Transition Bias and Substitution models Xuhua Xia xxiauottawa

Transition Bias and Substitution models Xuhua Xia xxia@uottawa. ca http: //dambe. bio. uottawa. ca

Transitions and Transversions A G Purine C T Pyrimidine A G C T Transition: the substitution of a purine for a purine or a pyrimidine for a pyrimidine. Symbolized by s. Transversion: the substitution of a purine for a pyrimidine or vice versa. Symbolized by v. What is transition bias? A G C T Xuhua Xia Transition bias refers to the degree by which the s/v ratio deviates from the expected 1/2. The observed s/v ratio is almost always much larger than 1/2.
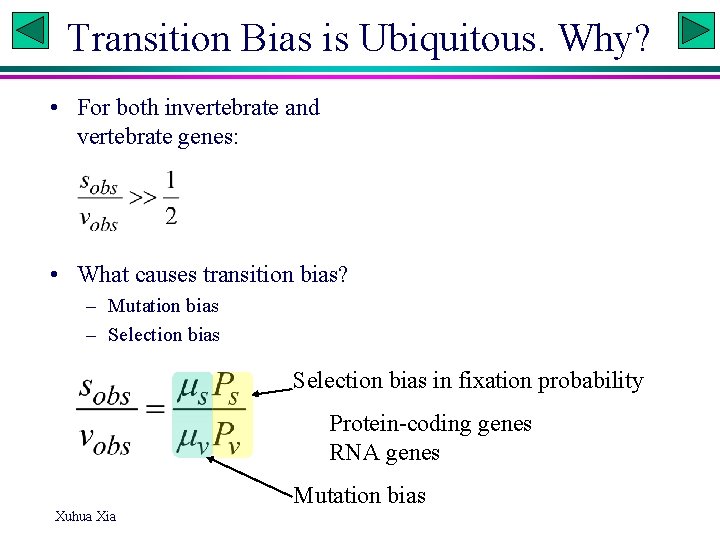
Transition Bias is Ubiquitous. Why? • For both invertebrate and vertebrate genes: • What causes transition bias? – Mutation bias – Selection bias in fixation probability Protein-coding genes RNA genes Mutation bias Xuhua Xia
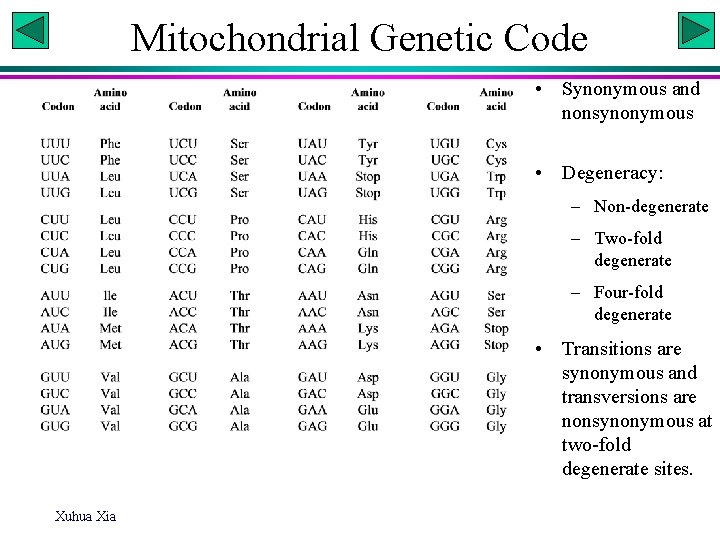
Mitochondrial Genetic Code • Synonymous and nonsynonymous • Degeneracy: – Non-degenerate – Two-fold degenerate – Four-fold degenerate • Transitions are synonymous and transversions are nonsynonymous at two-fold degenerate sites. Xuhua Xia
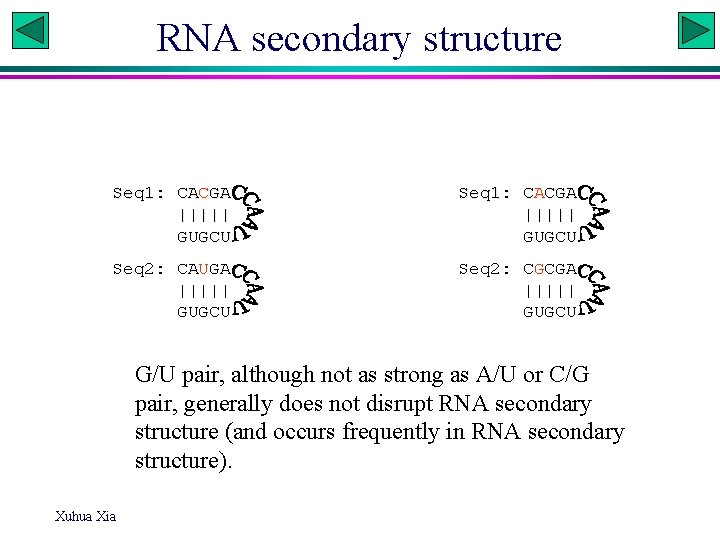
RNA secondary structure Seq 1: CACGA ||||| GUGCU Seq 2: CAUGA ||||| GUGCU Seq 2: CGCGA ||||| GUGCU G/U pair, although not as strong as A/U or C/G pair, generally does not disrupt RNA secondary structure (and occurs frequently in RNA secondary structure). Xuhua Xia

Causes of transition bias I often say that when you can measure what you are speaking about, and express it in numbers, you know something about it; but when you cannot measure it, when you cannot express it in numbers, your knowledge is of a meagre and unsatisfactory kind; it may be the beginning of knowledge, but you have scarcely in your thoughts advanced to the state of Science, whatever the matter may be. " Lord Kelvin: Phys. Letter A, vol. 1, "Electrical Units of Measurement", 1883 -05 -03 Xuhua Xia

At Four-fold Degenerate Sites At four-fold degenerate sites, all nucleotide substitutions are synonymous and subject to roughly the same selection pressure (similar fixation probabilities) Gly Asn Lys Gly Asp Lys Ala Pro Ala Cys. . . Fold 4 2 2 4 4 4 2 S 1 GGA AAU AAA GGA GAC AAA GCC CCU GCG UGU. . . S 2 GGG AAC AAA GAU AAG GCC GCU CCA GGG UGG. . . s s v Glu Gly Trp Glycine codon: GGA GGC GGG GGT Four-fold degenerate site Xuhua Xia

At Nondegenerate Sites At nondegenerate sites, all nucleotide substitutions are nonsynonymous and subject to roughly the same selection pressure (similar fixation probabilities) S 1 S 2 Glycine codon: GGA GGC Gly Asn Lys Gly Asp Lys Ala Pro Ala Cys. . . GGA AAU AAA GGA GAC AAA GCC CCU GCG UGU. . . GGG AAC AAA GAU AAG GCC GCU CCA GGG UGG. . . s v Glu Gly Trp GGG GGT nondegenerate site Xuhua Xia

At Two-fold Degenerate Sites At two-fold degenerate sites, all transitional substitutions are synonymous, and all transversional substitutions are nonsynonymous Gly Asn Lys Gly Asp Lys Ala Pro Ala Cys. . . Fold 4 2 2 4 4 4 2 S 1 GGA AAU AAA GGA GAC AAA GCC CCU GCG UGU. . . S 2 GGG AAC AAA GAU AAG GCC GCU CCA GGG UGG. . . s s s v Glu Gly Trp GAA His GAG His GAC Gln GAT Gln 2 -fold degenerate site A transition is about 40 time as like to become fixed as a transversion. Xuhua Xia
Methylation and deamination H 3 CDonor Xuhua Xia Methyltransferase + Acceptor H 3 C-
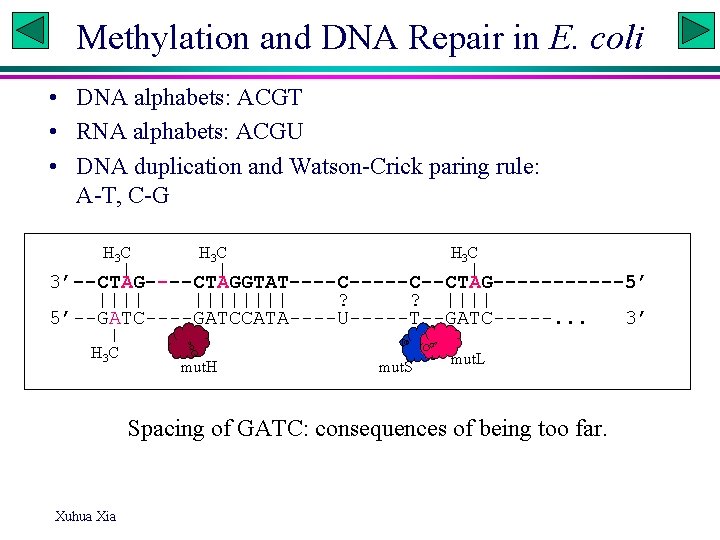
Methylation and DNA Repair in E. coli • DNA alphabets: ACGT • RNA alphabets: ACGU • DNA duplication and Watson-Crick paring rule: A-T, C-G H 3 C 3’--CTAG----CTAGGTAT----C--CTAG------5’ |||||||| ? ? |||| 5’--GATC----GATCCATA----U-----T--GATC-----. . . 3’ H 3 C mut. H mut. S mut. L Spacing of GATC: consequences of being too far. Xuhua Xia
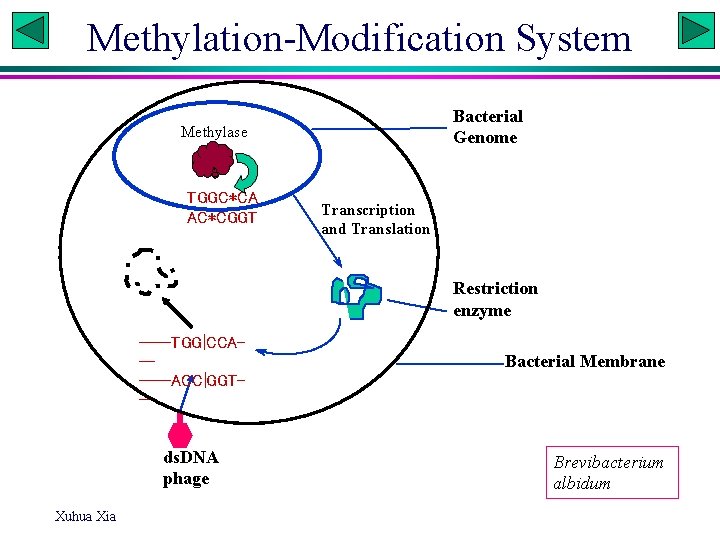
Methylation-Modification System Bacterial Genome Methylase TGGC*CA AC*CGGT Transcription and Translation Restriction enzyme ----TGG|CCA-----ACC|GGT-- ds. DNA phage Xuhua Xia Bacterial Membrane Brevibacterium albidum
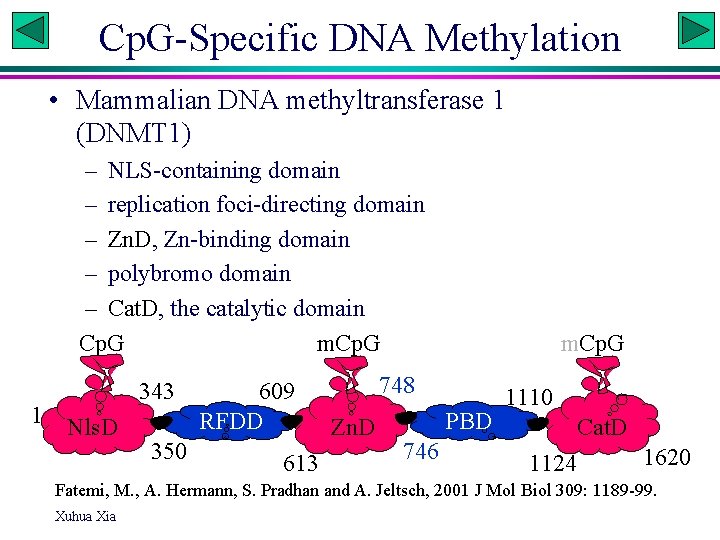
Cp. G-Specific DNA Methylation • Mammalian DNA methyltransferase 1 (DNMT 1) – NLS-containing domain – replication foci-directing domain – Zn. D, Zn-binding domain – polybromo domain – Cat. D, the catalytic domain Cp. G m. Cp. G 1 Nls. D 343 350 609 RFDD 613 m. Cp. G 748 Zn. D PBD 746 1110 Cat. D 1124 1620 Fatemi, M. , A. Hermann, S. Pradhan and A. Jeltsch, 2001 J Mol Biol 309: 1189 -99. Xuhua Xia
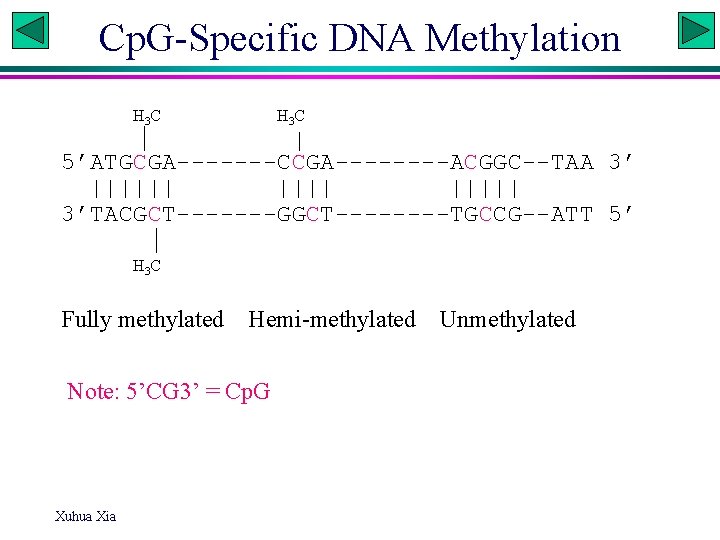
Cp. G-Specific DNA Methylation H 3 C 5’ATGCGA-------CCGA----ACGGC--TAA 3’ |||||| 3’TACGCT-------GGCT----TGCCG--ATT 5’ H 3 C Fully methylated Hemi-methylated Note: 5’CG 3’ = Cp. G Xuhua Xia Unmethylated

Methylation and Gene Regulation • Proteins with a methyl-Cp. G binding domain (MBD) – MBD 1, MBD 2, and MBD 3 – Me. CP 2 • • Deacetylases: An enzyme that removes an acetyl group Histone deacetylases: deacetylate lysyl residues in histones (the half life of an acetyl group is ~10 min). Acetylation removes a positive charge on the lysine amino group and promote nucleosome melting (and gene expression). Deacetylation tend to decrease or turn off gene expression. Histone deacetylase MBD ---m. Cp. G--------- Condensed DNA with repressed transcription Wade, P. A. , and A. P. Wolffe, 2001 Nat Struct Biol 8: 575 -7. Xuhua Xia Lysine demethylation
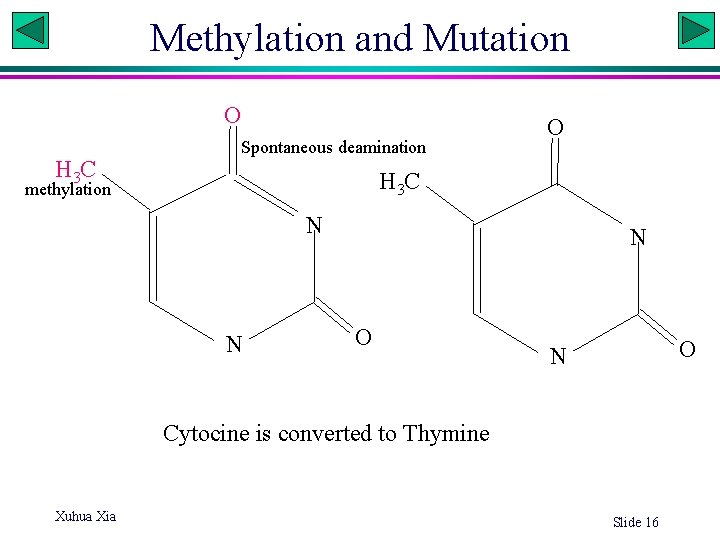
Methylation and Mutation NH O 2 H 3 C Spontaneous deamination O H 3 C methylation N N N O O N Cytocine is converted to Thymine Xuhua Xia Slide 16
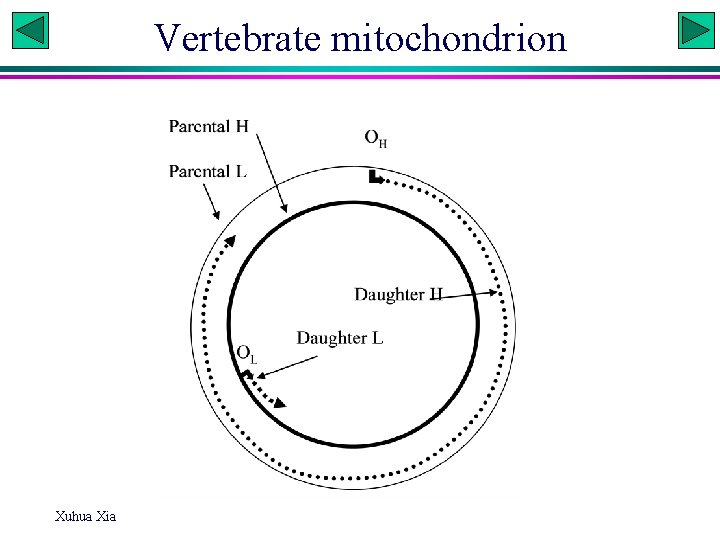
Vertebrate mitochondrion Xuhua Xia
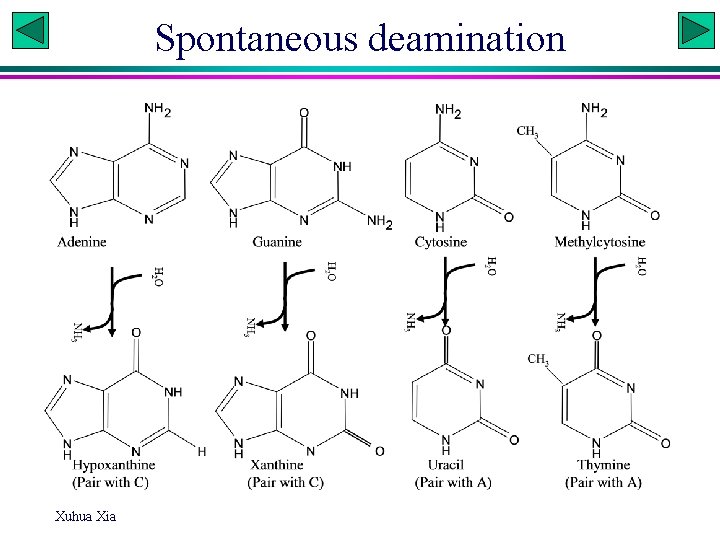
Spontaneous deamination Xuhua Xia
Transversion can erase transitions Transitions can erase transitions, and transversions can erase transversions. However, a transversion can erase many transitions occurring before it, and subsequent transitions cannot erase the transversion: AACGCTTGACG AACGCTTAACGCTTGACG AACGCTTCACG AACGCTTGACG AACGCTTTACG AACGCTTAACGCTTGACG AACGCTTTACG Although a transition could also erase 2 n transversions occurring before it, this is rare because transversions are in generally much rarer than transitions. Transitions tend to be missed in counting much more frequently than transversions. Xuhua Xia

Summary • Selection: Transitions are tolerated more than transversion by natural selection because – they are more likely synonymous in protein-coding sequences than transversions – they are less likely to disrupt RNA secondary structure than transversions. • Mutation: Transitional mutation occurs more frequently than transversions because – Misincorporation during DNA replication occur more frequently between two purines or between two pyrimidines than between a purine and a pyrimidine – A purine is more likely to mutate chemically to another purine than to a pyrimidine (e. g. , through spontaneous deamination). The same for pyrimidine. • Bias in counting: Transitions tend to be missed in counting much more frequently than transversions (which necessitates the substitution models) Xuhua Xia
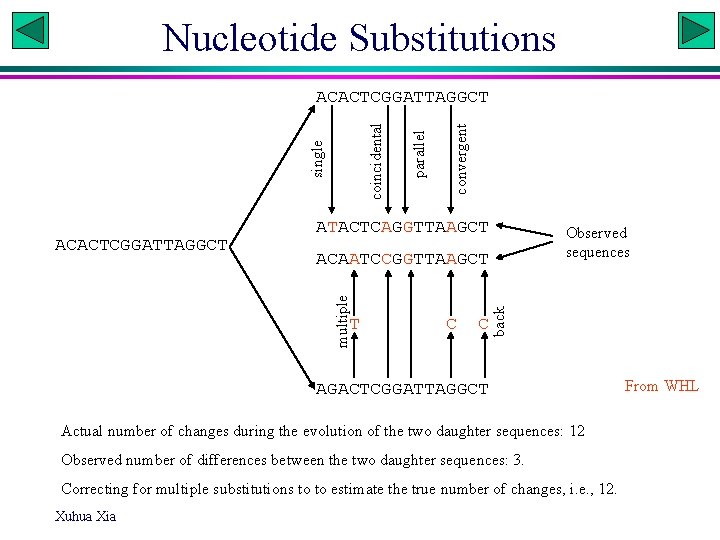
Nucleotide Substitutions convergent ATACTCAGGTTAAGCT Observed sequences T C C back ACAATCCGGTTAAGCT multiple ACACTCGGATTAGGCT parallel single coincidental ACACTCGGATTAGGCT AGACTCGGATTAGGCT Actual number of changes during the evolution of the two daughter sequences: 12 Observed number of differences between the two daughter sequences: 3. Correcting for multiple substitutions to to estimate the true number of changes, i. e. , 12. Xuhua Xia From WHL

Substitution models and phylogenetics • A substitution model is to model the evolutonary process so as to correct for multiple hits. • A phylogenetic reconstruction method implicitly or explicitly assumes a substitution model. • A phylogenetic method assuming a wrong substitution model will typically lead to wrong trees produced. • An alignment with an inappropriate substitution score matrix will typically lead to inaccurate alignment (e. g. , strong transition bias among A sequences but a substitution score matrix without strong penalty against transversion) C Xuhua Xia G T
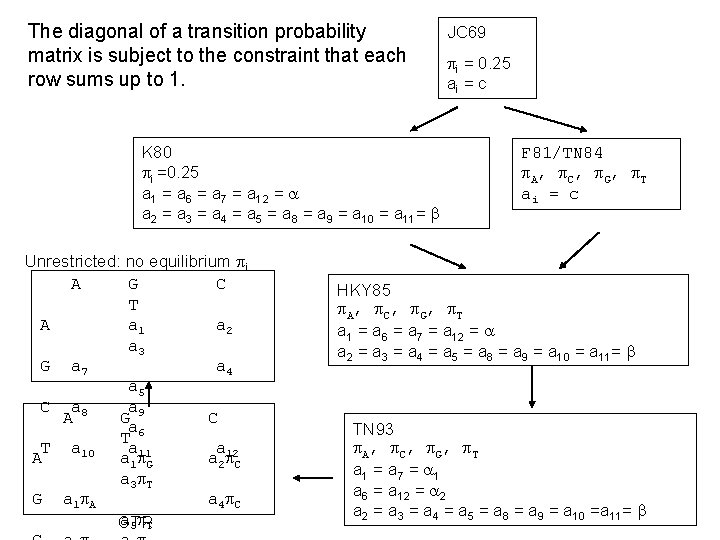
The diagonal of a transition probability matrix is subject to the constraint that each row sums up to 1. K 80 i =0. 25 a 1 = a 6 = a 7 = a 12 = a 2 = a 3 = a 4 = a 5 = a 8 = a 9 = a 10 = a 11= Unrestricted: no equilibrium i A G C T A a 1 a 2 a 3 G a 7 a 4 a 5 C a 8 a A G 9 C a 6 T T a 10 a a A a 1 11 G a 2 12 C a 3 T G a 1 A a 4 C a 5 T GTR JC 69 i = 0. 25 ai = c F 81/TN 84 A, C, G, T ai = c HKY 85 A, C, G, T a 1 = a 6 = a 7 = a 12 = a 2 = a 3 = a 4 = a 5 = a 8 = a 9 = a 10 = a 11= TN 93 A, C, G, T a 1 = a 7 = 1 a 6 = a 12 = 2 a 2 = a 3 = a 4 = a 5 = a 8 = a 9 = a 10 =a 11=

The TN 93 model as an example T A G C T C A G - frequency parameters k - rate ratio parameters In addition to illustrated assumptions, it also assumes that the frequency and rate ratio parameters do not change over time, i. e. , the substitution process is stationary. Xuhua Xia

Substitution Models • There are three types of substitution models in molecular evolution – Nucleotide-based – Amino acid-based – Codon-based • Substitution models are characterized by two categories of parameters: the frequency parameters and the ratio parameters, and different models differ by their assumptions concerning these two categories of parameters. • Substitution models, substitution score matrix and sequence alignment. Xuhua Xia
- Slides: 25