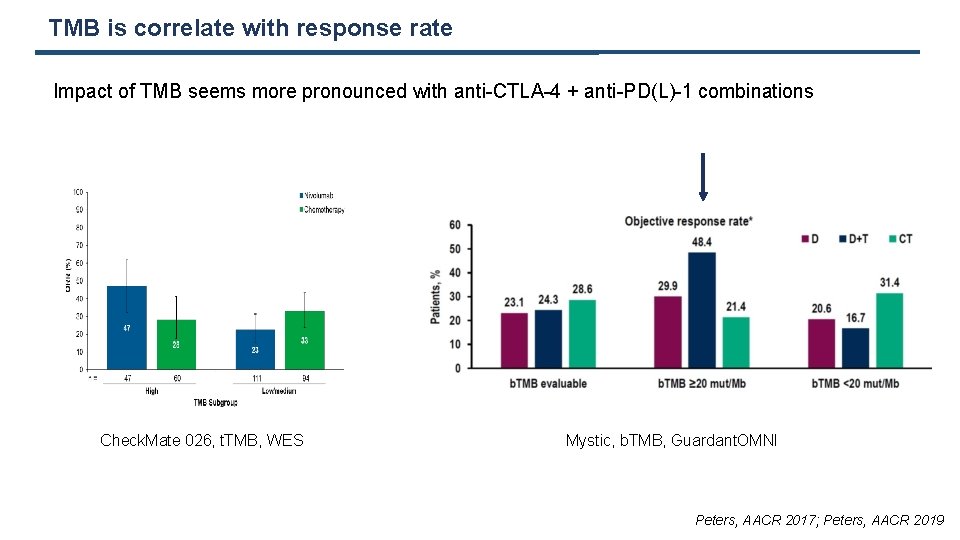
TMB is correlate with response rate Impact of TMB seems more pronounced with anti-CTLA-4 + anti-PD(L)-1 combinations Check. Mate 026, t. TMB, WES Mystic, b. TMB, Guardant. OMNI Peters, AACR 2017; Peters, AACR 2019
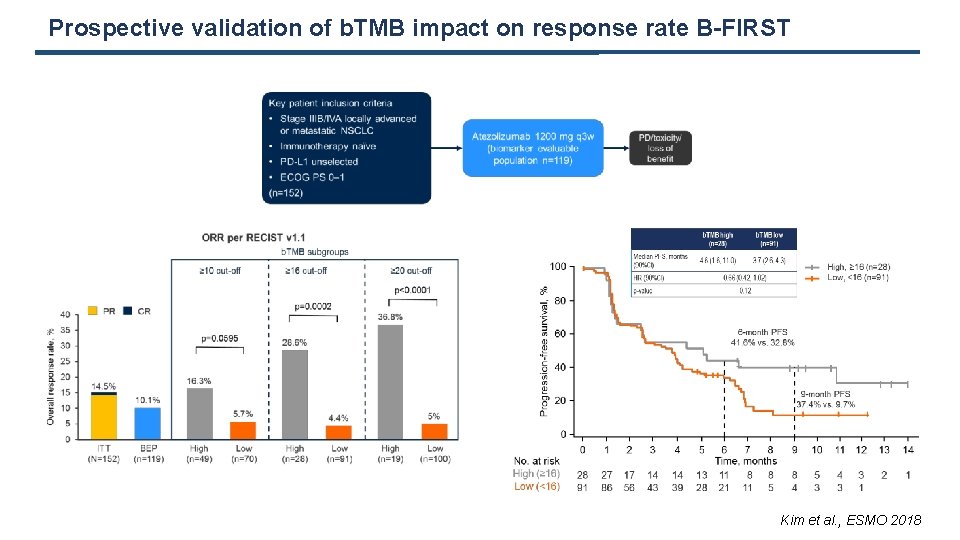
Prospective validation of b. TMB impact on response rate B-FIRST Kim et al. , ESMO 2018
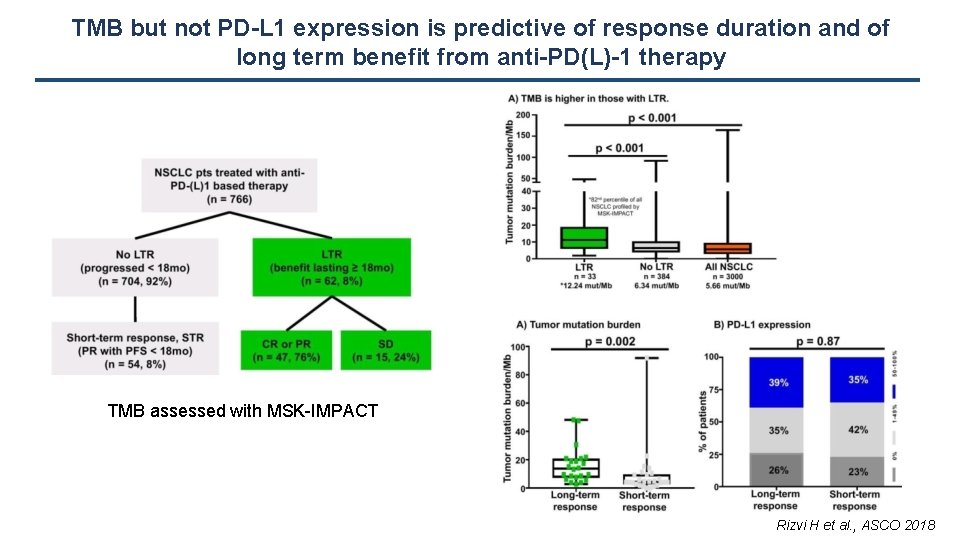
TMB but not PD-L 1 expression is predictive of response duration and of long term benefit from anti-PD(L)-1 therapy TMB assessed with MSK-IMPACT Rizvi H et al. , ASCO 2018
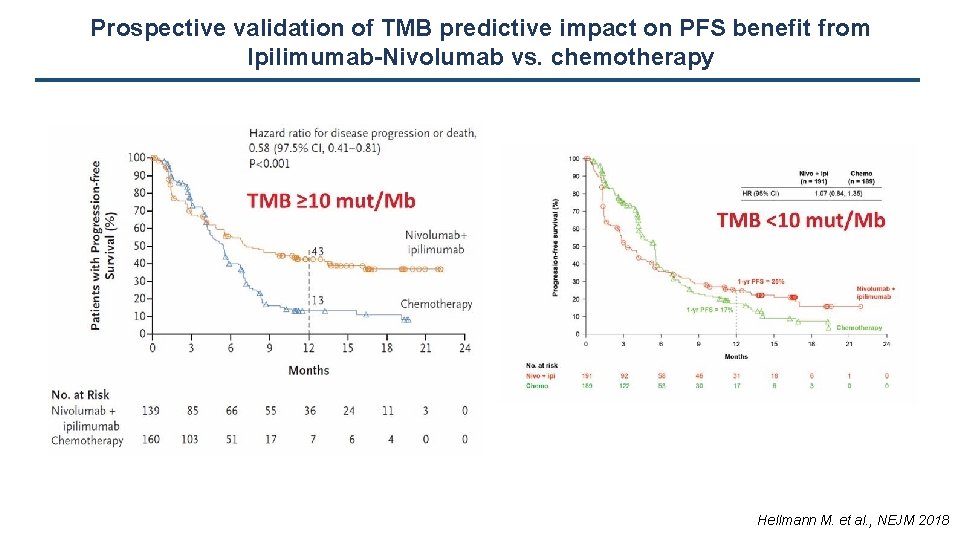
Prospective validation of TMB predictive impact on PFS benefit from Ipilimumab-Nivolumab vs. chemotherapy Hellmann M. et al. , NEJM 2018
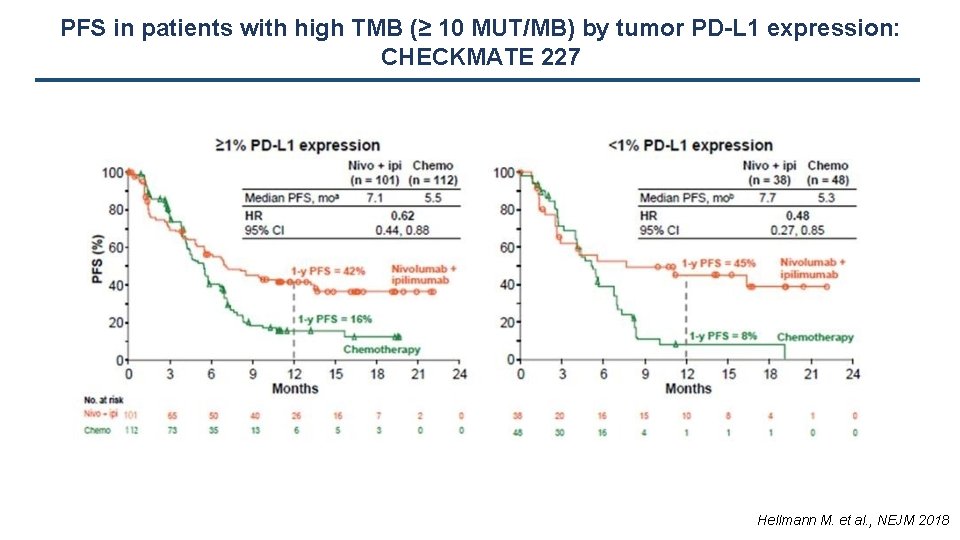
PFS in patients with high TMB (≥ 10 MUT/MB) by tumor PD-L 1 expression: CHECKMATE 227 Hellmann M. et al. , NEJM 2018
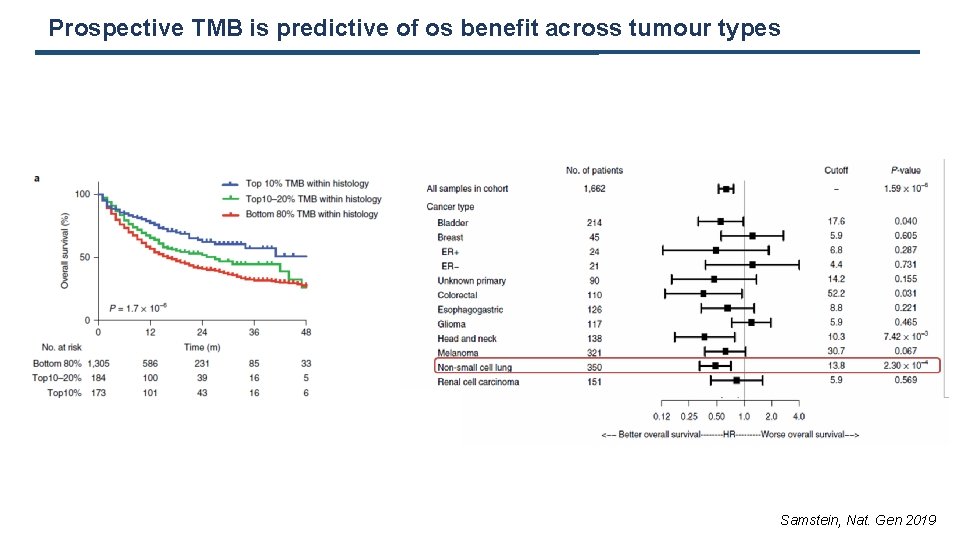
Prospective TMB is predictive of os benefit across tumour types Samstein, Nat. Gen 2019
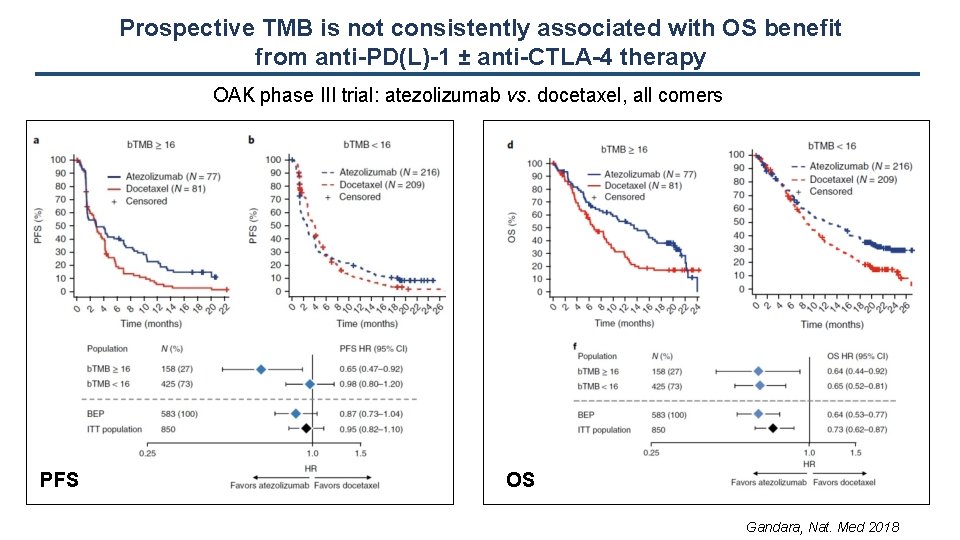
Prospective TMB is not consistently associated with OS benefit from anti-PD(L)-1 ± anti-CTLA-4 therapy OAK phase III trial: atezolizumab vs. docetaxel, all comers PFS OS Gandara, Nat. Med 2018
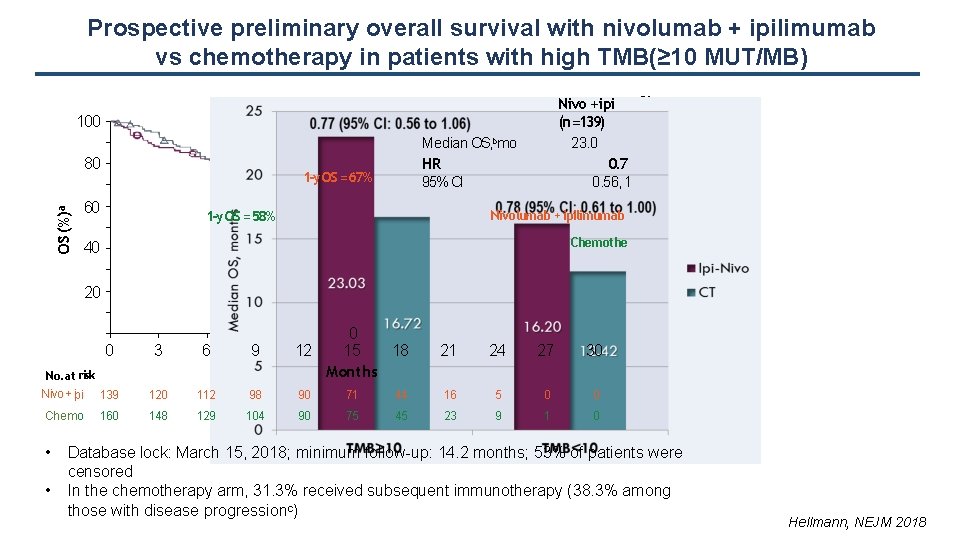
Prospective preliminary overall survival with nivolumab + ipilimumab vs chemotherapy in patients with high TMB(≥ 10 MUT/MB) Chemo Nivo +ipi (n = 160) (n =139) 23. 0 16. 4 0. 79 0. 56, 1. 10 100 OS (%)a 80 Median OS, bmo HR 95% CI 1 -y OS = 67% 60 Nivolumab + ipilimumab 1 -y OS = 58% Chemothe rapy 40 20 3 6 9 No. at risk Nivo + ipi 139 0 12 15 18 Months 120 112 98 90 71 44 16 5 0 0 Chemo 148 129 104 90 75 45 23 9 1 0 0 • • 160 21 24 27 30 Database lock: March 15, 2018; minimum follow-up: 14. 2 months; 53% of patients were censored In the chemotherapy arm, 31. 3% received subsequent immunotherapy (38. 3% among those with disease progressionc) Hellmann, NEJM 2018
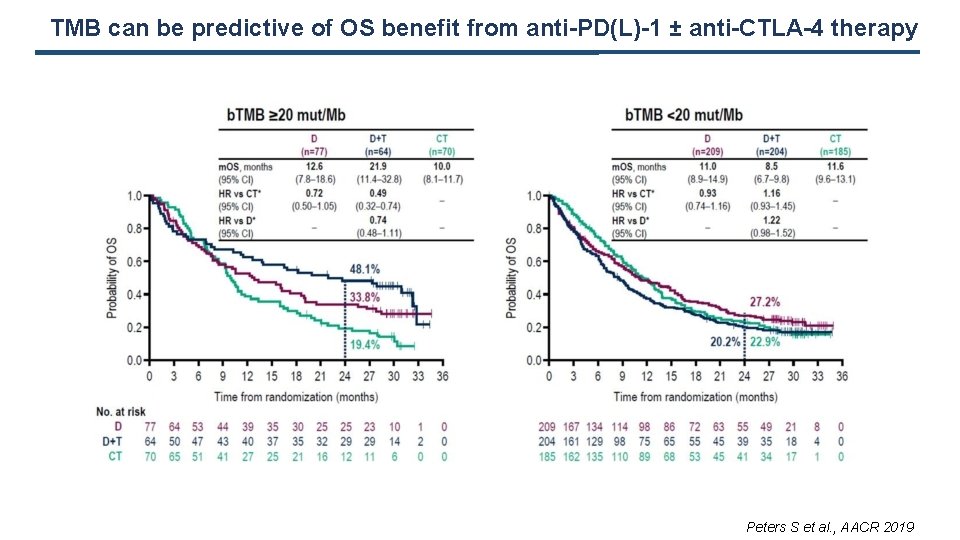
TMB can be predictive of OS benefit from anti-PD(L)-1 ± anti-CTLA-4 therapy Peters S et al. , AACR 2019

Why TMB would not predict OS benefit from ICIs in NSCLS whereas it is more consistently predictive of PFS benefit? Prognostic role? Selection of the cut-off? Role of cross over? Similar than EGFR TKI vs. chemotherapy in EGFR mutated tumors? High TMB tumours might generate more rapidly acquired resistance to ICIs due to genomic instability Role of subclonal mutations Not consistent with correlation of TMB with long term responses No data from trials assessing ICIs + chemotherapy combinations yet (ASCO 2019) Better correlation of PFS and OS benefits

TMB: the next biomarker in addition to PD-L 1? A way to refine treatment algorithm for 1 st line treatment of advanced NSCLC
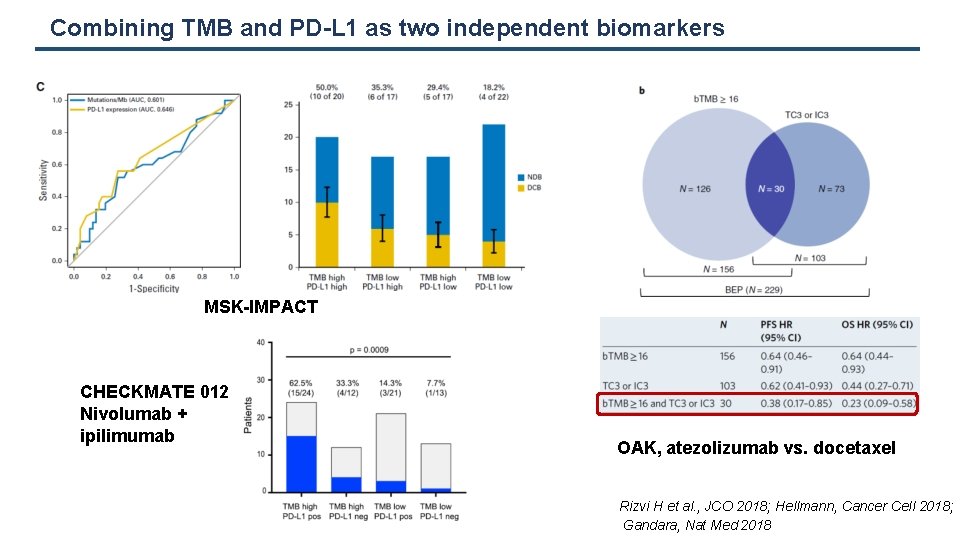
Combining TMB and PD-L 1 as two independent biomarkers MSK-IMPACT CHECKMATE 012 Nivolumab + ipilimumab OAK, atezolizumab vs. docetaxel Rizvi H et al. , JCO 2018; Hellmann, Cancer Cell 2018; Gandara, Nat Med 2018
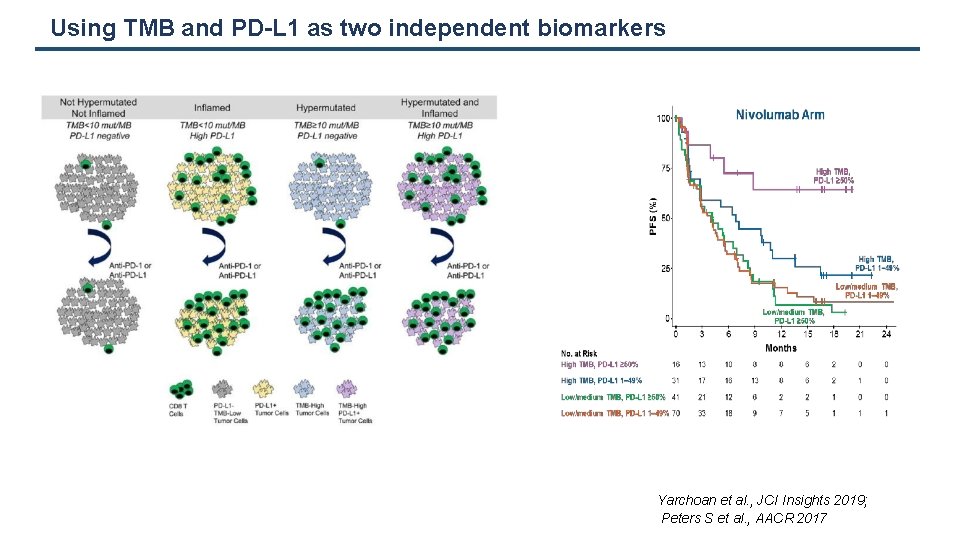
Using TMB and PD-L 1 as two independent biomarkers Yarchoan et al. , JCI Insights 2019; Peters S et al. , AACR 2017
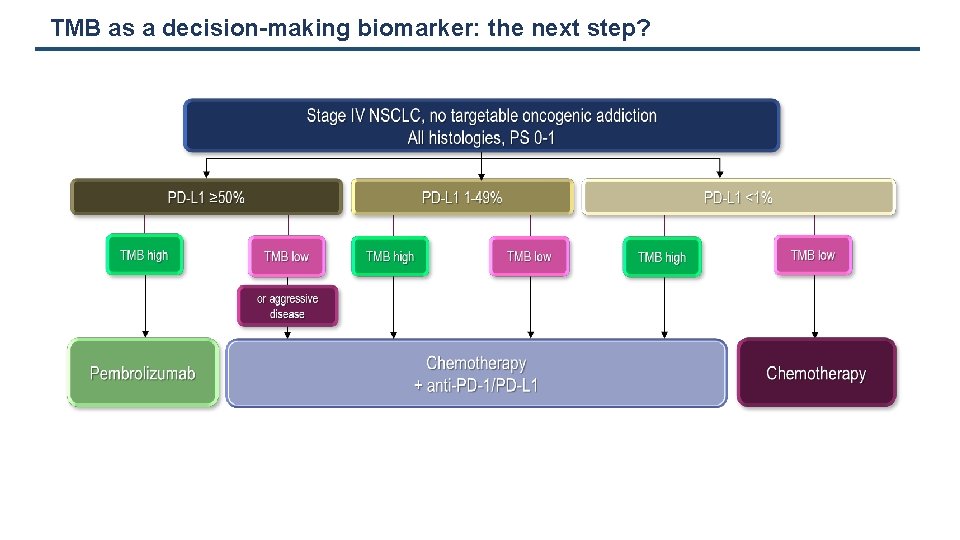
TMB as a decision-making biomarker: the next step?

TMB assessment: the next biomarker? The available data support the hypothesis that the number of nonsynonymous mutations (and ultimately neoantigens) significantly correlates with clinical response to immune checkpoint blockade TMB is a continuous biomarker, appearing as an acceptable surrogate of neoantigen load Consistent correlation with response rates, duration of response, PFS benefit with anti-PD(L)-1 in NSCLC and across tumour types Correlation with OS benefit from ICIs is more variable in randomized trials vs. chemotherapy Predictive impact on ICI benefit seems more pronounced with anti-PD(L)-1 + anti-CTLA-4 combinations Independent of PD-L 1 expression: TMB will not replace PD-L 1 but will supplement it Could be used in addition to PD-L 1 expression to refine 1 st line treatment algorithm and selection of patients Many remaining issues but advances in technology make possible to expect using soon TMB in clinical practice

Measuring TMB Whole genome/exome sequencing with “paired normal” Number of variants per megabase - count and divide by coverage Can refine the count by excluding select alterations: Intronic/intragenic Germline Synonymous Chan, Ann Oncol 2019 Chalmers, Genome Med 2017

Measuring TMB “Large” targeted panels (without “paired normal”) Same assumptions ; HOWEVER: Use predictive approach to remove germline variants Include synonymous variants Exclude known “pathogenic” variants to account for presumed bias (use COSMIC filter) Chan, Ann Oncol 2019 Chalmers, Genome Med 2017
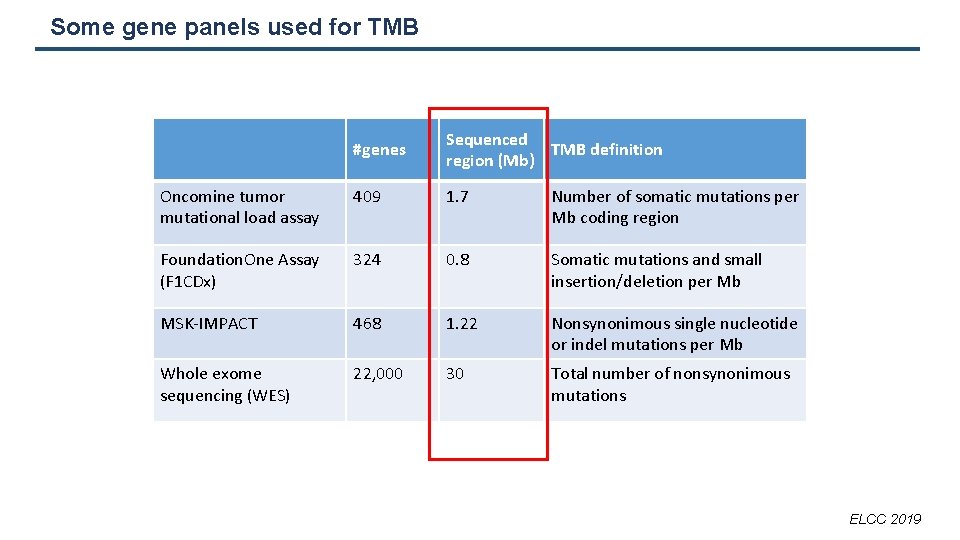
Some gene panels used for TMB #genes Sequenced TMB definition region (Mb) Oncomine tumor mutational load assay 409 1. 7 Number of somatic mutations per Mb coding region Foundation. One Assay (F 1 CDx) 324 0. 8 Somatic mutations and small insertion/deletion per Mb MSK-IMPACT 468 1. 22 Nonsynonimous single nucleotide or indel mutations per Mb Whole exome sequencing (WES) 22, 000 30 Total number of nonsynonimous mutations ELCC 2019
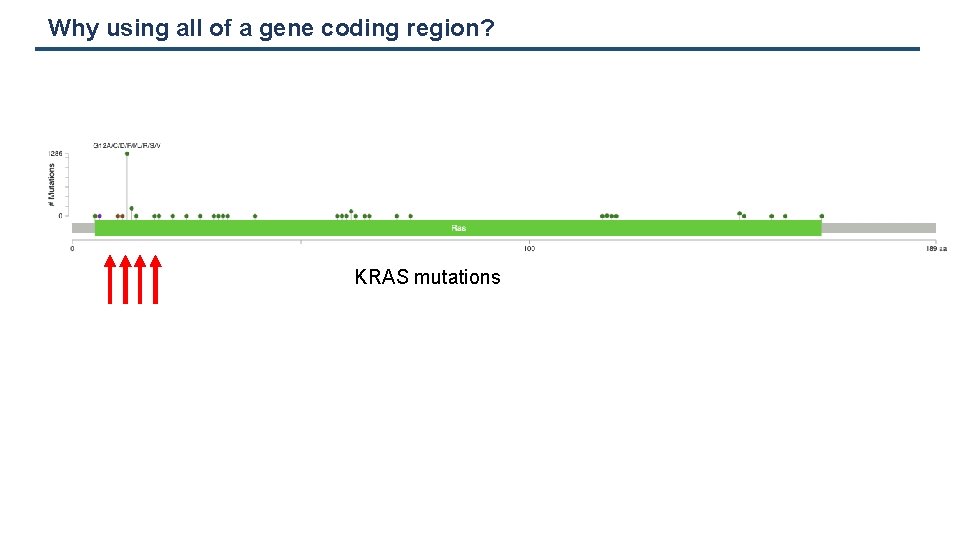
Why using all of a gene coding region? KRAS mutations
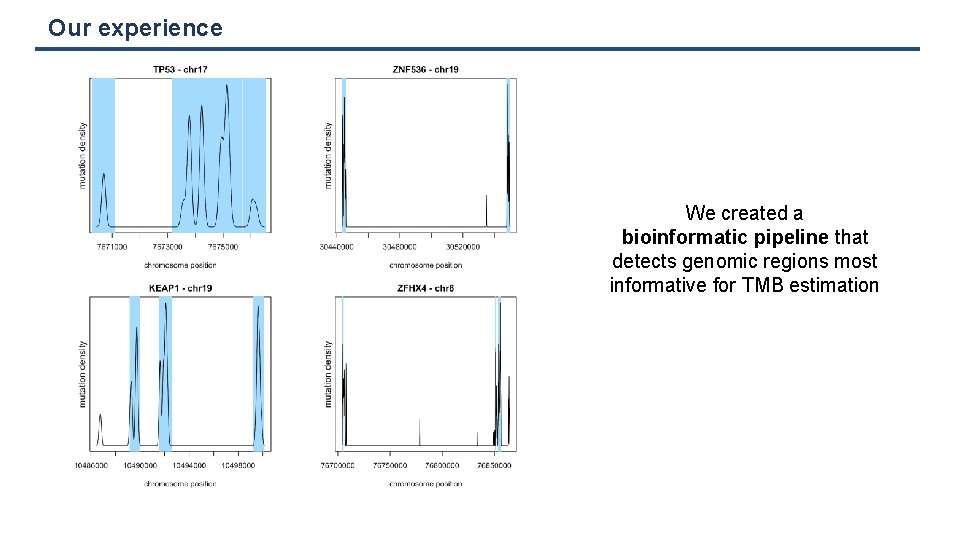
Our experience We created a bioinformatic pipeline that detects genomic regions most informative for TMB estimation
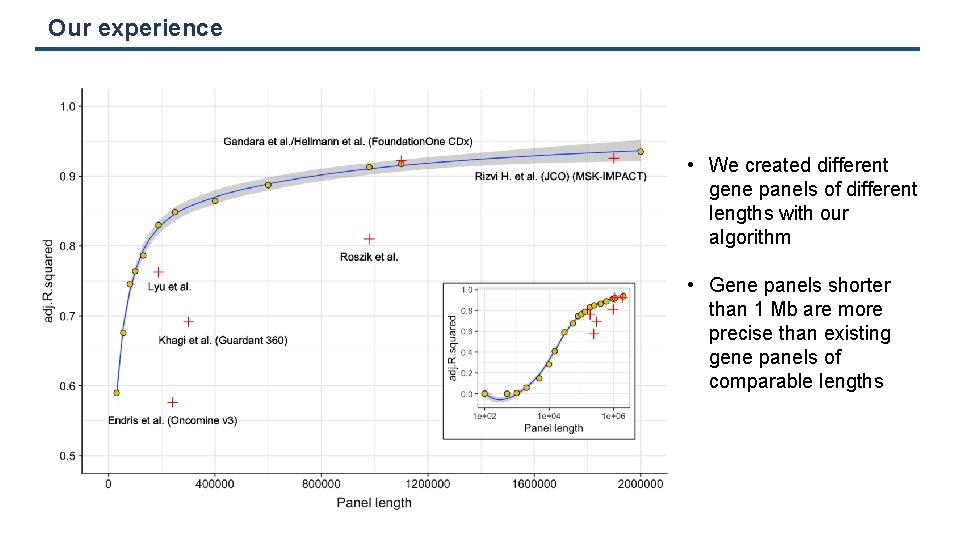
Our experience • We created different gene panels of different lengths with our algorithm • Gene panels shorter than 1 Mb are more precise than existing gene panels of comparable lengths
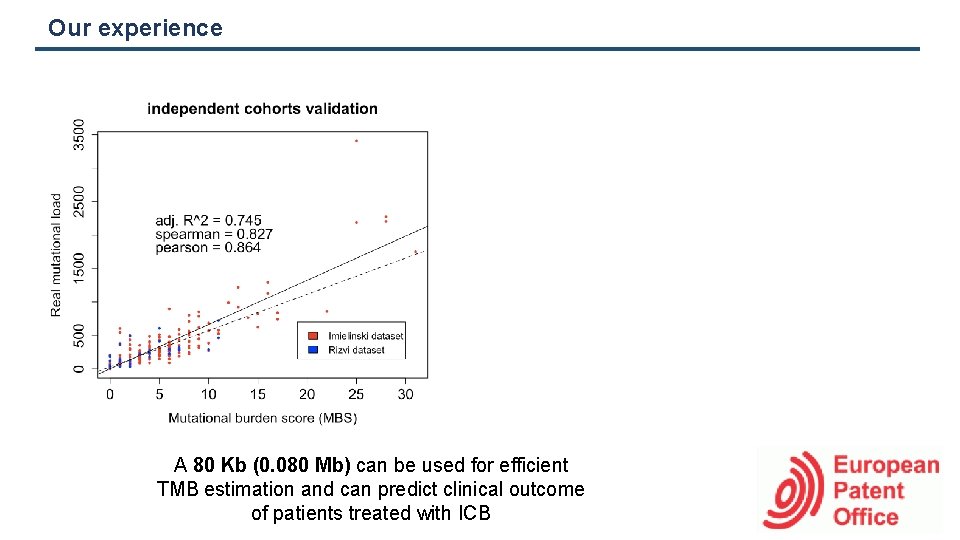
Our experience A 80 Kb (0. 080 Mb) can be used for efficient TMB estimation and can predict clinical outcome of patients treated with ICB

Conclusions Shrinking the amount of DNA needed for TMB estimation might: Reduce turn around times Lower Costs Enable blood-TMB estimation

Special thanks to: Prof. Giuseppe Tonini Prof. Daniele Santini Prof. Bruno Vincenzi Dr. Paolo Manca Translational Oncology Lab: Dr. Michele Iuliani Dr. Giulia Ribelli Dr. Sonia Simonetti 31
- Slides: 31