TRANSCRIPTIONSYNTHESIS OF RNA Dr Zakira Naureen Previously we





























- Slides: 69

TRANSCRIPTION--SYNTHESIS OF RNA Dr. Zakira Naureen

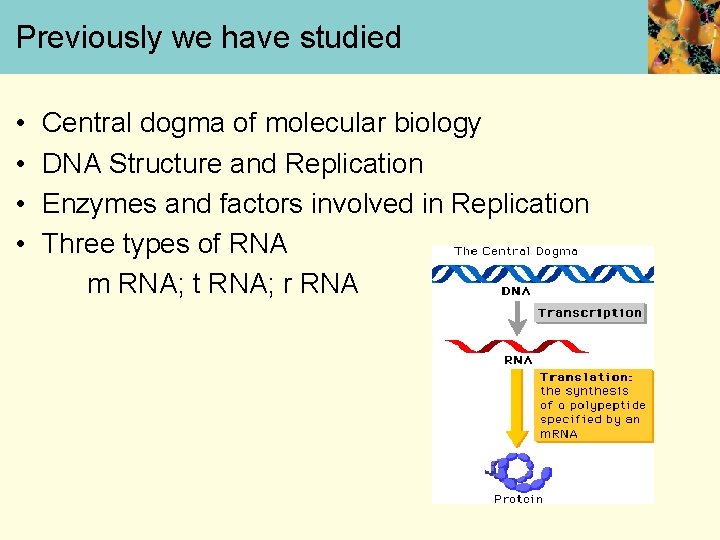
Previously we have studied • • Central dogma of molecular biology DNA Structure and Replication Enzymes and factors involved in Replication Three types of RNA m RNA; t RNA; r RNA

Learning objectives • What is gene ? • What is Transcription? • Structure of RNA polymerases

Genes • A gene is a region of DNA that controls a discrete hereditary characteristic, usually corresponding to a single m. RNA which will be translated into a protein. • In eukaryotes, the genes have their coding sequences (exons) interrupted by non-coding sequences (introns). • In humans, genes constitute only about 2 -3% of DNA, the rest is “junk” DNA.

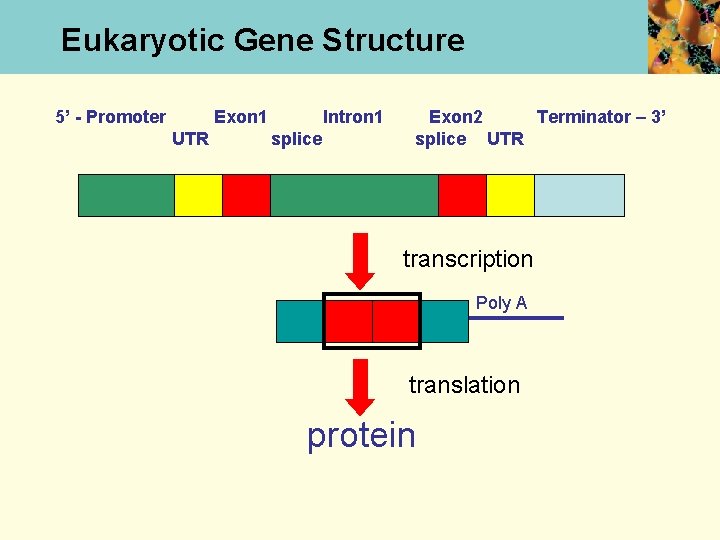
Eukaryotic Gene Structure 5’ - Promoter Exon 1 UTR Intron 1 splice Exon 2 Terminator – 3’ splice UTR transcription Poly A translation protein

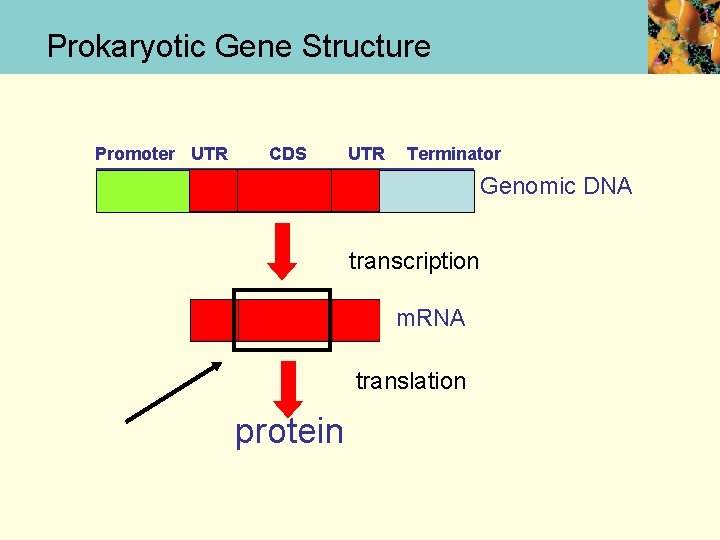
Prokaryotic Gene Structure Promoter UTR CDS UTR Terminator Genomic DNA transcription m. RNA translation protein

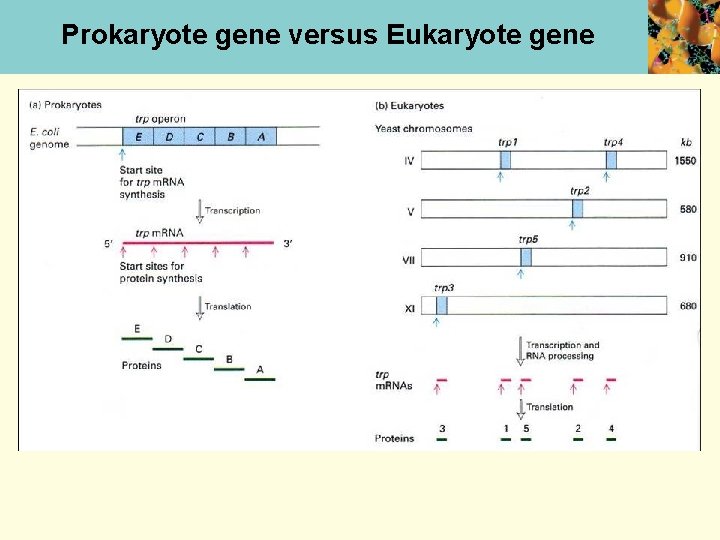
Prokaryote gene versus Eukaryote gene

What is Transcription ? • • It is the process of copying DNA to RNA. It differs from DNA synthesis in that only one strand of DNA, the template strand, is used to make m. RNA. Unlike DNA replication, it does not need a primer to start. It can involve multiple RNA polymerases. Transcription is divided into 3 stages Initiation • Promoter recognition and binding • Initiation of polymerization of NTPs Elongation Termination

Transcription Tools • • • Template DNA-dependent RNA polymerase ATP, GTP, CTP, and UTP are required, Mg 2+ Transcription factors ATP

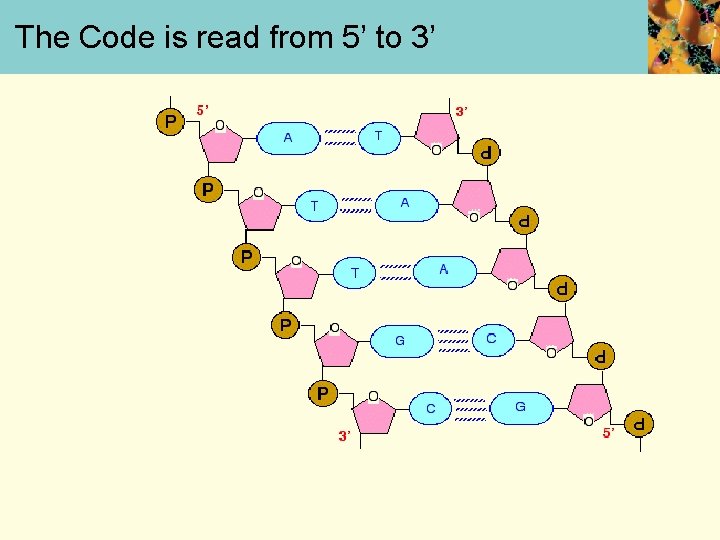
The coding strand the template strand of DNA The important thing to realize is that the genetic information is carried on only one of the two strands of the DNA. This is known as the coding strand. The other strand is known as the template strand, and is complementary to the coding strand. If you took the template strand built a new DNA strand on it (as happens in DNA replication), you would get an exact copy of the original DNA coding strand formed. Almost exactly the same thing happens when you make RNA. If you build an RNA strand on the template strand, you will get a copy of the information on the DNA coding strand - but with one important difference.

The Code is read from 5’ to 3’

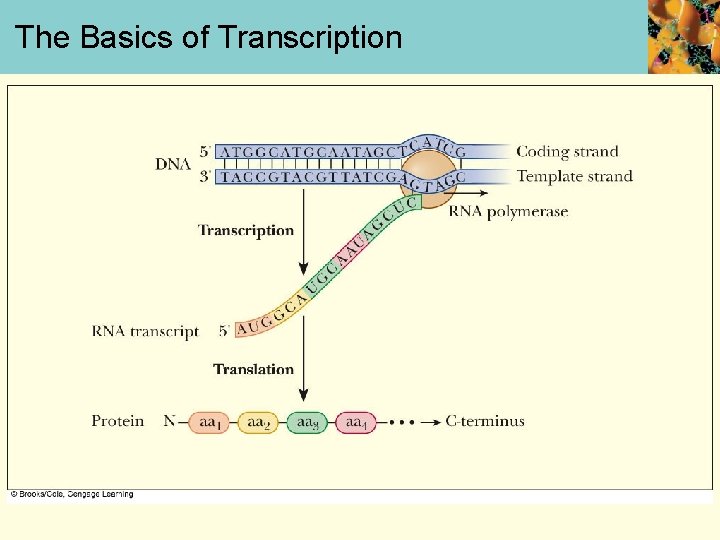
The Basics of Transcription

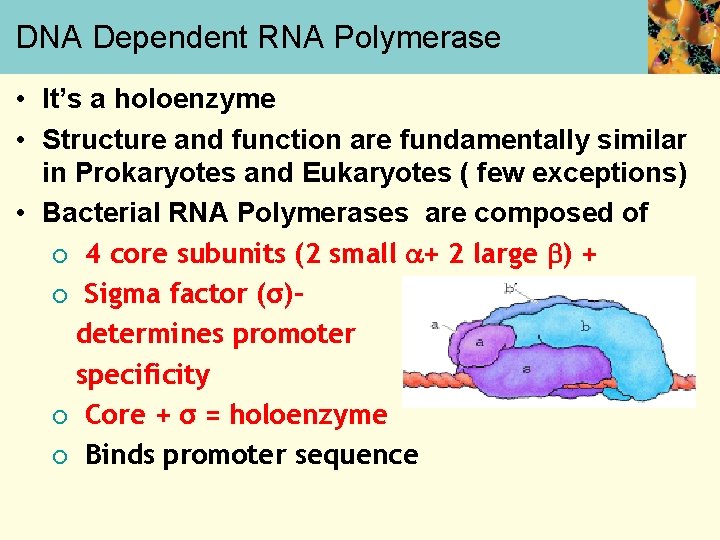
DNA Dependent RNA Polymerase • It’s a holoenzyme • Structure and function are fundamentally similar in Prokaryotes and Eukaryotes ( few exceptions) • Bacterial RNA Polymerases are composed of ¡ 4 core subunits (2 small + 2 large ) + ¡ Sigma factor (σ)– determines promoter specificity ¡ Core + σ = holoenzyme ¡ Binds promoter sequence

continued • In eukaryotes the DNA dependent RNA Polymerase has several other binding factors i. e. Transcription factors • More than one polymerases are required to make all different types of RNA unlike bacterial RNA Polymerase that can catalyze formation of all three RNA molecules. • Three polymerases for 3 RNA molecules • RNA pol I----r. RNA • RNA pol II-------m. RNA • RNA pol III------t. RNA

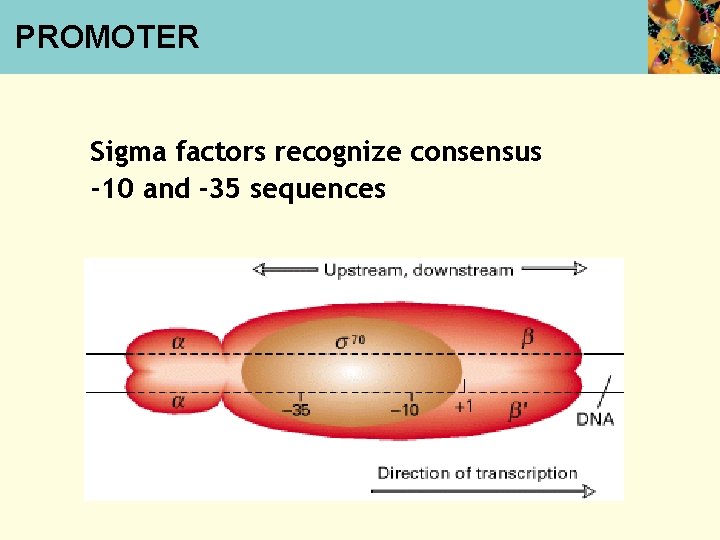
PROMOTER Sigma factors recognize consensus -10 and -35 sequences

continued • Promoter determines: 1. Which strand will serve as a template. 2. Transcription starting point. 3. Strength of polymerase binding. 4. Frequency of polymerase binding.

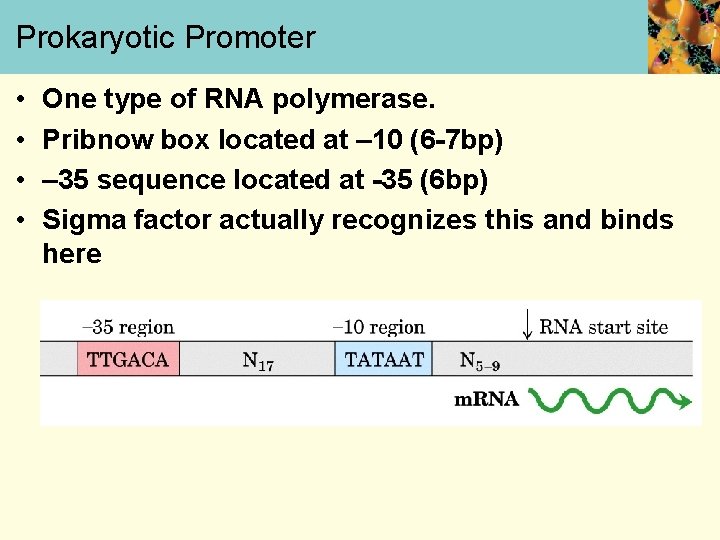
Prokaryotic Promoter • • One type of RNA polymerase. Pribnow box located at – 10 (6 -7 bp) – 35 sequence located at -35 (6 bp) Sigma factor actually recognizes this and binds here

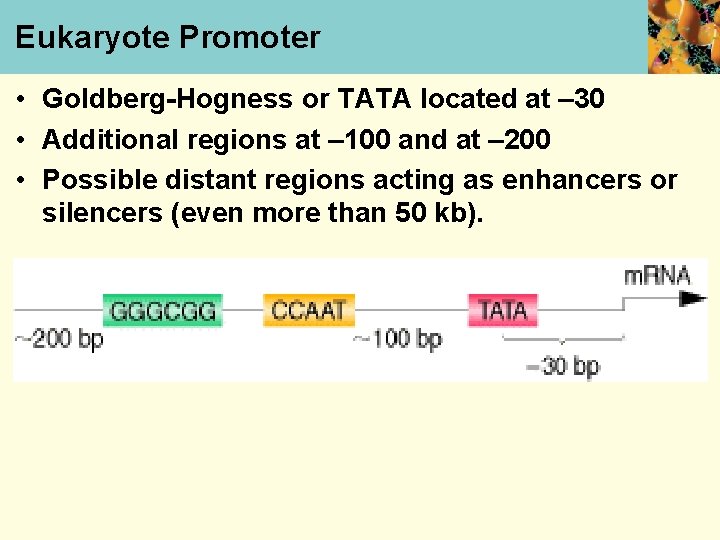
Eukaryote Promoter • Goldberg-Hogness or TATA located at – 30 • Additional regions at – 100 and at – 200 • Possible distant regions acting as enhancers or silencers (even more than 50 kb).

Promoter Sequence

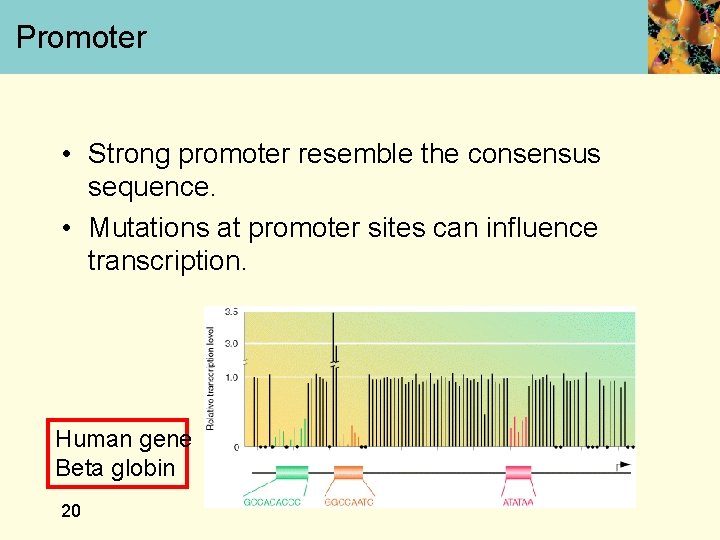
Promoter • Strong promoter resemble the consensus sequence. • Mutations at promoter sites can influence transcription. Human gene Beta globin 20

Stages of Transcription 1) Initiation – the RNA polymerase enzyme binds to a promoter site on the DNA and unzips the double helix. 2) Elongation – free nucleotides bind to their complementary pairs on the template strand of the DNA elongating the RNA chain which is identical to the informational strand of DNA, except that the nucleotide thymine in DNA is replaced by uracil in RNA. The polymerase moves along the DNA in the 3’ to 5’ direction, extending the RNA 5’ to 3’. 3) Termination – specific sequences in the DNA signal termination of transcription; when one of these is encountered by the polymerase, the RNA transcript is released from the DNA and the double helix can zip up again.

Transcription video

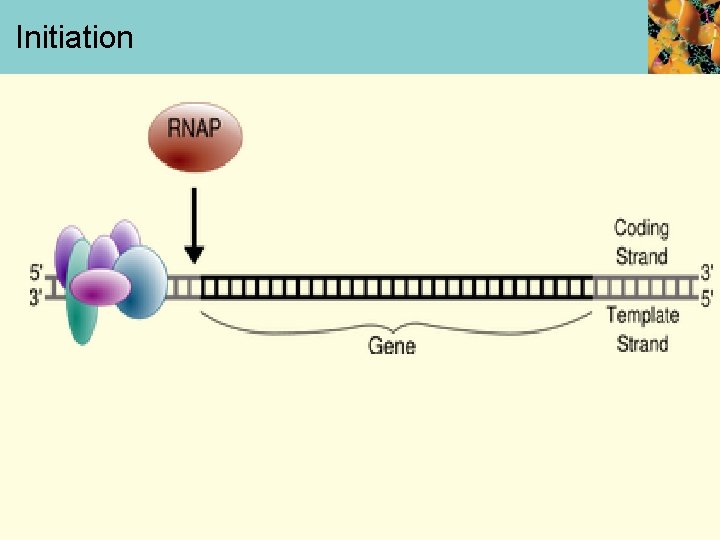
Chain Initiation • First phase of transcription is initiation • Initiation begins when RNA polymerase binds to promoter and forms closed complex • After this, DNA unwinds at promoter to form open complex, which is required for chain initiation

Initiation

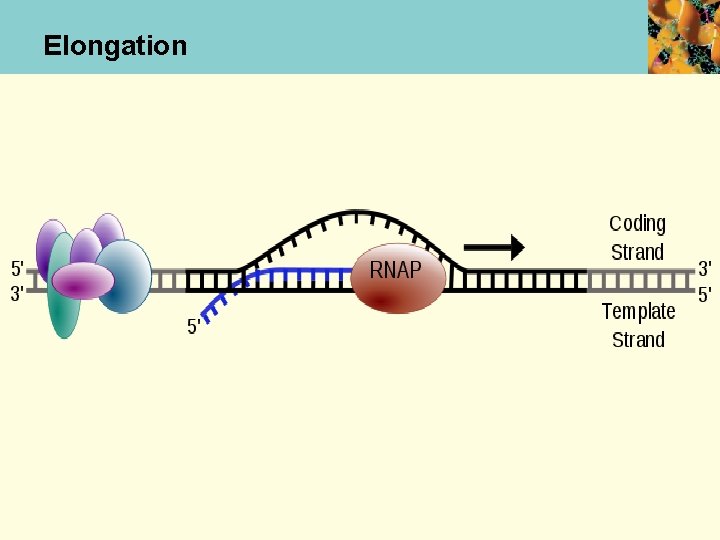
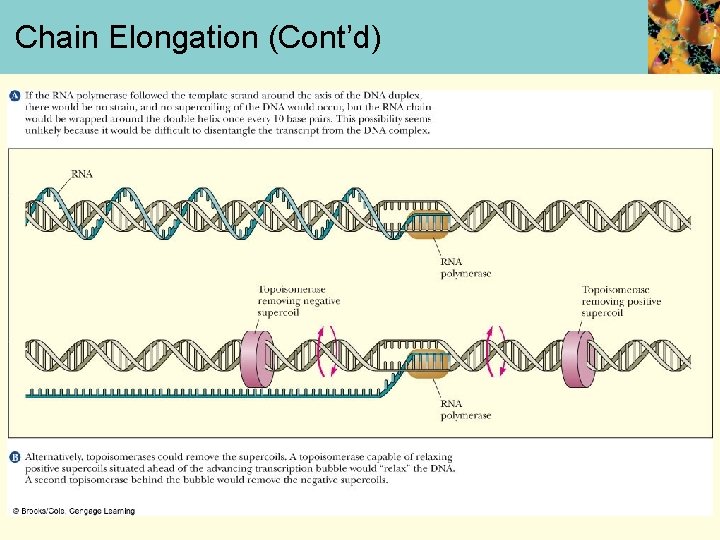
Chain Elongation • After strands separated, transcription bubble of ~17 bp moves down the DNA sequence to be transcribed • RNA polymerase catalyzes formation of phosphodiester bonds between the incorp. ribonucleotides • Topoisomerases relax supercoils in front of and behind transcription bubble

Elongation

Chain Elongation (Cont’d)

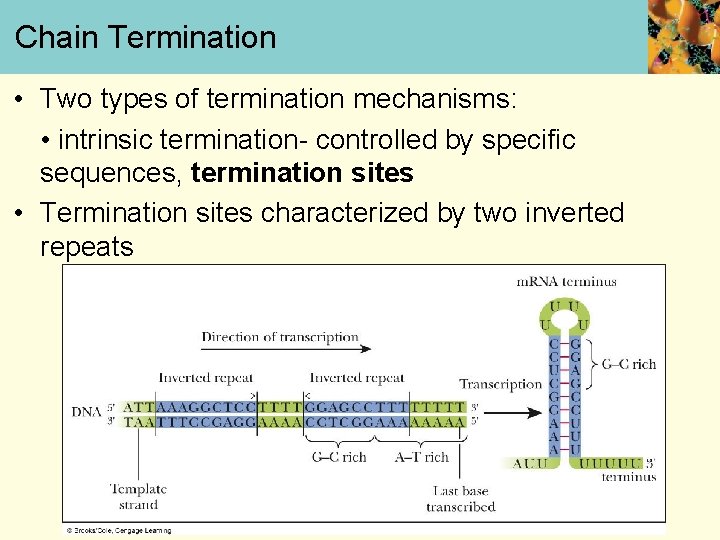
Chain Termination • Two types of termination mechanisms: • intrinsic termination- controlled by specific sequences, termination sites • Termination sites characterized by two inverted repeats

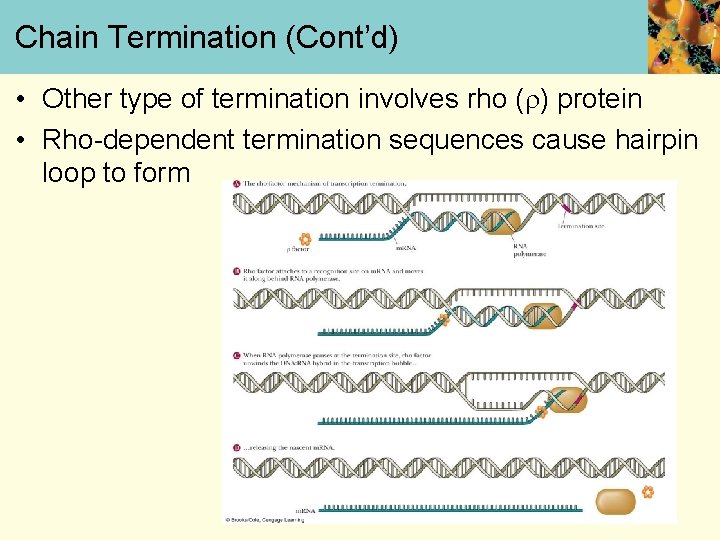
Chain Termination (Cont’d) • Other type of termination involves rho ( ) protein • Rho-dependent termination sequences cause hairpin loop to form

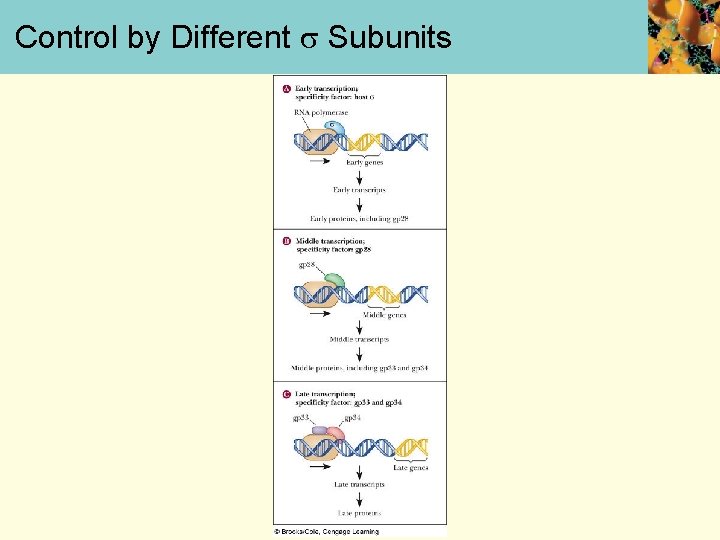
Transcription Regulation in Prokaryotes • In prokaryotes, transcription regulated by: • alternative factors • enhancers • operons • transcription attenuation • Alternative s factors • Viruses and bacteria exert control over which genes are expressed by producing different -subunits that direct the RNA polymerase to different genes.

Control by Different Subunits

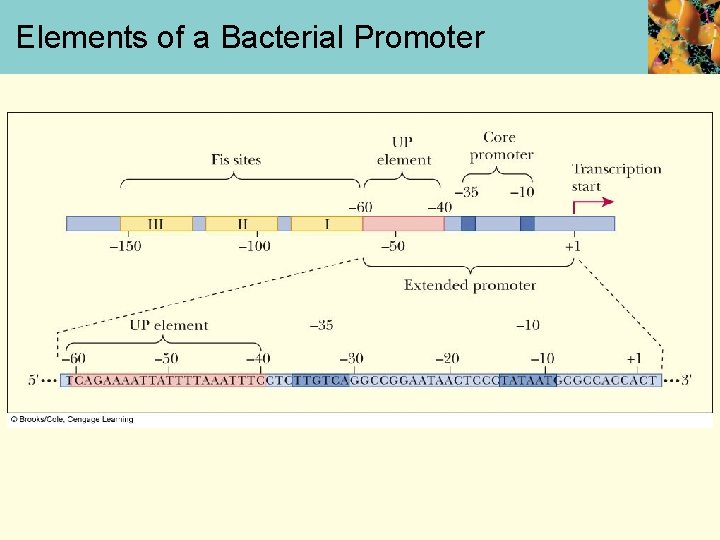
Enhancers • Certain genes include sequences upstream of extended promoter region • These genes for ribosomal production have 3 upstream sites, Fis sites • Class of DNA sequences that do this are called enhancers • Bound by proteins called transcription factors

Elements of a Bacterial Promoter

Operon • Operon: a group of operator, promoter, and structural genes that codes for proteins • the control sites, promoter, and operator genes are physically adjacent to the structural gene in the DNA • the regulatory gene can be quite far from the operon • operons are usually not transcribed all the time • -Galactosidase, an inducible protein • • coded for by a structural gene, lac. Z structural gene lac. Y codes for lactose permease structural gene lac. A codes for transacetylase expression of these three structural genes is controlled by the regulatory gene lac. I that codes for a repressor

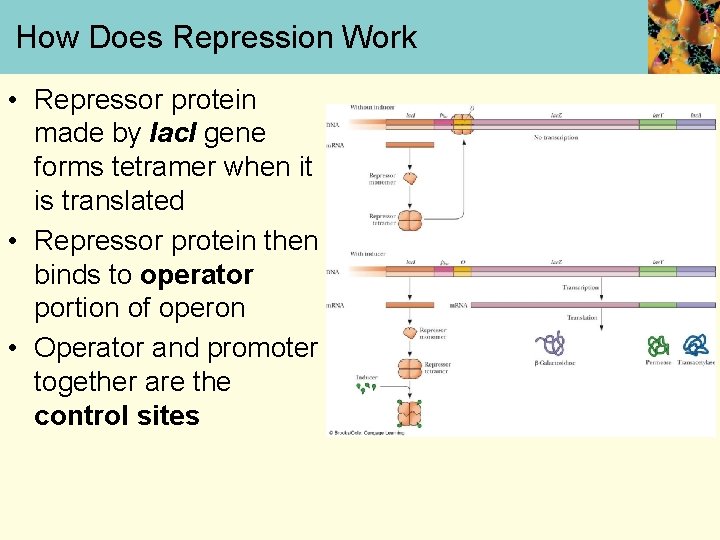
How Does Repression Work • Repressor protein made by lac. I gene forms tetramer when it is translated • Repressor protein then binds to operator portion of operon • Operator and promoter together are the control sites

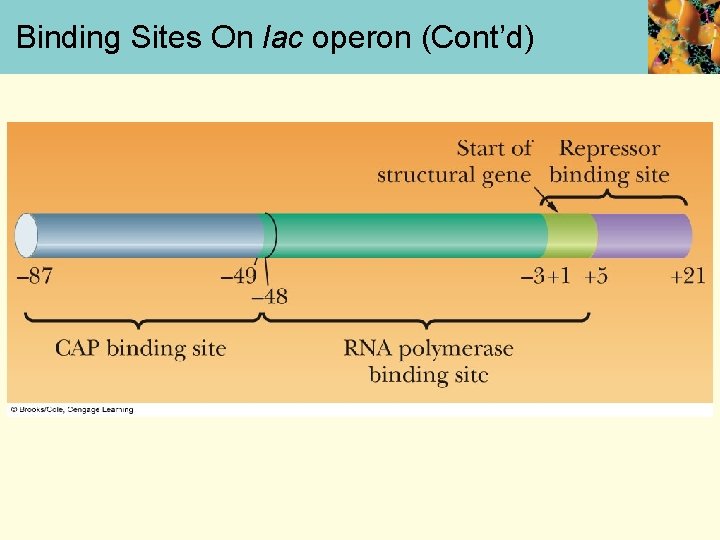
Binding Sites On the lac operon • Lac operon is induced when E. coli has lactose as the carbon source • Lac protein synthesis repressed by glucose (catabolite repression) • E. coli recognizes presence of glucose by promoter as it has 2 regions: RNA polymerase binding site, catabolite activator protein (CAP) binding site

Binding Sites On lac operon (Cont’d)

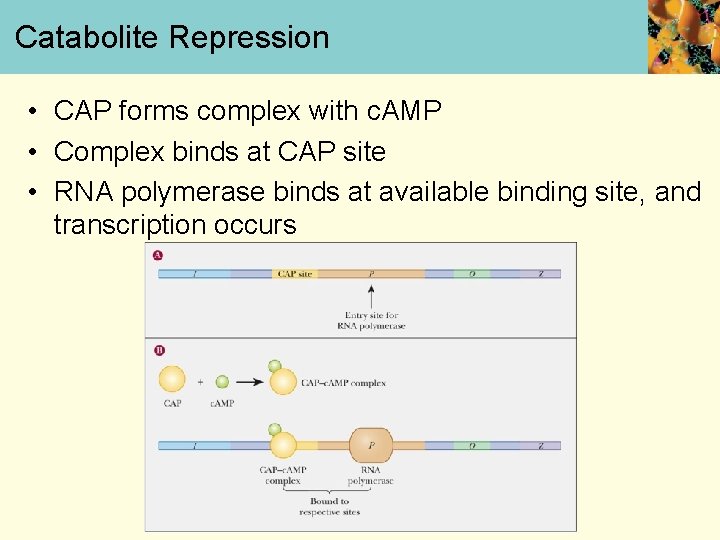
Catabolite Repression • CAP forms complex with c. AMP • Complex binds at CAP site • RNA polymerase binds at available binding site, and transcription occurs

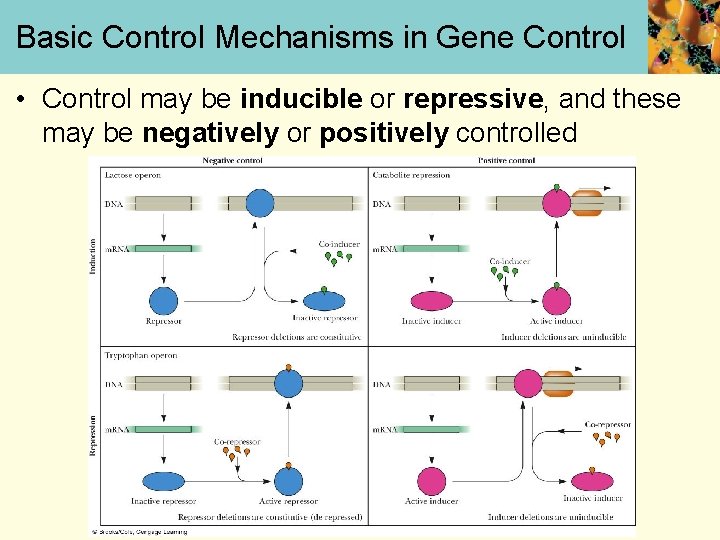
Basic Control Mechanisms in Gene Control • Control may be inducible or repressive, and these may be negatively or positively controlled

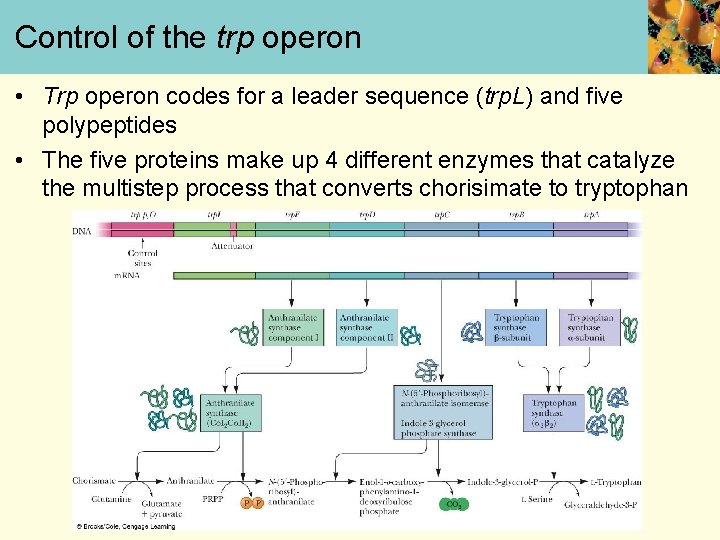
Control of the trp operon • Trp operon codes for a leader sequence (trp. L) and five polypeptides • The five proteins make up 4 different enzymes that catalyze the multistep process that converts chorisimate to tryptophan

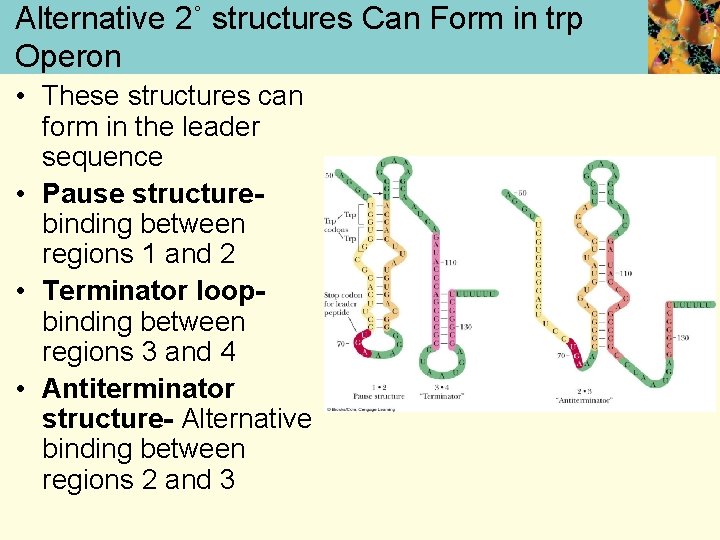
Alternative 2˚ structures Can Form in trp Operon • These structures can form in the leader sequence • Pause structurebinding between regions 1 and 2 • Terminator loopbinding between regions 3 and 4 • Antiterminator structure- Alternative binding between regions 2 and 3

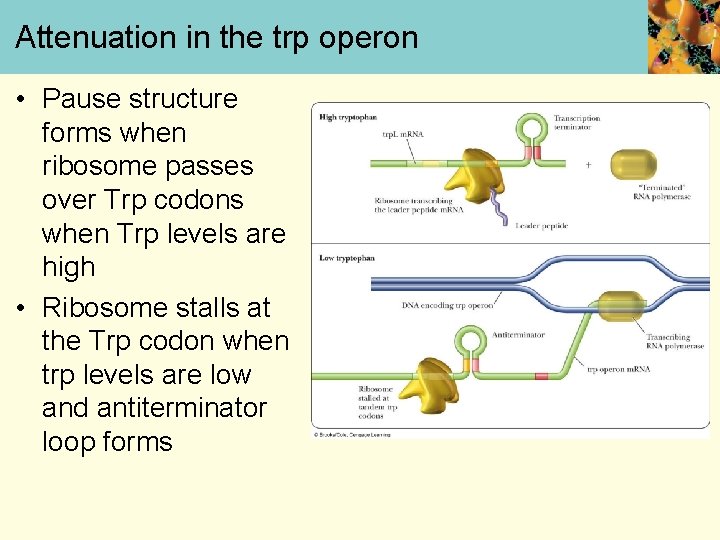
Attenuation in the trp operon • Pause structure forms when ribosome passes over Trp codons when Trp levels are high • Ribosome stalls at the Trp codon when trp levels are low and antiterminator loop forms

Transcription in Eukaryotes • Three RNA polymerases are known; each transcribes a different set of genes and recognizes a different set of promoters: • RNA Polymerase I- found in the nucleolus and synthesizes precursors of most r. RNAs • RNA Polymerase II- found in the nucleoplasm and synthesizes m. RNA precursors • RNA Polymerase III- found in the nucleoplasm and synthesizes t. RNAs, other RNA molecules involved in m. RNA processing and protein transport

RNA Polymerase II • Most studied on the polymerases • Consists of 12 subunits • RPB- RNA Polymerase B

How does Pol II Recognize the Correct DNA? • Four elements of the Pol II promoter allow for this phenomenon

Initiation of Transcription • Any protein regulator of transcription that is not itself a subunit of Pol II is a transcription factor • Initiation begins by forming the preinitiation complex • Transcription control is based here

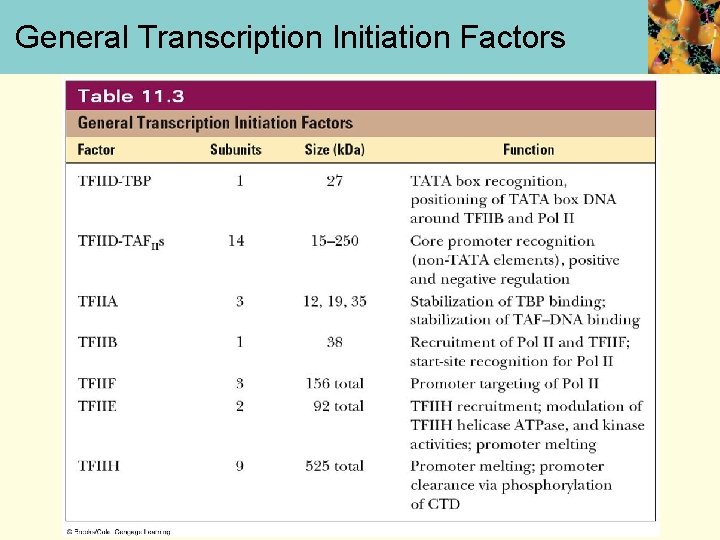
General Transcription Initiation Factors

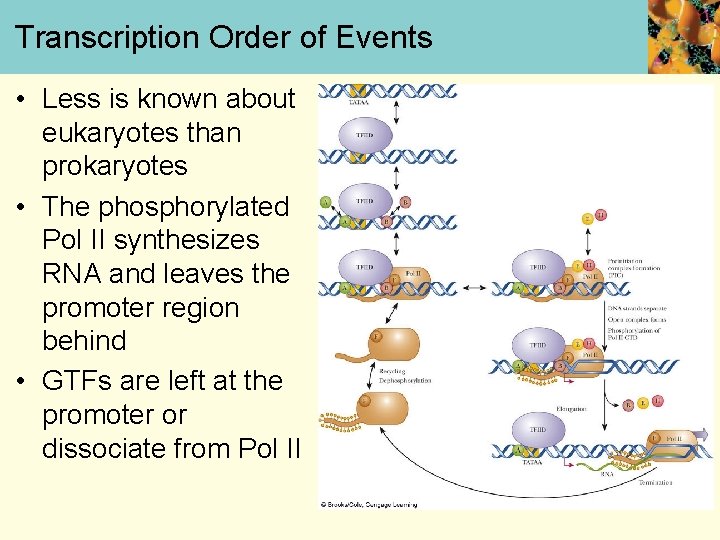
Transcription Order of Events • Less is known about eukaryotes than prokaryotes • The phosphorylated Pol II synthesizes RNA and leaves the promoter region behind • GTFs are left at the promoter or dissociate from Pol II

Elongation and Termination • Elongation is controlled by: • pause sites, where RNA Pol will hesitate • anti-termination, which proceeds past the normal termination point • positive transcription elongation factor (P-TEF) and negative transcription elongation factor (N-TEF) • Termination • begins by stopping RNA Pol; the eukaryotic consensus sequence for termination is AAUAAA

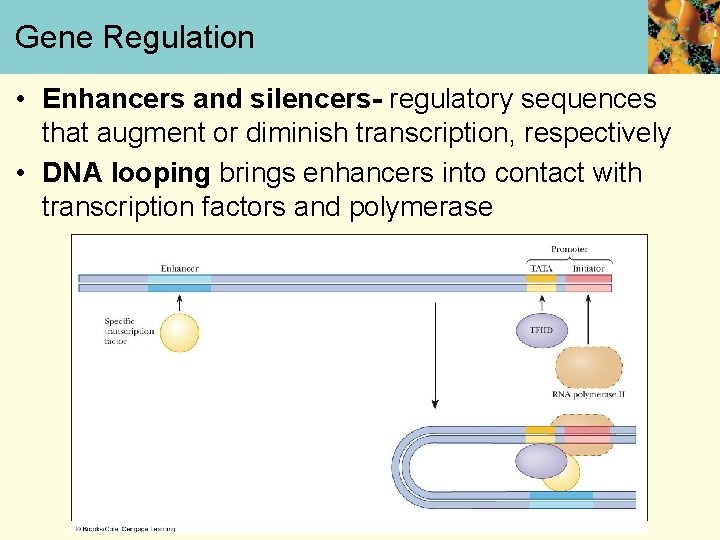
Gene Regulation • Enhancers and silencers- regulatory sequences that augment or diminish transcription, respectively • DNA looping brings enhancers into contact with transcription factors and polymerase

Eukaryotic Gene Regulation • Response elements are enhancers that respond to certain metabolic factors • heat shock element (HSE) • glucocorticoid response element (GRE) • metal response element (MRE) • cyclic-AMP response element (CRE) • Response elements all bind proteins (transcription factors) that are produced under certain cell conditions

Response Elements

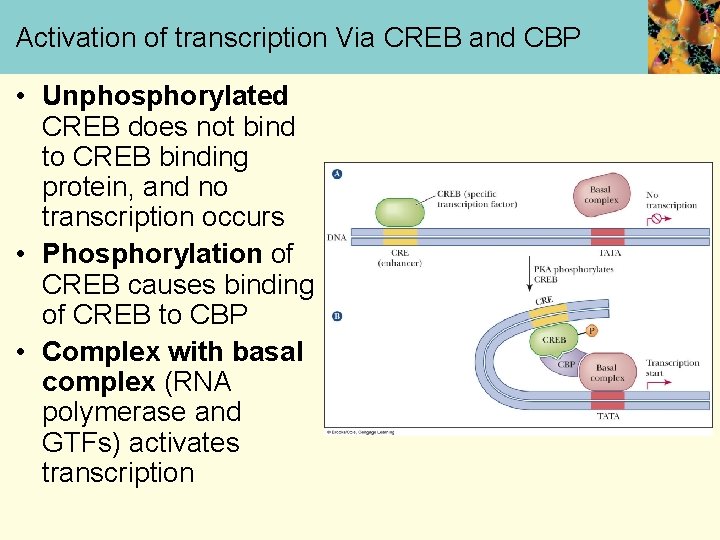
Activation of transcription Via CREB and CBP • Unphosphorylated CREB does not bind to CREB binding protein, and no transcription occurs • Phosphorylation of CREB causes binding of CREB to CBP • Complex with basal complex (RNA polymerase and GTFs) activates transcription

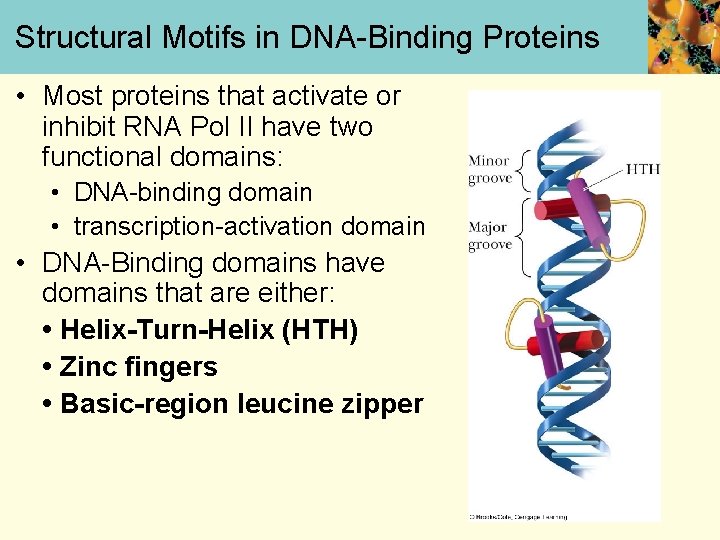
Structural Motifs in DNA-Binding Proteins • Most proteins that activate or inhibit RNA Pol II have two functional domains: • DNA-binding domain • transcription-activation domain • DNA-Binding domains have domains that are either: • Helix-Turn-Helix (HTH) • Zinc fingers • Basic-region leucine zipper

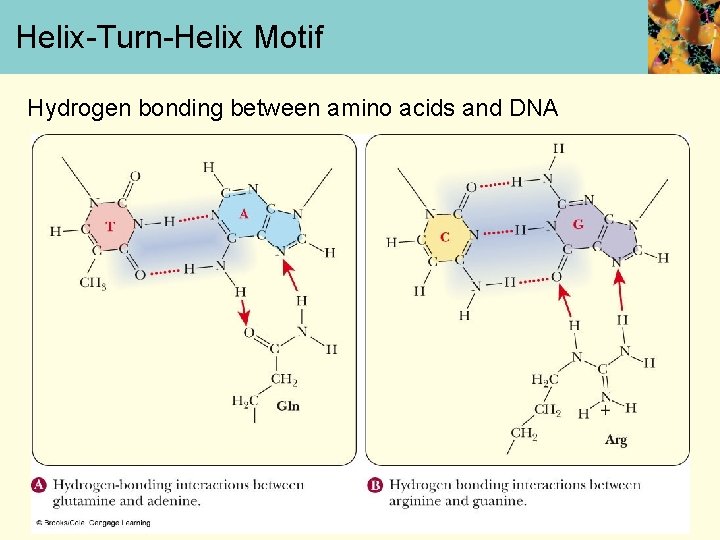
Helix-Turn-Helix Motif Hydrogen bonding between amino acids and DNA

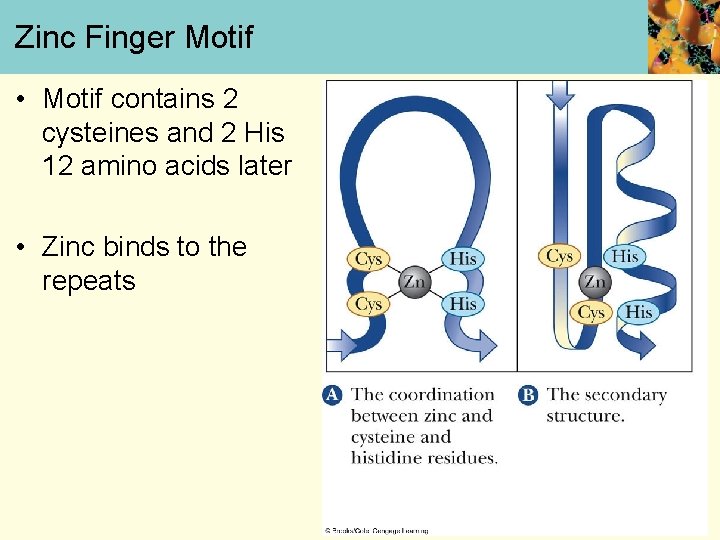
Zinc Finger Motif • Motif contains 2 cysteines and 2 His 12 amino acids later • Zinc binds to the repeats

Basic Region Leucine Zipper Motif • Many transcription factors contain this motif, such as CREB (Biochemical Connections, page 315) • Half of the protein composed of basic region of conserved Lys, Arg, and His • Half contains series of Leu • Leu line up on one side, forming hydrophobic pocket

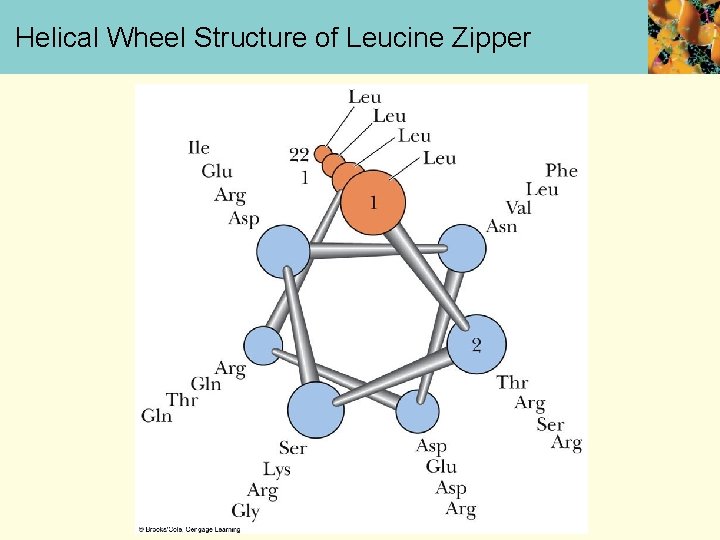
Helical Wheel Structure of Leucine Zipper

Transcription Activation Domains • acidic domains- rich in Asp and Glu. Gal 4 has domain of 49 amino acids, 11 are acidic • glutamine-rich domains- Seen in several transcription factors. Sp 1 has 2 glutamine-rich domains, one with 39 Glu in 143 amino acids • proline-rich domains- Seen in CTF-1 (an activator). It has 84 amino acid domain, of which 19 are Pro

Post Transcriptional RNA Modification • t. RNA, r. RNA, and m. RNA are all modified after transcription to give the functional form • the initial size of the RNA transcript is greater than the final size because of the leader sequences at the 5’ end and the trailer sequences at the 3’ end • the types of processing in prokaryotes can differ greatly from that in eukaryotes, especially for m. RNA • Modifications • trimming of leader and trailer sequences • addition of terminal sequences (after transcription) • modification of the structure of specific bases (particularly in t. RNA)

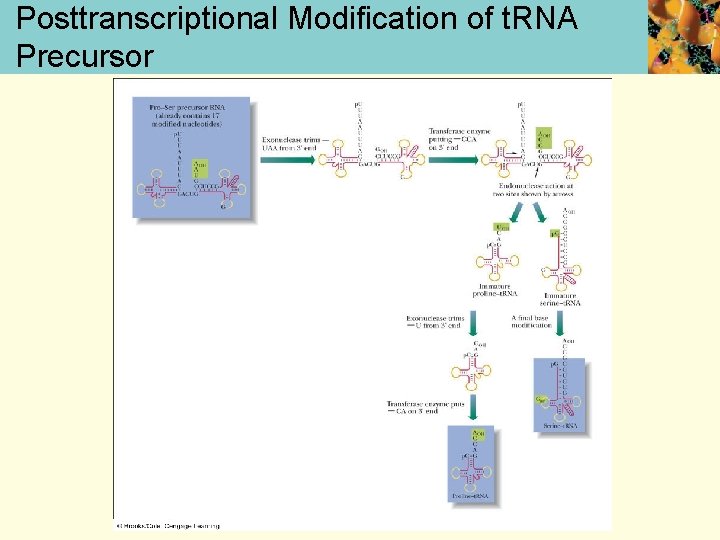
Posttranscriptional Modification of t. RNA Precursor

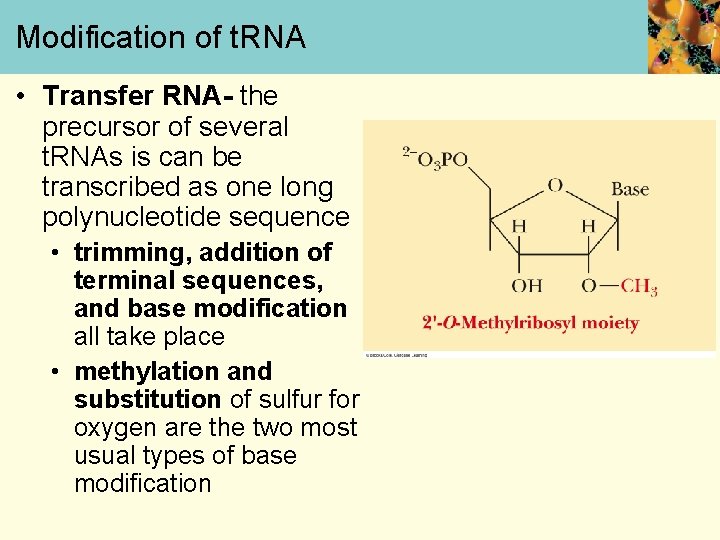
Modification of t. RNA • Transfer RNA- the precursor of several t. RNAs is can be transcribed as one long polynucleotide sequence • trimming, addition of terminal sequences, and base modification all take place • methylation and substitution of sulfur for oxygen are the two most usual types of base modification

Modification of r. RNA • Ribosomal RNA • processing of r. RNA is primarily a matter of methylation and trimming to the proper size • in prokaryotes, 3 r. RNAs in one intact ribosome • in Eukaryotes, ribosomes have 80 s, 60 s, and 40 s subunits • base modification in both prokaryotes and eukaryotes is primarily by methylation

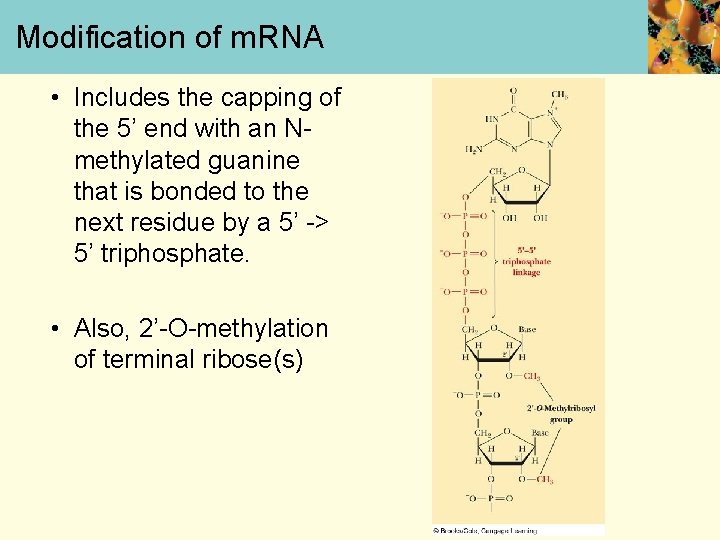
Modification of m. RNA • Includes the capping of the 5’ end with an Nmethylated guanine that is bonded to the next residue by a 5’ -> 5’ triphosphate. • Also, 2’-O-methylation of terminal ribose(s)

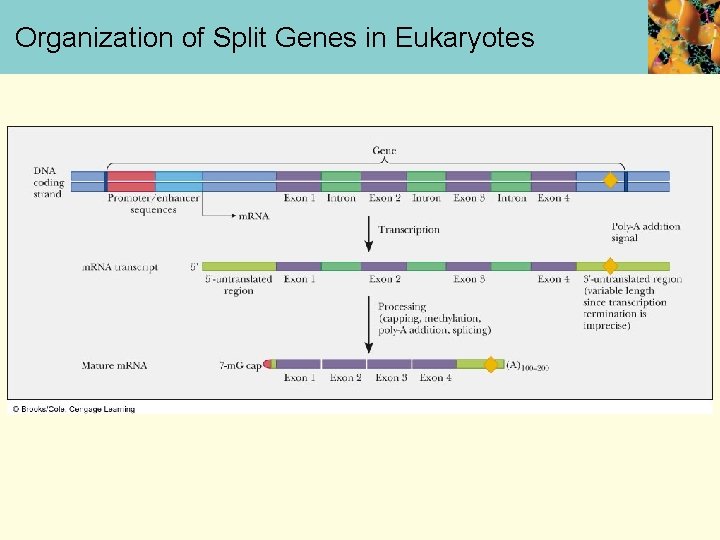
m. RNA Modification • A polyadenylate “tail” that is usually 100 -200 nucleotides long, is added to the 3’ end before the m. RNA leaves the nucleus • This tail protects the m. RNA from nucleases and phosphatases • Eukaryote genes frequently contain intervening base sequences that do not appear in the final m. RNA of that gene product • Expressed DNA sequences are called exons • Intervening DNA sequences that are not expressed are called introns • These genes are often referred to as split genes

Organization of Split Genes in Eukaryotes

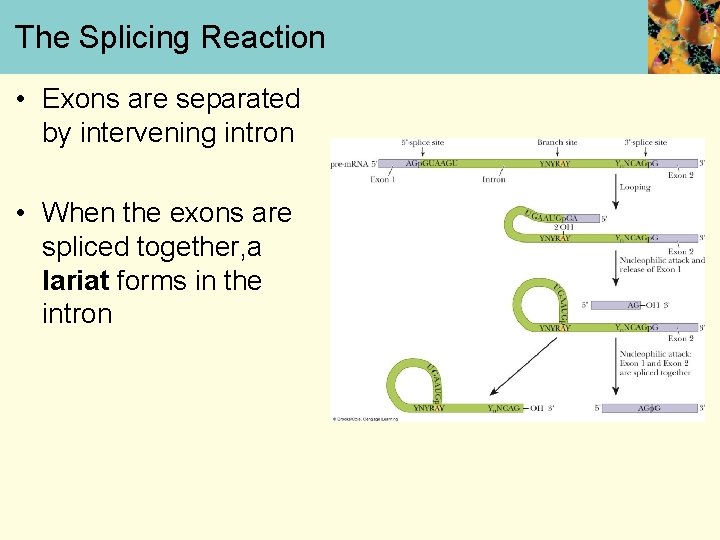
The Splicing Reaction • Exons are separated by intervening intron • When the exons are spliced together, a lariat forms in the intron

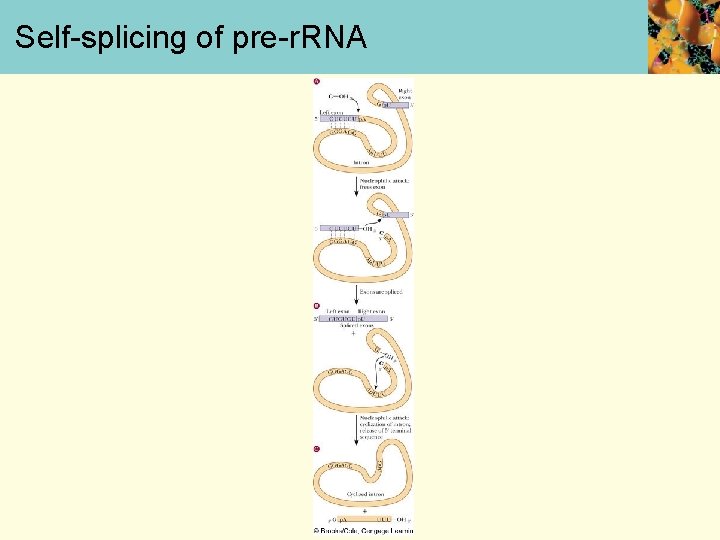
Ribozymes • The first ribozymes discovered included those that catalyze their own self-splicing • More recently, ribozymes have been discovered that are involved in protein synthesis • Group I ribozymes • require an external guanosine • example: pre-r. RNA of the protozoan Tetrahymena (next screen) • Group II ribozymes • display a lariat mechanism similar to m. RNA splicing • no requirement for an external nucleotide

Self-splicing of pre-r. RNA