Transcription DNA RNA Gene Translation at ribosome m
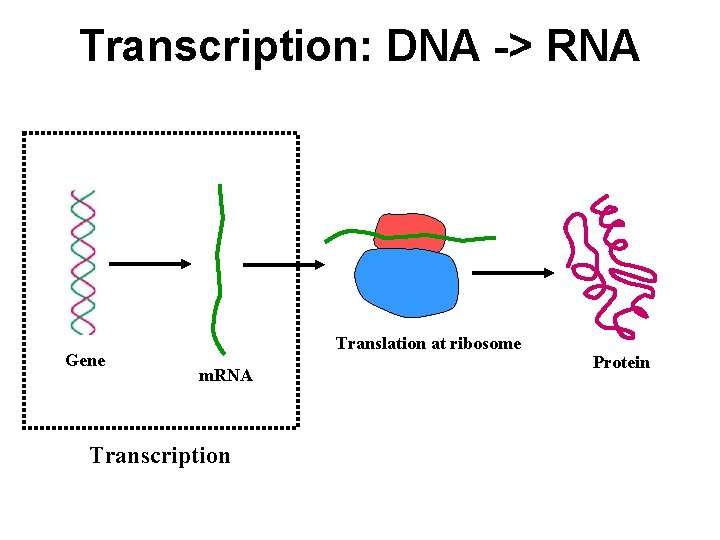
Transcription: DNA -> RNA Gene Translation at ribosome m. RNA Transcription Protein
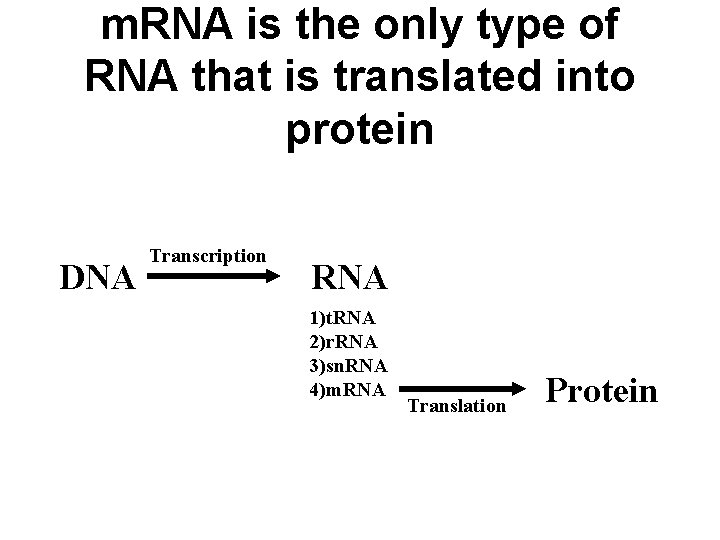
m. RNA is the only type of RNA that is translated into protein DNA Transcription RNA 1)t. RNA 2)r. RNA 3)sn. RNA 4)m. RNA Translation Protein
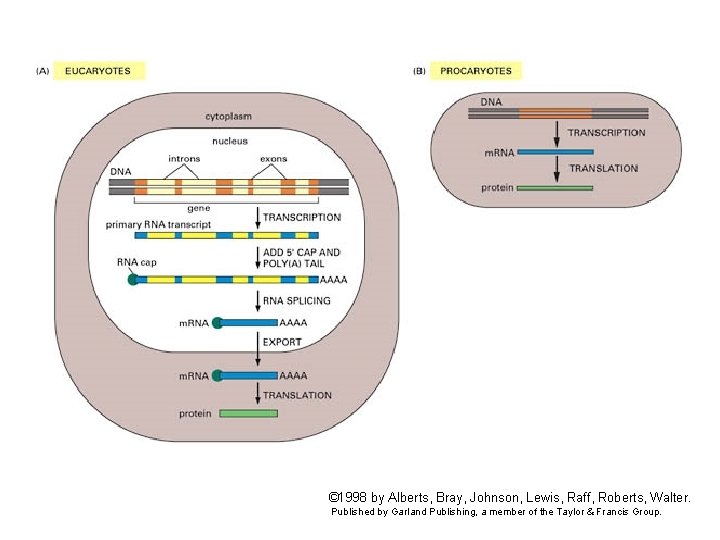
© 1998 by Alberts, Bray, Johnson, Lewis, Raff, Roberts, Walter. Published by Garland Publishing, a member of the Taylor & Francis Group.

The Population of m. RNA Molecules in a Typical Mammalian Cell -----------------------------------------------------Copies/cell#/class Total m. RNA molecules/class Abundant 12, 000 4 = 48, 000 Intermediate 300 500 = 150, 000 Scarce 15 11, 000 = 165, 000 -----------------------------------------------------This division of m. RNAs into just three discrete classes is somewhat arbitrary, and in many cells a more continuous spread in abundances is seen. However, a total of 10, 000 to 20, 000 different m. RNA species is normally observed in each cell, most species being present at a low level (5 to 15 molecules per cell). Most of the total cytoplasmic RNA is r. RNA, and only 3% to 5% is m. RNA, a ratio consistent with the presence of about 10 ribosomes per m. RNA molecule. This particular cell type contains a total of about 360, 000 m. RNA molecules in its cytoplasm. -----------------------------------------------------
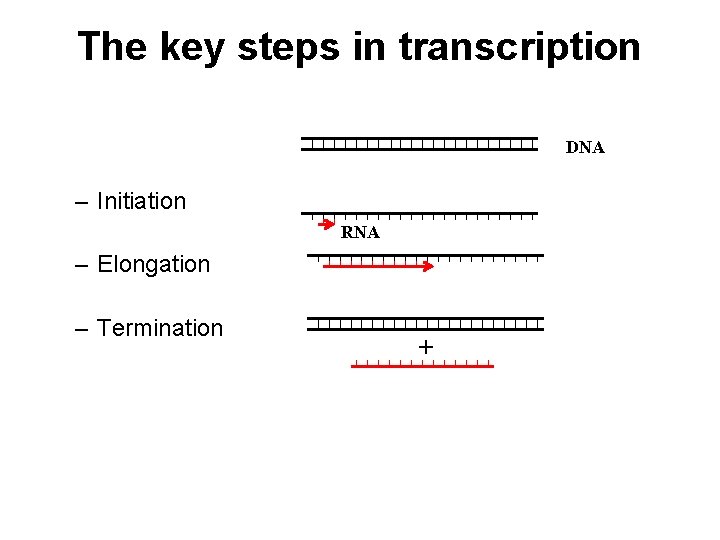
The key steps in transcription DNA – Initiation RNA – Elongation – Termination +
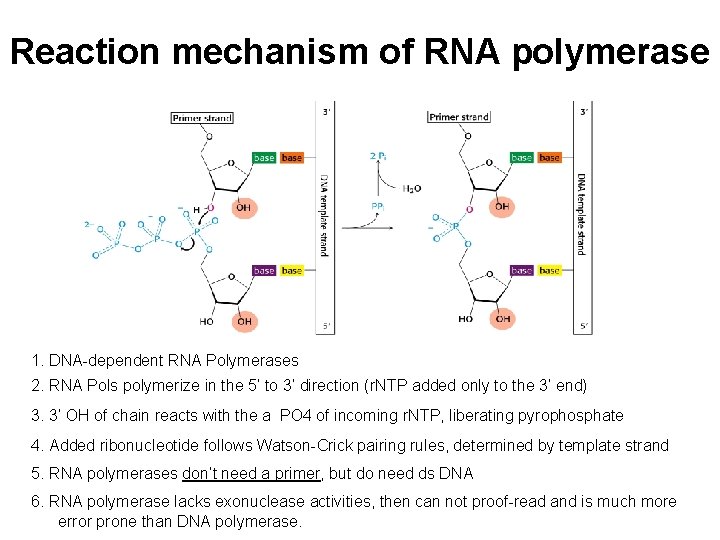
Reaction mechanism of RNA polymerase 1. DNA-dependent RNA Polymerases 2. RNA Pols polymerize in the 5’ to 3’ direction (r. NTP added only to the 3’ end) 3. 3’ OH of chain reacts with the a PO 4 of incoming r. NTP, liberating pyrophosphate 4. Added ribonucleotide follows Watson-Crick pairing rules, determined by template strand 5. RNA polymerases don’t need a primer, but do need ds DNA 6. RNA polymerase lacks exonuclease activities, then can not proof-read and is much more error prone than DNA polymerase.
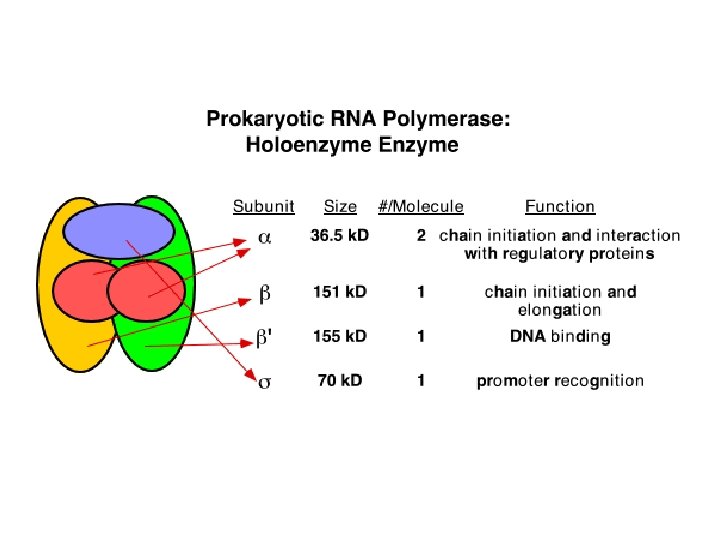
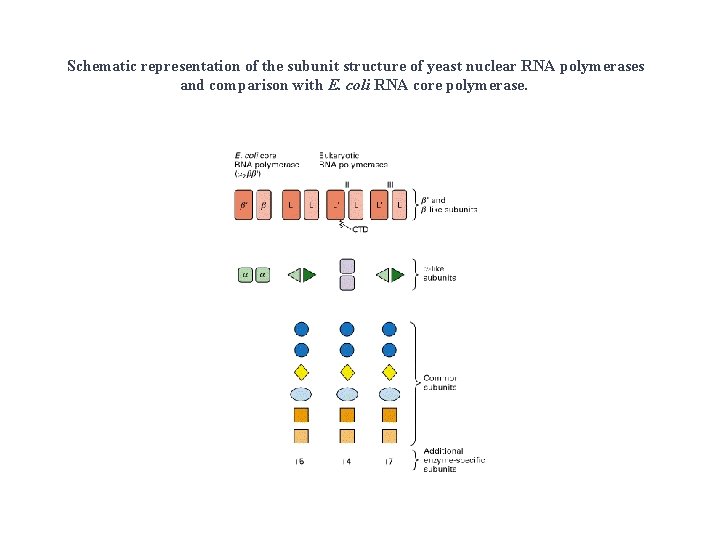
Schematic representation of the subunit structure of yeast nuclear RNA polymerases and comparison with E. coli RNA core polymerase.

Subunit structure of purified nuclear RNA polymerases (n. RNAP) • All 3 have 10 -14 subunits. • Subunits range from 10 to 220 k. Da. • All 3 have 2 very large (>125 k. D) subunits and several smaller ones. • Several of the smaller subunits (5 in yeast) are common to all 3 Pol.
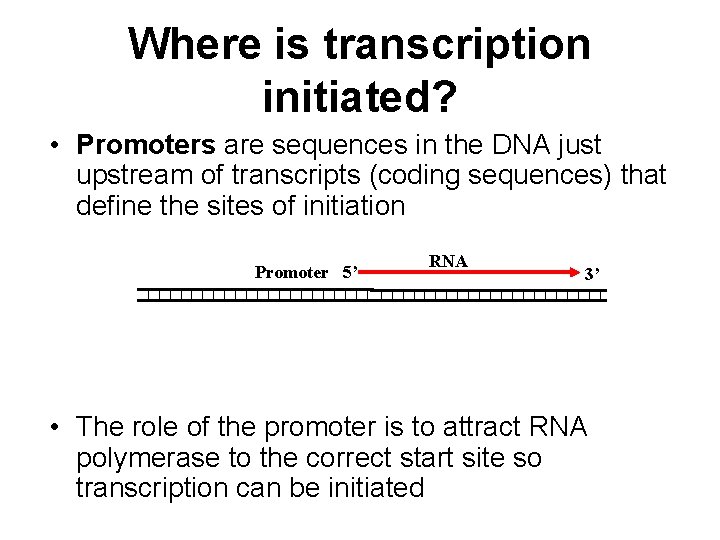
Where is transcription initiated? • Promoters are sequences in the DNA just upstream of transcripts (coding sequences) that define the sites of initiation Promoter 5’ RNA 3’ • The role of the promoter is to attract RNA polymerase to the correct start site so transcription can be initiated
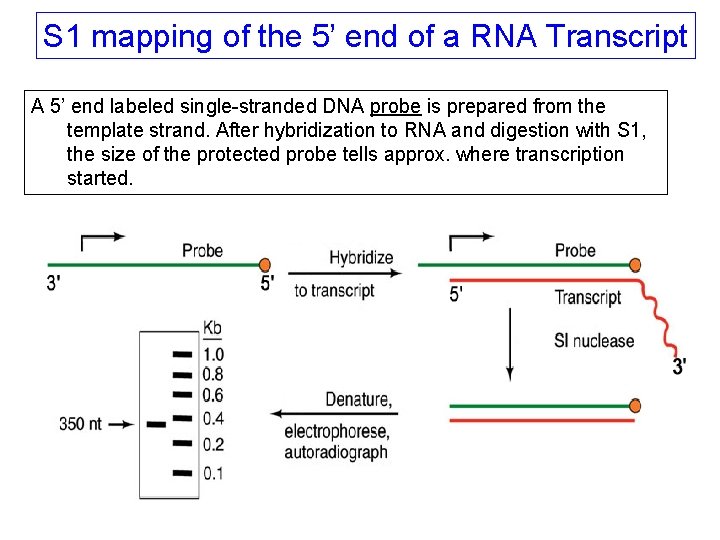
S 1 mapping of the 5’ end of a RNA Transcript A 5’ end labeled single-stranded DNA probe is prepared from the template strand. After hybridization to RNA and digestion with S 1, the size of the protected probe tells approx. where transcription started.
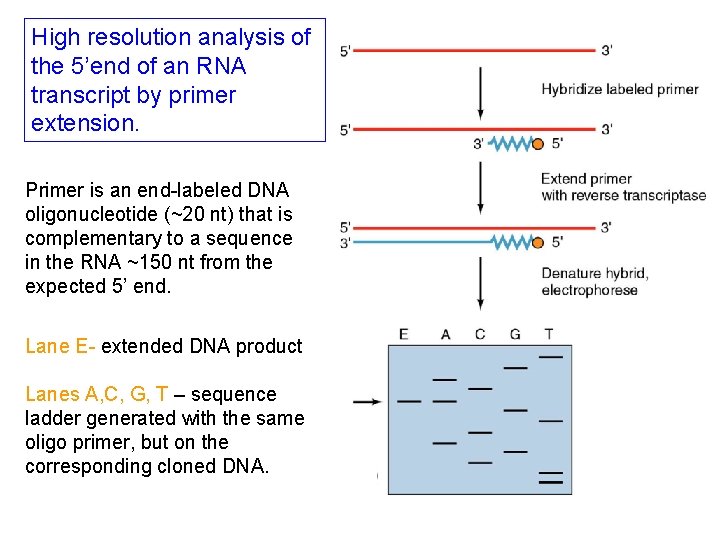
High resolution analysis of the 5’end of an RNA transcript by primer extension. Primer is an end-labeled DNA oligonucleotide (~20 nt) that is complementary to a sequence in the RNA ~150 nt from the expected 5’ end. Lane E- extended DNA product Lanes A, C, G, T – sequence ladder generated with the same oligo primer, but on the corresponding cloned DNA.

Mapping DNA-Protein interactions

Biochemical approaches to defining promoter sites • DNAse footprinting can be used to identify sites where RNA polymerase is in close contact with DNA. How we can generate this end labeled DNA?
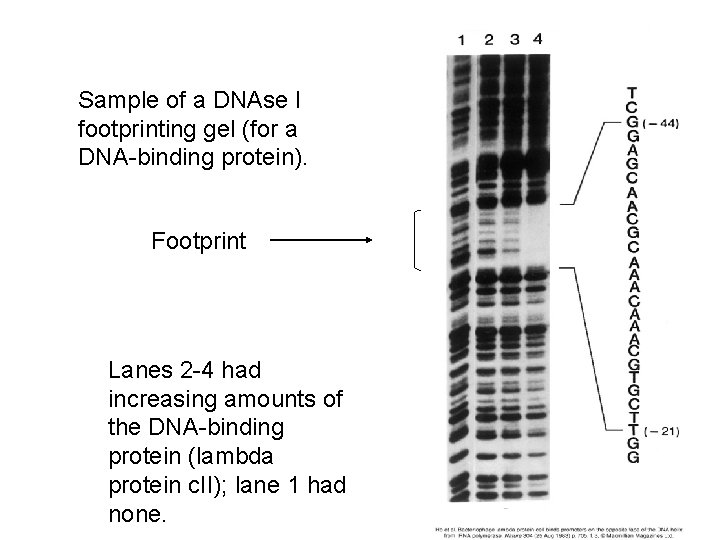
Sample of a DNAse I footprinting gel (for a DNA-binding protein). Footprint Lanes 2 -4 had increasing amounts of the DNA-binding protein (lambda protein c. II); lane 1 had none.

Dimethylsulfate (DMS) Footprinting 1. End-label DNA fragment. 2. Bind protein. 3. Treat with dimethylsulfate, which methylates purine bases. 4. Partially cleave DNA by depurinating the methylated bases (piperidine) 5. Separate DNA fragments on DNA sequencing gels.
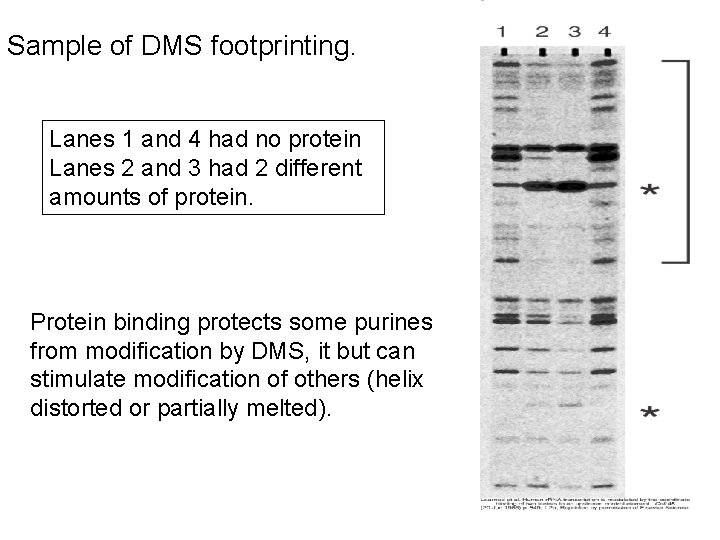
Sample of DMS footprinting. Lanes 1 and 4 had no protein Lanes 2 and 3 had 2 different amounts of protein. Protein binding protects some purines from modification by DMS, it but can stimulate modification of others (helix distorted or partially melted).

An electrophorectic mobility shift assay (EMSA) The principle of the assay is shown schematically in (A). In this example an extract of an antibody-producing cell line is mixed with a radioactive DNA fragment containing about 160 nucleotides of a regulatory DNA sequence from a gene encoding the light chain of the antibody made by the cell line. The effect of the proteins in the extract on the mobility of the DNA fragment is analyzed by polyacrylamide-gel electrophoresis followed by autoradiography. The free DNA fragments run rapidly to the bottom of the gel, while those fragments bound to proteins are retarded; the finding of six retarded bands suggests that the extract contains six different sequence-specific DNA-binding proteins (indicated as C 1 C 6) that bind to this DNA sequence. (For simplicity, any DNA fragments with more than one protein bound have been omitted from the figure. ) In (B) the extract was fractionated by a standard chromatographic technique (top), and each fraction was mixed with the radio-active DNA fragment, applied to one lane of a polyacrylamide gel, and analyzed as in (A). (B, modified from C. Scheidereit, A. Heguy, and R. G. Roeder, Cell 51: 783 -793, 1987. © Cell Press. ) Navigation
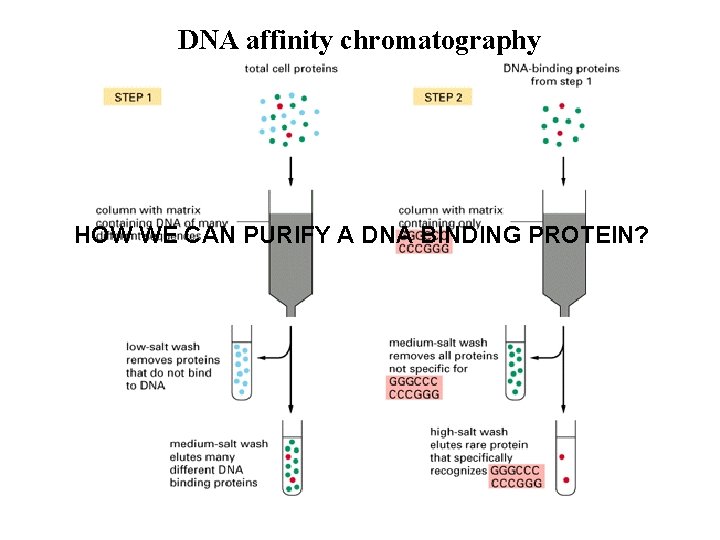
DNA affinity chromatography HOW WE CAN PURIFY A DNA BINDING PROTEIN?

Bioinformatics approaches to defining promoter sites Promoter 5’ RNA 3’ • Comparison of known start sites to identify consensus sequences:

TRASCRIPTION IN BACTERIA
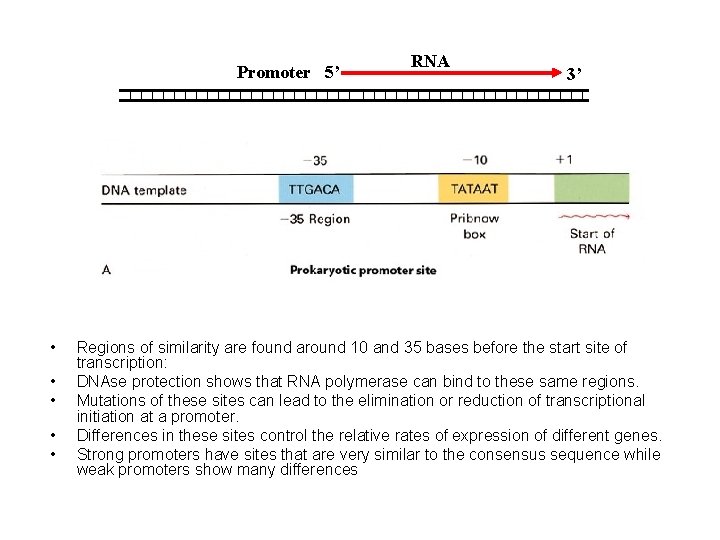
Promoter 5’ • • • RNA 3’ Regions of similarity are found around 10 and 35 bases before the start site of transcription: DNAse protection shows that RNA polymerase can bind to these same regions. Mutations of these sites can lead to the elimination or reduction of transcriptional initiation at a promoter. Differences in these sites control the relative rates of expression of different genes. Strong promoters have sites that are very similar to the consensus sequence while weak promoters show many differences

Schematic diagram of the steps in the initiation of RNA synthesis (DNA transcription) catalyzed by RNA How does s promote binding of RNA polymerase to polymerase. promoters? 70 • s 70 lowers the general affinity of RNA polymerase for DNA. The enzyme– first forms a closed complex in As a result, RNA polymerase is able which the two to move quickly along DNA strands remain fully basescanning for promoter sites. paired. In the next step the enzyme catalyzes the • s 70 can bind specifically to promoters (the opening of-10 and -35 regions). a little more than one turn of the This allows the holoenzyme to bind DNA helix –to form an open -35 complex, -10 in which tightly to promoters when they are the template DNA strand is exposed for the encountered. initiation of an RNA chain. The polymerase • RNA polymerase searches for promoter containingsites by moving along the DNA rather than the bound s subunit, however, by searching randomly throughout the cell. behaves as though it is tethered to the promoter site: it seems unable to proceed with the elongation of the RNA chain and on its own frequently synthesizes and releases short RNA chains. As indicated, the conversion to an actively elongating polymerase requires the release of initiation factors (the sigma subunit in the case of the E. coli enzyme) and generally involves the binding of other proteins that serve as elongation factors.
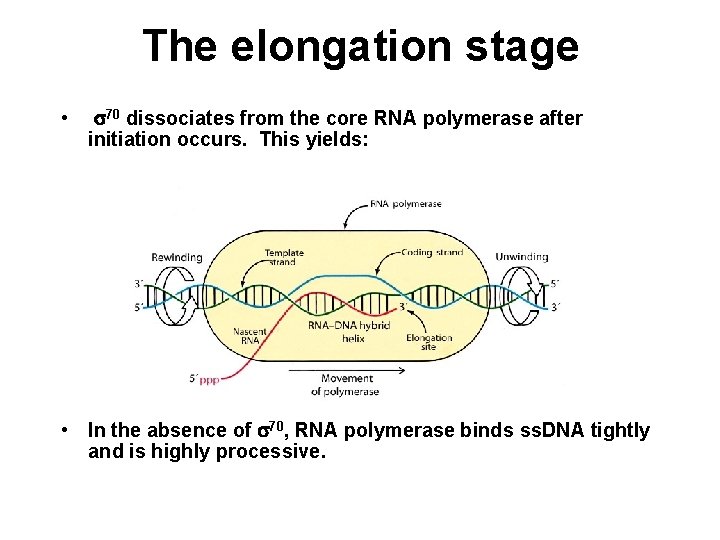
The elongation stage • s 70 dissociates from the core RNA polymerase after initiation occurs. This yields: • In the absence of s 70, RNA polymerase binds ss. DNA tightly and is highly processive.

4. Termination Two types of termination events in E. coli – Rho independent – Rho dependent
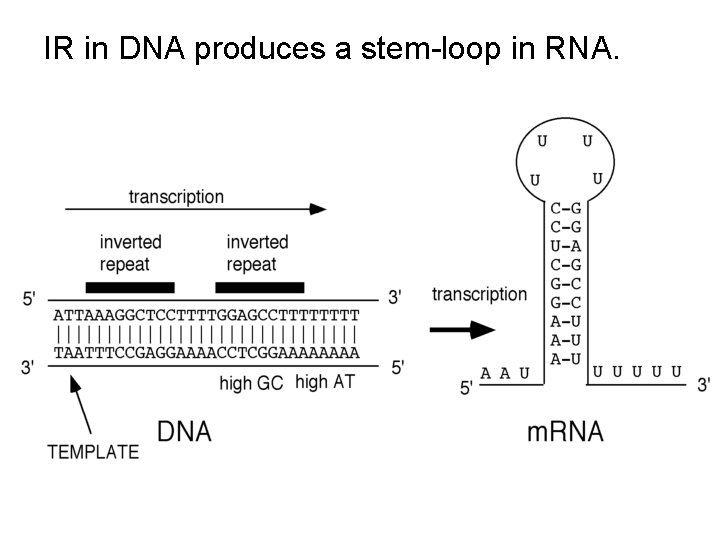
IR in DNA produces a stem-loop in RNA.
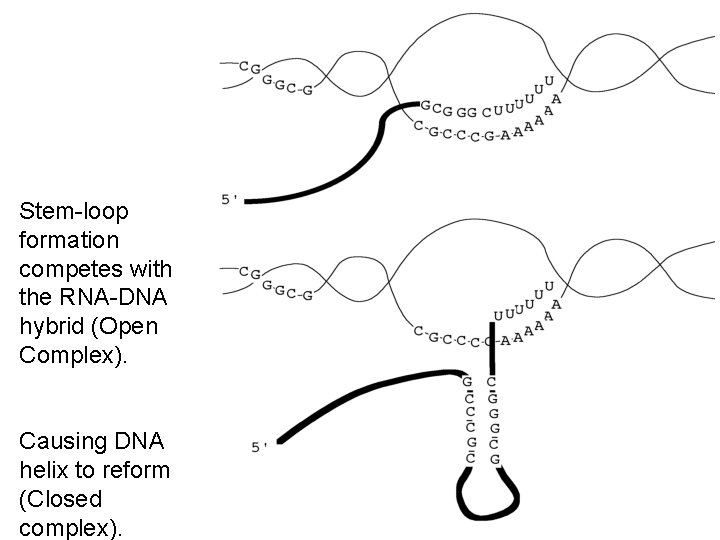
Stem-loop formation competes with the RNA-DNA hybrid (Open Complex). Causing DNA helix to reform (Closed complex).
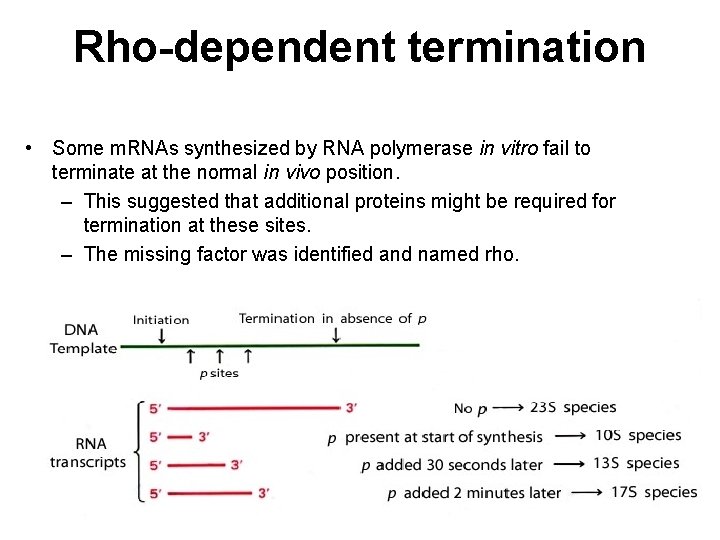
Rho-dependent termination • Some m. RNAs synthesized by RNA polymerase in vitro fail to terminate at the normal in vivo position. – This suggested that additional proteins might be required for termination at these sites. – The missing factor was identified and named rho.
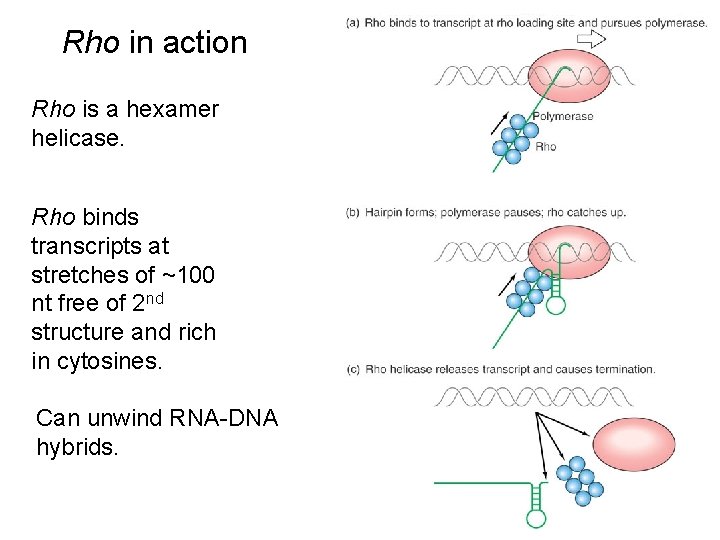
Rho in action Rho is a hexamer helicase. Rho binds transcripts at stretches of ~100 nt free of 2 nd structure and rich in cytosines. Can unwind RNA-DNA hybrids.

TRANSCRIPTION IN EUKARYOTES

Studies of RNA synthesis by isolated nuclei • RNA synthesis by isolated nuclei indicated that there were at least 2 polymerases; one of which was in the nucleolus and synthesized r. RNA – r. RNA often has a higher G-C content than other RNAs; a G-C rich RNA fraction was preferentially synthesized with low ionic strength and Mg 2+ – Another less G-C rich RNA fraction was preferentially synthesized at higher ionic strength with Mn 2+
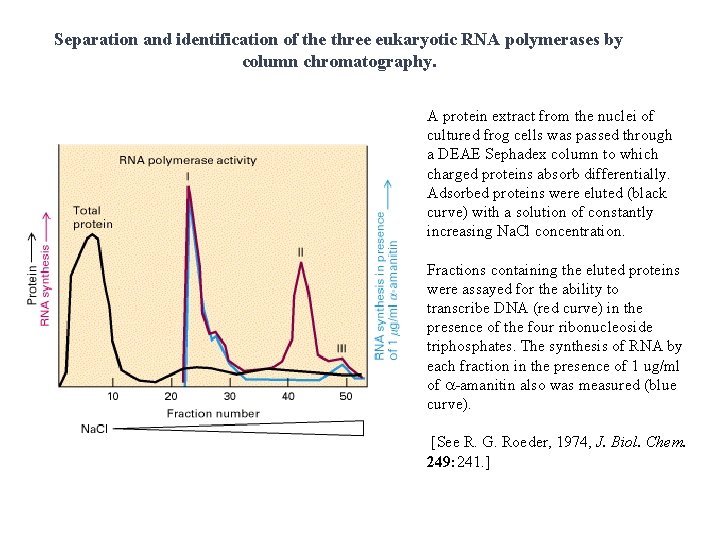
Separation and identification of the three eukaryotic RNA polymerases by column chromatography. A protein extract from the nuclei of cultured frog cells was passed through a DEAE Sephadex column to which charged proteins absorb differentially. Adsorbed proteins were eluted (black curve) with a solution of constantly increasing Na. Cl concentration. Fractions containing the eluted proteins were assayed for the ability to transcribe DNA (red curve) in the presence of the four ribonucleoside triphosphates. The synthesis of RNA by each fraction in the presence of 1 ug/ml of a-amanitin also was measured (blue curve). [See R. G. Roeder, 1974, J. Biol. Chem. 249: 241. ]

Determining roles for each polymerase • Purified polymerases don’t transcribe DNA specifically – so used nuclear fractions. • Also useful were two transcription inhibitors 1. a-aminitin – from a mushroom, inhibits Pol II, and Pol III at higher concentrations. 2. Actinomycin D - general transcription inhibitor, binds DNA and intercalates into helix, prefers G-C rich regions (like r. RNA genes).
Drugs that inhibit RNA polymerases a-amanitin: actinomycin D:
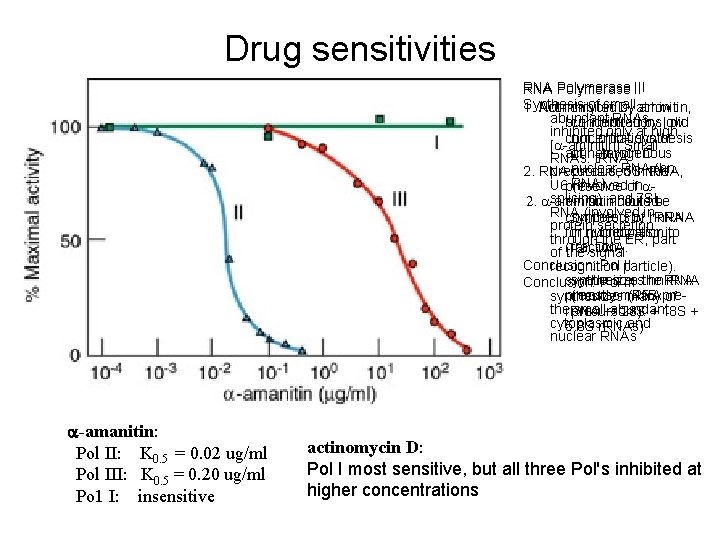
Drug sensitivities RNA Polymerase III RNA Polymerase I: RNA Polymerase II Synthesis of small 1. Not inhibited by aminitin, 1. Actinomycin D, at low abundant RNAs but inhibited by low concentrations, did inhibited only at high concentrations of not inhibit synthesis [a-aminitin] Small actinomycin D. of heterogenous RNAs: t. RNA nuclear RNA (hn 2. RNA produced in the precursors, 5 S r. RNA, RNA). U 6 (involved in presence of asplicing), and 7 SL 2. a-aminitin inhibited aminitin could be RNA (involved in competed by r. RNA synthesis of hn. RNA protein secretion for hybridization to in nucleoplasmic through the ER, part (rat) DNA. fraction. of the signal Conclusion: Pol II recognition particle). synthesizes the r. RNA synthesizes hn. RNA Conclusion: Pol III precursor (45 S pre(mostly m. RNA synthesizes many of the small abundant r. RNA 28 S + 18 S + precursors). cytoplasmic and 5. 8 S r. RNAs) nuclear RNAs a-amanitin: Pol II: K 0. 5 = 0. 02 ug/ml Pol III: K 0. 5 = 0. 20 ug/ml Po 1 I: insensitive actinomycin D: Pol I most sensitive, but all three Pol's inhibited at higher concentrations

HOW WE CAN MEASURE TRANSCRIPTIONAL ACTIVITY IN VIVO?
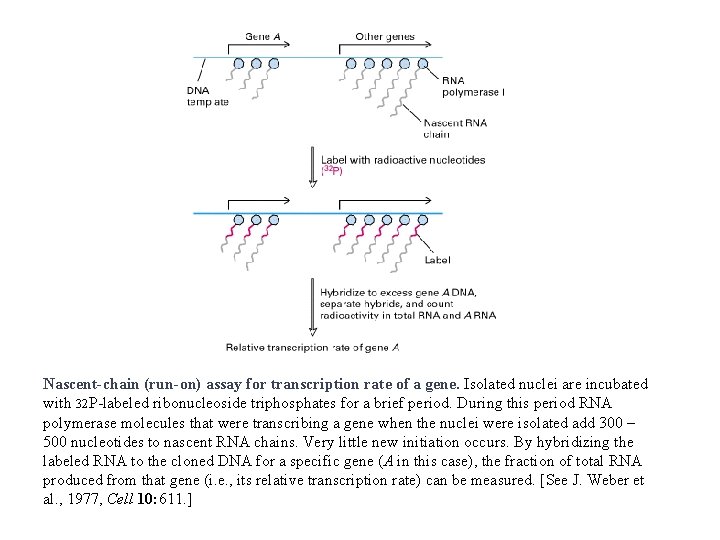
Nascent-chain (run-on) assay for transcription rate of a gene. Isolated nuclei are incubated with 32 P-labeled ribonucleoside triphosphates for a brief period. During this period RNA polymerase molecules that were transcribing a gene when the nuclei were isolated add 300 – 500 nucleotides to nascent RNA chains. Very little new initiation occurs. By hybridizing the labeled RNA to the cloned DNA for a specific gene (A in this case), the fraction of total RNA produced from that gene (i. e. , its relative transcription rate) can be measured. [See J. Weber et al. , 1977, Cell 10: 611. ]
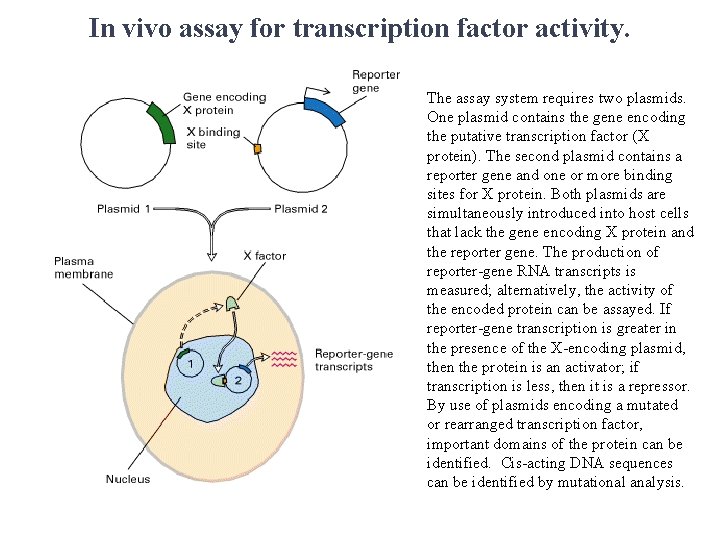
In vivo assay for transcription factor activity. The assay system requires two plasmids. One plasmid contains the gene encoding the putative transcription factor (X protein). The second plasmid contains a reporter gene and one or more binding sites for X protein. Both plasmids are simultaneously introduced into host cells that lack the gene encoding X protein and the reporter gene. The production of reporter-gene RNA transcripts is measured; alternatively, the activity of the encoded protein can be assayed. If reporter-gene transcription is greater in the presence of the X-encoding plasmid, then the protein is an activator; if transcription is less, then it is a repressor. By use of plasmids encoding a mutated or rearranged transcription factor, important domains of the protein can be identified. Cis-acting DNA sequences can be identified by mutational analysis.
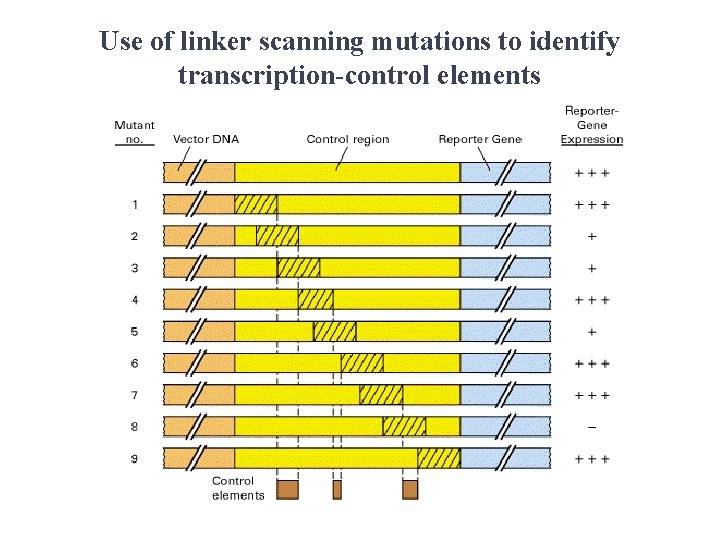
Use of linker scanning mutations to identify transcription-control elements
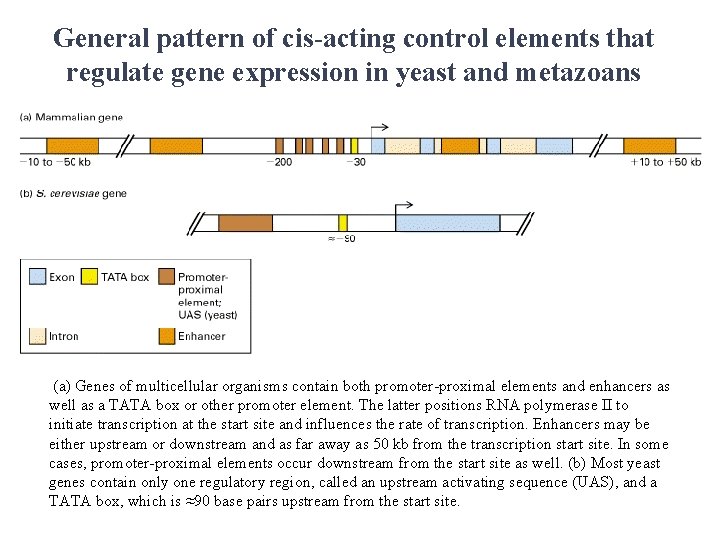
General pattern of cis-acting control elements that regulate gene expression in yeast and metazoans (a) Genes of multicellular organisms contain both promoter-proximal elements and enhancers as well as a TATA box or other promoter element. The latter positions RNA polymerase II to initiate transcription at the start site and influences the rate of transcription. Enhancers may be either upstream or downstream and as far away as 50 kb from the transcription start site. In some cases, promoter-proximal elements occur downstream from the start site as well. (b) Most yeast genes contain only one regulatory region, called an upstream activating sequence (UAS), and a TATA box, which is ≈90 base pairs upstream from the start site.
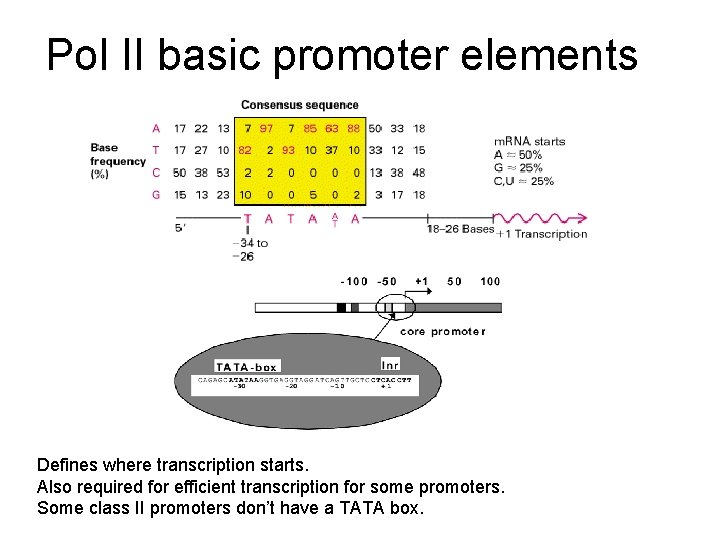
Pol II basic promoter elements Defines where transcription starts. Also required for efficient transcription for some promoters. Some class II promoters don’t have a TATA box.
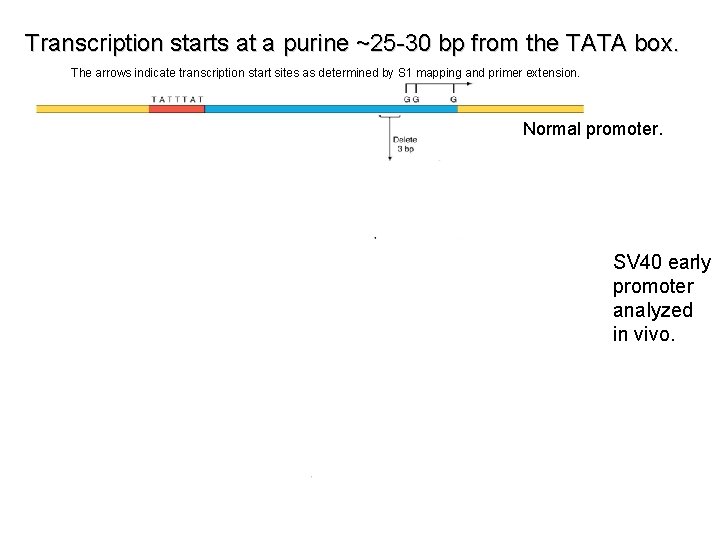
Transcription starts at a purine ~25 -30 bp from the TATA box. The arrows indicate transcription start sites as determined by S 1 mapping and primer extension. Normal promoter. SV 40 early promoter analyzed in vivo.
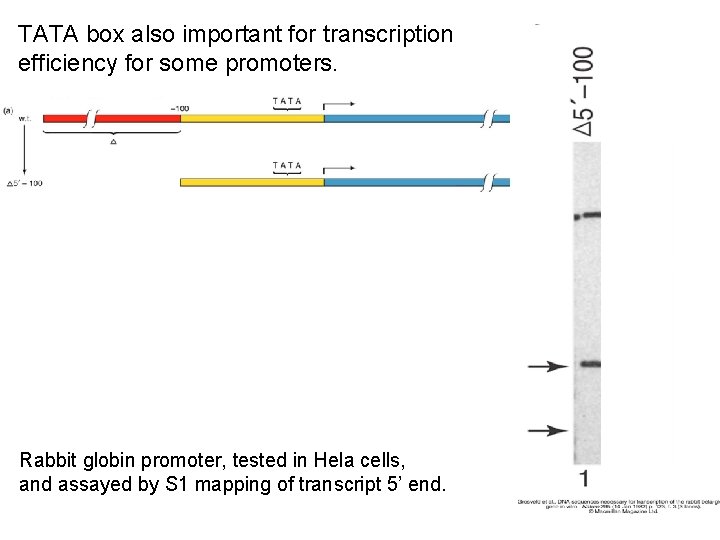
TATA box also important for transcription efficiency for some promoters. Rabbit globin promoter, tested in Hela cells, and assayed by S 1 mapping of transcript 5’ end.
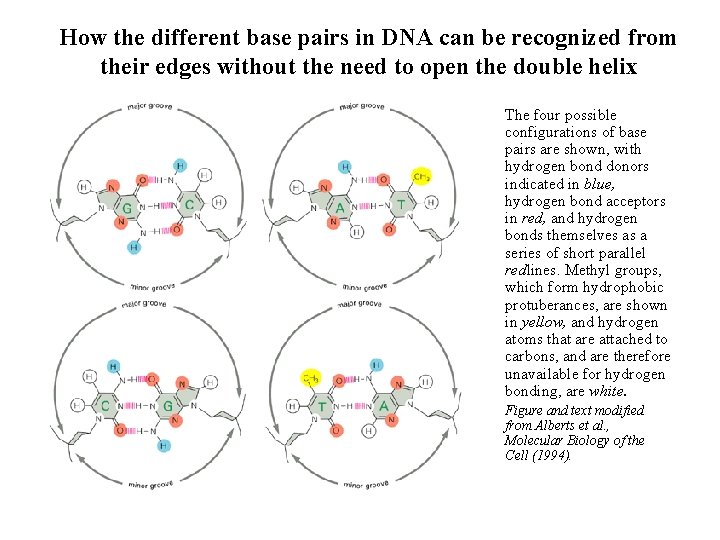
How the different base pairs in DNA can be recognized from their edges without the need to open the double helix The four possible configurations of base pairs are shown, with hydrogen bond donors indicated in blue, hydrogen bond acceptors in red, and hydrogen bonds themselves as a series of short parallel redlines. Methyl groups, which form hydrophobic protuberances, are shown in yellow, and hydrogen atoms that are attached to carbons, and are therefore unavailable for hydrogen bonding, are white. Figure and text modified from Alberts et al. , Molecular Biology of the Cell (1994).
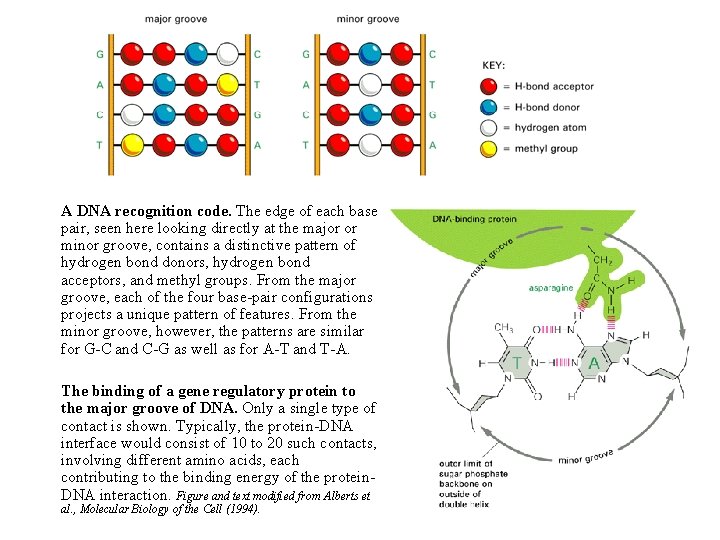
A DNA recognition code. The edge of each base pair, seen here looking directly at the major or minor groove, contains a distinctive pattern of hydrogen bond donors, hydrogen bond acceptors, and methyl groups. From the major groove, each of the four base-pair configurations projects a unique pattern of features. From the minor groove, however, the patterns are similar for G-C and C-G as well as for A-T and T-A. The binding of a gene regulatory protein to the major groove of DNA. Only a single type of contact is shown. Typically, the protein-DNA interface would consist of 10 to 20 such contacts, involving different amino acids, each contributing to the binding energy of the protein. DNA interaction. Figure and text modified from Alberts et al. , Molecular Biology of the Cell (1994).
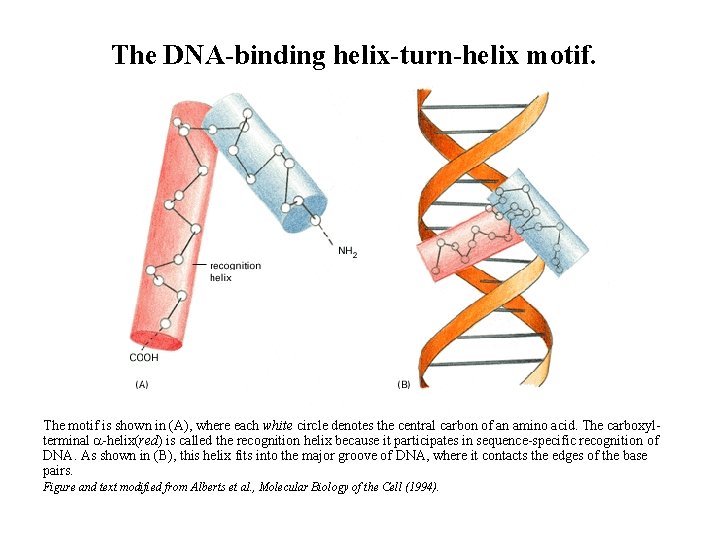
The DNA-binding helix-turn-helix motif. The motif is shown in (A), where each white circle denotes the central carbon of an amino acid. The carboxylterminal a-helix(red) is called the recognition helix because it participates in sequence-specific recognition of DNA. As shown in (B), this helix fits into the major groove of DNA, where it contacts the edges of the base pairs. Figure and text modified from Alberts et al. , Molecular Biology of the Cell (1994).
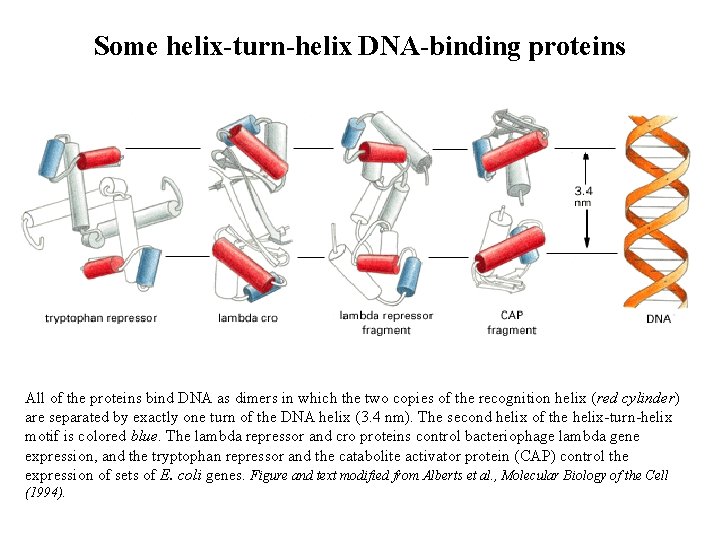
Some helix-turn-helix DNA-binding proteins All of the proteins bind DNA as dimers in which the two copies of the recognition helix (red cylinder) are separated by exactly one turn of the DNA helix (3. 4 nm). The second helix of the helix-turn-helix motif is colored blue. The lambda repressor and cro proteins control bacteriophage lambda gene expression, and the tryptophan repressor and the catabolite activator protein (CAP) control the expression of sets of E. coli genes. Figure and text modified from Alberts et al. , Molecular Biology of the Cell (1994).
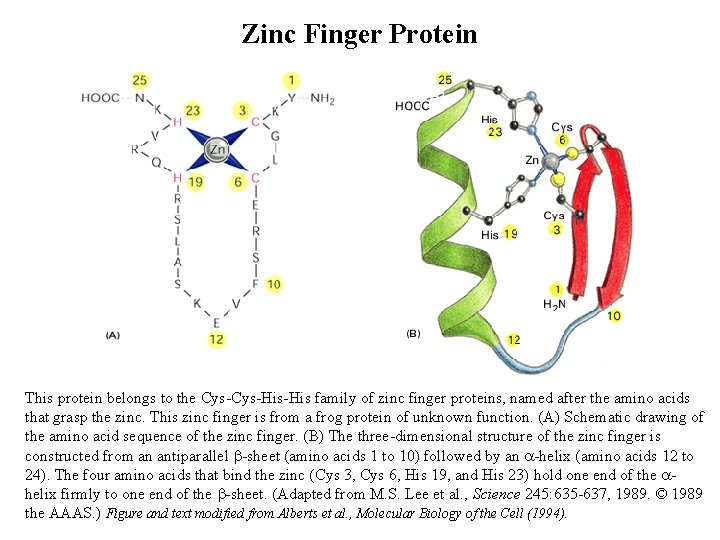
Zinc Finger Protein This protein belongs to the Cys-His-His family of zinc finger proteins, named after the amino acids that grasp the zinc. This zinc finger is from a frog protein of unknown function. (A) Schematic drawing of the amino acid sequence of the zinc finger. (B) The three-dimensional structure of the zinc finger is constructed from an antiparallel b-sheet (amino acids 1 to 10) followed by an a-helix (amino acids 12 to 24). The four amino acids that bind the zinc (Cys 3, Cys 6, His 19, and His 23) hold one end of the ahelix firmly to one end of the b-sheet. (Adapted from M. S. Lee et al. , Science 245: 635 -637, 1989. © 1989 the AAAS. ) Figure and text modified from Alberts et al. , Molecular Biology of the Cell (1994).
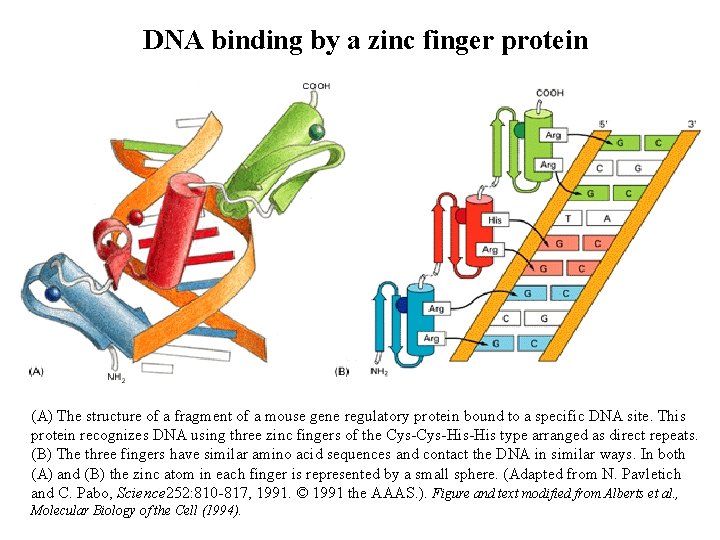
DNA binding by a zinc finger protein (A) The structure of a fragment of a mouse gene regulatory protein bound to a specific DNA site. This protein recognizes DNA using three zinc fingers of the Cys-His-His type arranged as direct repeats. (B) The three fingers have similar amino acid sequences and contact the DNA in similar ways. In both (A) and (B) the zinc atom in each finger is represented by a small sphere. (Adapted from N. Pavletich and C. Pabo, Science 252: 810 -817, 1991. © 1991 the AAAS. ). Figure and text modified from Alberts et al. , Molecular Biology of the Cell (1994).

Required reading: Blau et al. , Mol Cell Biol 1996, 16 (5): 204 Three functional Classes of Transcriptional Activation Domains
- Slides: 52