Trich Laboratory Testing Standardization Trichomonas foetus DNA testing

Trich Laboratory Testing Standardization Trichomonas foetus DNA testing Proper & Confidential Proprietary & Confidential 1

Trich Laboratory Testing Standardization Objectives § To build confidence in Trich testing § To increase consistency in test results § To minimize sample quality influence variables § To minimize laboratory testing variables § To reduce the burden and cost to man and beast Proper & Confidential Proprietary & Confidential 2

Trich Laboratory Testing Standardization Opportunities § Veterinarian sample collection § Veterinarian sample handling & shipment recommended by Laboratory § Laboratory sample preparation method § Laboratory extract volume used § Laboratory Taq enzyme type used § Laboratory analysis threshold level § Laboratory use of IPC (Internal Positive Control) § Laboratory use of USDA licensed kits § Laboratory pooling of samples Proprietary & Confidential Proper & Confidential 3

Pooling of cultured samples and comparison of multistate laboratory workflows with the Mag. MAX sample preparation system and Vet. MAX quantitative polymerase chain reaction reagents for detection of Tritrichomonas foetus–colonized bulls Lee Effinger Lalitha Peddireddi Marilyn Simunich Richard Oberst Catherine O’Connell Ivan Leyva-Baca Oregon Department of Agriculture, Animal Health and Identification Division, Animal Health Laboratory, Salem, OR (Effinger) Department of Diagnostic Medicine/Pathobiology (Peddireddi), Kansas State University, Manhattan, KS Kansas State Veterinary Diagnostic Laboratory (Oberst), Kansas State University, Manhattan, KS Animal Health Laboratory, Idaho State Department of Agriculture, Boise, ID (Simunich) Animal Health and Food Safety Group at Life Technologies, Austin, TX (Leyva-Baca, O’Connell) JVDI, 2014, Vol. 26(1) 72 -87 Proper & Confidential Proprietary & Confidential The world leader in serving science Proprietary & Confidential 4

Study Background 2010 AAVLD parasitology committee • Proposed a study to determine whether T. foetus samples can be pooled in order to reduce the costs for testing • Lee Effinger from Oregon State Department of Agriculture led Experimental Design for the project • Marilyn Simunich Idaho State Department of Agriculture served as Study Coordinator & Data Keeper • The Life Technologies Animal Health & Food Safety Group agreed to support the study Proper & Confidential Proprietary & Confidential 5

Study Objectives 1. Determine the effect of pooling a single positive sample having various CT ranges with four negative samples (1: 5). If a negative effect was seen, a 1: 3 pooling study would then be conducted 2. Compare different sample preparation systems and various real-time PCR (feeder lab workflows) with the 5 X Mag. MAXTM-pathogen RNA/DNA purification kit and amplification with Vet. MAXTM T. foetus reagents (Life Technologies workflow) 3. Assess the specificity of the Vet. MAXTM T. foetus reagents by sequencing all positive samples with CT values less than 38 and suspect sample CT values between 38 and less than 40 cycles Proprietary & Confidential Proper & Confidential 6

Materials and Methods • Sample collection (Cultured Smegma Samples) • 5 Feeder labs provided 803 samples • 1 on the West Coast • 1 in the Southwest • 1 in the Central States • 2 in the South • Each feeder lab ran their own protocol including sample preparation system and real-time PCR § 1 Central Study lab (KSVDL) § Sample preparation with Mag. MAXTM § Real-time PCR with Vet. MAXTM T. foetus reagents Proprietary & Confidential Proper & Confidential 7
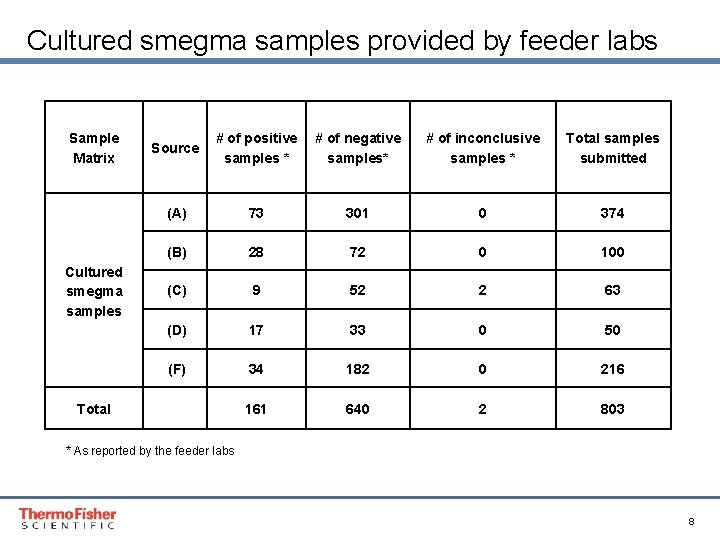
Cultured smegma samples provided by feeder labs Sample Matrix Cultured smegma samples Source # of positive # of negative samples * samples* # of inconclusive samples * Total samples submitted (A) 73 301 0 374 (B) 28 72 0 100 (C) 9 52 2 63 (D) 17 33 0 50 (F) 34 182 0 216 161 640 2 803 Total * As reported by the feeder labs Proprietary & Confidential Proper & Confidential 8
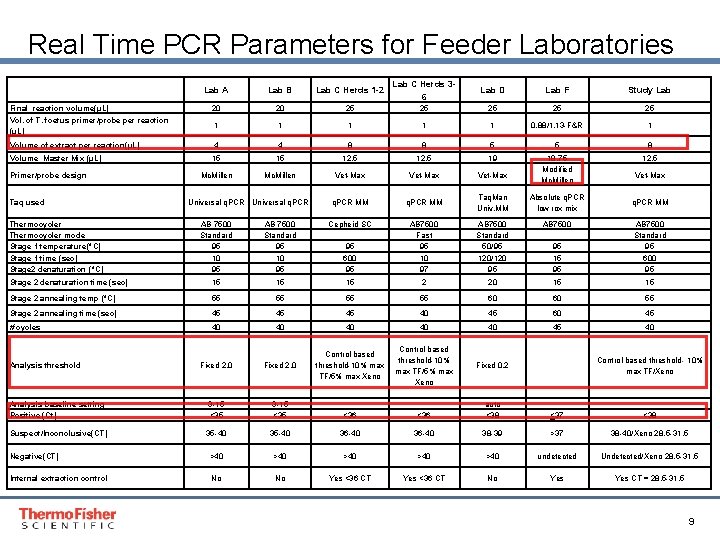
Real Time PCR Parameters for Feeder Laboratories Lab A Lab B Lab C Herds 1 -2 Lab C Herds 36 Lab D Lab F Study Lab Final reaction volume(µL) Vol. of T. foetus primer/probe per reaction (µL) 20 20 25 25 25 1 1 1 0. 88/1. 13 -F&R 1 Volume of extract per reaction(µL) 4 4 8 8 5 5 8 Volume Master Mix (µL) 15 15 12. 5 19 12. 5 Mc. Millen Vet-Max 18. 75 Modified Mc. Millen q. PCR MM Taq. Man Univ. MM Absolute q. PCR low rox mix AB 7500 Standard 50/95 120/120 95 AB 7500 95 15 95 AB 7500 Standard 95 600 95 Primer/probe design Taq used Thermocycler mode Stage 1 temperature(°C) Stage 1 time (sec) Stage 2 denaturation (°C) Universal q. PCR Vet-Max q. PCR MM AB 7500 Standard 95 10 95 Cepheid SC 95 600 95 AB 7500 Fast 95 10 97 Stage 2 denaturation time (sec) 15 15 15 2 20 15 15 Stage 2 annealing temp (°C) 55 55 60 60 55 Stage 2 annealing time (sec) 45 45 45 40 45 60 45 # cycles 40 40 40 45 40 Fixed 2. 0 Control based threshold-10% max TF/5% max Xeno Fixed 0. 2 Analysis baseline setting Positive (Ct) 3 -15 <35 <36 auto <38 <37 <38 Suspect/Inconclusive(CT) 35 -40 36 -40 38 -39 >37 38 -40/Xeno 28. 5 -31. 5 Negative(CT) >40 >40 >40 undetected Undetected/Xeno 28. 5 -31. 5 Internal extraction control No No Yes <36 CT No Yes CT = 28. 5 -31. 5 Analysis threshold Control based threshold- 10% max TF/Xeno Proprietary & Confidential Proper & Confidential 9
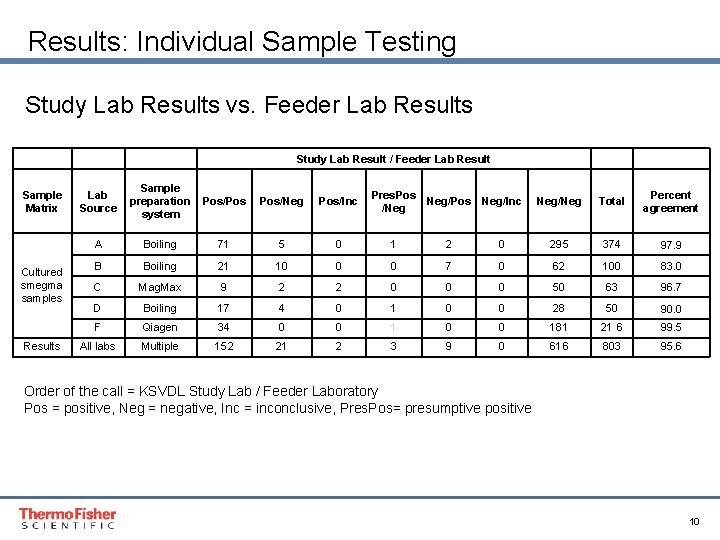
Results: Individual Sample Testing Study Lab Results vs. Feeder Lab Results Sample Matrix Lab Source Cultured smegma samples Results Study Lab Result / Feeder Lab Result Sample preparation Pos/Pos system Pos/Neg Pos/Inc Pres. Pos /Neg Neg/Pos Neg/Inc Neg/Neg Total Percent agreement A Boiling 71 5 0 1 2 0 295 374 97. 9 B Boiling 21 10 0 0 7 0 62 100 83. 0 C Mag. Max 9 2 2 0 0 0 50 63 96. 7 D Boiling 17 4 0 1 0 0 28 50 90. 0 F Qiagen 34 0 0 181 21 6 99. 5 All labs Multiple 152 21 2 3 9 0 616 803 95. 6 Order of the call = KSVDL Study Lab / Feeder Laboratory Pos = positive, Neg = negative, Inc = inconclusive, Pres. Pos= presumptive positive Proprietary & Confidential Proper & Confidential 10

Conclusions Individual Testing • 803 smegma samples were provided by feeder labs (FL) • All the samples were tested by study laboratory with Life Technologies workflow systems: • Mag. MAXTM • Vet. MAXTM T. foetus reagents • Agreement of 95. 6% was reached with 768/803 samples between feeder labs and study lab • Interestingly, Lab F reached almost 100% agreement using a different sample prep system and a modified Mc. Millen’s assay • Study laboratory (KSVDL) with LT protocol identified 24 more positives than the feeder laboratories. On retesting, one of the feeder labs missed 9 samples reported as positives. Proprietary & Confidential Proper & Confidential 11
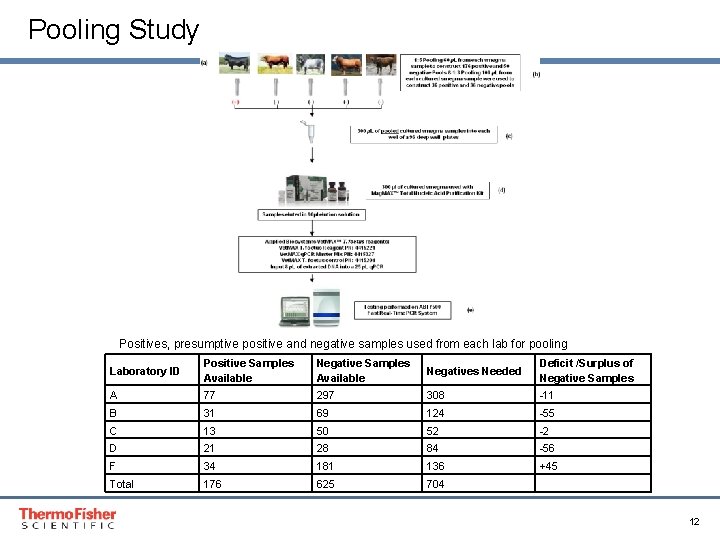
Pooling Study Positives, presumptive positive and negative samples used from each lab for pooling Laboratory ID Positive Samples Available Negatives Needed Deficit /Surplus of Negative Samples A 77 297 308 -11 B 31 69 124 -55 C 13 50 52 -2 D 21 28 84 -56 F 34 181 136 +45 Total 176 625 704 Proprietary & Confidential Proper & Confidential 12
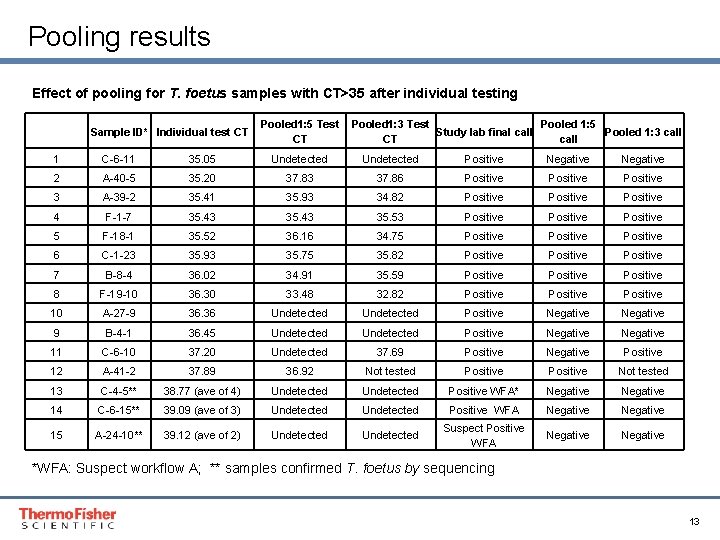
Pooling results Effect of pooling for T. foetus samples with CT>35 after individual testing Sample ID* Individual test CT Pooled 1: 5 Test Pooled 1: 3 Test Pooled 1: 5 Study lab final call Pooled 1: 3 call CT CT call 1 C-6 -11 35. 05 Undetected Positive Negative 2 A-40 -5 35. 20 37. 83 37. 86 Positive 3 A-39 -2 35. 41 35. 93 34. 82 Positive 4 F-1 -7 35. 43 35. 53 Positive 5 F-18 -1 35. 52 36. 16 34. 75 Positive 6 C-1 -23 35. 93 35. 75 35. 82 Positive 7 B-8 -4 36. 02 34. 91 35. 59 Positive 8 F-19 -10 36. 30 33. 48 32. 82 Positive 10 A-27 -9 36. 36 Undetected Positive Negative 9 B-4 -1 36. 45 Undetected Positive Negative 11 C-6 -10 37. 20 Undetected 37. 69 Positive Negative Positive 12 A-41 -2 37. 89 36. 92 Not tested Positive Not tested 13 C-4 -5** 38. 77 (ave of 4) Undetected Positive WFA* Negative 14 C-6 -15** 39. 09 (ave of 3) Undetected Positive WFA Negative 15 A-24 -10** 39. 12 (ave of 2) Undetected Suspect Positive WFA Negative *WFA: Suspect workflow A; ** samples confirmed T. foetus by sequencing Proprietary & Confidential Proper & Confidential 13

Pooling results 1: 5 Pools • 1: 5 pooling of positive samples with a CT of 35 and below were all detected • Only 3 of 9 positive samples with CTs between 36 -39. 9 were detected in 1: 5 pools • Pooling at 1: 5 missed 4% (7/176) of T. foetus positive samples 1: 3 Pools • Only 8 of 15 positive samples with CTs between 36 -39. 9 were detected in the 1: 3 pools • 1: 3 pooling missed 3. 5% (6/176) of T. foetus positive samples Proprietary & Confidential Proper & Confidential 14
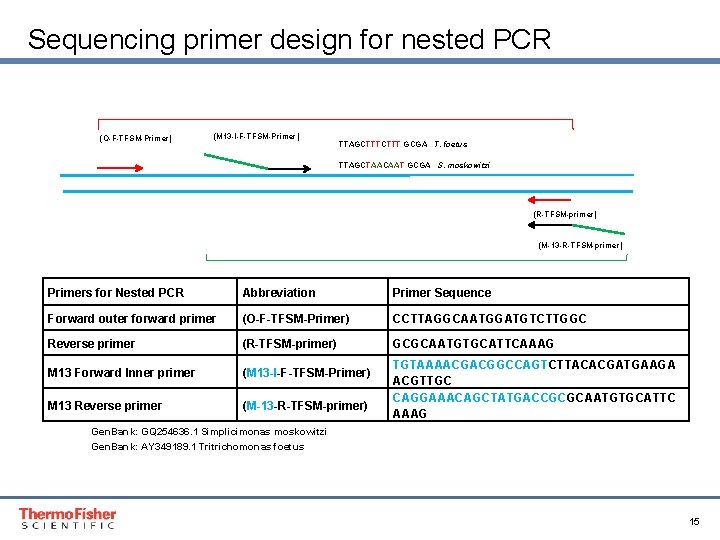
Sequencing primer design for nested PCR (O-F-TFSM-Primer) (M 13 -I-F-TFSM-Primer) TTAGCTTT GCGA T. foetus TTAGCTAACAAT GCGA S. moskowitzi (R-TFSM-primer) (M-13 -R-TFSM-primer) Primers for Nested PCR Abbreviation Primer Sequence Forward outer forward primer (O-F-TFSM-Primer) CCTTAGGCAATGGATGTCTTGGC Reverse primer (R-TFSM-primer) GCGCAATGTGCATTCAAAG M 13 Forward Inner primer (M 13 -I-F-TFSM-Primer) M 13 Reverse primer (M-13 -R-TFSM-primer) TGTAAAACGACGGCCAGTCTTACACGATGAAGA ACGTTGC CAGGAAACAGCTATGACCGCGCAATGTGCATTC AAAG Gen. Bank: GQ 254636. 1 Simplicimonas moskowitzi Gen. Bank: AY 349189. 1 Tritrichomonas foetus Proprietary & Confidential Proper & Confidential 15

Sequencing results for 175 T. foetus positives 175/176 T. foetus positive samples, including three late risers, were confirmed T. foetus by DNA sequencing Proprietary & Confidential Proper & Confidential 16
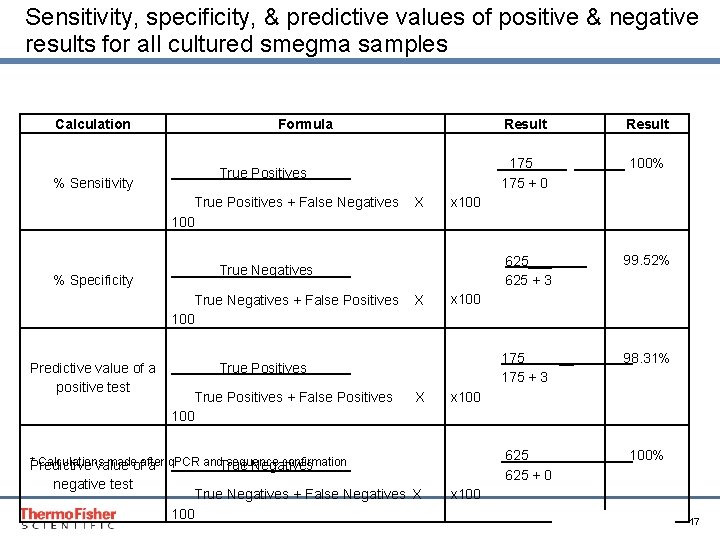
Sensitivity, specificity, & predictive values of positive & negative results for all cultured smegma samples Calculation % Sensitivity Formula True Positives + False Negatives X 100 % Specificity True Negatives + False Positives X 100 Predictive value of a positive test True Positives + False Positives X 100 * Calculations made after q. PCR and sequence confirmation Predictive value of a True Negatives negative test True Negatives + False Negatives X 100 Result 175 _ 100% 175 + 0 x 100 625___ 625 + 3 x 100 99. 52% 98. 31% 175 __ 175 + 3 x 100% 625 __ 625 + 0 x 100 Proprietary & Confidential Proper & Confidential 17

Sequencing Results • 175/176 positive samples by q. PCR were able to be sequenced • 1 sample (A-7 -25) with a CT 33. 95 was not able to be sequenced. • It is possible that there are point mutations in this positive sample in the sequencing primer regions, which were designed based on a few T. foetus and a single S. moskowitzi sequences from Gen. Bank • Most importantly, none of the samples reported S. moskowitzi DNA sequences Proprietary & Confidential Proper & Confidential 18

Overall Study Results • 95. 6 % agreement was reached between Study Lab (KSVDL) using Life technologies Mag. MAXTM and Vet. MAXTM T. foetus reagents and the feeder laboratories • 1: 5 Pooling it is likely to miss 4% of the positives • 1: 3 Pooling it is likely to miss 3. 5% of the positives • DNA sequencing • 175/176 positive samples were confirmed to be T. foetus, the 176 th sample could not be sequenced with the primers designed for this study Proprietary & Confidential Proper & Confidential 19

Acknowledgements Lalitha Peddireddi, KSVDL – performed the study at KSVDL Lee Effinger, ODA-Animal Health Laboratory Marilyn Simunich, Idaho State Dept. of Agriculture Cate O’Connell, Life Technologies Mangkey Bounpheng, Texas Veterinary Medical Diagnostic Laboratory Dawn Bueschel, NMDA Veterinary Diagnostic Services Muthu Chengappa, Kansas State Veterinary Diagnostic Laboratory Alfonso Clavijo, Texas Veterinary Medical Diagnostic Laboratory Kris A. Clothier, California Animal Health & Food Safety Lab System Hemant K. Naikare, Texas Veterinary Medical Diagnostics Laboratory Jeff Zinza, Life Technologies Mary Anne Williams, Life Technologies Proprietary & Confidential Proper & Confidential 20

Trich Laboratory Testing Standardization Objectives § To build confidence in Trich testing § To increase consistency in test results § To minimize sample quality influence variables § To minimize laboratory testing variables § To reduce the burden and cost to man and beast Proper & Confidential Proprietary & Confidential 21

Trich Laboratory Testing Standardization Trichomonas foetus DNA testing Proper & Confidential Proprietary & Confidential 22
- Slides: 22