Transposition Xuhua Xia xxiauottawa ca http dambe bio

Transposition Xuhua Xia xxia@uottawa. ca http: //dambe. bio. uottawa. ca

Causes of genomic evolution • Variation in genome size and gene content – sequence duplication (genome, segmental, gene) – horizontal gene transfer – transposition (Barbara Mc. Clintock) • Genomic mutation – – – replication fidelity DNA repair strand bias DNA modification functional constraints Slide 2
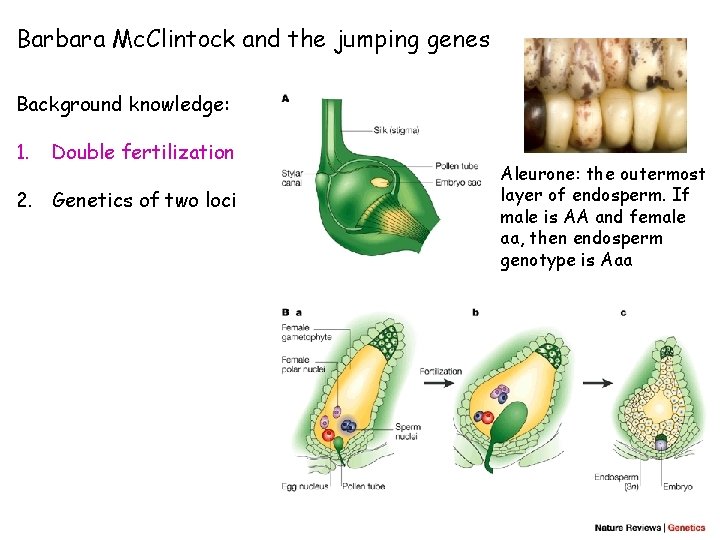
Barbara Mc. Clintock and the jumping genes Background knowledge: 1. Double fertilization 2. Genetics of two loci Aleurone: the outermost layer of endosperm. If male is AA and female aa, then endosperm genotype is Aaa
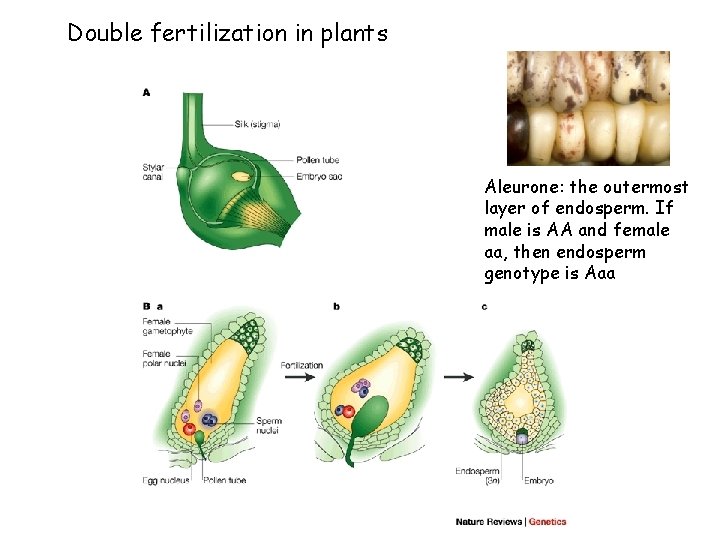
Double fertilization in plants Aleurone: the outermost layer of endosperm. If male is AA and female aa, then endosperm genotype is Aaa
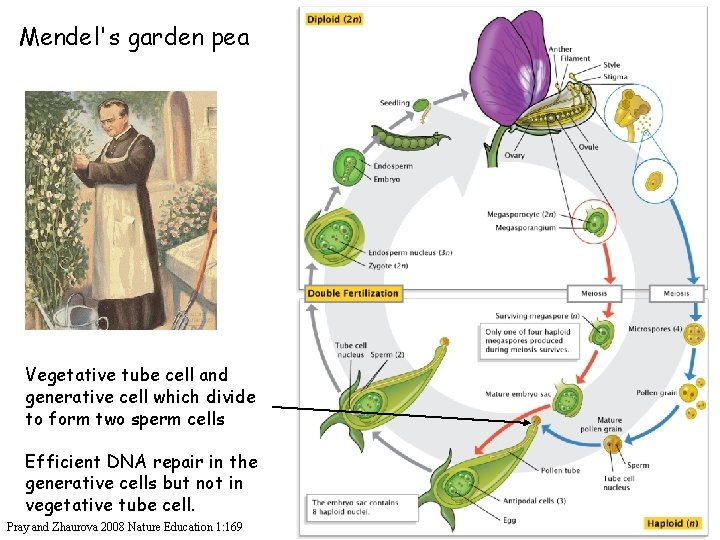
Mendel's garden pea Vegetative tube cell and generative cell which divide to form two sperm cells Efficient DNA repair in the generative cells but not in vegetative tube cell. Pray and Zhaurova 2008 Nature Education 1: 169
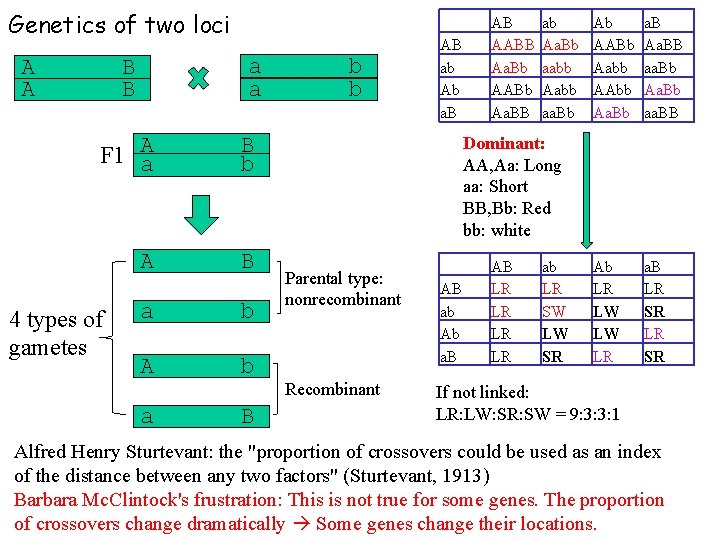
Genetics of two loci B B a a A F 1 a B b A B A A 4 types of gametes a b A b a B b b AB ab Ab a. B AB AABB Aa. Bb AABb Aa. BB ab Aa. Bb aabb Aabb aa. Bb Ab AABb Aabb AAbb Aa. Bb a. B Aa. BB aa. Bb Aa. Bb aa. BB Ab LR LW LW LR a. B LR SR Dominant: AA, Aa: Long aa: Short BB, Bb: Red bb: white Parental type: nonrecombinant Recombinant AB ab Ab a. B AB LR LR ab LR SW LW SR If not linked: LR: LW: SR: SW = 9: 3: 3: 1 Alfred Henry Sturtevant: the "proportion of crossovers could be used as an index of the distance between any two factors" (Sturtevant, 1913) Barbara Mc. Clintock's frustration: This is not true for some genes. The proportion of crossovers change dramatically Some genes change their locations.
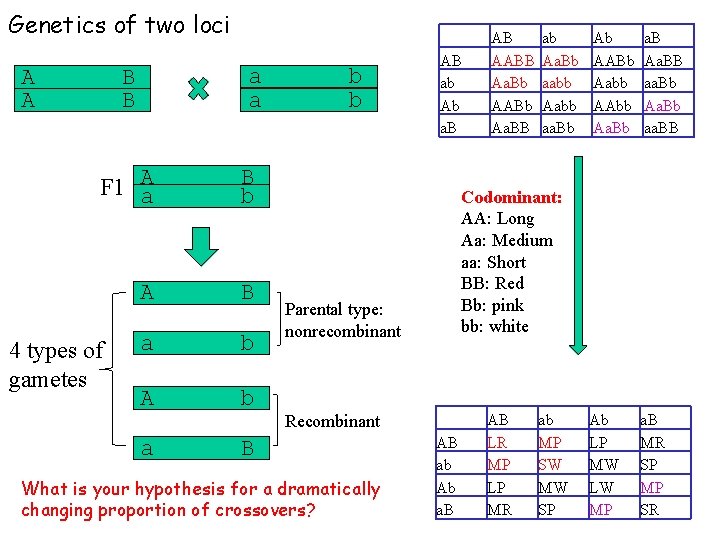
Genetics of two loci B B a a A F 1 a B b A B a b A b a B A A 4 types of gametes b b AB ab Ab a. B ab Aa. Bb aabb Aabb aa. Bb Ab AABb Aabb AAbb Aa. Bb a. B Aa. BB aa. Bb Aa. Bb aa. BB Ab LP MW LW MP a. B MR SP MP SR Codominant: AA: Long Aa: Medium aa: Short BB: Red Bb: pink bb: white Parental type: nonrecombinant Recombinant What is your hypothesis for a dramatically changing proportion of crossovers? AB AABB Aa. Bb AABb Aa. BB AB ab Ab a. B AB LR MP LP MR ab MP SW MW SP

Table 1: Maize Genes Studied by Barbara Mc. Clintock Gene Description C' Dominant allele on the short arm of Chr 9 that prevents color from being expressed in the aleurone layer of the maize kernel, causing a so-called "colorless" phenotype (which is actually white or yellow in color). This is also known as the inhibitor allele. C Recessive allele on the short arm of Chr 9 that leads to color development. Bz Dominant allele on the short arm of Chr 9 that leads to a purple phenotype. bz Recessive allele on the short arm of Chr 9 that leads to a dark brown phenotype. Ds "Dissociation gene": Genetic location on the short arm of chromosome 9 at which chromosomal breakage occurs. Ac Activator gene: A factor of unknown location (at least when Mc. Clintock was conducting her research) that impacts the expression of Ds. Mostly colorless, but some have dark brown or purple spots Inferences: 1. C'? ? Ds genotype has colors: C' is disabled by Ds 2. CCCBzbzbz. Ds is dark brown instead of purple: Bz disabled by Ds, allowing bz to express 3. The genotype regains purple: Ds has moved away from original Ds 4. Ds effect depends on Ac, both defying mapping: they are jumping genes, and Ds needs As to jump Further observations: 1. CCCBzbzbz. Ds-- may be dark brown instead of purple 2. The genotype above may regain the purple color 3. Ds effect depends on Ac, both defying mapping Pray and Zhaurova 2008 Nature Education 1: 169 CCbzbz-- C'C'Bz. Ds. Ds C'CCBzbzbz. Ds--
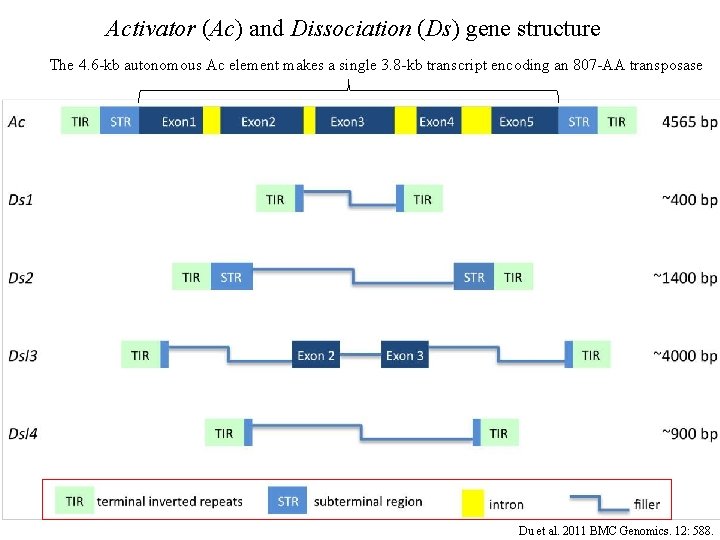
Activator (Ac) and Dissociation (Ds) gene structure The 4. 6 -kb autonomous Ac element makes a single 3. 8 -kb transcript encoding an 807 -AA transposase Du et al. 2011 BMC Genomics. 12: 588.
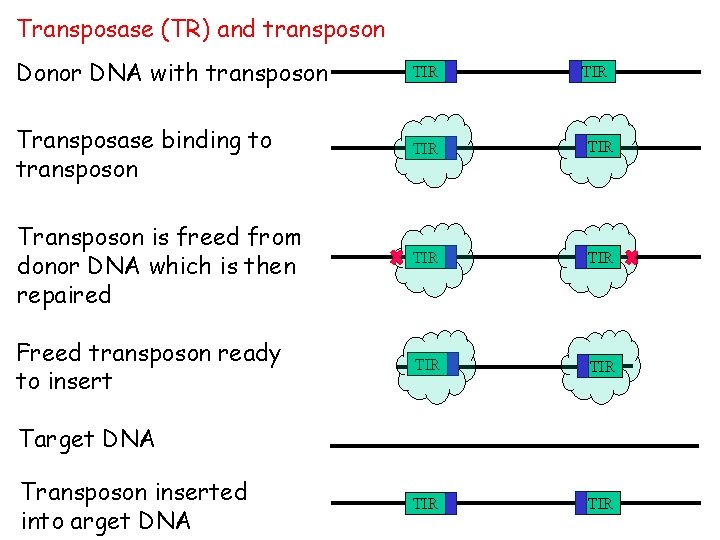
Transposase (TR) and transposon Donor DNA with transposon TIR Transposase binding to transposon TIR Transposon is freed from donor DNA which is then repaired TIR Freed transposon ready to insert TIR TIR TIR Target DNA Transposon inserted into arget DNA
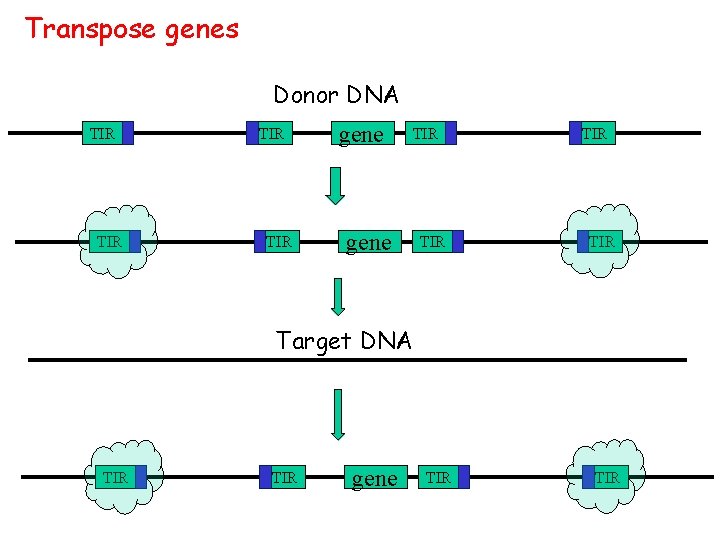
Transpose genes Donor DNA TIR TIR gene TIR TIR Target DNA TIR gene TIR
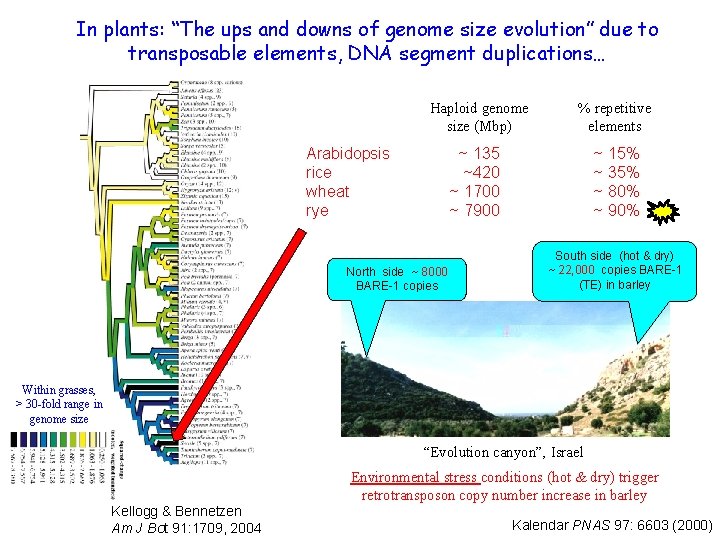
In plants: “The ups and downs of genome size evolution” due to transposable elements, DNA segment duplications… Haploid genome size (Mbp) % repetitive elements ~ 135 ~420 ~ 1700 ~ 7900 ~ 15% ~ 35% ~ 80% ~ 90% Arabidopsis rice wheat rye North side ~ 8000 BARE-1 copies South side (hot & dry) ~ 22, 000 copies BARE-1 (TE) in barley Within grasses, > 30 -fold range in genome size “Evolution canyon”, Israel Kellogg & Bennetzen Am J Bot 91: 1709, 2004 Environmental stress conditions (hot & dry) trigger retrotransposon copy number increase in barley Kalendar PNAS 97: 6603 (2000)
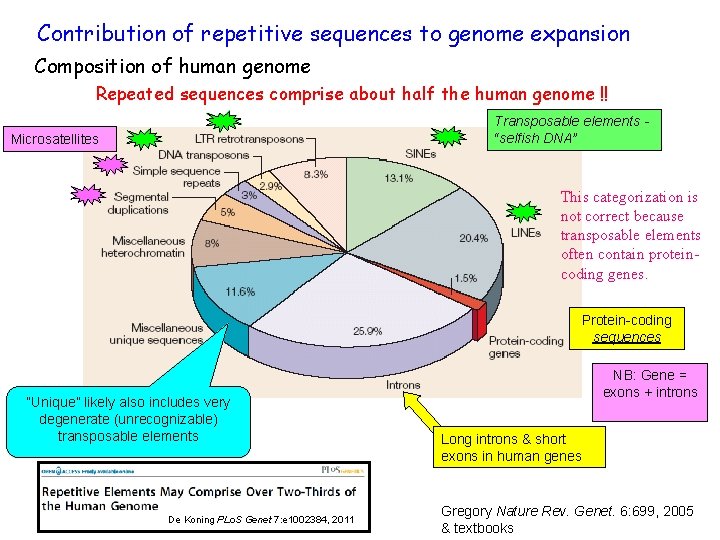
Contribution of repetitive sequences to genome expansion Composition of human genome Repeated sequences comprise about half the human genome !! Transposable elements “selfish DNA” Microsatellites This categorization is not correct because transposable elements often contain proteincoding genes. Protein-coding sequences “Unique” likely also includes very degenerate (unrecognizable) transposable elements De Koning PLo. S Genet 7: e 1002384, 2011 NB: Gene = exons + introns Long introns & short exons in human genes Gregory Nature Rev. Genet. 6: 699, 2005 & textbooks

Classification of transposons • DNA transposons – cut-and-paste DNA transposons (e. g. , Ac) – rolling-circle DNA transposons (Helitrons) – self-synthesizing DNA transposons (Polintons) • Retrotransposons – long terminal repeat (LTR) retrotransposons: Ty 1 copia, Ty 3 -gypsy, BEL-Pao-like and DIRS; ERV 1, ERV 2 and ERV 3 (of endogenous retroviruses) – non-LTR retrotransposons: • LINEs (Long INterspersed Elements): with genes encoding all functions for transposition (autonomous retroelements) • SINEs (Short INterspersed Elements): nonautonomous retroelements Wicker et al. 2007. Nature Reviews Genetics 8: 973– 982 Kapitonov & Jurka 2008. Nature Reviews Genetics 9: 411– 412 (used here)
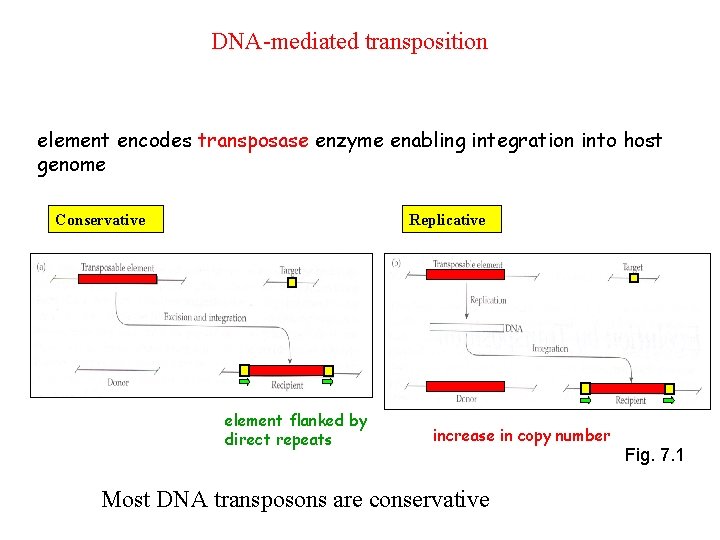
DNA-mediated transposition element encodes transposase enzyme enabling integration into host genome Conservative Replicative element flanked by direct repeats increase in copy number Most DNA transposons are conservative Fig. 7. 1
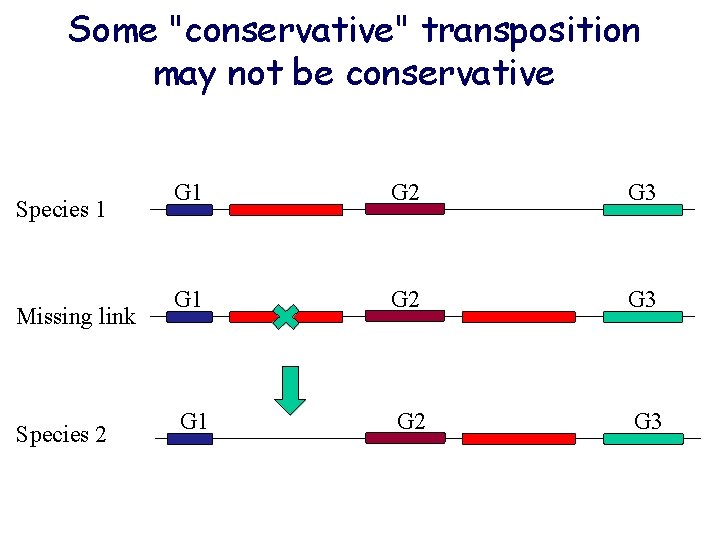
Some "conservative" transposition may not be conservative Species 1 Missing link Species 2 G 1 G 2 G 3
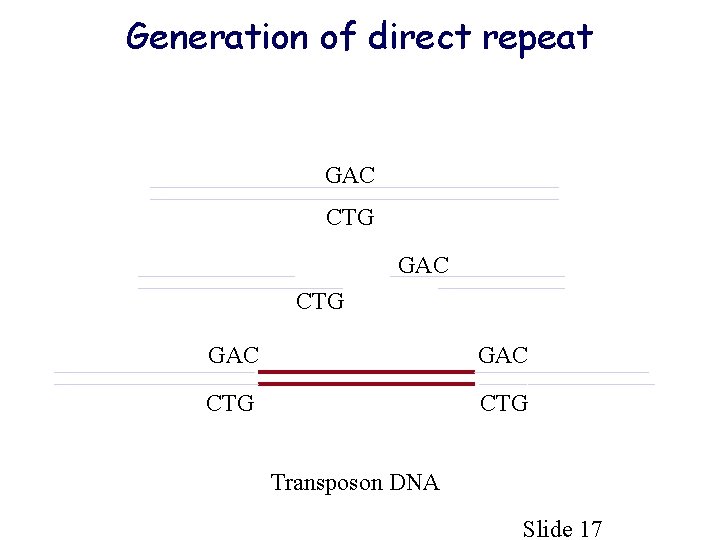
Generation of direct repeat GAC CTG Transposon DNA Slide 17
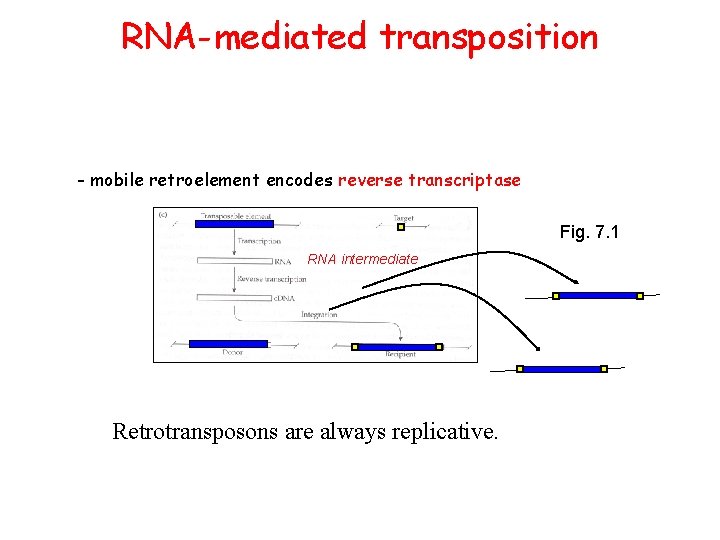
RNA-mediated transposition - mobile retroelement encodes reverse transcriptase Fig. 7. 1 RNA intermediate Retrotransposons are always replicative.
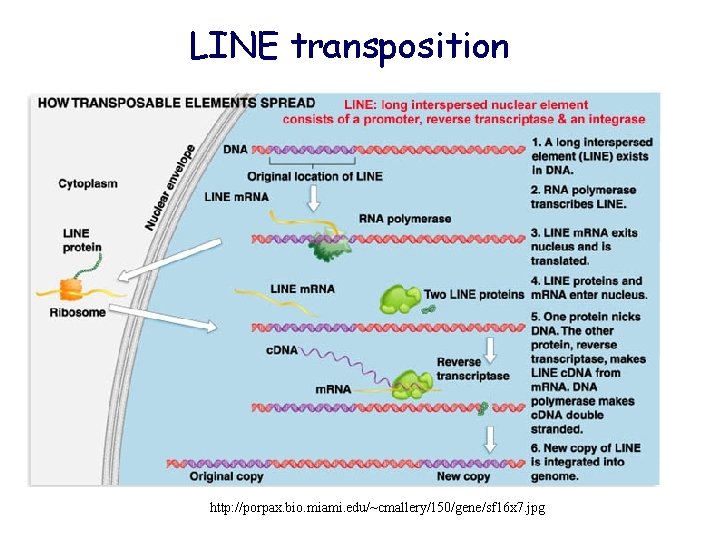
LINE transposition http: //porpax. bio. miami. edu/~cmallery/150/gene/sf 16 x 7. jpg
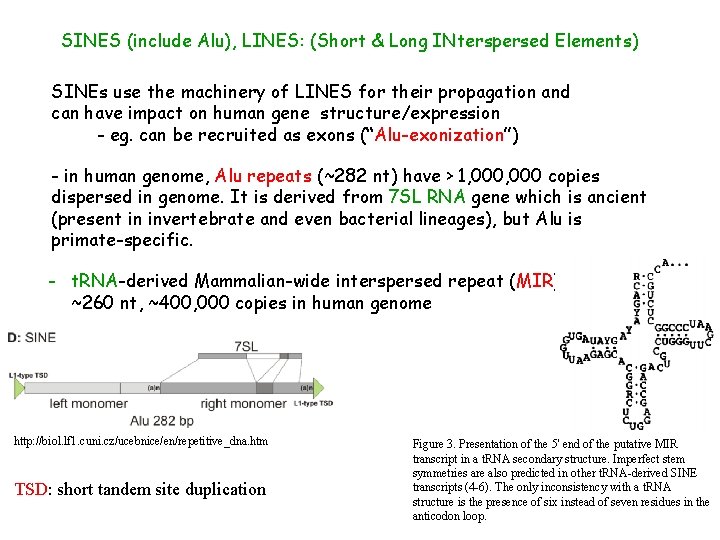
SINES (include Alu), LINES: (Short & Long INterspersed Elements) SINEs use the machinery of LINES for their propagation and can have impact on human gene structure/expression - eg. can be recruited as exons (“Alu-exonization”) - in human genome, Alu repeats (~282 nt) have > 1, 000 copies dispersed in genome. It is derived from 7 SL RNA gene which is ancient (present in invertebrate and even bacterial lineages), but Alu is primate-specific. - t. RNA-derived Mammalian-wide interspersed repeat (MIR), ~260 nt, ~400, 000 copies in human genome http: //biol. lf 1. cuni. cz/ucebnice/en/repetitive_dna. htm TSD: short tandem site duplication Figure 3. Presentation of the 5' end of the putative MIR transcript in a t. RNA secondary structure. Imperfect stem symmetries are also predicted in other t. RNA-derived SINE transcripts (4 -6). The only inconsistency with a t. RNA structure is the presence of six instead of seven residues in the anticodon loop.
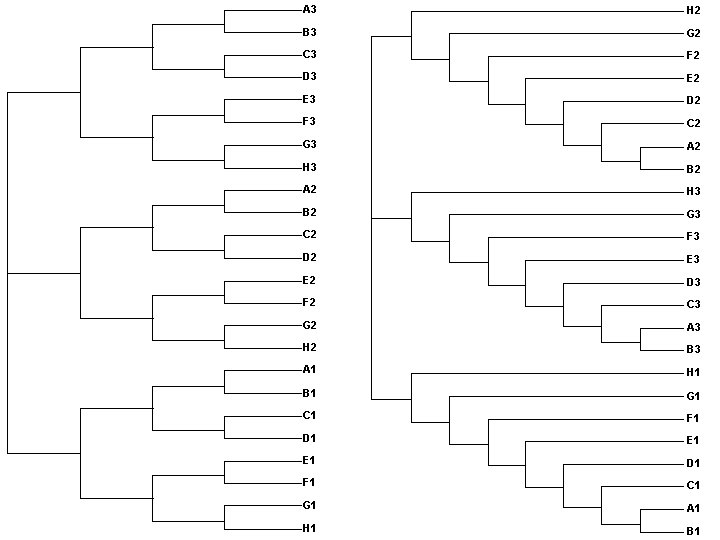
A 3 H 2 B 3 G 2 C 3 F 2 D 3 E 2 E 3 D 2 F 3 C 2 G 3 A 2 H 3 B 2 G 3 C 2 F 3 D 2 E 3 E 2 D 3 F 2 C 3 G 2 A 3 H 2 B 3 A 1 H 1 B 1 G 1 C 1 F 1 D 1 E 1 D 1 F 1 C 1 G 1 A 1 H 1 B 1
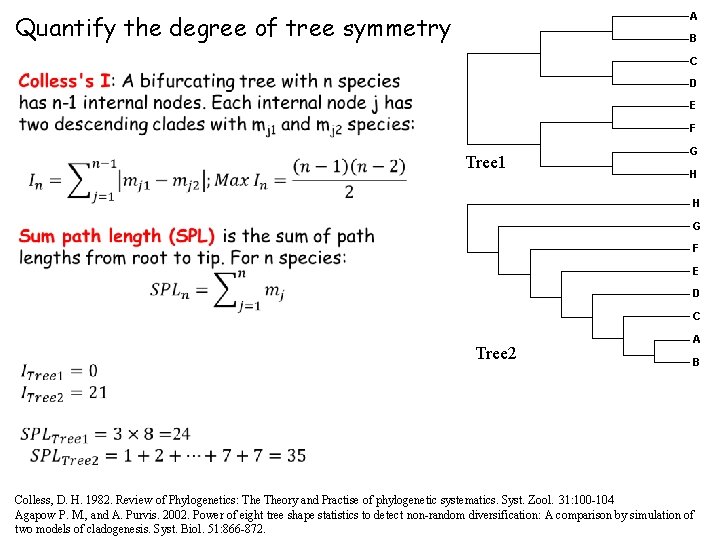
Quantify the degree of tree symmetry A B C D E F Tree 1 G H H G F E D C Tree 2 A B Colless, D. H. 1982. Review of Phylogenetics: Theory and Practise of phylogenetic systematics. Syst. Zool. 31: 100 -104 Agapow P. M. , and A. Purvis. 2002. Power of eight tree shape statistics to detect non-random diversification: A comparison by simulation of two models of cladogenesis. Syst. Biol. 51: 866 -872.
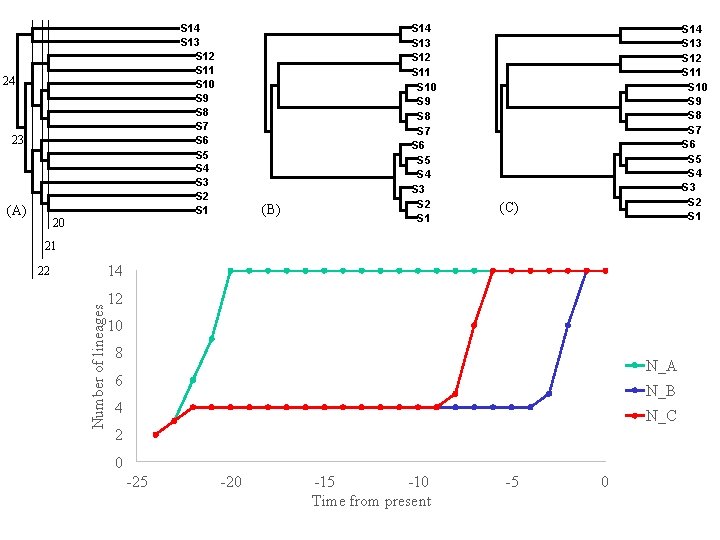
S 14 S 13 S 12 S 11 S 10 S 9 S 8 S 7 S 6 S 5 S 4 S 3 S 2 S 1 24 23 (A) 20 (B) S 14 S 13 S 12 S 11 S 10 S 9 S 8 S 7 S 6 S 5 S 4 S 3 S 2 S 1 (C) 21 14 Number of lineages 22 12 10 8 N_A 6 N_B 4 N_C 2 0 -25 -20 -15 -10 Time from present -5 0
- Slides: 23