Transposons Mechanisms of Transposition Donnie Pickel Elizabeth Jensen
Transposons & Mechanisms of Transposition Donnie Pickel, Elizabeth Jensen and Dasmeet Kaur

Mobile DNA element or transposable element v Interspaced repeats have the capability to “move around” in the genome v “jumping genes” v “selfish DNA”-- exist only to maintain themselves ? v “junk DNA”-- ? ? ? v Transposition -- The process by which these sequences are copied and inserted into a new site in the genome v Significant role in evolution -- Positive and negative impacts on genome *gene inactivation *modulate gene expression *induce illegitimate recombination v Selective transposition in germ cells are passed down to successive generations for their accumulation in the genome. v ~25% of human promoter regions and ~ 4% of human exons derived

§ 1940 s, Barbara Mc. Clintock discovered the first transposable element in maize § Eventually earned vindication when she became the only woman to independently win the Nobel Prize in medicine (1983). § Late 1960 s, transposition was also found in Bacteria. http: //www. bestvalueschools. com/stem-careers-for-women/

DNA Transposons and Retrotransposons �Transposons excise themselves and reintegrate elsewhere �Retrotransposons follow replicative mechanism and reintegrate themselves
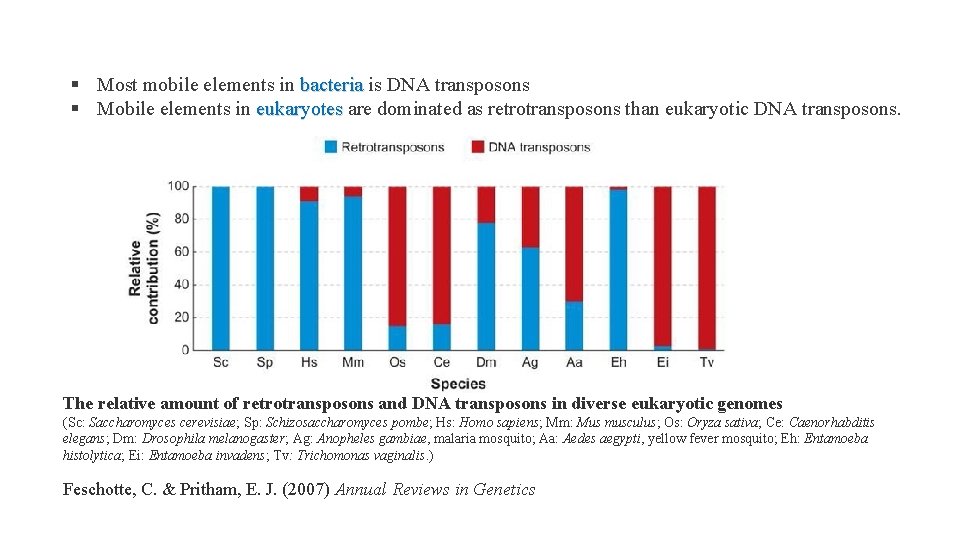
§ Most mobile elements in bacteria is DNA transposons bacteria § Mobile elements in eukaryotes are dominated as retrotransposons than eukaryotic DNA transposons. eukaryotes The relative amount of retrotransposons and DNA transposons in diverse eukaryotic genomes (Sc: Saccharomyces cerevisiae; Sp: Schizosaccharomyces pombe; Hs: Homo sapiens; Mm: Mus musculus; Os: Oryza sativa; Ce: Caenorhabditis elegans; Dm: Drosophila melanogaster; Ag: Anopheles gambiae, malaria mosquito; Aa: Aedes aegypti, yellow fever mosquito; Eh: Entamoeba histolytica; Ei: Entamoeba invadens; Tv: Trichomonas vaginalis. ) Feschotte, C. & Pritham, E. J. (2007) Annual Reviews in Genetics
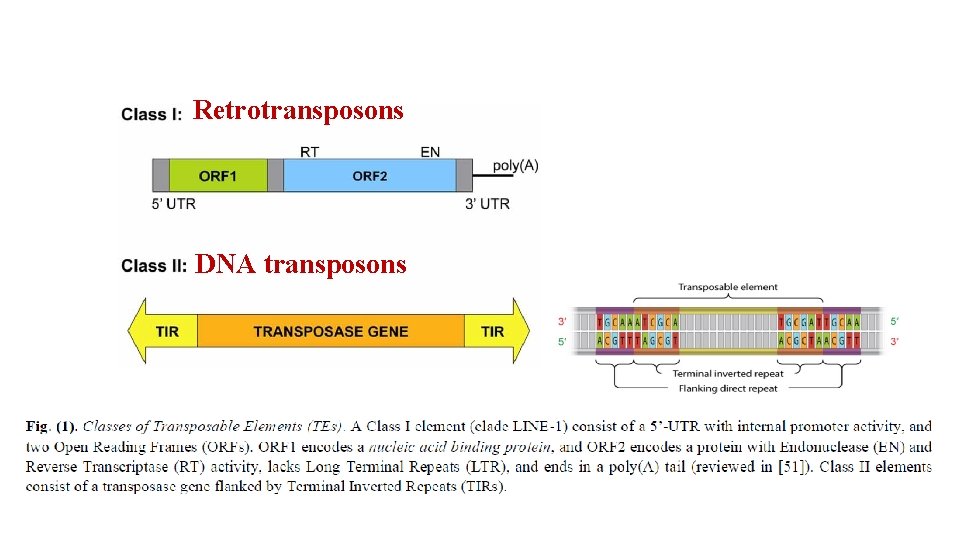
Retrotransposons DNA transposons
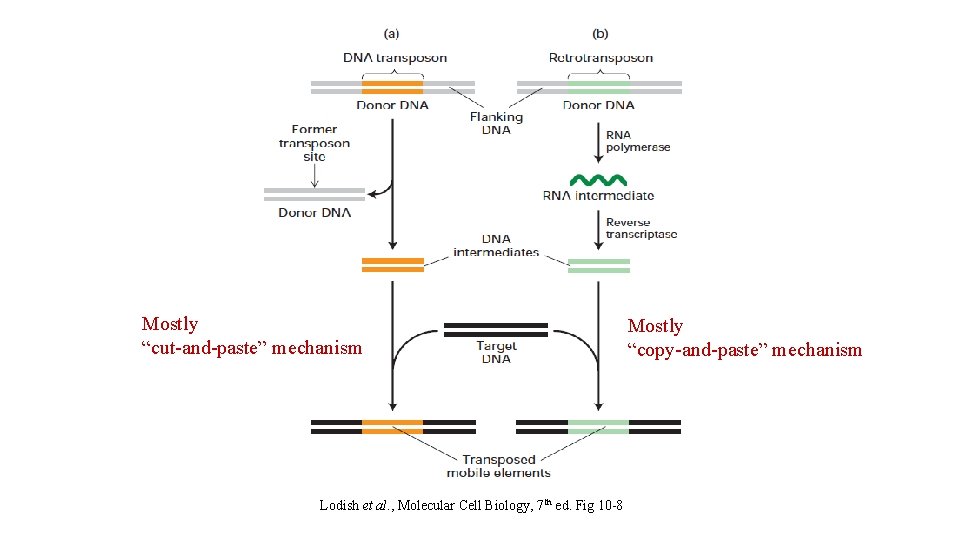
Mostly “cut-and-paste” mechanism Lodish et al. , Molecular Cell Biology, 7 th ed. Fig 10 -8 Mostly “copy-and-paste” mechanism
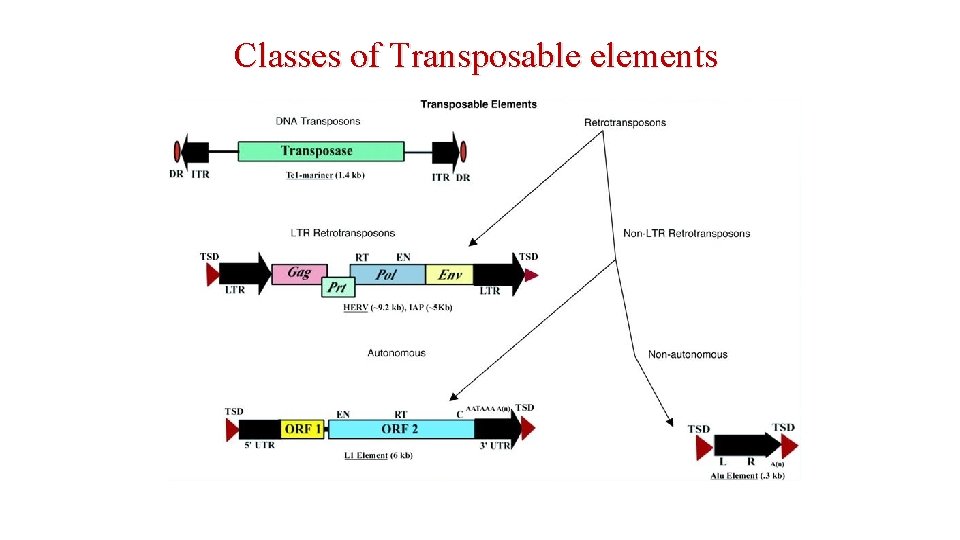
Classes of Transposable elements
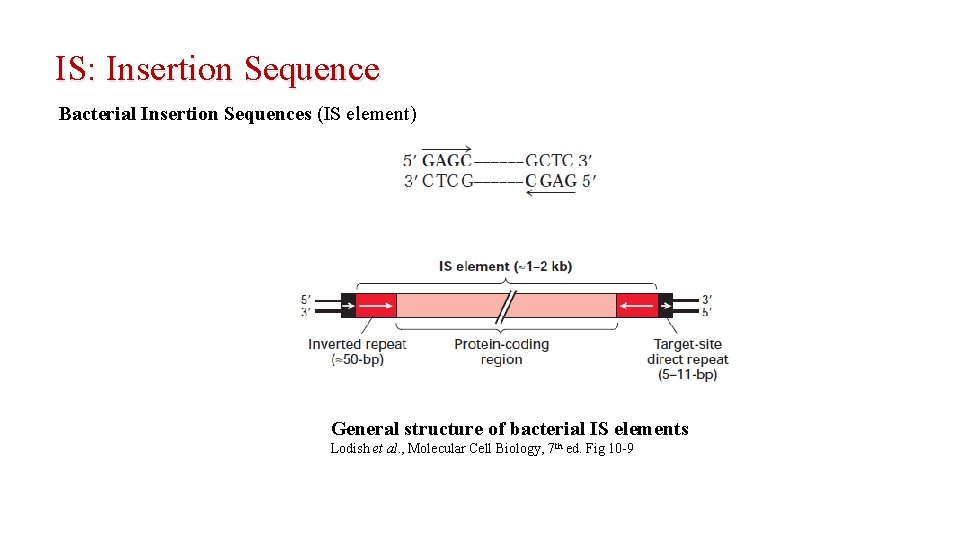
IS: Insertion Sequence Bacterial Insertion Sequences (IS element) General structure of bacterial IS elements Lodish et al. , Molecular Cell Biology, 7 th ed. Fig 10 -9
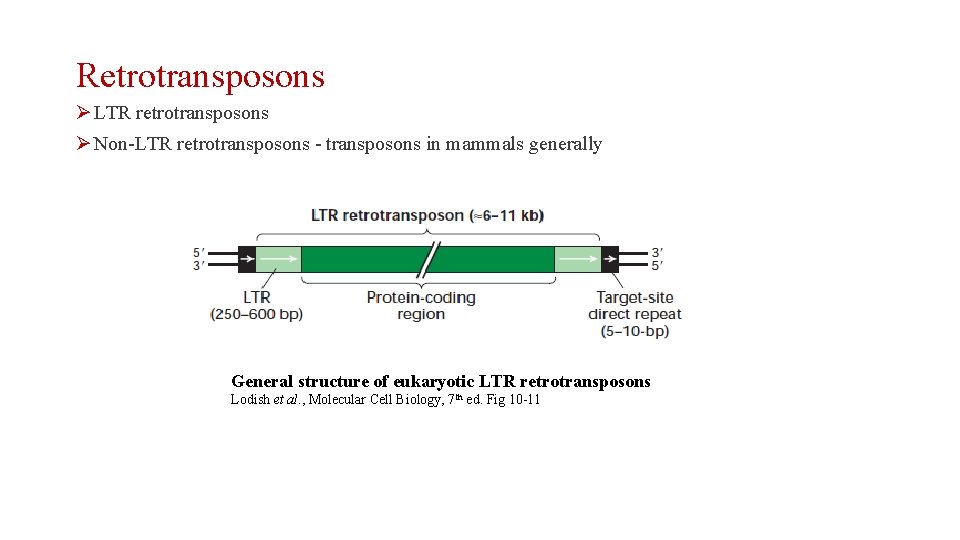
Retrotransposons Ø LTR retrotransposons Ø Non-LTR retrotransposons - transposons in mammals generally General structure of eukaryotic LTR retrotransposons Lodish et al. , Molecular Cell Biology, 7 th ed. Fig 10 -11

Autonomous & Nonautonomous • Autonomous (activator elements) are similar in structure and function to bacterial IS elements example-Ac elements • Nonautonomous (dissociation elements) lack transposase gene and cannot move by themselves -Must have a cis transposable element with transposase gene to move example-Miniature Inverted-repeat Transposable Elements (MITEs) -Ds elements

DNA transposons families -depending on their sequence, TIRs, and/or TSDs: Subclass I: Tc 1/mariner PIF/Harbinger h. AT piggy. Bac Subclass II: Helitron Maverick
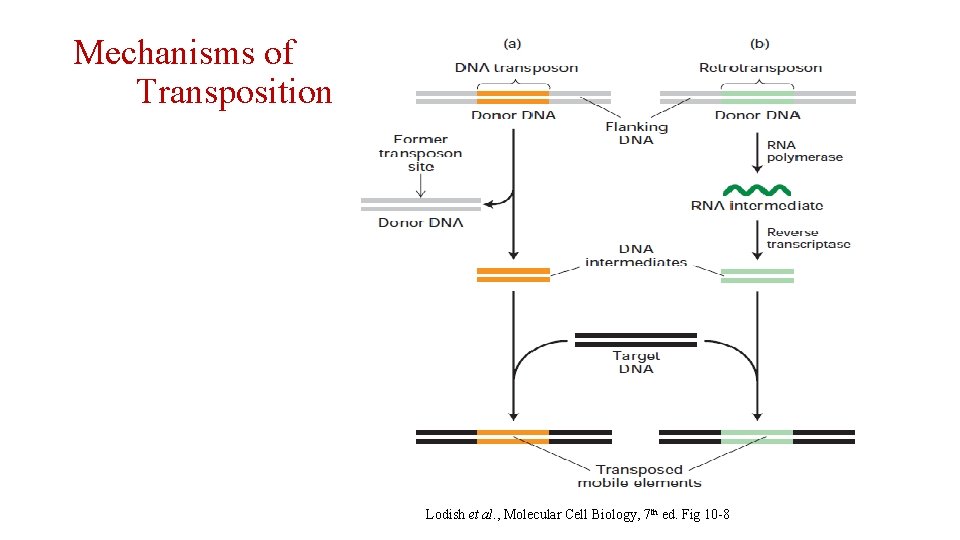
Mechanisms of Transposition Lodish et al. , Molecular Cell Biology, 7 th ed. Fig 10 -8
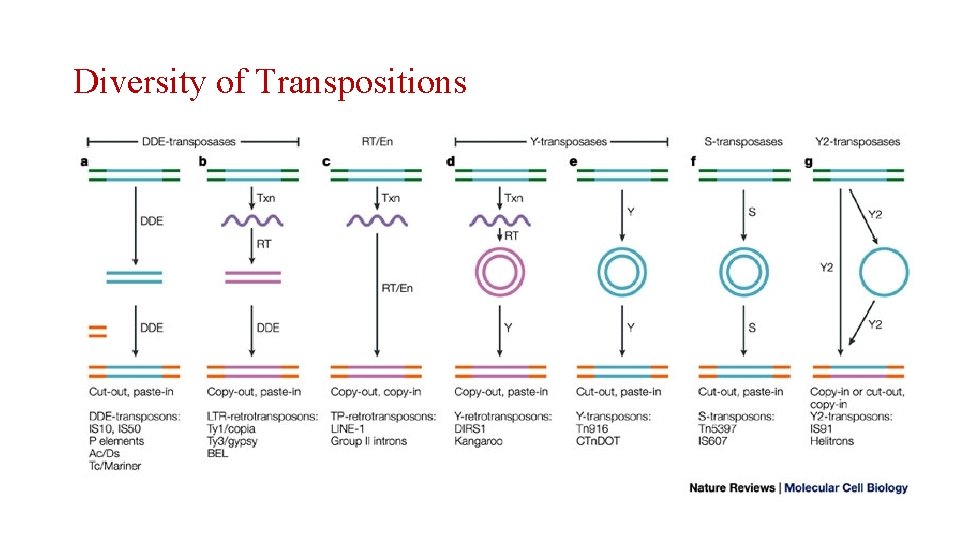
Diversity of Transpositions
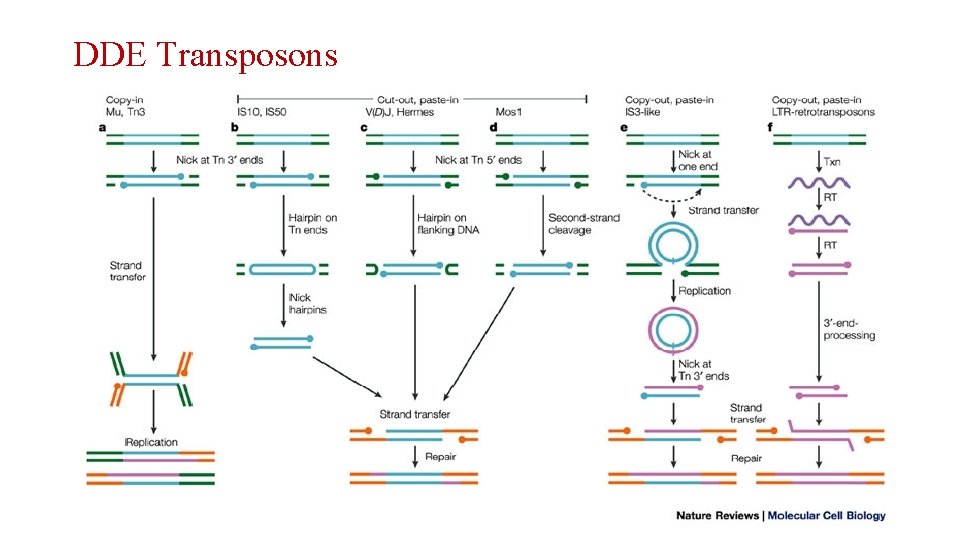
DDE Transposons
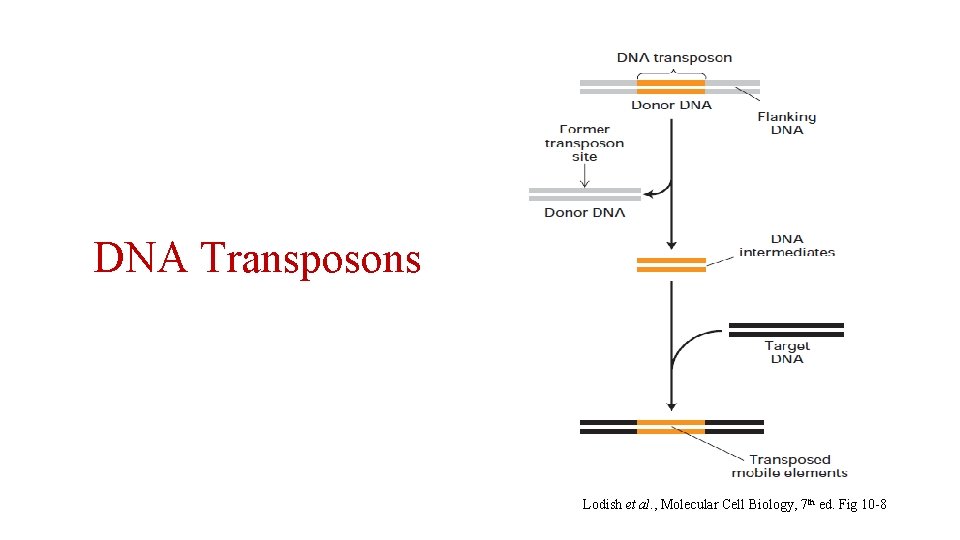
DNA Transposons Lodish et al. , Molecular Cell Biology, 7 th ed. Fig 10 -8
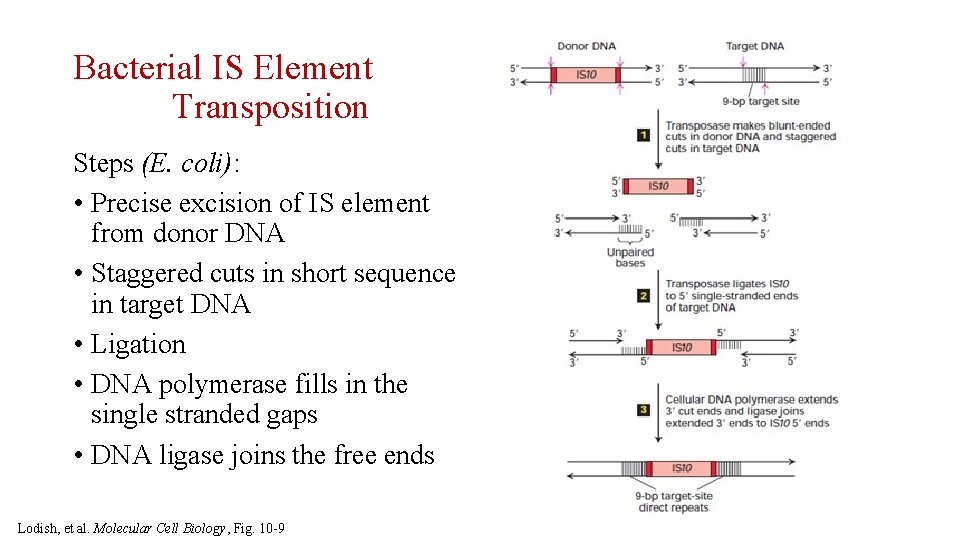
Bacterial IS Element Transposition Steps (E. coli): • Precise excision of IS element from donor DNA • Staggered cuts in short sequence in target DNA • Ligation • DNA polymerase fills in the single stranded gaps • DNA ligase joins the free ends Lodish, et al. Molecular Cell Biology, Fig. 10 -9
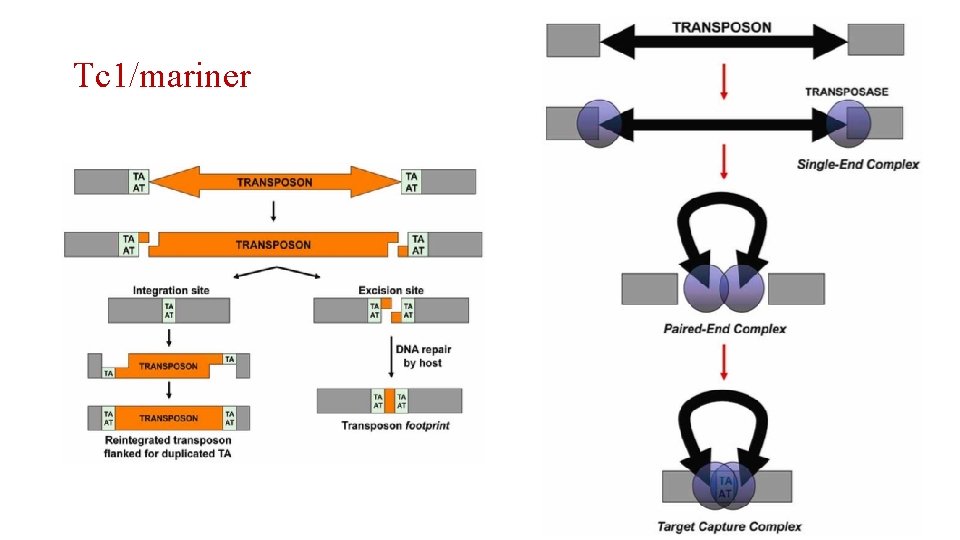
Tc 1/mariner
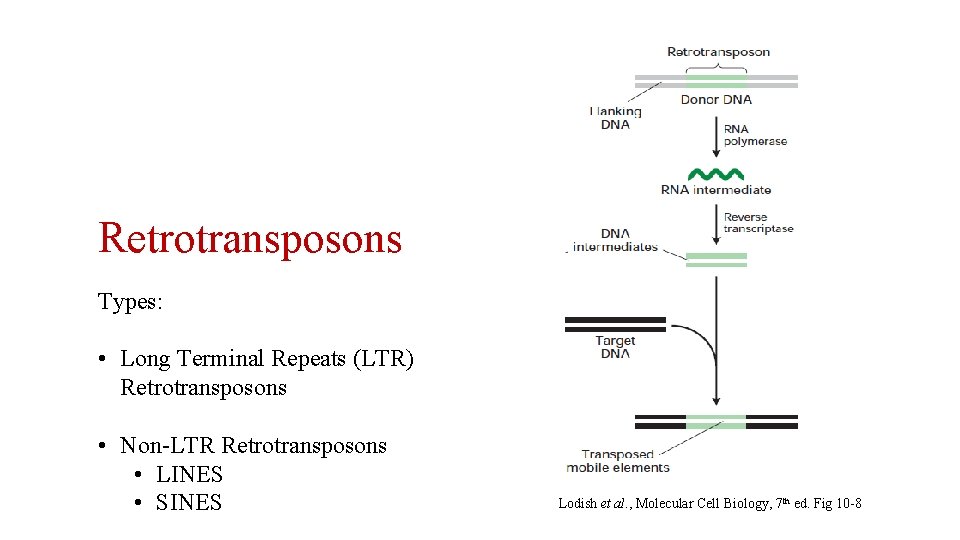
Retrotransposons Types: • Long Terminal Repeats (LTR) Retrotransposons • Non-LTR Retrotransposons • LINES • SINES Lodish et al. , Molecular Cell Biology, 7 th ed. Fig 10 -8
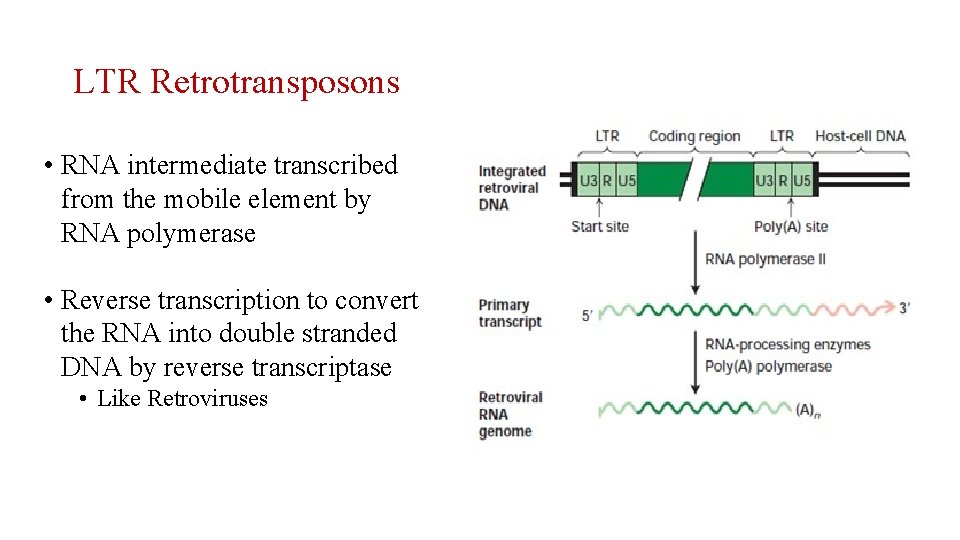
LTR Retrotransposons • RNA intermediate transcribed from the mobile element by RNA polymerase • Reverse transcription to convert the RNA into double stranded DNA by reverse transcriptase • Like Retroviruses
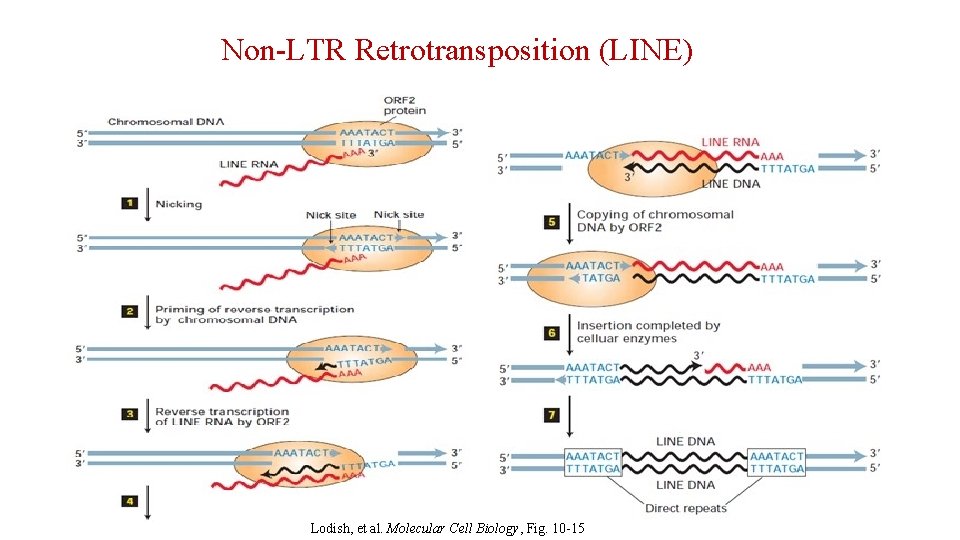
Non-LTR Retrotransposition (LINE) Lodish, et al. Molecular Cell Biology, Fig. 10 -15

Transposon-Seq (Tn-Seq) • Essential Gene Identification • Genetic Interaction Discovery • Genetic Factors in Biological Processes • Regulatory Factor Discovery
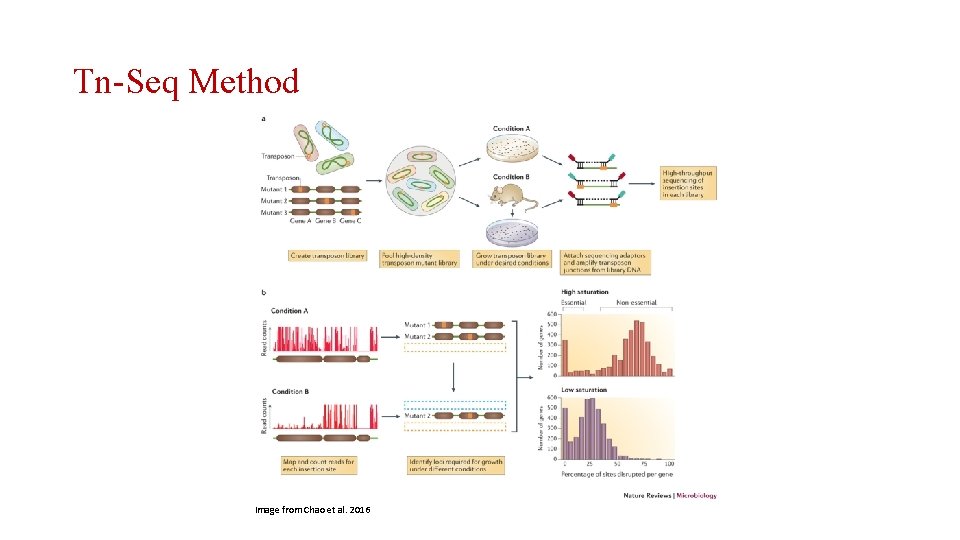
Tn-Seq Method Image from Chao et al. 2016

Transposons in Gene Therapy • Lower Production Costs • Increased Biosafety • Low Immunogenecity
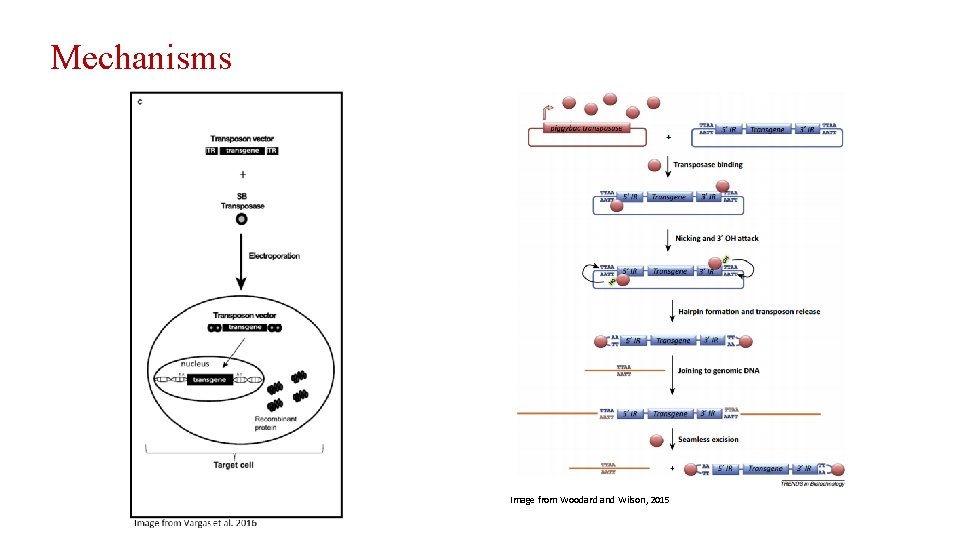
Mechanisms Image from Woodard and Wilson, 2015

Piggybac Sleeping Beauty • Tc 1/Mariner-like Elements • Relatively Small Carrier (6 -18 kb) • Limiting Conc. Of Transposon • Overproduction Inhibition • TA Dinucleotides (random) • Low Promoter/Enhancer Activity • Better Transgenesis Efficiency • • • Tc 1/Mariner-like Elements Relatively Large Carrier (14 -100 kb) TTAA Incoporation Sites 3 -5 Nucleotide Mobilization Footprint Overproduction Inhibition Transient Expression Inducible Vector Non-Specific Incorporation Preferential Incorporation at Transcription Start Sites

References § http: //www. nature. com/scitable/topicpage/transposons-the-jumping-genes-518 § Feschotte, C. & Pritham, E. J. (2007) DNA transposons and the evolution of eukaryotic genomes. Annual Reviews in Genetics 41, 331– 348. § Lodish et al. , Molecular Cell Biology, 7 th ed. § Pray, L. (2008) Transposons: The jumping genes. Nature Education 1(1): 204 § López, M. M. and García-Pérez, J. L. (2010) DNA Transposons: Nature and Applications in Genomics Curr Genomics 11(2): 115– 128. doi: 10. 2174/138920210790886871 § Curcio MJ and Derbyshire, KM. (2003). The outs and ins of transposition: from Mu to Kangaroo. Nature Reviews Molecular Cell Biology, 4, 865 -877. http: //www. nature. com/nrm/journal/v 4/n 11/full/nrm 1241. html § Lodish et al. , Molecular Cell Biology, 7 th ed. § Morris, ER et al. (2016) A bend, flip and trap mechanism for transposon integration. e. LIFE; 5: e 15537. DOI: 10. 7554/e. Life. 15537. § Munoz-Lopez M. and Garcia-Perez JL. (2010). DNA Transposons: Nature and Applications in Genomics. Current Genomics, 115 -128. § Chao, M. C. , Abel, S. , Davis, B. M. , & Waldor, M. K. (2016). The design and analysis of transposon insertion sequencing experiments. Nature Reviews: Microbiology, 119 -128. § Kwon, Y. M. , Ricke, S. C. , & Mandal, R. K. (2016). Transposon sequencing: methods and expanding applications. Applications of Microbiology in Biotechnology, 31 -43. § Vargas, J. E. , Chicaybam, L. , Stein, R. T. , Tanuri, A. , Delgado-Canedo, A. , & Bonamino, M. H. (2016). Retroviral vectors and transposons for stable gene therapy: advances, current challenges, and perspectives. Journal of Translational Medicine. § Woodard, L. E. , & Wilson, M. H. (2015). piggy. Bac-ing models and new therapeutic strategies. Trends in Biotechnology, 525 -533.
- Slides: 27