TRANSCRIPTION IN PROKARYOTES PRESENTED BY FLORENCE PGT BIOLOGY

TRANSCRIPTION IN PROKARYOTES PRESENTED BY— FLORENCE PGT BIOLOGY KV NO -I NARIMEDU MADURAI
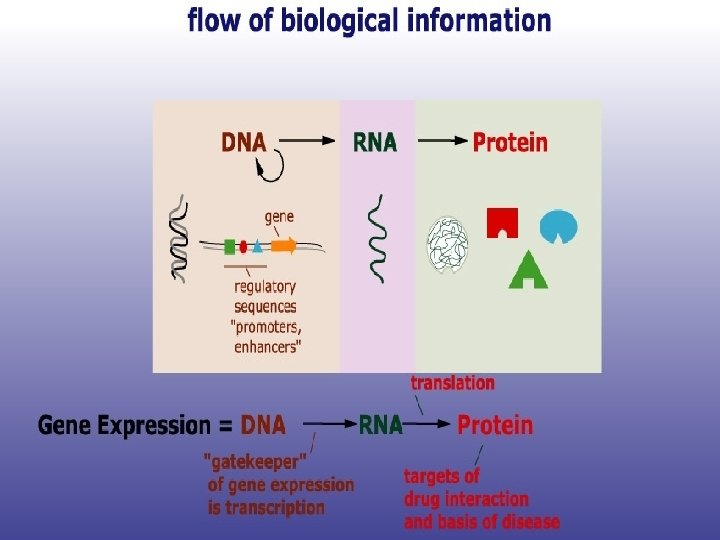
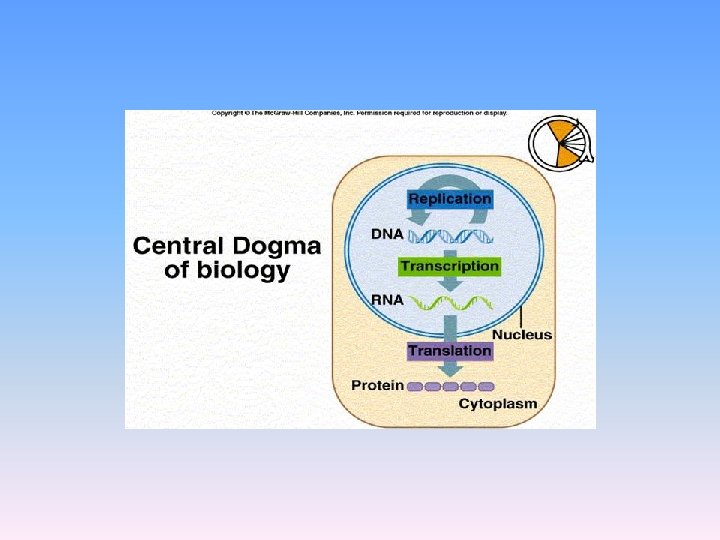

CDMB • In the words of Crick- “The central dogma of molecular biology deals with the detailed residue-by-residue transfer of sequential information. It states that such information cannot be transferred back from protein to either protein or nucleic acid. “ the word Dogma means a belief without a proof

TRANSCRIPTION • The process of copying genetic information from one strand of the DNA into RNA is termed as transcription. • The principle of complementary governs the process of transcription. • Adenine forms base pair with uracil instead of thymine. Guanine pairs with cytosine.

• Transcriptional unit helps in transcription. • It contains promoter , terminator and structural genes. • Promoter is located on DNA towards upstream of coding strand (5’ polarity) and terminator is found in down stream (3’ polarity of coding strand).

• Cistron is a segment of DNA(coding sequence), codes a polypeptide(a chain of amino acids ). • Structural genes in a transcription unit of eukaryote is called monocistronic while in prokaryotes is called polycistronic. • Due to small size of prokaryotic DNA, it contains cistron while Eukaryotic DNA contains both coding and non coding sequences. • Coding sequences are called as exons and non coding called introns in eukaryotes.

• In replication both strands of DNA are used while in transcription only one strand of DNA is used in the formation of RNA. • Template strand of DNA plays role in the formation of RNA while other strand is called as coding strand. * What will happen if both strands of DNA are used in transcription ? • Give answer

• In bacteria, three types of RNA are found. • m. RNA , r. RNA and t. RNA • All three RNAs are synthesized by single DNA dependent RNA polymerase in bacteria. • These RNAs are required for synthesis of protein in bacterial cell. • Transcription in prokaryotes occurs in the cytosol or cytoplasm.

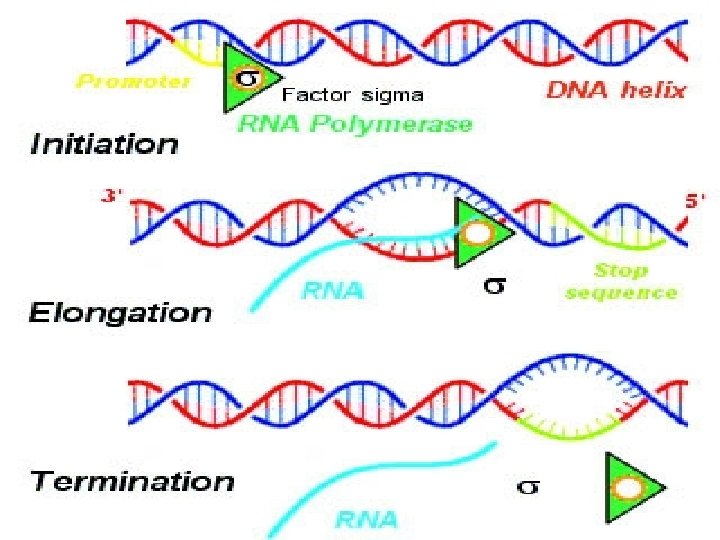
STEPS OF TRANSCRIPTION • 1 - INITIATION • 2 - ELONGATION • 3 -TERMINATION

INITIATION • RNA polymerase binds to promoter site of DNA with the help of sigma factor. • Sigma factor recognizes the promoter site. • RNA polymerase starts the formation of RNA on template strand. • It uses nucleoside triphosphates as substrate and polymerises in 5’— 3’ direction on template strand of polarity 3’— 5’ (complementary base pairing). • After initiation, sigma factor dissociates from RNA polymerase. • RNA polymerase forms small stretch of RNA.

• DNA-dependent RNA polymerase catalyses the formation of RNA on template strand. • RNA polymerase structure of prokaryotic cell

The dissociable sigma subunit gives promoter specificity to prokaryotic RNA polymerase (RNAP) sigma is called initiation factor ’ + Core enzyme ’ Holoenzyme Sigma Factor Promoters Recognized Promoter Consensus -35 Region TTGACAT Most genes -10 Region TATAAT (Region in Pribnow Box)

Transcription initiation by prokaryotic RNA polymerase Holoenzyme “sliding and scanning” Promoter -35 -10 ’ Closed complex ’ Sigma separates from the core once a few phosphodiester bonds are formed NTPs 2 Pi Core enzyme Open complex; initiation 5’ppp. A m. RNA ’

ELONGATION • RNA polymerase catalyses the formation on RNA continuously. • Synthesized stretch of RNA remains bound to RNA polymerase and template strand of DNA. • Enzyme it self opens DNA helix.

Transcriptional elongation: Movement of transcription bubble

TERMINATION • In terminator region nascent RNA and RNA polymerase dissociates from DNA. • Rho factor is known as termination factor which binds to enzyme and leads termination. • Terminator region forms a hair pin loop like structure in RNA which indicates signal of termination.
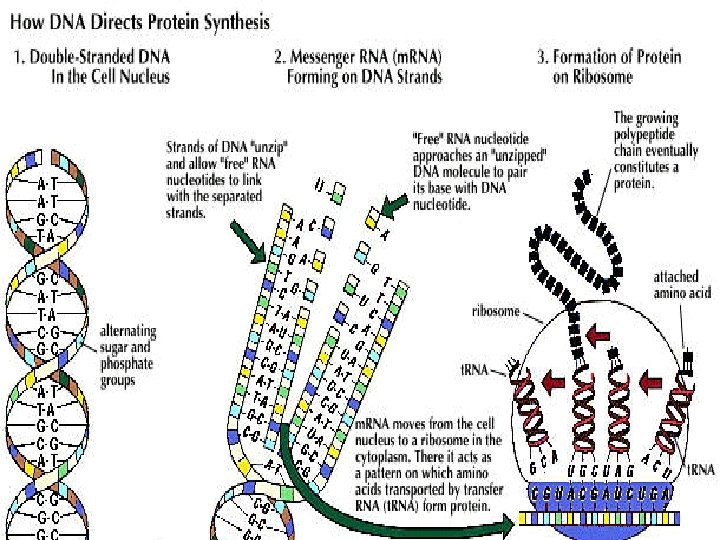
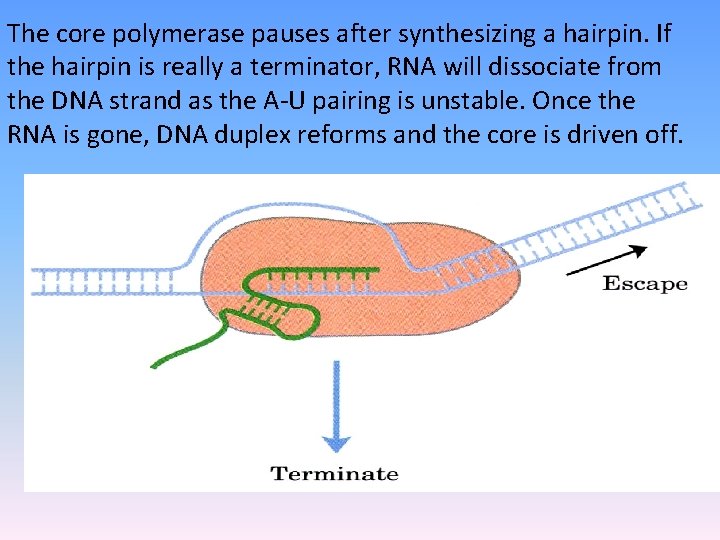
The core polymerase pauses after synthesizing a hairpin. If the hairpin is really a terminator, RNA will dissociate from the DNA strand as the A-U pairing is unstable. Once the RNA is gone, DNA duplex reforms and the core is driven off.
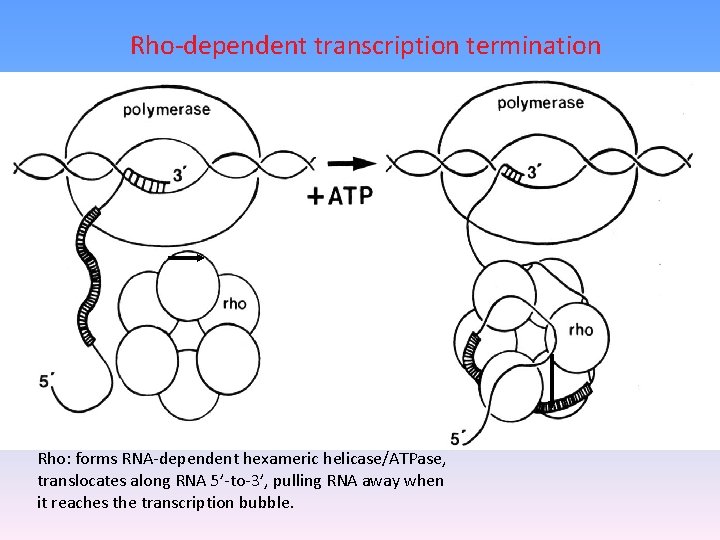
Rho-dependent transcription termination Rho: forms RNA-dependent hexameric helicase/ATPase, translocates along RNA 5’-to-3’, pulling RNA away when it reaches the transcription bubble.

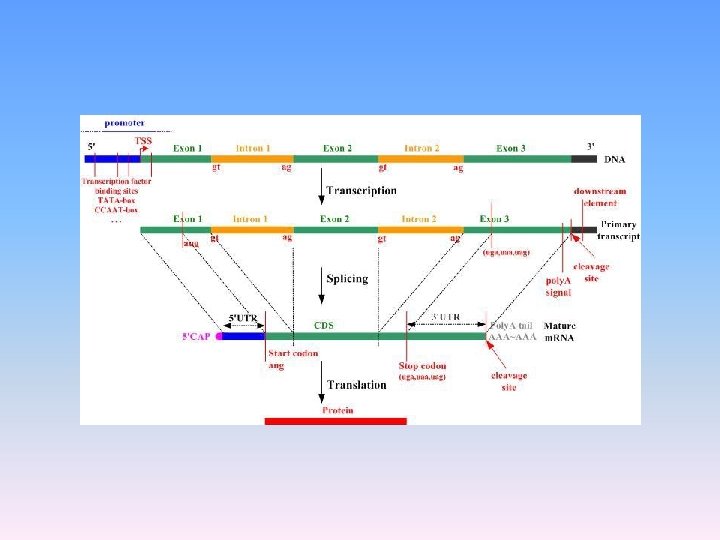
PROCESSING OF NASCENT/ hn RNA/IMMATURE RNA

. Splicing -- it is the process by which the introns are removed and Exons are joined by using ligase. 2. Capping -- Process of adding methyl guanosine triphosphate in the 5’ end. 3. Tailing -- Process of adding adenylate residues (200 -300) at the 3’end



Eukaryotes have three nuclear RNA polymerases, each with distinct roles and properties: Name &Transcribed Product RNA Polymerase I (Pol I, Pol A) nucleolus Larger ribosomal RNA (r. RNA) (28 S, 18 S, 5. 8 S) RNA Polymerase II (Pol II, Pol B) nucleus messenger RNA (m. RNA) and most small nuclear RNAs (sn. RNAs) RNA Polymerase III (Pol III, Pol C) nucleus (and possibly the nucleolus-nucleoplasm interface) transfer RNA (t. RNA) and other small RNAs (including the small 5 S r. RNA) Other organisms utilize RNA polymerase I to transcribe certain protein-coding genes in addition to r. RNAs.

- Slides: 29