Trasporto dellm RNA Transport through the nuclear pores









- Slides: 46









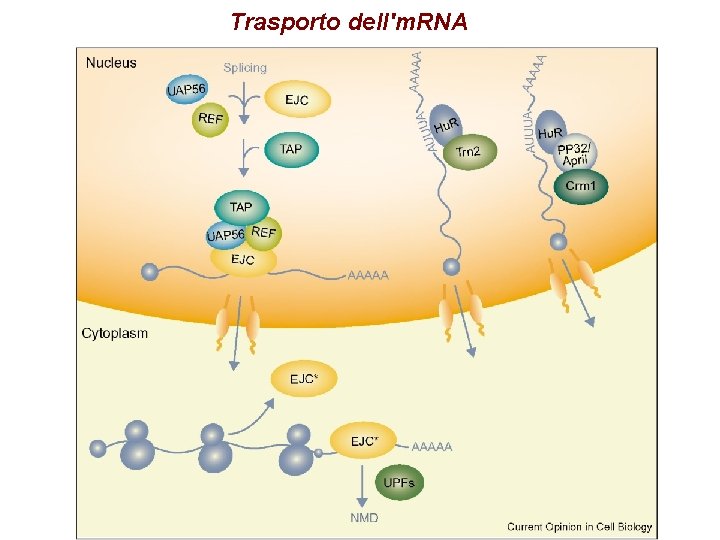
Trasporto dell'm. RNA





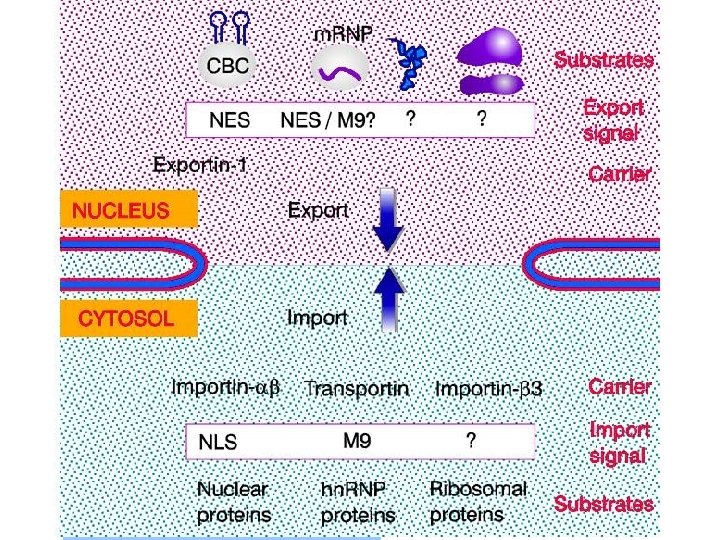
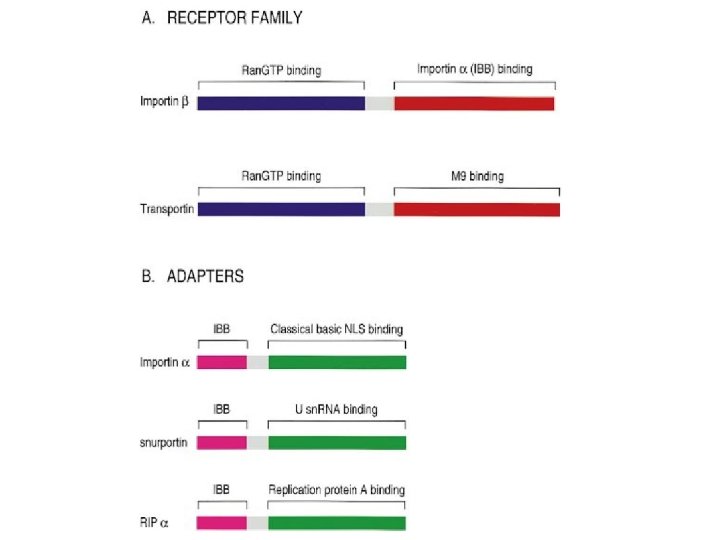
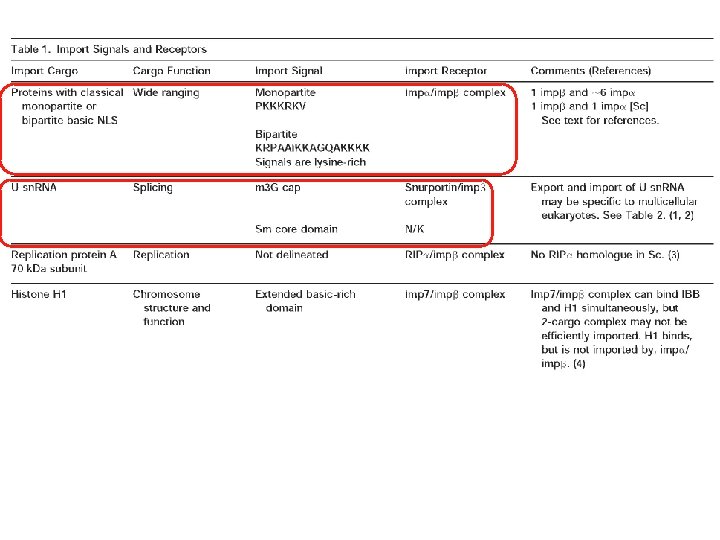
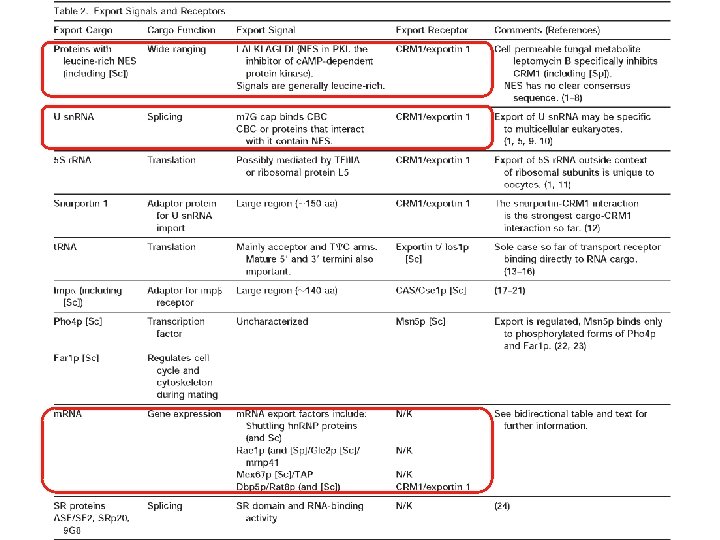
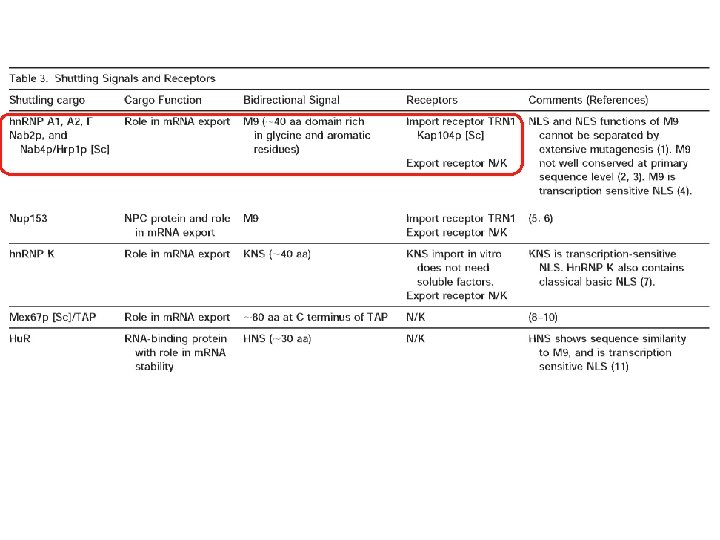
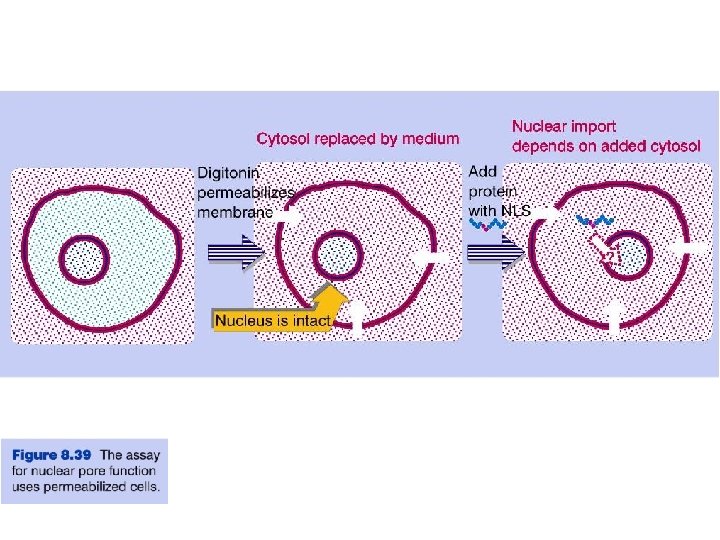
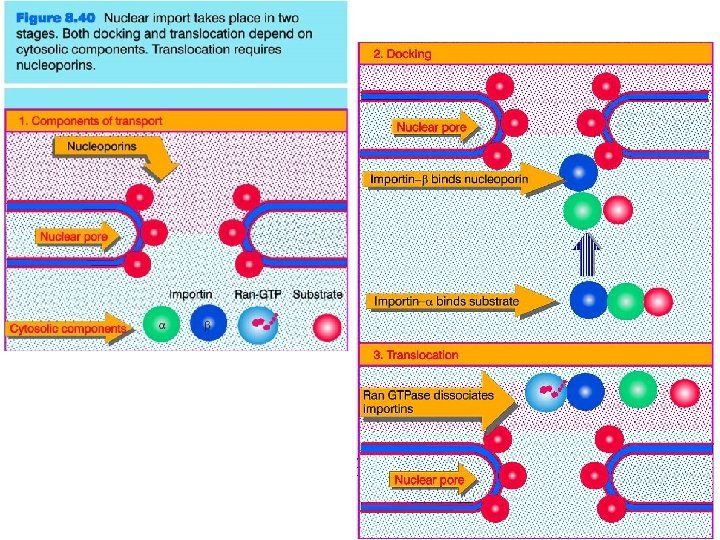
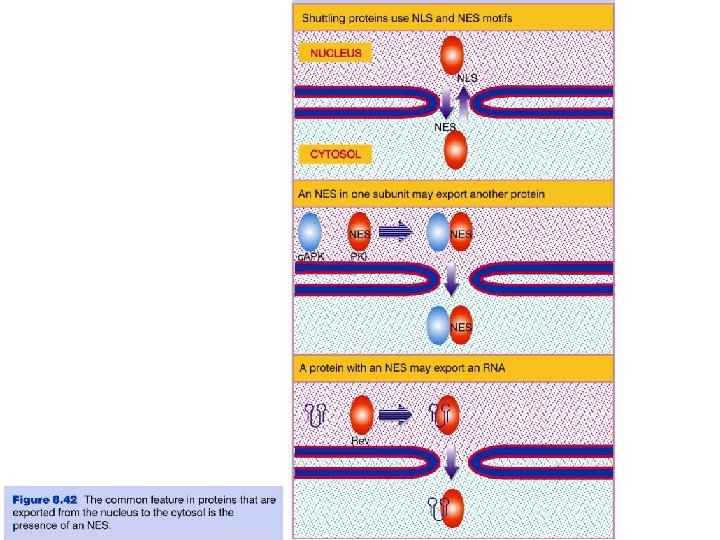
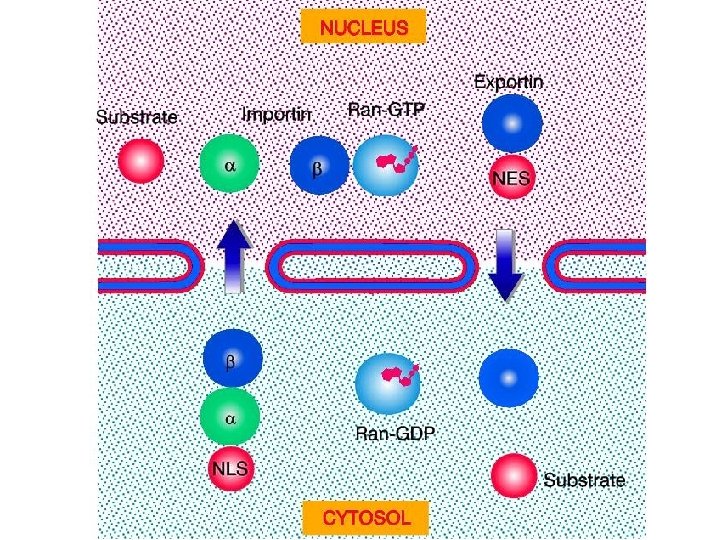
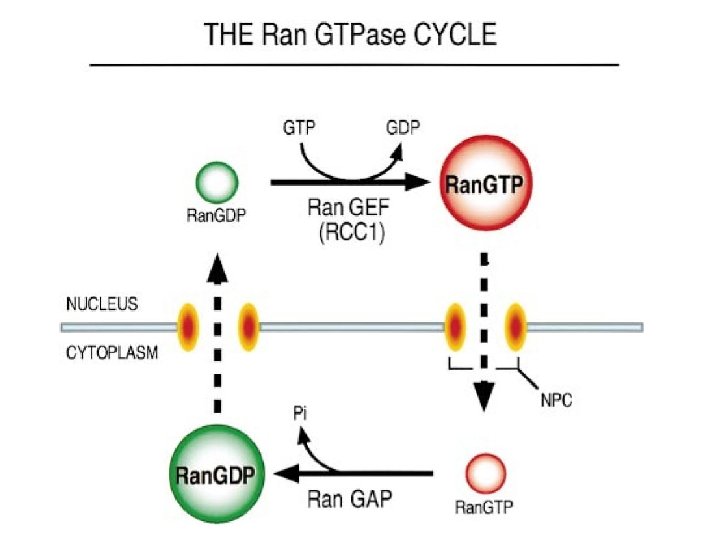
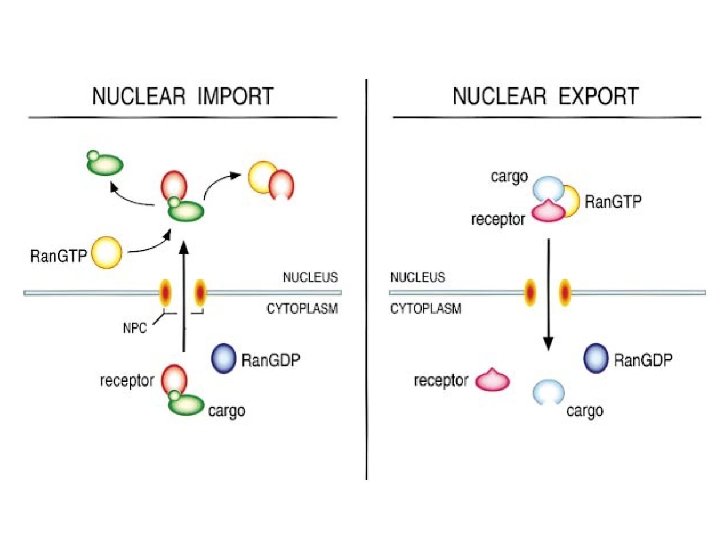
Transport through the nuclear pores • The NLS and NES consist of short sequences that are necessary and sufficient for proteins to be transported through the nuclear pores. • Transport receptors have the dual properties of recognizing NLS or NES sequences and binding to the nuclear pore. • The direction of transport is controlled by the state of the monomeric G protein, Ran. • The nucleus contains Ran-GTP, which stabilizes export complexes, while the cytosol contains Ran-GDP, which stabilizes import complexes. • The mechanism of movement does not involve a motor.

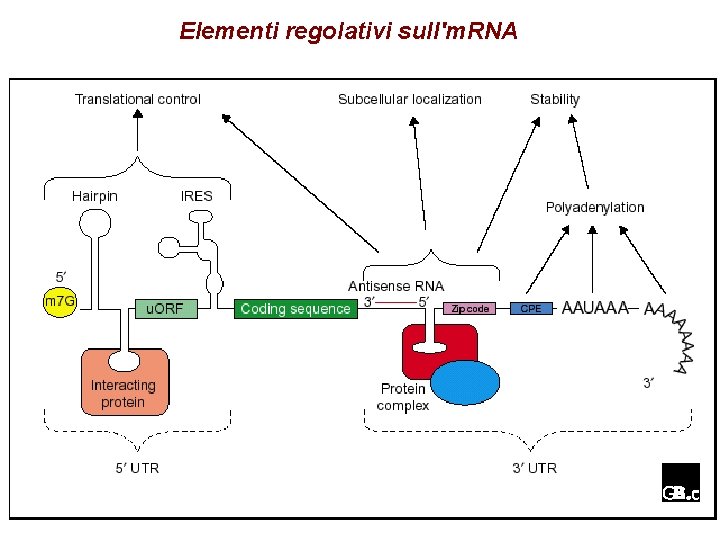
Elementi regolativi sull'm. RNA


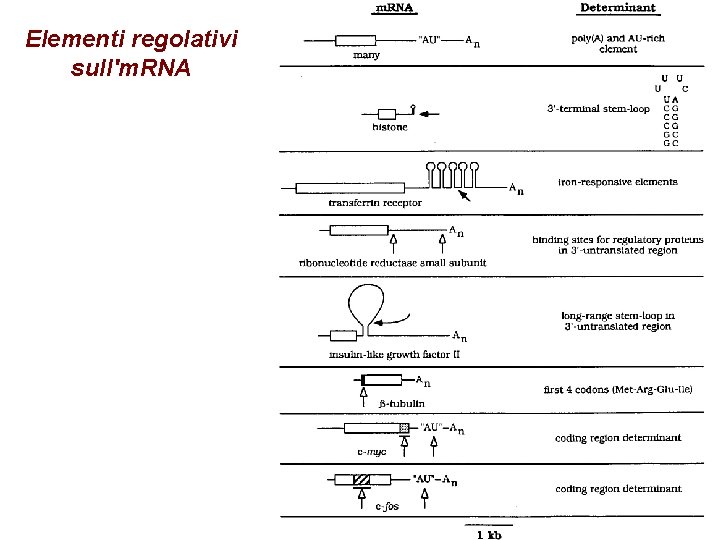
Elementi regolativi sull'm. RNA

Fattori che influenzano la stabilità dell'm. RNA

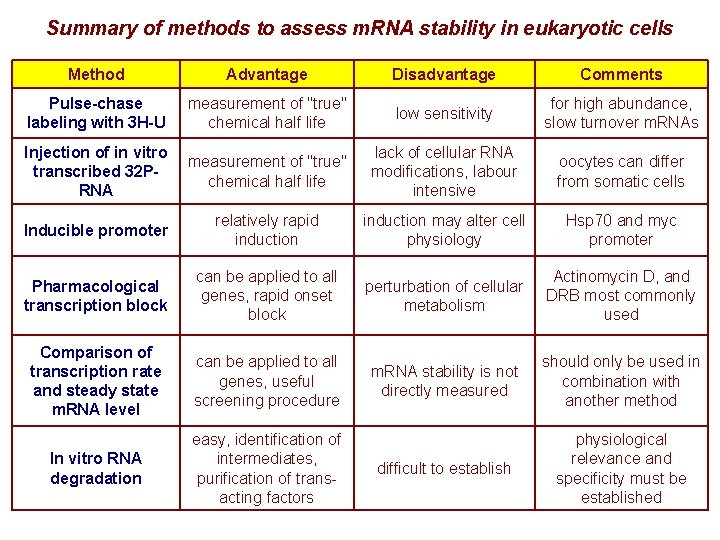
Summary of methods to assess m. RNA stability in eukaryotic cells Method Advantage Disadvantage Comments Pulse-chase labeling with 3 H-U measurement of "true" chemical half life low sensitivity for high abundance, slow turnover m. RNAs Injection of in vitro transcribed 32 PRNA measurement of "true" chemical half life lack of cellular RNA modifications, labour intensive oocytes can differ from somatic cells Inducible promoter relatively rapid induction may alter cell physiology Hsp 70 and myc promoter Pharmacological transcription block can be applied to all genes, rapid onset block perturbation of cellular metabolism Actinomycin D, and DRB most commonly used Comparison of transcription rate and steady state m. RNA level can be applied to all genes, useful screening procedure m. RNA stability is not directly measured should only be used in combination with another method In vitro RNA degradation easy, identification of intermediates, purification of transacting factors difficult to establish physiological relevance and specificity must be established

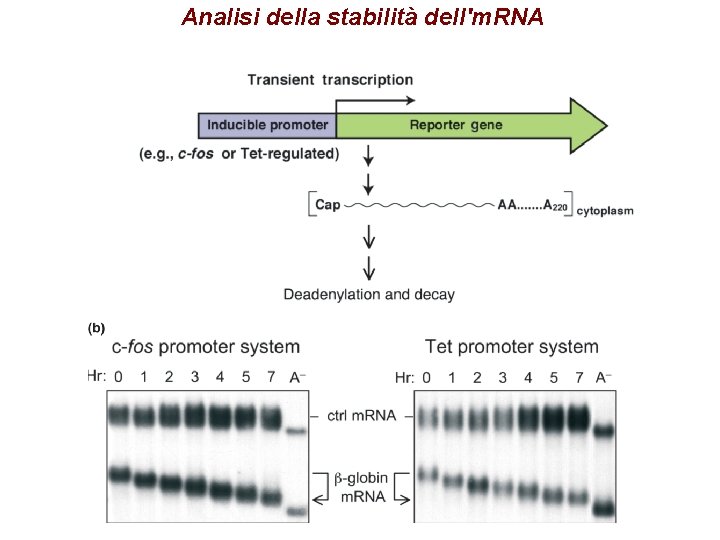
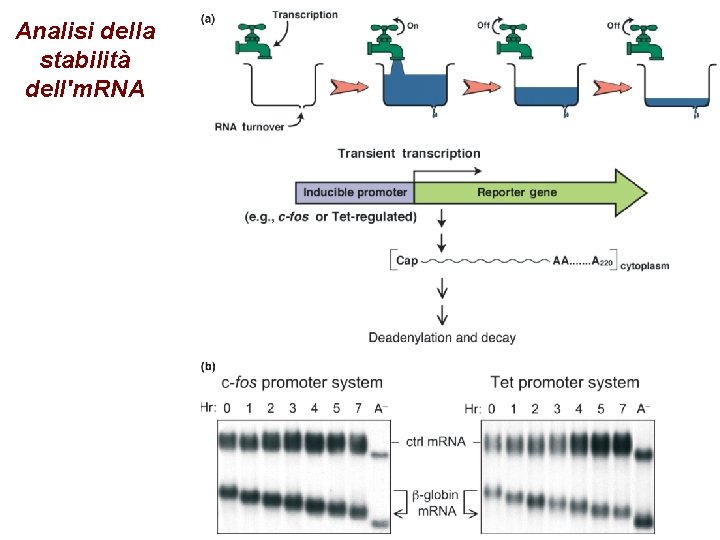
Analisi della stabilità dell'm. RNA

Degradazione dipendente da deadenilazione

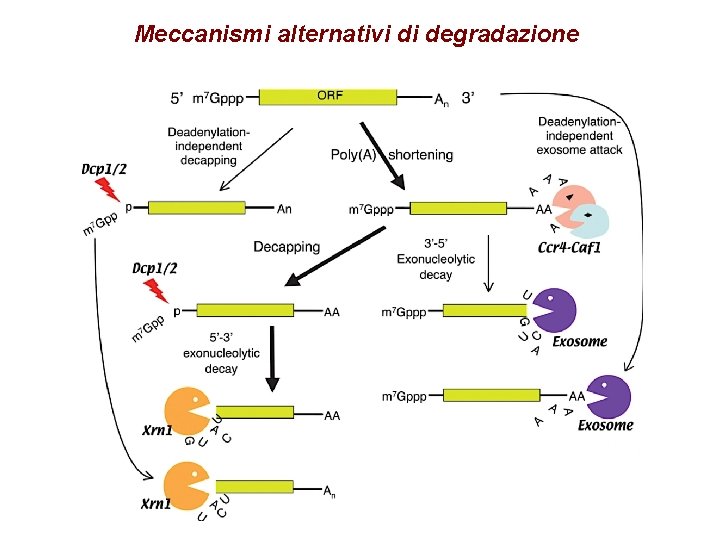
Meccanismi alternativi di degradazione

m. RNA degradative activities in mammalian cells Deadenylation • PARN • CCR 4/CAF 1/NOT complex • PAN 2/PAN 3 complex • Decapping • DCP 2 which binds RNA as a prerequisite for cap recognition. • DCP 1 augments DCP 2 activity • LSM (SM-LIKE) PROTEINS augment DCP 2 activity 5’ -to-3’ exonuclease activity • XRN 1 is a proven 5’ -to-3’ exonuclease that localizes to the cytoplasm. • RAT 1/XRN 2 is only thought to be a 5’ -to-3’ exonuclease on the basis of its similarity to the yeast orthologue. 3’ -to-5’ exonuclease activity • Exosome (six RNase-PH-DOMAIN components, PM/SCL 75, MTR 3, RRP 41, RRP 42, RRP 43 and RRP 46; three S 1 and KH RNA-binding components, RRP 40 and CSL 4; the RNASE D-like components PM/SCL 100; the putative helicase. KIAA 0053; and a protein that is phosphorylated in the M phase of the cell) PMR 1 -like activity • Polysomal ribonuclease 1 (PMR 1) is a polysome-associated m. RNA endonuclease

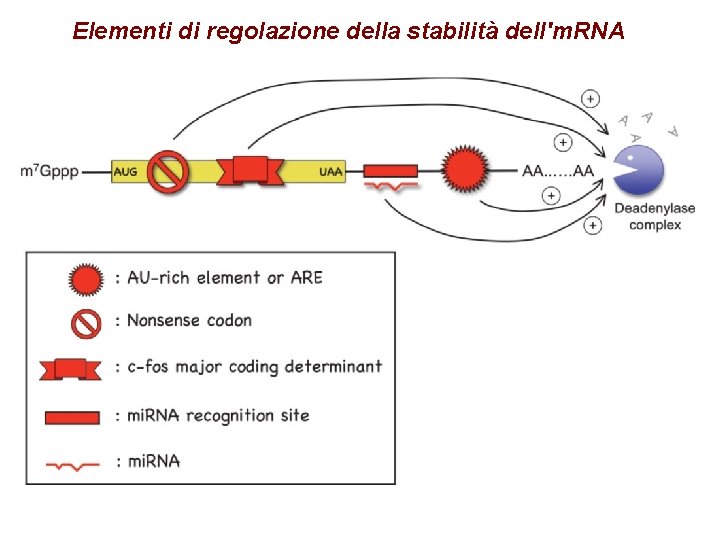
Elementi di regolazione della stabilità dell'm. RNA

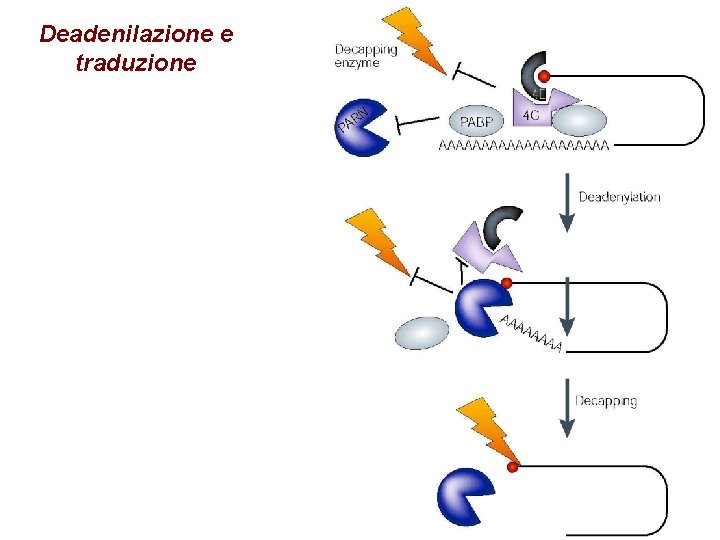
Deadenilazione e traduzione

Elementi ARE (AU rich element)

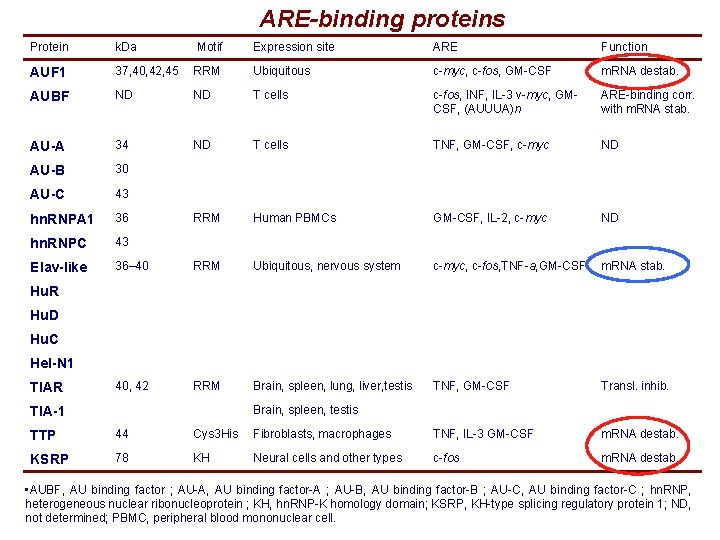
ARE-binding proteins Protein k. Da Motif Expression site ARE Function AUF 1 37, 40, 42, 45 RRM Ubiquitous c-myc, c-fos, GM-CSF m. RNA destab. AUBF ND ND T cells c-fos, INF, IL-3 v-myc, GMCSF, (AUUUA)n ARE-binding corr. with m. RNA stab. AU-A 34 ND T cells TNF, GM-CSF, c-myc ND AU-B 30 AU-C 43 hn. RNPA 1 36 RRM Human PBMCs GM-CSF, IL-2, c-myc ND hn. RNPC 43 Elav-like 36– 40 RRM Ubiquitous, nervous system c-myc, c-fos, TNF-a, GM-CSF m. RNA stab. 40, 42 RRM Brain, spleen, lung, liver, testis TNF, GM-CSF Transl. inhib. Hu. R Hu. D Hu. C Hel-N 1 TIAR Brain, spleen, testis TIA-1 TTP 44 Cys 3 His Fibroblasts, macrophages TNF, IL-3 GM-CSF m. RNA destab. KSRP 78 KH Neural cells and other types c-fos m. RNA destab. • AUBF, AU binding factor ; AU-A, AU binding factor-A ; AU-B, AU binding factor-B ; AU-C, AU binding factor-C ; hn. RNP, heterogeneous nuclear ribonucleoprotein ; KH, hn. RNP-K homology domain; KSRP, KH-type splicing regulatory protein 1; ND, not determined; PBMC, peripheral blood mononuclear cell.

ARE-binding proteins

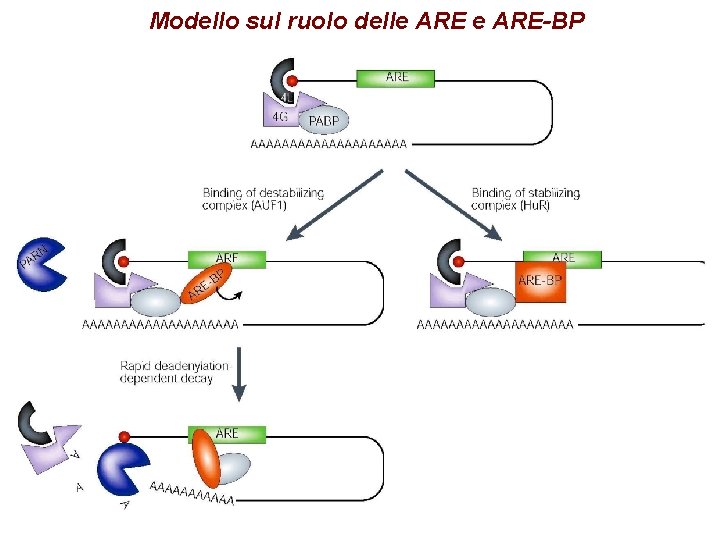
Modello sul ruolo delle ARE-BP

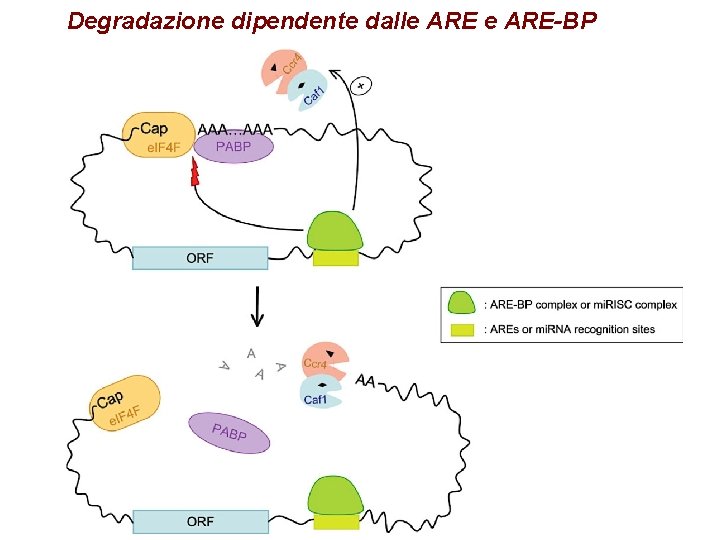
Degradazione dipendente dalle ARE-BP

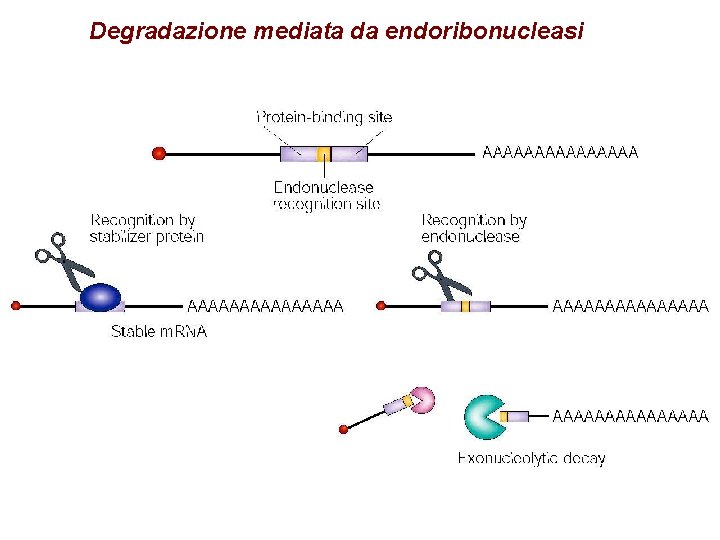
Degradazione mediata da endoribonucleasi

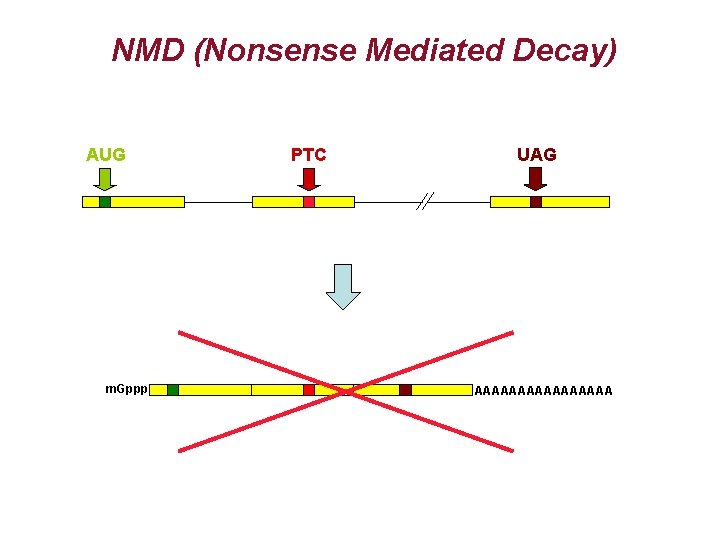
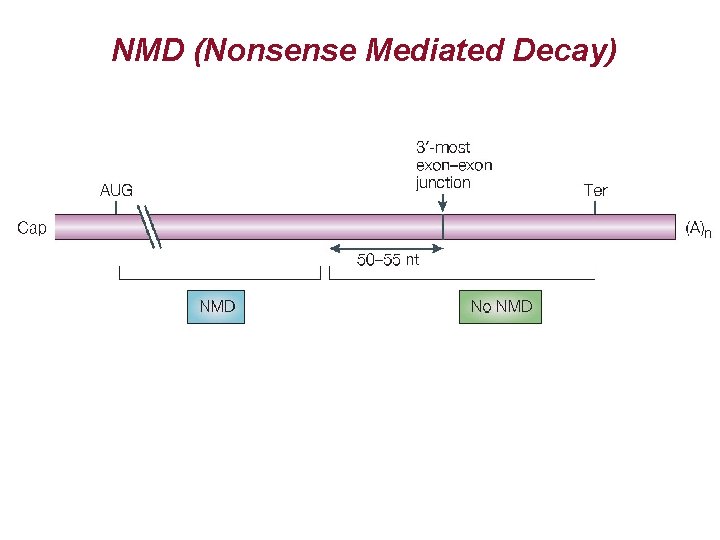
NMD (Nonsense Mediated Decay) AUG m. Gppp PTC UAG AAAAAAAA

NMD (Nonsense Mediated Decay)

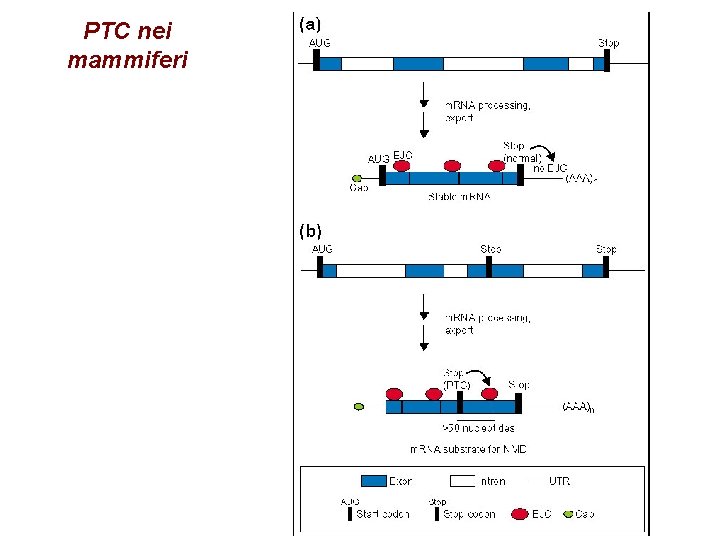
PTC nei mammiferi

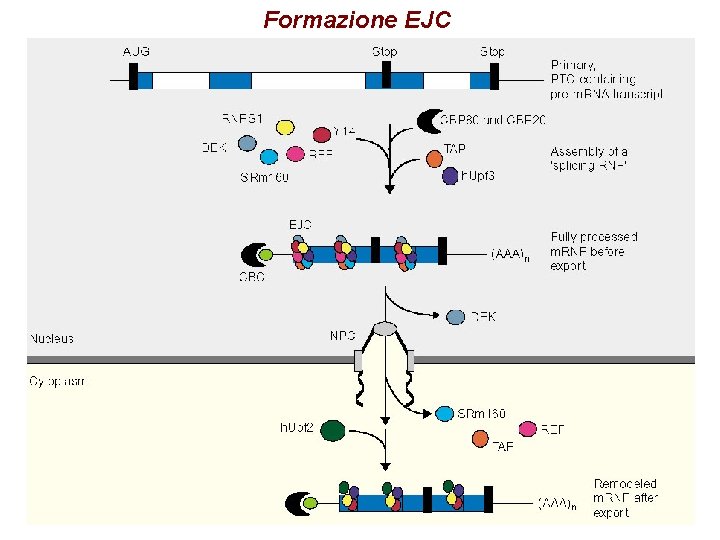
Formazione EJC

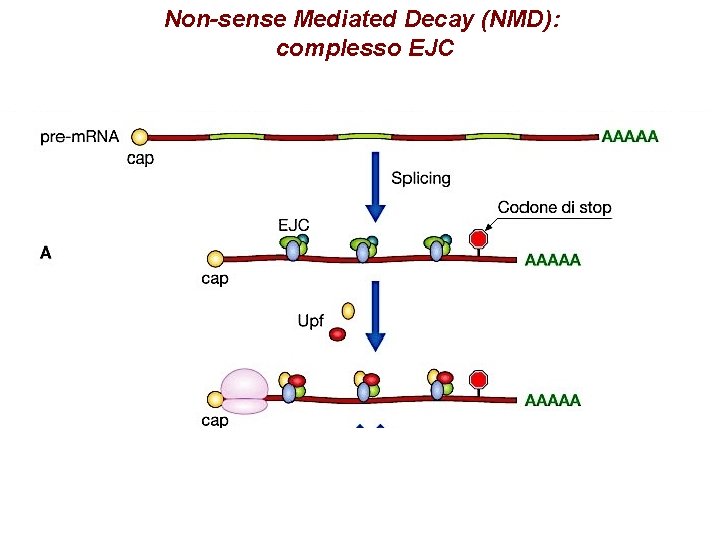
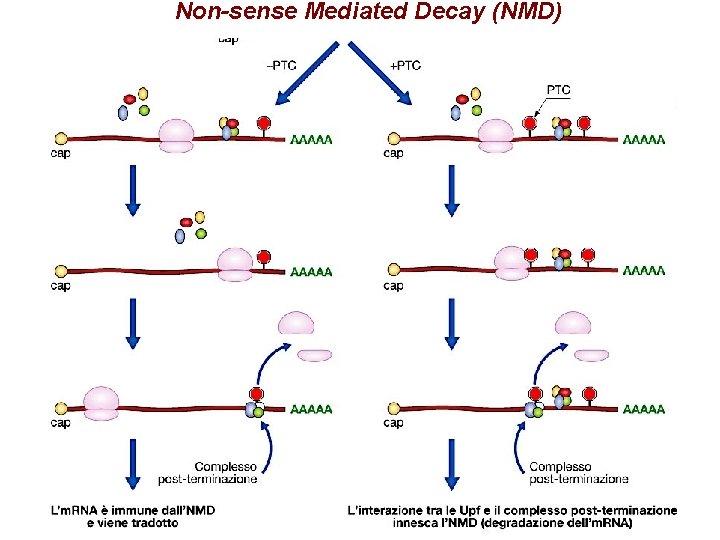
Non-sense Mediated Decay (NMD): complesso EJC

Non-sense Mediated Decay (NMD)


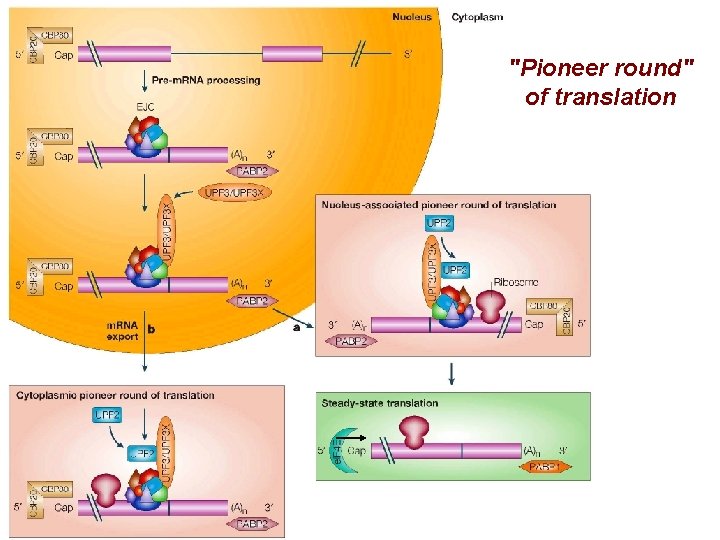
"Pioneer round" of translation


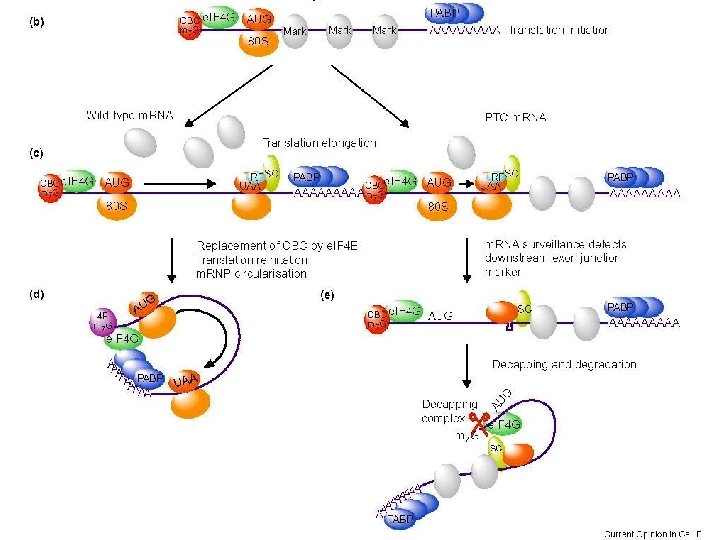
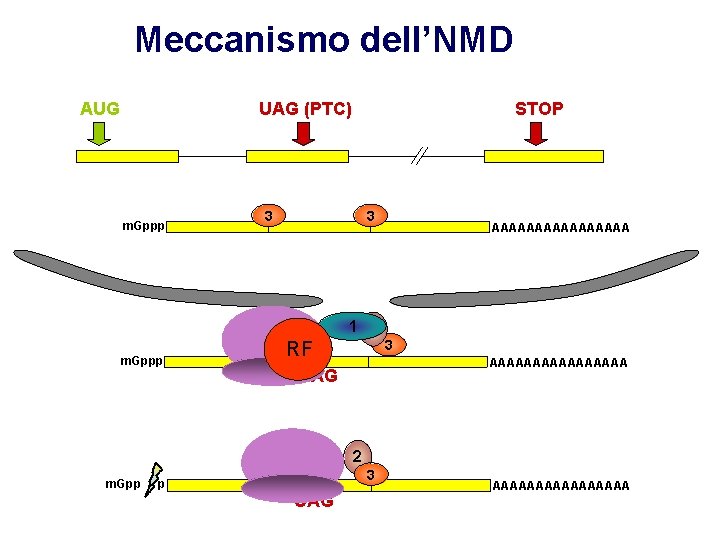
Meccanismo dell’NMD AUG UAG (PTC) m. Gppp 3 22 m. Gppp STOP 3 3 3 AAAAAAAA 1 2 RF 3 AAAAAAAA UAG 2 m. Gpp p 3 UAG AAAAAAAA

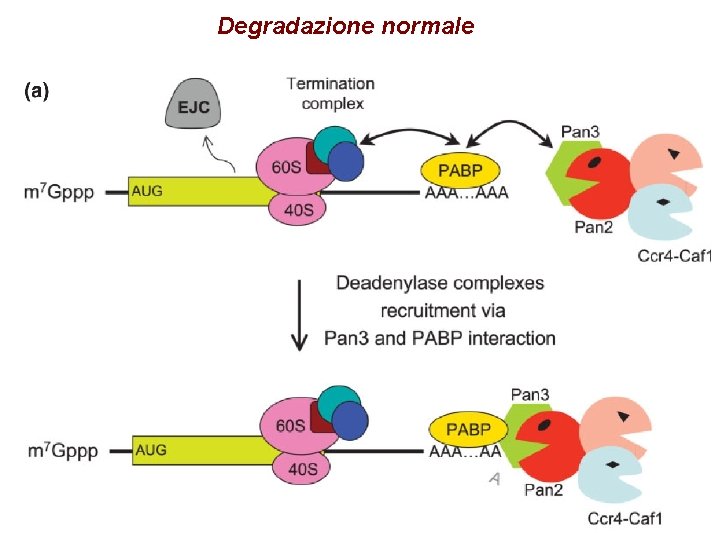
Degradazione normale

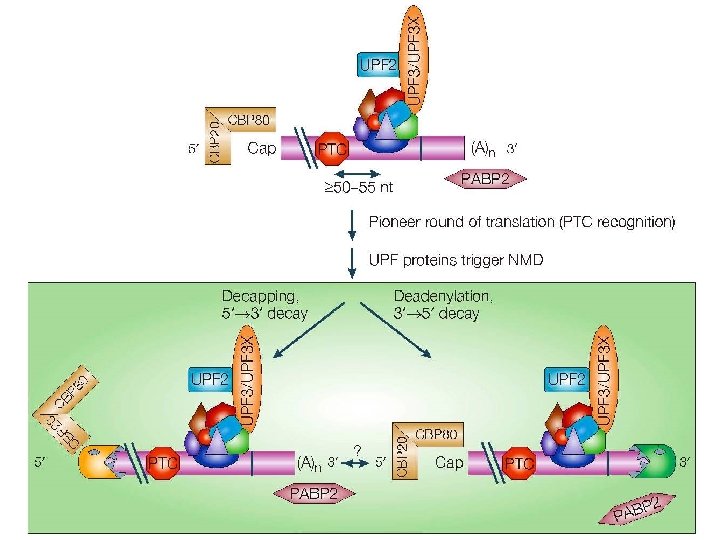
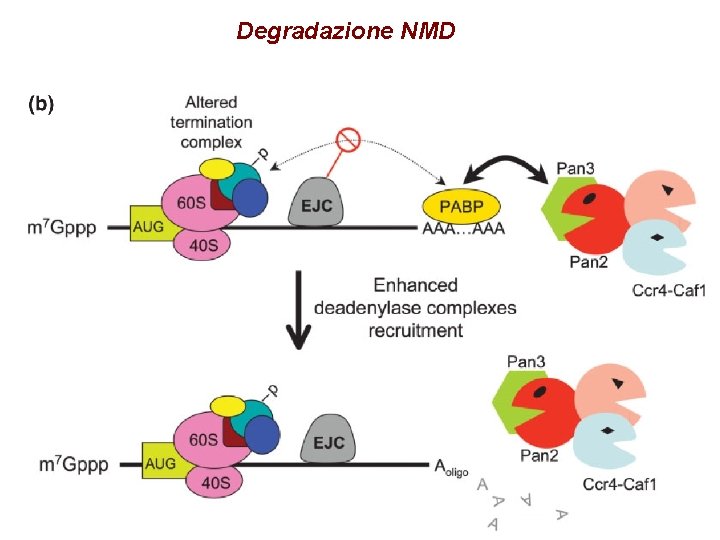
Degradazione NMD

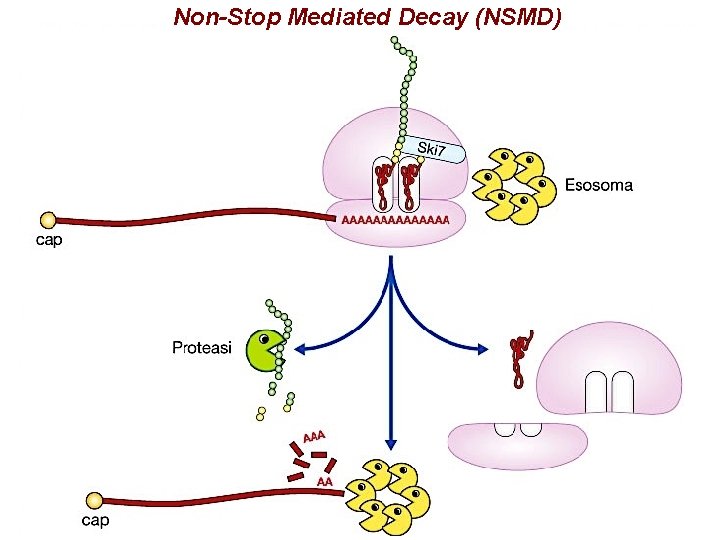
Non-Stop Mediated Decay (NSMD)


Analisi della stabilità dell'm. RNA