Towards understanding the SouthAsian maternal lineages tree Outline




- Slides: 22

Towards understanding the South-Asian maternal lineages tree

Outline of the presentation • Emerging topology of the Indian M and R lineages • The spread of East-Eurasian, West-Eurasian and Indian specific mt. DNA lineages in South- and Southeast-Asia • mt. DNA haplotype sharing within India and beyond

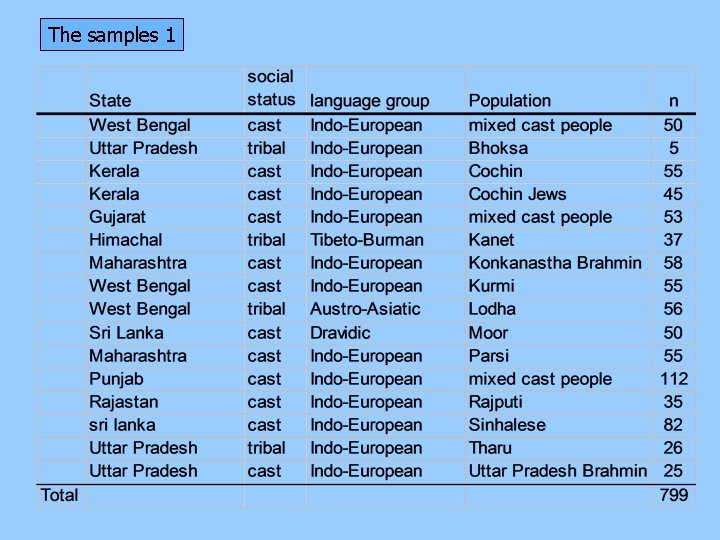
The samples 1

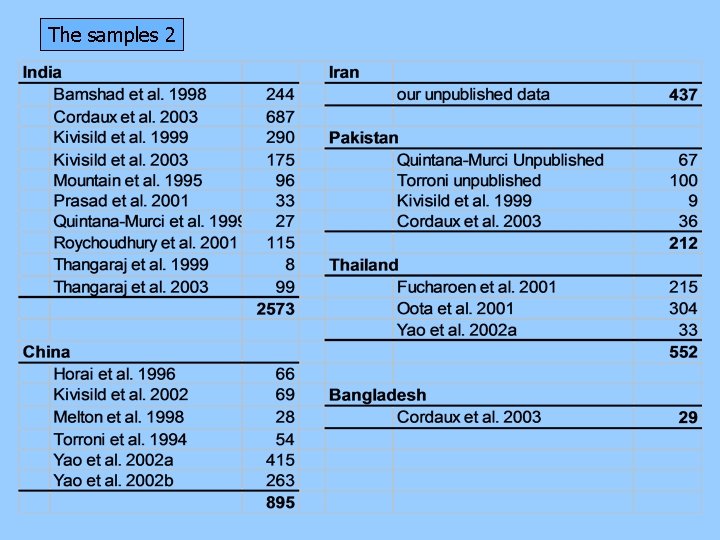
The samples 2

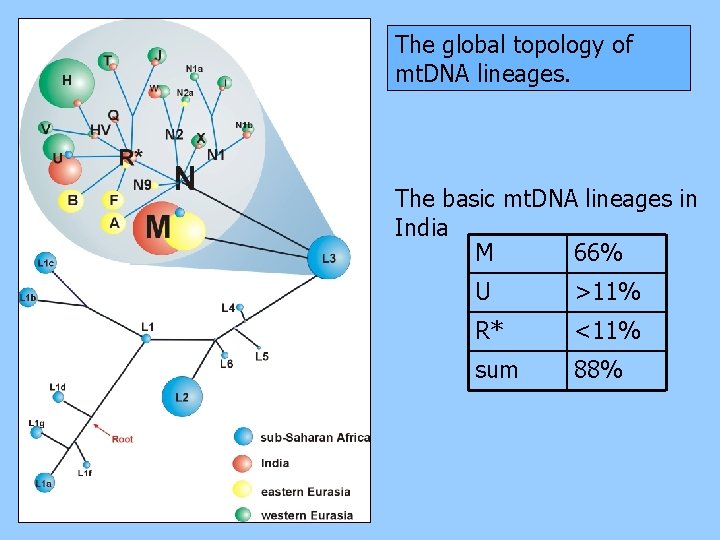
The global topology of mt. DNA lineages. The basic mt. DNA lineages in India M 66% U >11% R* <11% sum 88%

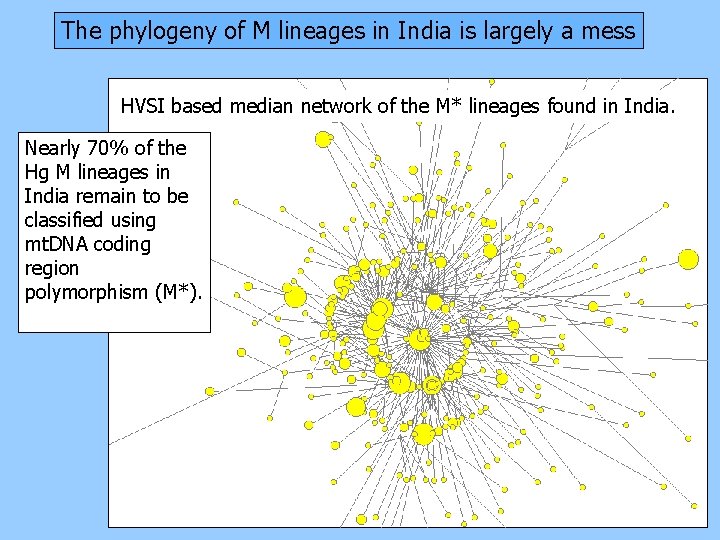
The phylogeny of M lineages in India is largely a mess HVSI based median network of the M* lineages found in India. Nearly 70% of the Hg M lineages in India remain to be classified using mt. DNA coding region polymorphism (M*).

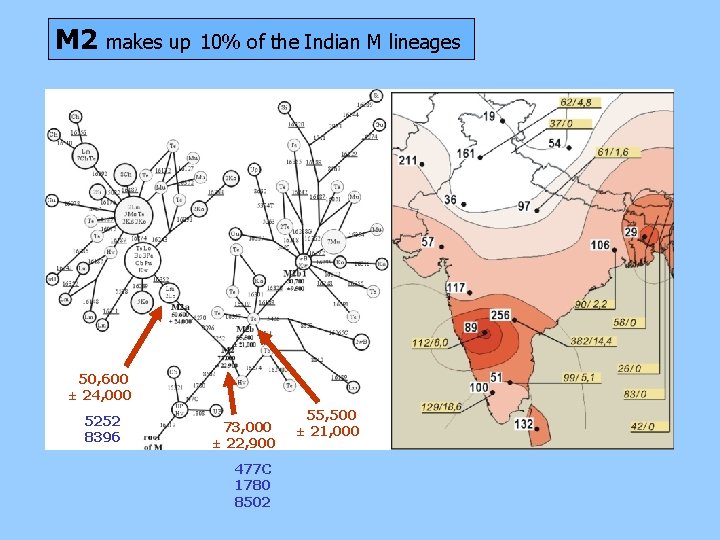
M 2 makes up 10% of the Indian M lineages 50, 600 ± 24, 000 5252 8396 73, 000 ± 22, 900 477 C 1780 8502 55, 500 ± 21, 000

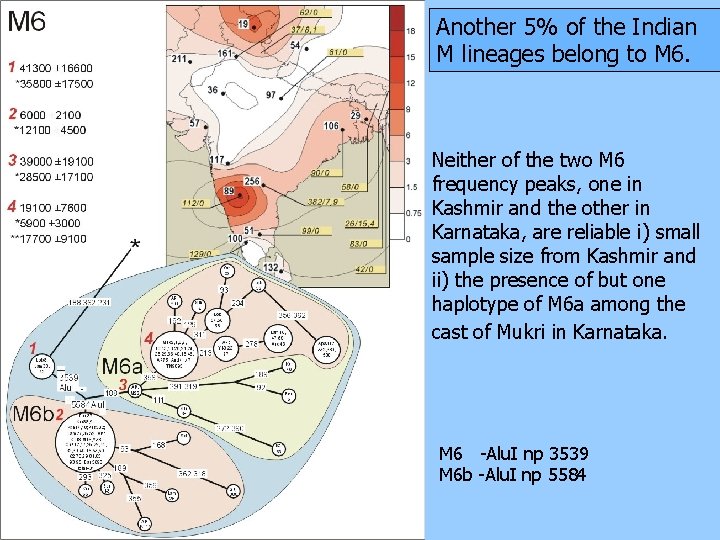
Another 5% of the Indian M lineages belong to M 6. Neither of the two M 6 frequency peaks, one in Kashmir and the other in Karnataka, are reliable i) small sample size from Kashmir and ii) the presence of but one haplotype of M 6 a among the cast of Mukri in Karnataka. M 6 -Alu. I np 3539 M 6 b -Alu. I np 5584

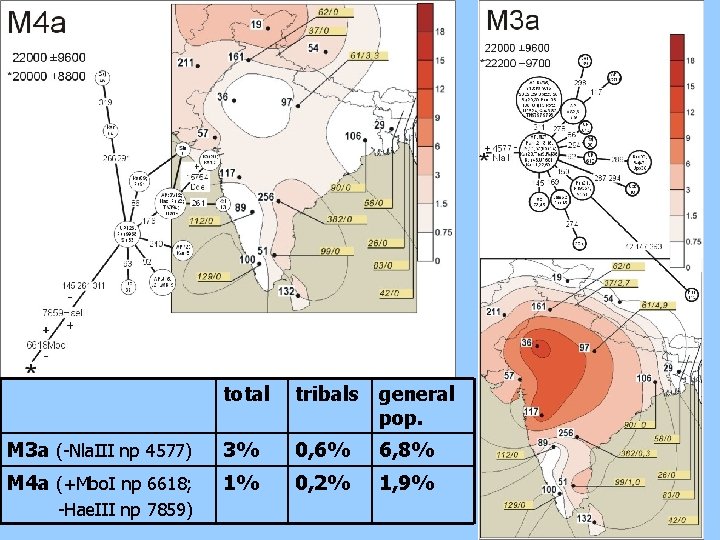
total tribals general pop. M 3 a (-Nla. III np 4577) 3% 0, 6% 6, 8% M 4 a (+Mbo. I np 6618; 1% 0, 2% 1, 9% -Hae. III np 7859)

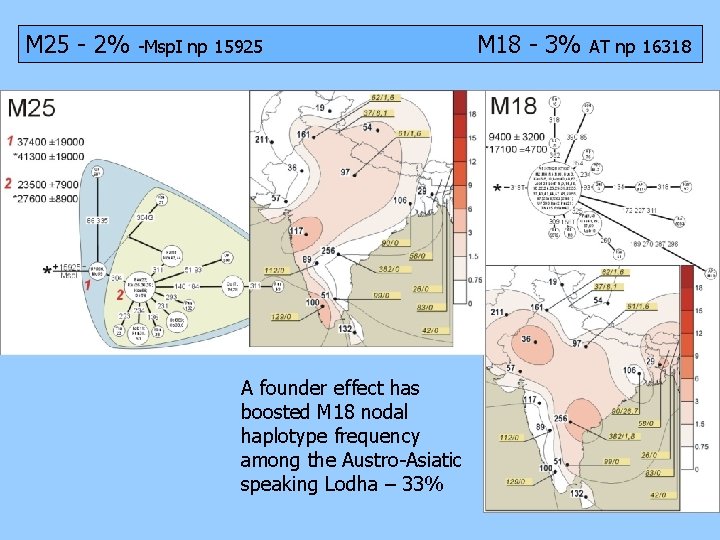
M 25 - 2% -Msp. I np 15925 A founder effect has boosted M 18 nodal haplotype frequency among the Austro-Asiatic speaking Lodha – 33% M 18 - 3% AT np 16318

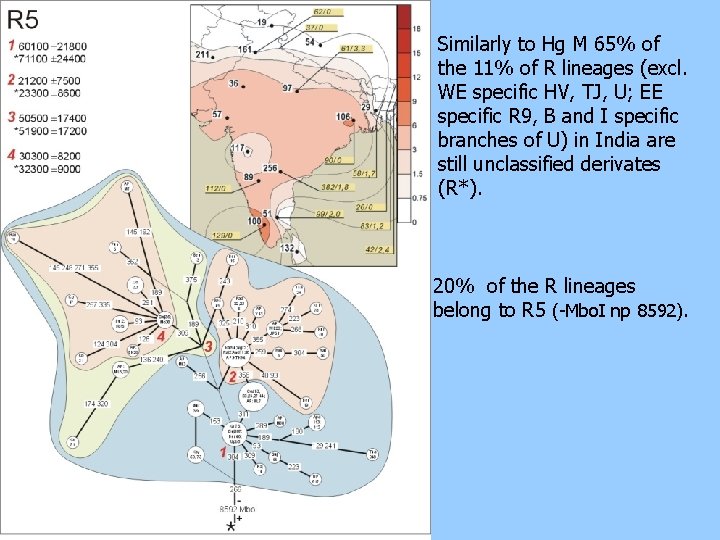
Similarly to Hg M 65% of the 11% of R lineages (excl. WE specific HV, TJ, U; EE specific R 9, B and I specific branches of U) in India are still unclassified derivates (R*). 20% of the R lineages belong to R 5 (-Mbo. I np 8592).

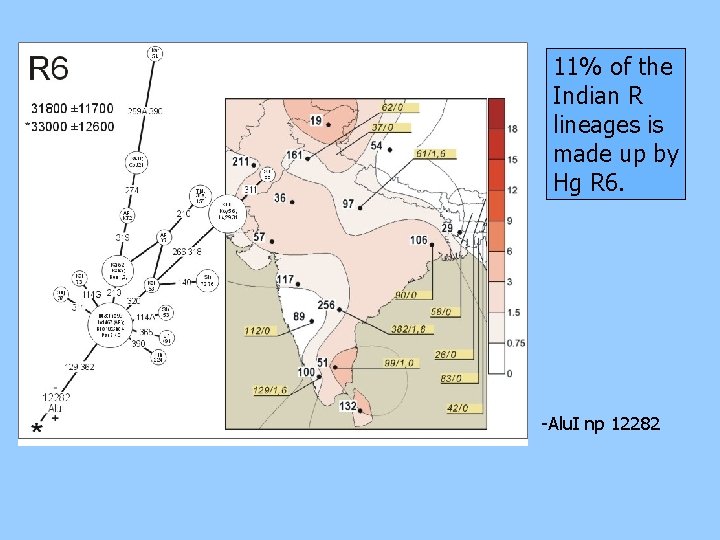
11% of the Indian R lineages is made up by Hg R 6. -Alu. I np 12282

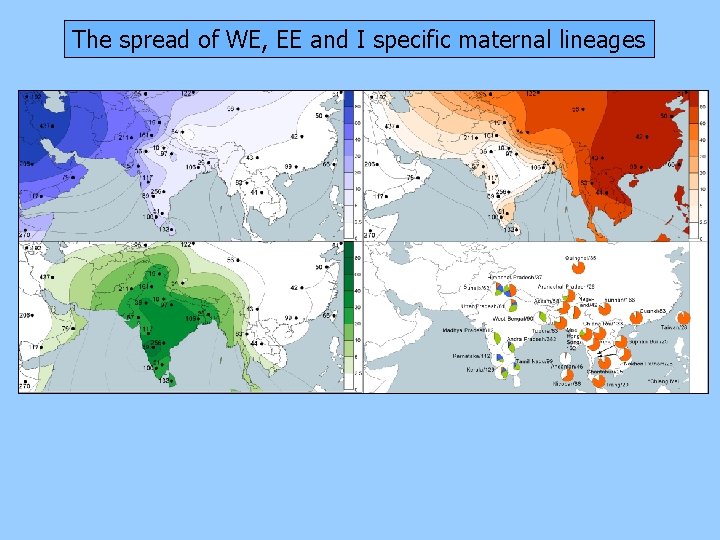
The spread of WE, EE and I specific maternal lineages

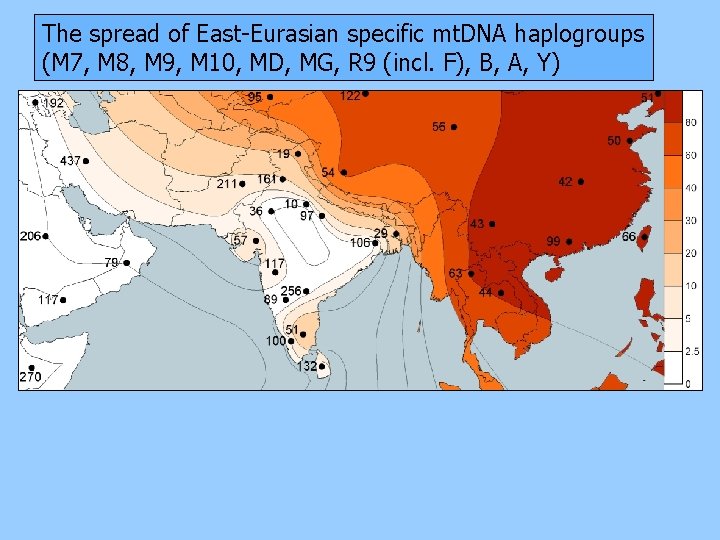
The spread of East-Eurasian specific mt. DNA haplogroups (M 7, M 8, M 9, M 10, MD, MG, R 9 (incl. F), B, A, Y)

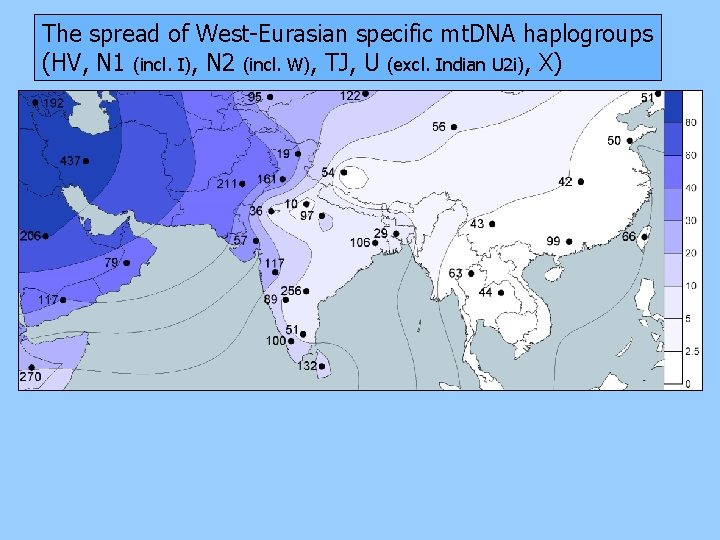
The spread of West-Eurasian specific mt. DNA haplogroups (HV, N 1 (incl. I), N 2 (incl. W), TJ, U (excl. Indian U 2 i), X)

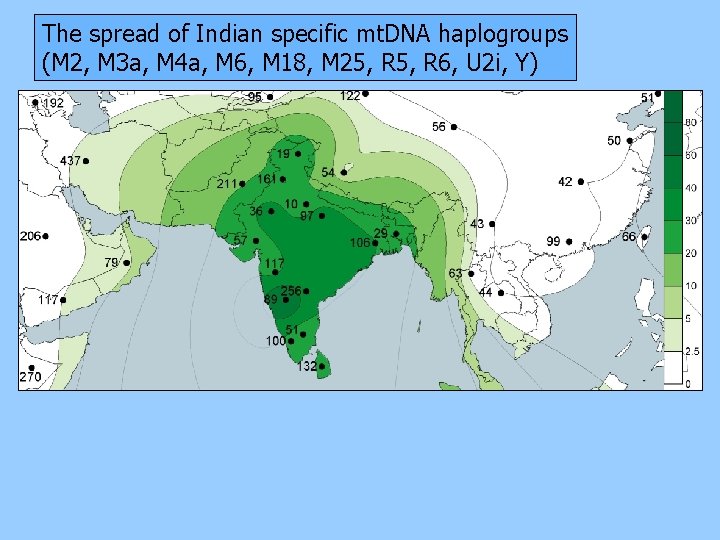
The spread of Indian specific mt. DNA haplogroups (M 2, M 3 a, M 4 a, M 6, M 18, M 25, R 6, U 2 i, Y)

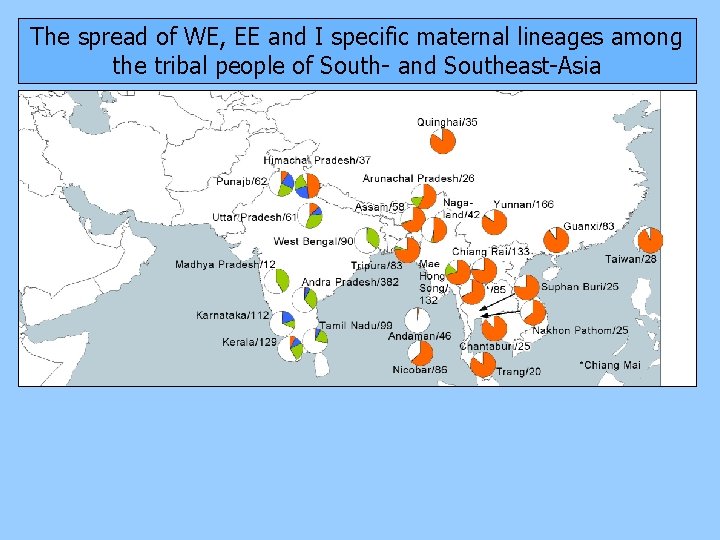
The spread of WE, EE and I specific maternal lineages among the tribal people of South- and Southeast-Asia

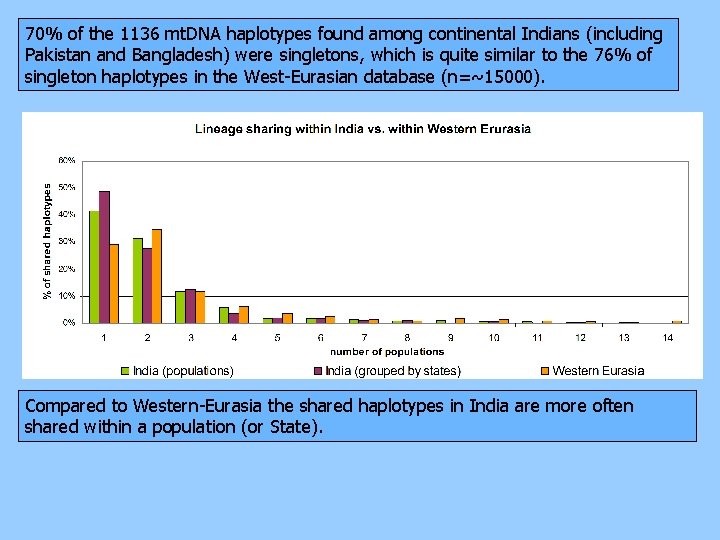
70% of the 1136 mt. DNA haplotypes found among continental Indians (including Pakistan and Bangladesh) were singletons, which is quite similar to the 76% of singleton haplotypes in the West-Eurasian database (n=~15000). Compared to Western-Eurasia the shared haplotypes in India are more often shared within a population (or State).

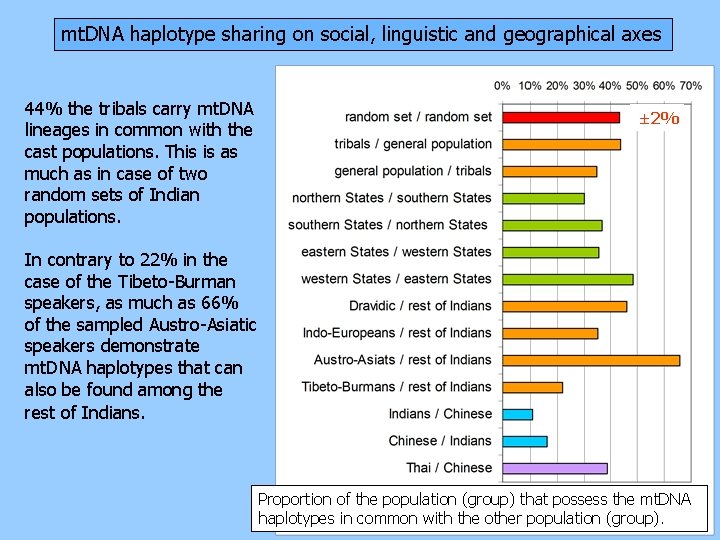
mt. DNA haplotype sharing on social, linguistic and geographical axes 44% the tribals carry mt. DNA lineages in common with the cast populations. This is as much as in case of two random sets of Indian populations. ± 2% In contrary to 22% in the case of the Tibeto-Burman speakers, as much as 66% of the sampled Austro-Asiatic speakers demonstrate mt. DNA haplotypes that can also be found among the rest of Indians. Proportion of the population (group) that possess the mt. DNA haplotypes in common with the other population (group).

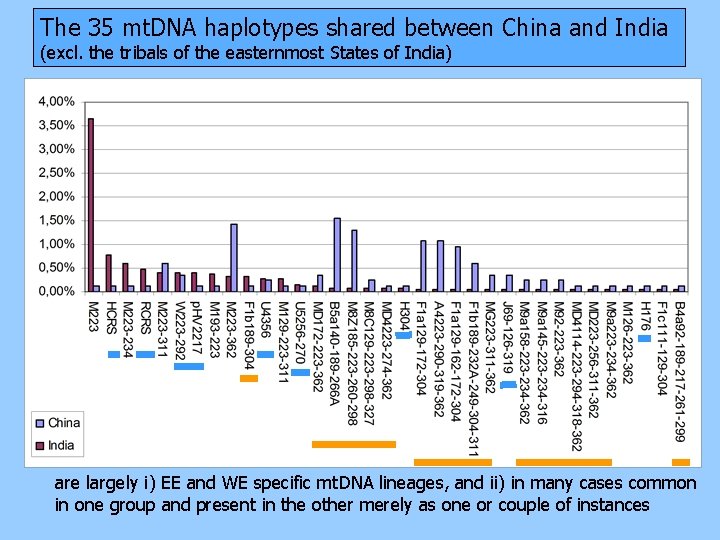
The 35 mt. DNA haplotypes shared between China and India (excl. the tribals of the easternmost States of India) are largely i) EE and WE specific mt. DNA lineages, and ii) in many cases common in one group and present in the other merely as one or couple of instances

Conclusions: • 30% of the Indian M and R (excl. U) lineages have been assigned to Indian specific subhaplogroups that show coalescence times in early and late Upper Palaeolithic. • As expected, Western- and Eastern-Eurasian mt. DNA lineages are more dominant respectively in north-western and north-eastern India. Over half of the maternal lineages among the Tibeto-Burman speaking tribals of the eastern- and northernmost States of India belong to EE specific mt. DNA lineage clusters. • Mt. DNA haplotype sharing analysis did not support segregation of Indian populations based on linguistics and social status (tribal vs. cast pop. ) (except for the Tibeto-Burman speakers).

Acknowledgements Tartu, Estonia Jaan Lind Ille Hilpus Jüri Parik Eva-Liis Loogväli Katrin Kaldma Ene Metspalu Helle-Viivi Tolk Toomas Kivisild Richard Villems Loughborough, UK Sarabijt Mastana Newcastle-upon-Tyne, UK Surinder S. Papiha Pavia, Italy Chiara Rengo Antonio Torroni Haifa, Israel Doron Behar Paris, France Luis Quintana-Murci