Transposable Elements Presented by Anne Sternberger Yingnan Zhang

Transposable Elements Presented by Anne Sternberger, Yingnan Zhang & Yuxi Zhou

Transposable Elements (TEs) ● “Jumping genes” ○ Sequences of DNA that “jump” from one genome location to another ○ Discovered in 1940 s by maize geneticist Barbara Mc. Clintock ■ Initially dismissed as “junk DNA” Barbara Mc. Clintock, the “illuminator of transposons” ■ Functional roles that can be both beneficial and pathological From Hosaka and Kakutani, Curr Opin Genetics Dev. 49, 43 -48 (2018).

Transposable Elements (transposons, TE) ● Found in almost all organisms (prokaryotes and eukaryotes) and typically in large numbers. ○ E. g. comprise ~50% of human genome and ~90% of maize genome ● Two classes of TEs 1. Class 1 TEs: Retrotransposons 2. Class 2 TEs: DNA transposons ● Retrotransposons transpose via RNA intermediate ● Transposons transpose via DNA intermediate (cut-and-paste) From Hosaka and Kakutani, Curr Opin Genetics Dev. 49, 43 -48 (2018).
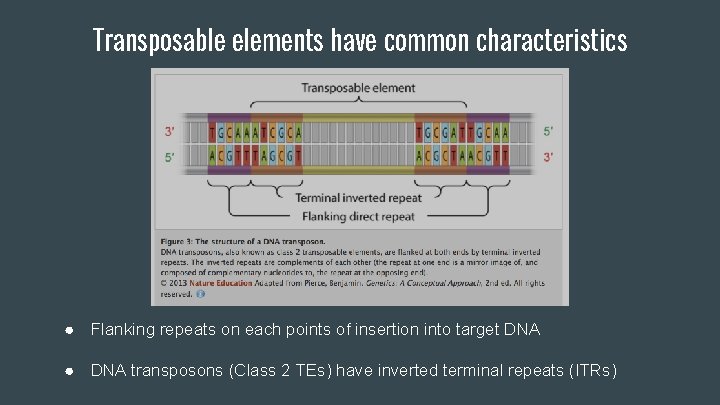
Transposable elements have common characteristics ● Flanking repeats on each points of insertion into target DNA ● DNA transposons (Class 2 TEs) have inverted terminal repeats (ITRs)
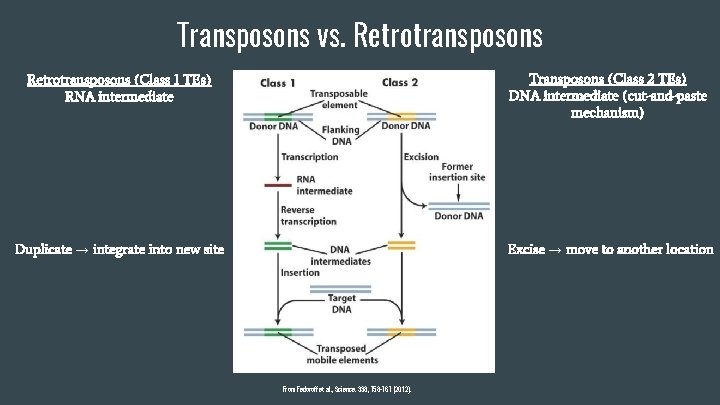
Transposons vs. Retrotransposons (Class 1 TEs) RNA intermediate Transposons (Class 2 TEs) DNA intermediate (cut-and-paste mechanism) Duplicate → integrate into new site Excise → move to another location From Fedoroff et al. , Science. 338, 758 -767 (2012).
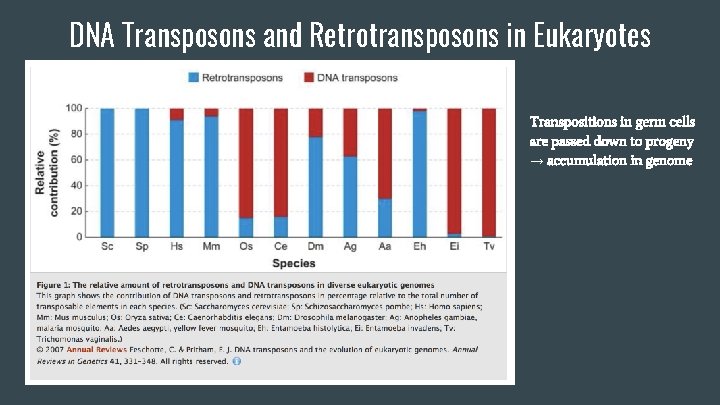
DNA Transposons and Retrotransposons in Eukaryotes Transpositions in germ cells are passed down to progeny → accumulation in genome

Autonomous vs. Non-autonomous TEs are further classified as autonomous or non-autonomous ● Autonomous TEs contain ORFs that encode proteins needed for retrotransposition ● Non-autonomous TEs lack reverse transcriptase (class 1) or transposase (Class 2) gene needed for transposition ○ “Borrow” proteins from other TEs (e. g. Ac/Ds elements) From Munoz-Lopez and Garcia-Perez. Curr Genomics. 11(2), 115 -128.
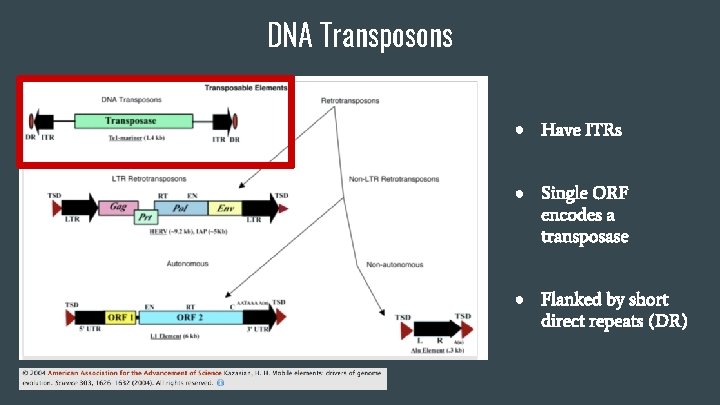
DNA Transposons ● Have ITRs ● Single ORF encodes a transposase ● Flanked by short direct repeats (DR)
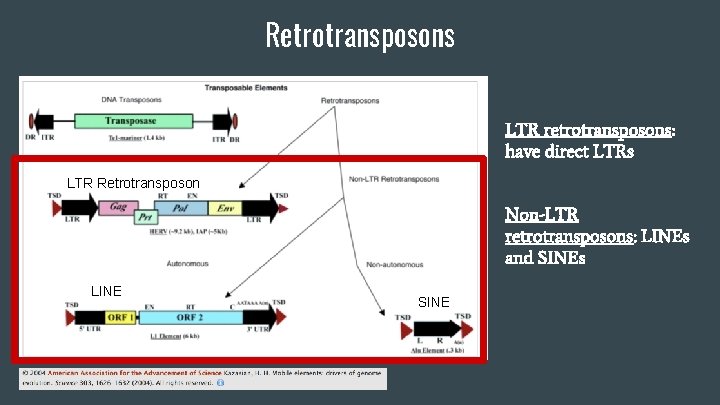
Retrotransposons LTR retrotransposons: have direct LTRs LTR Retrotransposon Non-LTR retrotransposons: LINEs and SINEs LINE SINE
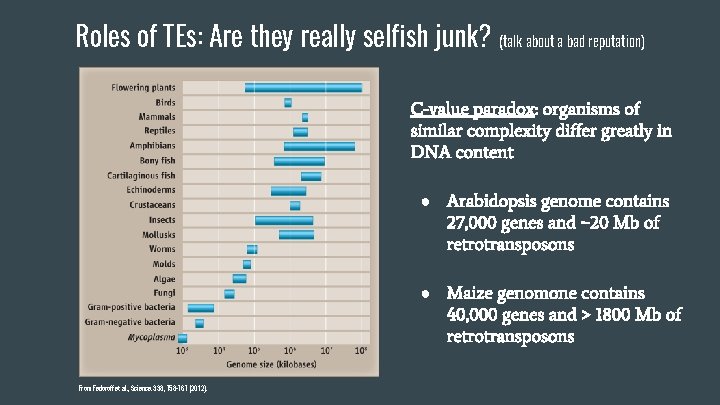
Roles of TEs: Are they really selfish junk? (talk about a bad reputation) C-value paradox: organisms of similar complexity differ greatly in DNA content ● Arabidopsis genome contains 27, 000 genes and ~20 Mb of retrotransposons ● Maize genomone contains 40, 000 genes and > 1800 Mb of retrotransposons From Fedoroff et al. , Science. 338, 758 -767 (2012).
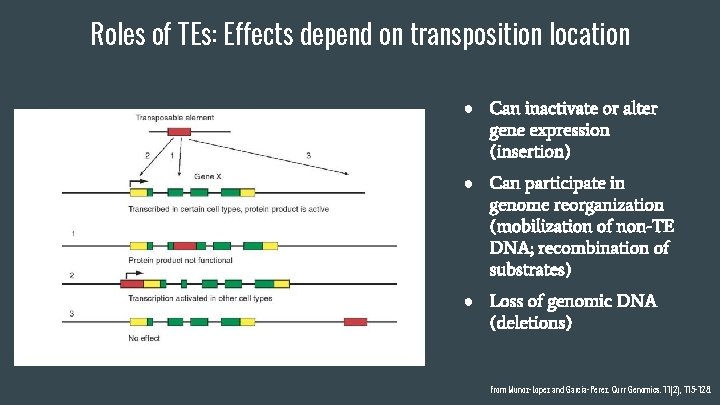
Roles of TEs: Effects depend on transposition location ● Can inactivate or alter gene expression (insertion) ● Can participate in genome reorganization (mobilization of non-TE DNA; recombination of substrates) ● Loss of genomic DNA (deletions) From Munoz-Lopez and Garcia-Perez. Curr Genomics. 11(2), 115 -128.
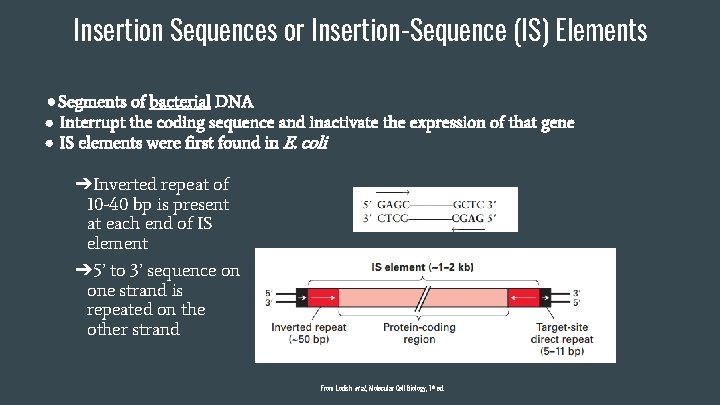
Insertion Sequences or Insertion-Sequence (IS) Elements ● Segments of bacterial DNA ● Interrupt the coding sequence and inactivate the expression of that gene ● IS elements were first found in E. coli ➔Inverted repeat of 10 -40 bp is present at each end of IS element ➔ 5’ to 3’ sequence on one strand is repeated on the other strand From Lodish et al. , Molecular Cell Biology, 7 th ed.
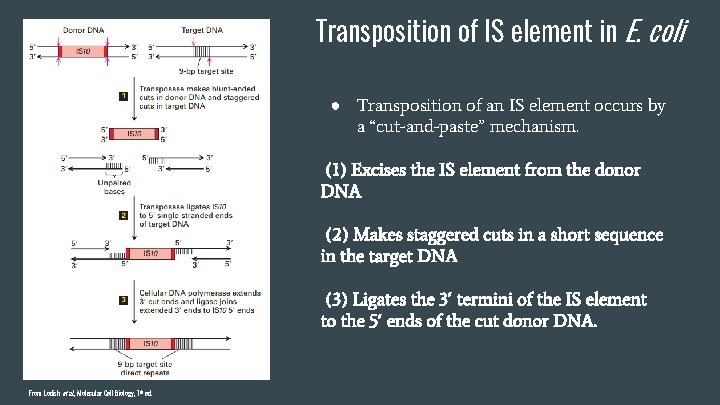
Transposition of IS element in E. coli ● Transposition of an IS element occurs by a “cut-and-paste” mechanism. (1) Excises the IS element from the donor DNA (2) Makes staggered cuts in a short sequence in the target DNA (3) Ligates the 3′ termini of the IS element to the 5′ ends of the cut donor DNA. From Lodish et al. , Molecular Cell Biology, 7 th ed.
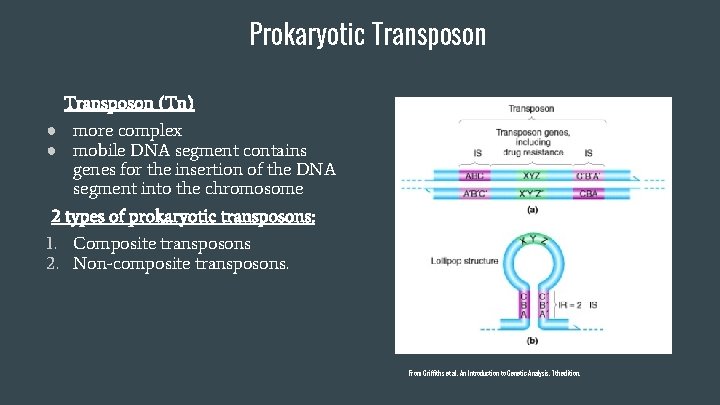
Prokaryotic Transposon (Tn) ● more complex ● mobile DNA segment contains genes for the insertion of the DNA segment into the chromosome 2 types of prokaryotic transposons: 1. Composite transposons 2. Non-composite transposons. From Griffiths et al. An Introduction to Genetic Analysis. 7 th edition.
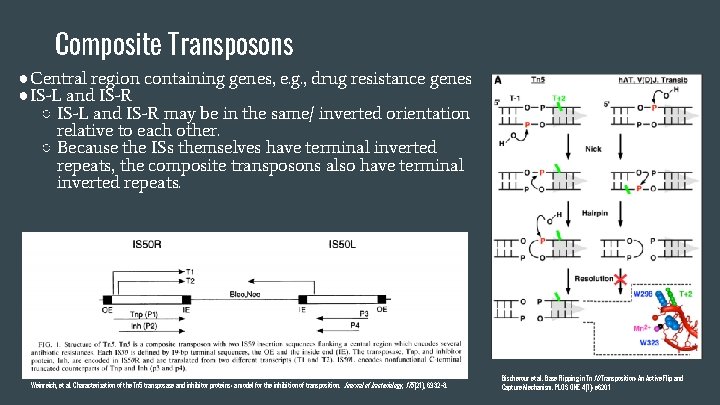
Composite Transposons ●Central region containing genes, e. g. , drug resistance genes ●IS-L and IS-R ○ IS-L and IS-R may be in the same/ inverted orientation relative to each other. ○ Because the ISs themselves have terminal inverted repeats, the composite transposons also have terminal inverted repeats. Weinreich, et al. Characterization of the Tn 5 transposase and inhibitor proteins: a model for the inhibition of transposition. Journal of bacteriology , 175(21), 6932 -8. Bischerour et al. Base Flipping in Tn 10 Transposition: An Active Flip and Capture Mechanism. PLOS ONE 4(7): e 6201
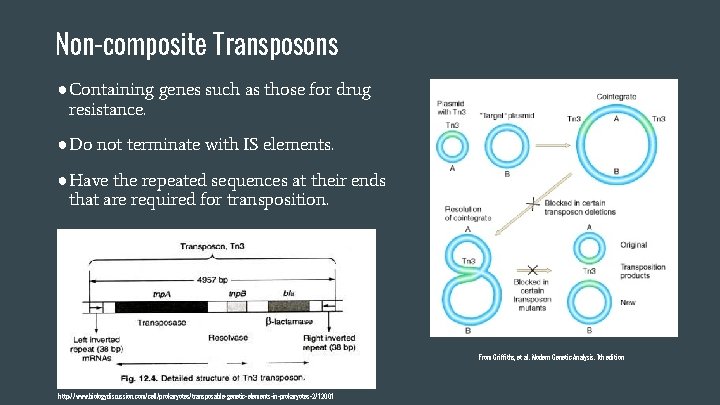
Non-composite Transposons ●Containing genes such as those for drug resistance. ●Do not terminate with IS elements. ●Have the repeated sequences at their ends that are required for transposition. From Griffiths, et al. Modern Genetic Analysis. 7 th edition http: //www. biologydiscussion. com/cell/prokaryotes/transposable-genetic-elements-in-prokaryotes-2/12001

Prokaryotic Summary • E. coli mutations caused by the spontaneous insertion of DNA sequence: insertion sequence/ IS element • Transposition of IS element is rare: 1 per 105 -107 cells per generation • Transpositions can inactivate essential genes, killing the host cell and IS elements it carries • Higher rates of transposition would result in too much mutation rate • IS elements transpose can enter nonessential regions and into plasmids or lysogenic viruses Lodish et al. , Molecular Cell Biology, 7 th ed.

Eukaryotic transposons: Transposable elements(TEs) occur in almost all eukaryotic genomes. In most situations, the transposons in a genome are epigenetically silenced(for example, silenced by histone modification). Helitrons are a group of Transposable elements. They are described as eukaryotic class 2 transposable element. Wang, Zhenxing, and Kunze, Reinhard(Jun 2015) Transposons in Eukaryotes (Part A): Structures, Mechanisms and Applications. In: e. LS. John Wiley & Sons Ltd, Chichester. http: //www. els. net [doi: 10. 1002/9780470015902. a 0026264]
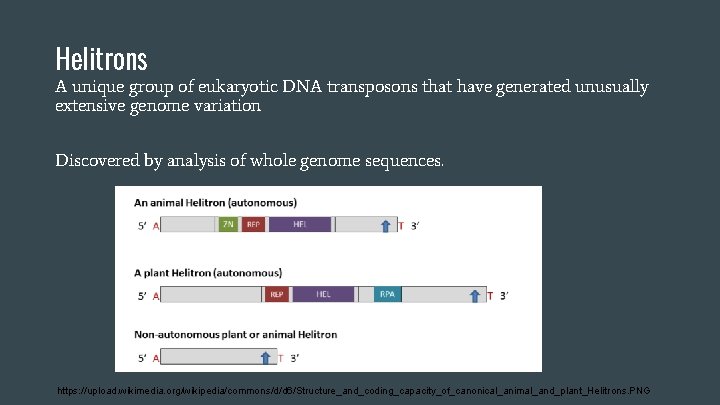
Helitrons A unique group of eukaryotic DNA transposons that have generated unusually extensive genome variation Discovered by analysis of whole genome sequences. https: //upload. wikimedia. org/wikipedia/commons/d/d 6/Structure_and_coding_capacity_of_canonical_animal_and_plant_Helitrons. PNG
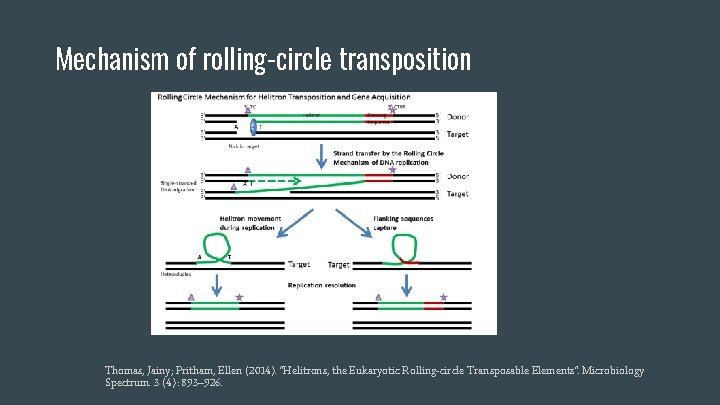
Mechanism of rolling-circle transposition Thomas, Jainy; Pritham, Ellen (2014). "Helitrons, the Eukaryotic Rolling-circle Transposable Elements". Microbiology Spectrum. 3 (4): 893– 926.
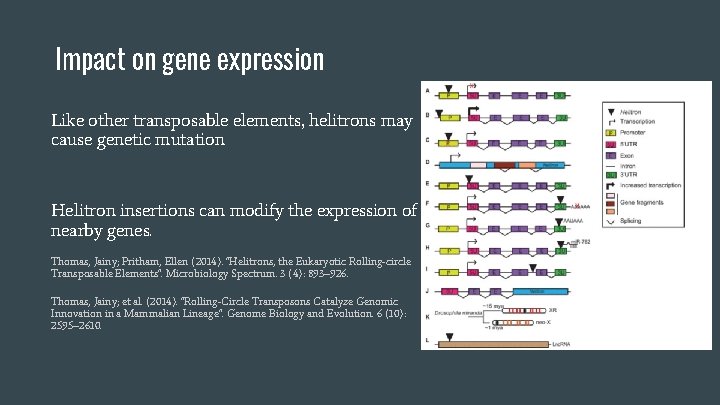
Impact on gene expression Like other transposable elements, helitrons may cause genetic mutation Helitron insertions can modify the expression of nearby genes. Thomas, Jainy; Pritham, Ellen (2014). "Helitrons, the Eukaryotic Rolling-circle Transposable Elements". Microbiology Spectrum. 3 (4): 893– 926. Thomas, Jainy; et al. (2014). "Rolling-Circle Transposons Catalyze Genomic Innovation in a Mammalian Lineage". Genome Biology and Evolution. 6 (10): 2595– 2610.
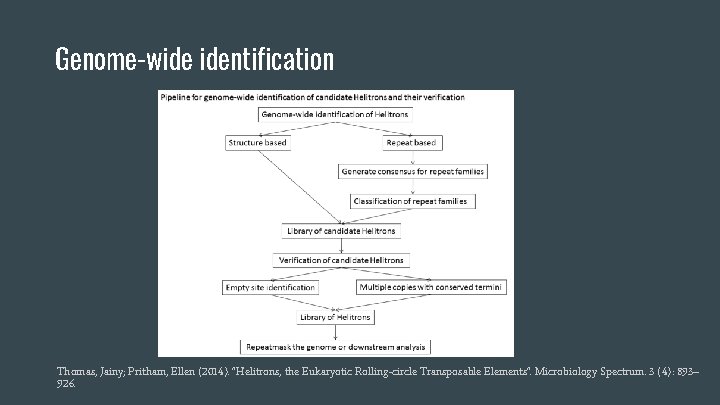
Genome-wide identification Thomas, Jainy; Pritham, Ellen (2014). "Helitrons, the Eukaryotic Rolling-circle Transposable Elements". Microbiology Spectrum. 3 (4): 893– 926.
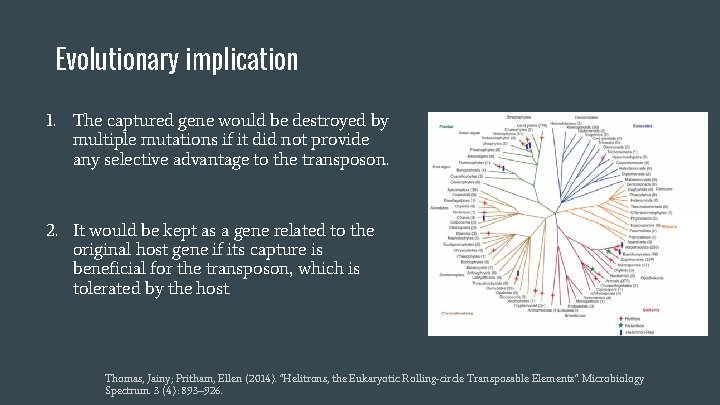
Evolutionary implication 1. The captured gene would be destroyed by multiple mutations if it did not provide any selective advantage to the transposon. 2. It would be kept as a gene related to the original host gene if its capture is beneficial for the transposon, which is tolerated by the host. Thomas, Jainy; Pritham, Ellen (2014). "Helitrons, the Eukaryotic Rolling-circle Transposable Elements". Microbiology Spectrum. 3 (4): 893– 926.
- Slides: 23