Tour of Viro BIKE Sequence comparison Viro BIKE

Tour of Viro. BIKE Sequence comparison Viro. BIKE (Biological Integrated Knowledge Environment) combines: Knowledge: All completed viral genomes known to NCBI Many viral metagenomes Analytical Tools: A powerful graphical language that permits creative expression to those with no programming experience It is accessible through a virological community web site: http: //ixion. csbc. vcu. edu/virobike This demonstration is best viewed as a slide show, enabling you to simulate a session and make changes in cursor more Click anywhere to position go on to theobvious. next slide To do this, click Slide Show on the top tool bar, then View show.

Tour of Viro. BIKE Sequence comparison In this tour, you'll see how to: Slide 4 • Log onto Viro. BIKE 8 • Speak Bio. BIKE (the language of Viro. BIKE) 17 • Display the sequence of a metagenome contig 24 • Find similar sequences amongst metagenomes 32 • Find similar sequences amongst known viruses 37 • Find similar sequences amongst everything in Gen. Bank 80 • Make a sequence alignment 85 • Make a phylogenetic tree 92 • Save your work session You can go to any slide in this tour at any time by typing the slide number and pressing Enter.

Coming Attractions! If you like this tour, you might also try Analysis of Metagenome Aggregates, where you'll see how to: • Find the number of contigs in a metagenome • Find the average contig size in a metagenome • Find the average GC content within a metagenome • Visualize the distribution of GC content amongst the contigs of a metagenome
URL: htpp: //ixion. csbc. vcu. edu/virobike The public can access everything at the community web site (except member names and e-mails), but only registered users can write to it. For now you're public. Access Viro. BIKE through the blue bar.
Click the link to the public access login screen.

Your name (no spaces) Enter anything you like as a login name, but no spaces or symbols. No password necessary. Click New Login
You can leave the blue bar to access other resources, but screen space is scarce, so grab it and move it offscreen to the left.

Function palette Workspace The Bio. BIKE environment is divided into three areas as shown. You'll bring functions down from the function palette to the workspace, execute them, and note the results in the results window Results window
Two very important buttons on the function palette: HELP! On-line help (general) PROBLEM Something went wrong? Tell us!
Two very important buttons in the workspace: Undo (return to workspace before last action) Redo (Get back the workspace you undid)
Our Story Suppose you have a special interest in a sequence, a contig, derived from the metagenome taken from the Arctic Ocean. The metagenome is called p-arct. The sequence is called C 60790. What does the sequence look like?
Clicking on any palette button brings down choices of functions or data to bring into the workspace. Click the function DISPLAYSEQUENCE-OF.
A DISPLAY-SEQUENCE-OF function box is now in the workspace. Before continuing with the problem, let's consider what function boxes mean.

General Syntax of Bio. BIKE Function-name Argument (object) Keyword object The basic unit of Bio. BIKE is the function box. It consists of the name of a function, perhaps one or more required arguments, and optional keywords and flags. A function may be thought of as a black box: you feed it information, it produces a product. Flag

General Syntax of Bio. BIKE Function-name Argument (object) Keyword object Flag Function boxes contain the following elements: • Function-name (e. g. SEQUENCE-OF or LENGTH-OF) • Argument: Required, acted on by function • Keyword clause: Optional, more information • Flag: Optional, more (yes/no) information

General Syntax of Bio. BIKE Function-name Argument (object) Keyword object Flag … and icons to help you work with functions: • Option icon: Brings up a menu of keywords and flags • Action icon: Brings up a menu enabling you to execute a function, copy and paste, information, get help, etc Clear/Delete icon: Removes information you entered or removes box entirely •

Back to our story… we were displaying the sequence of our favorite metagenome contig, C 60790. Click on the gray argument box to activate it for entry, either from the keyboard or by insertion.
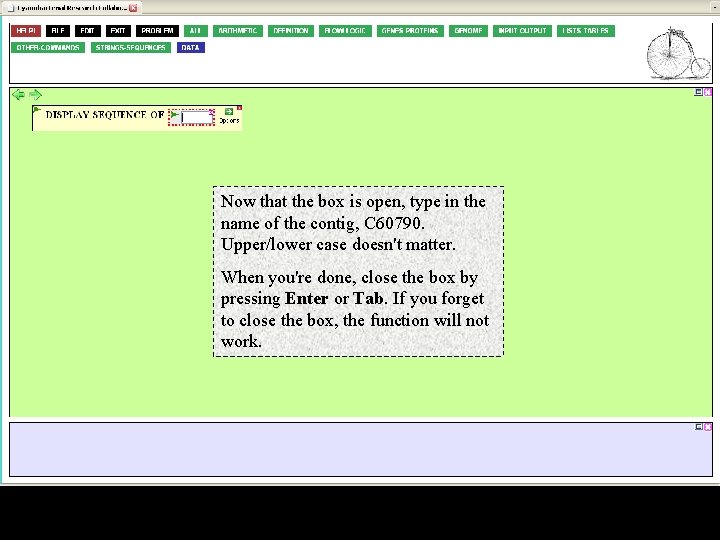
Now that the box is open, type in the name of the contig, C 60790. Upper/lower case doesn't matter. When you're done, close the box by pressing Enter or Tab. If you forget to close the box, the function will not work.
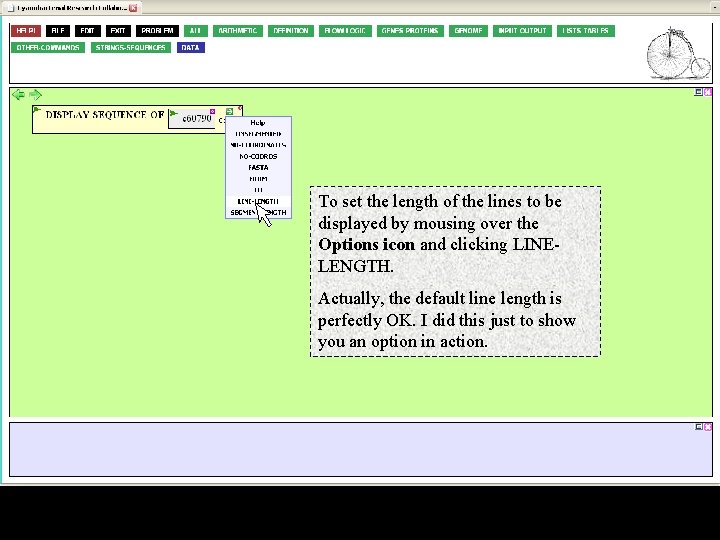
To set the length of the lines to be displayed by mousing over the Options icon and clicking LINELENGTH. Actually, the default line length is perfectly OK. I did this just to show you an option in action.
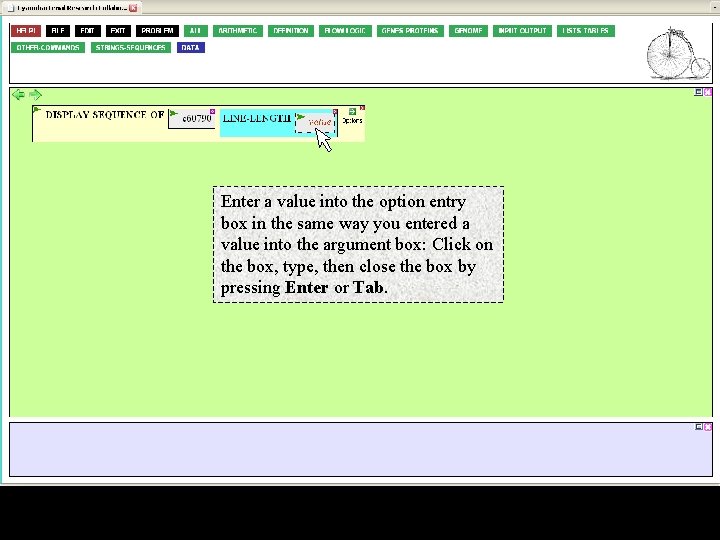
Enter a value into the option entry box in the same way you entered a value into the argument box: Click on the box, type, then close the box by pressing Enter or Tab.
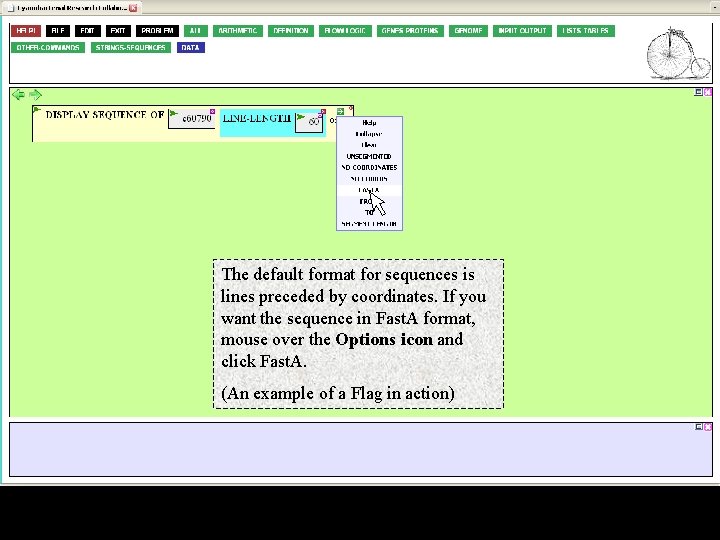
The default format for sequences is lines preceded by coordinates. If you want the sequence in Fast. A format, mouse over the Options icon and click Fast. A. (An example of a Flag in action)
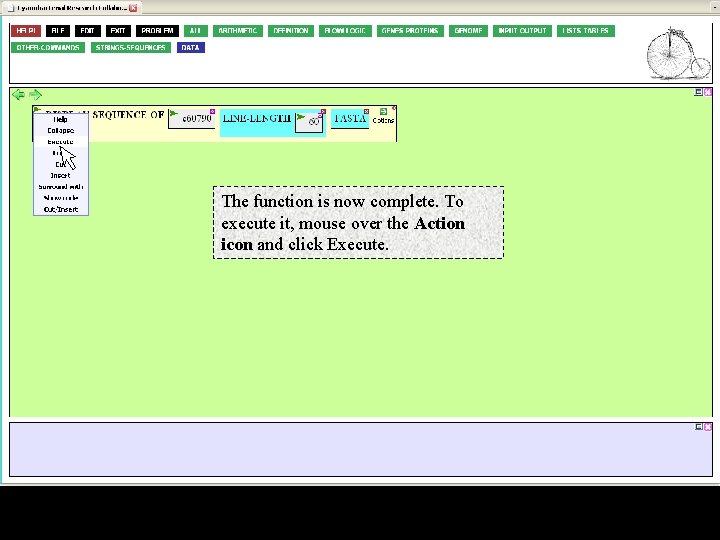
The function is now complete. To execute it, mouse over the Action icon and click Execute.
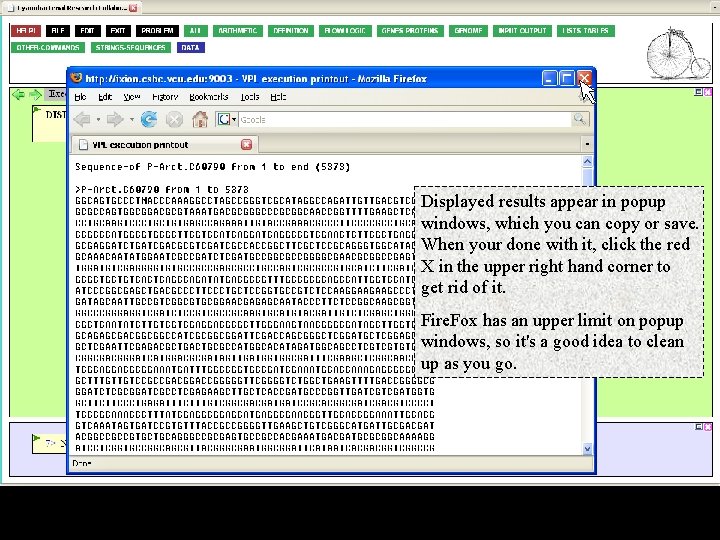
Displayed results appear in popup windows, which you can copy or save. When your done with it, click the red X in the upper right hand corner to get rid of it. Fire. Fox has an upper limit on popup windows, so it's a good idea to clean up as you go.
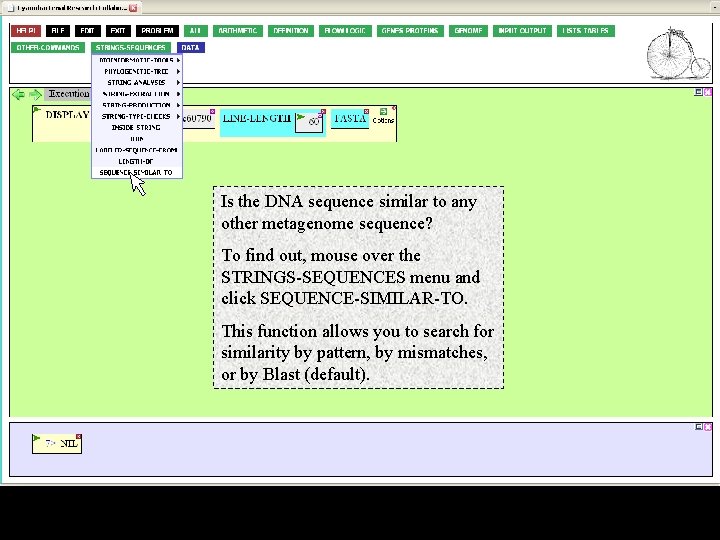
Is the DNA sequence similar to any other metagenome sequence? To find out, mouse over the STRINGS-SEQUENCES menu and click SEQUENCE-SIMILAR-TO. This function allows you to search for similarity by pattern, by mismatches, or by Blast (default).
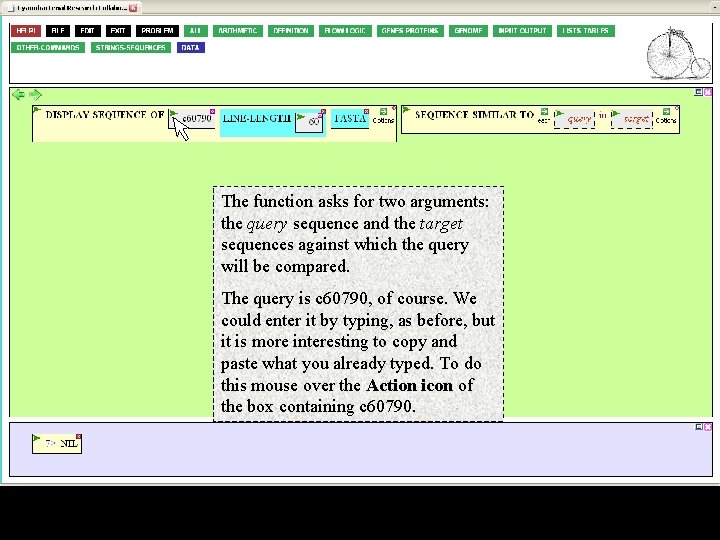
The function asks for two arguments: the query sequence and the target sequences against which the query will be compared. The query is c 60790, of course. We could enter it by typing, as before, but it is more interesting to copy and paste what you already typed. To do this mouse over the Action icon of the box containing c 60790.
Click Copy.
To paste, mouse over the Action icon of the box into which you're pasting and click Paste.
Now to enter the target sequences – the set of all metagenome sequences. Click on the target box to open it for entry. Once the box is open, you could specify by typing that you want to search metagenomic sequences… if you knew what to type.
If you don't know, then mouse over the DATA button, then Organisms, then Metagenomes. Clicking on Metagenomes transfers it to the open target box.

Execute the completed function as before, mousing over the Action icon of the function and clicking Execute. Doing so starts Blast, which may take several seconds to complete execution.
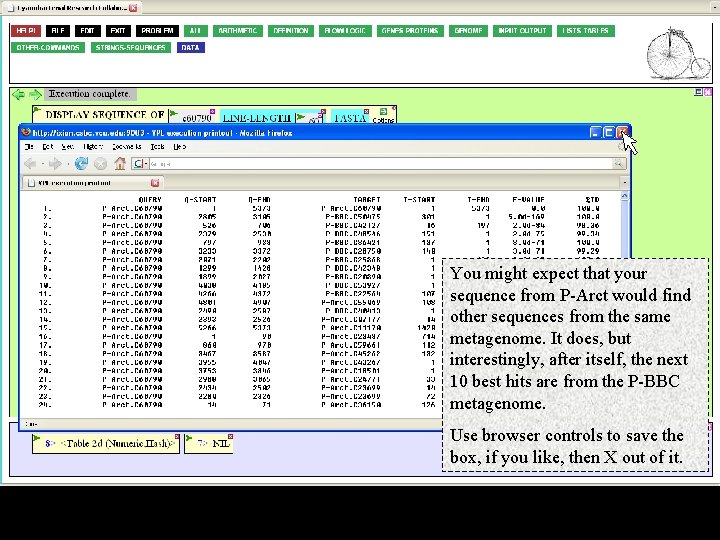
You might expect that your sequence from P-Arct would find other sequences from the same metagenome. It does, but interestingly, after itself, the next 10 best hits are from the P-BBC metagenome. Use browser controls to save the box, if you like, then X out of it.
Of course the metagenome sequences are not annotated. Perhaps you can learn more about your sequence by comparing it to sequences from known viruses. To do this, clear the target box, open it up again by clicking on it…
…and bring down Known Viruses into the box.
Protein searches will find more sequences, mouse over the Options icon and specify that your DNA sequence is to be translated and compared to viral proteins.
Execute the completed function. Again, execution may take several seconds.
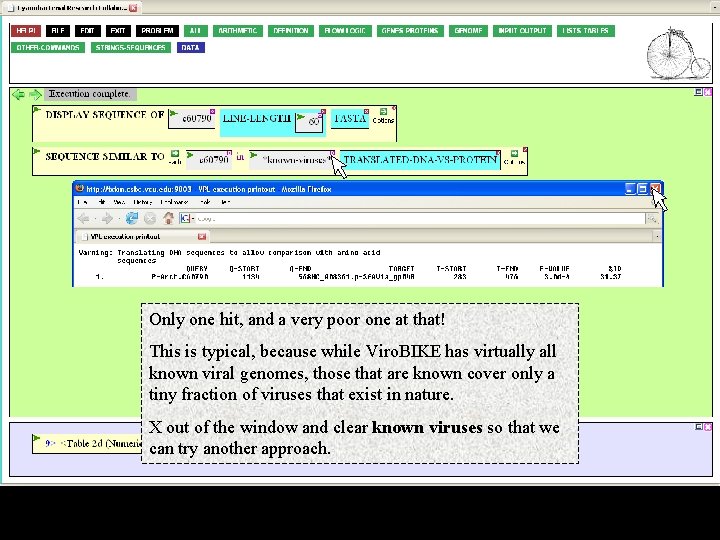
Only one hit, and a very poor one at that! This is typical, because while Viro. BIKE has virtually all known viral genomes, those that are known cover only a tiny fraction of viruses that exist in nature. X out of the window and clear known viruses so that we can try another approach.
There is a good deal more variety in organismal genomes than viral genomes, so let's search them. Viro. BIKE does not keep organismal genomes locally, so we need to go out to Gen. Bank. Click on the DATA button again.
…and this time click Gen. Bank.
Execute the function as usual. This time we will be at the mercy of NCBI, and depending on the time of day and the phase of the moon, execution may take a minute or longer. By default, Viro. BIKE times out execution at 40 seconds. If this occurs, you'll get a message like…

You can change the time limit, but let's say that fate is with us and you get your result. *** *** TIMEOUT ! TIMEOUT *** COMPUTATION ABORTED AFTER 40 SECONDS *** YOU CAN: - contact support for help: Bio. Lingua. Support@lists. Stanford. EDU *** - use the TOOLS -> PREFS menu or the SET-TIMELIMIT function to extend your timeout up to 1 hour *** - use RUNJOB to run your code in a separate process *** - type (explain-timeout) at the weblistener for detailed info.
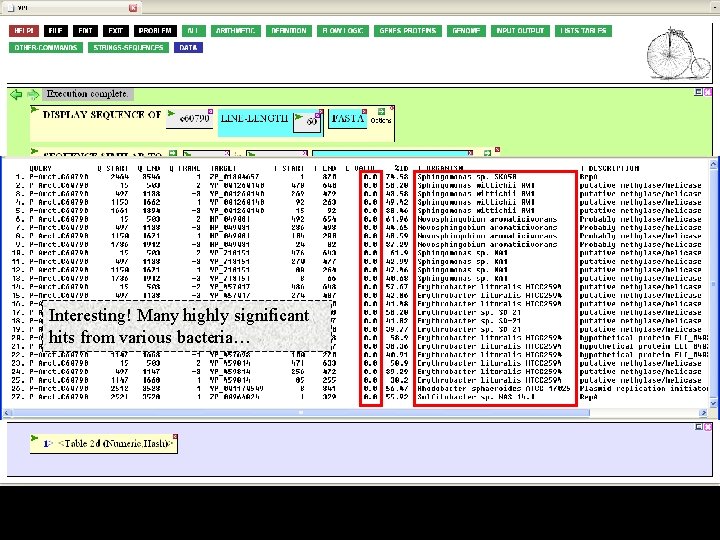
Interesting! Many highly significant hits from various bacteria…

…at different regions of your sequence. At NCBI, that would be the end of the story. In Viro. BIKE, it's the beginning, since you can work with your Blast results. First, we'll want to give the result a name.
To name a result, mouse over the DEFINITION menu and click DEFINE.
The DEFINE function asks for two arguments: the name of the variable and the value that will be assigned to it. Click on the variable entry box.
You can name the result anything you like, so long as the name does not contain spaces (hyphens and underscores are OK). I chose c 67090 -vs-NR. Press Tab after typing a name.
Tabbing opens up the next argument, the value box. The value to be assigned is the Blast table. There are many ways to retrieve that result. One way is to recognize that it is the result of the previous function. Click the OTHER-COMMAND button. . .

…and click Previous-Result.
Executing the function will cause the variable you named to spring into existence, accessible through a new button. Watch for it!

We'll be using that VARIABLES button in a moment. For now, mouse over STRINGS-SEQUENCES, then SEARCH/COMPARE, and…

Click on BLAST-VALUE. This function allows you to extract values from the Blast table.
What values do we want to extract? Recall…
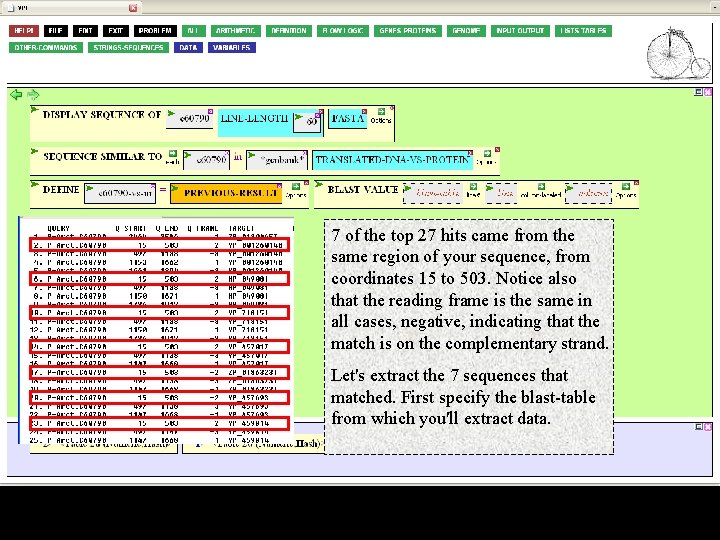
7 of the top 27 hits came from the same region of your sequence, from coordinates 15 to 503. Notice also that the reading frame is the same in all cases, negative, indicating that the match is on the complementary strand. Let's extract the 7 sequences that matched. First specify the blast-table from which you'll extract data.
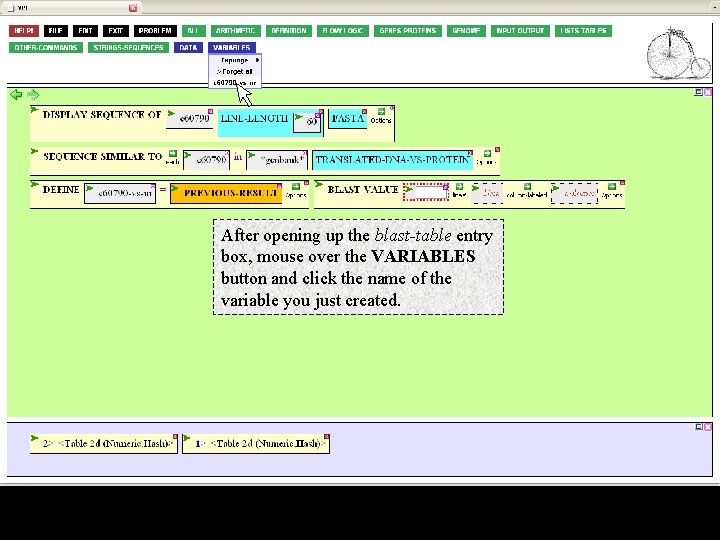
After opening up the blast-table entry box, mouse over the VARIABLES button and click the name of the variable you just created.

This brings the variable into the open box. Now specify the cells you want, by row numbers (lines) and column. Click to open the line box
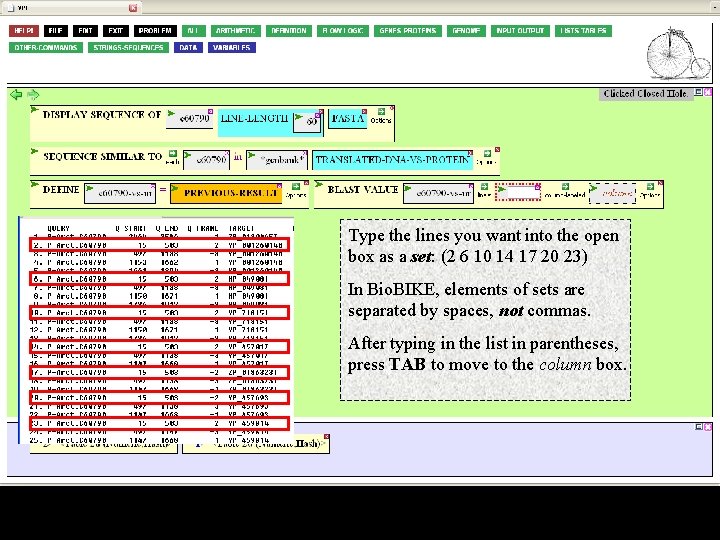
Type the lines you want into the open box as a set: (2 6 10 14 17 20 23) In Bio. BIKE, elements of sets are separated by spaces, not commas. After typing in the list in parentheses, press TAB to move to the column box.

You can enter any column shown in the Blast table plus several other fields that are normally not displayed. One of these fields is the sequence of the target ("T-SEQ"). Type this into the column box and press Enter.
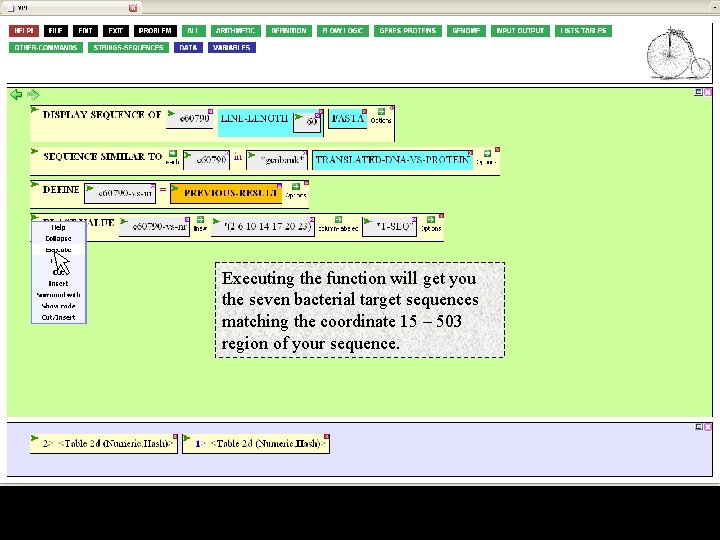
Executing the function will get you the seven bacterial target sequences matching the coordinate 15 – 503 region of your sequence.
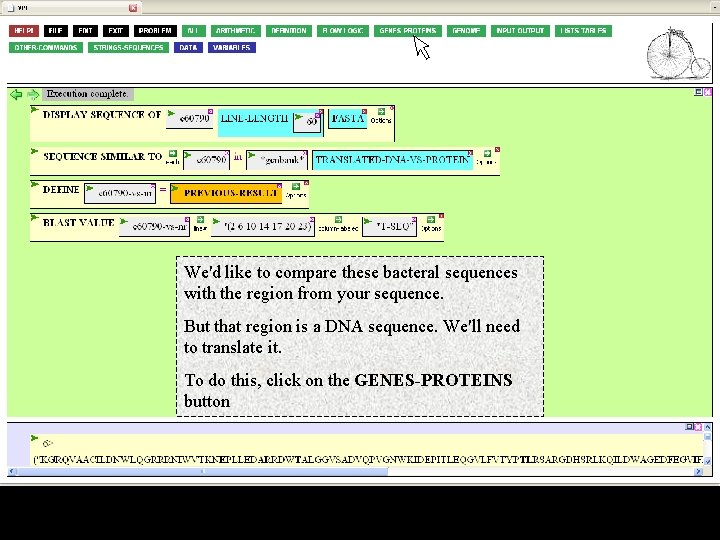
We'd like to compare these bacteral sequences with the region from your sequence. But that region is a DNA sequence. We'll need to translate it. To do this, click on the GENES-PROTEINS button
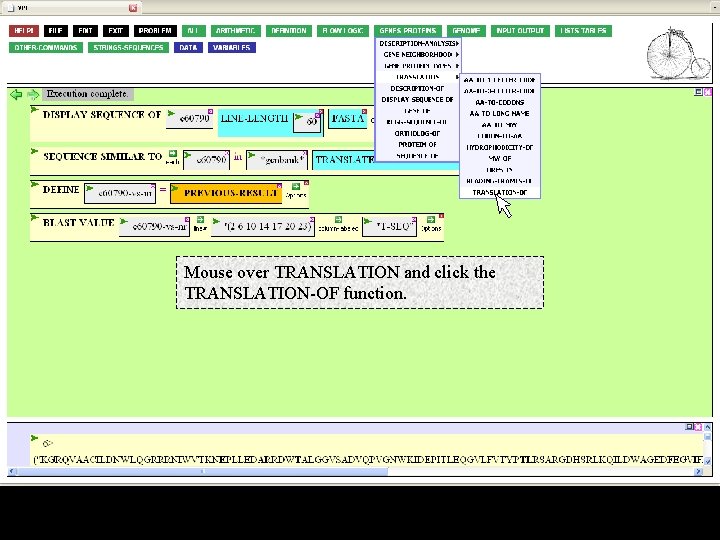
Mouse over TRANSLATION and click the TRANSLATION-OF function.
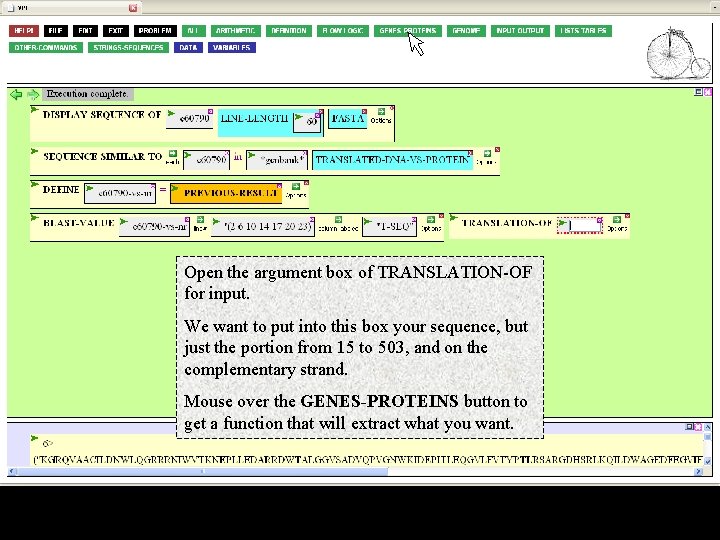
Open the argument box of TRANSLATION-OF for input. We want to put into this box your sequence, but just the portion from 15 to 503, and on the complementary strand. Mouse over the GENES-PROTEINS button to get a function that will extract what you want.

Click the SEQUENCE-OF function.
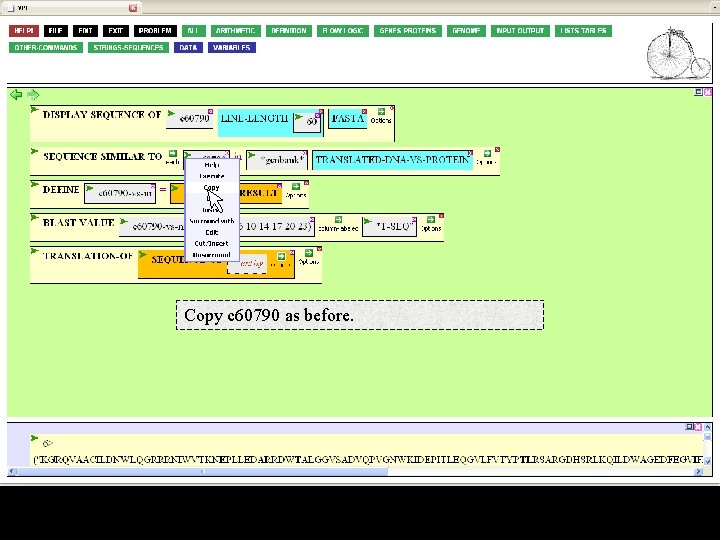
Copy c 60790 as before.

And paste it into the argument of SEQUENCE-OF. Executing now will translate the entire sequence. But we want only part of the sequence.
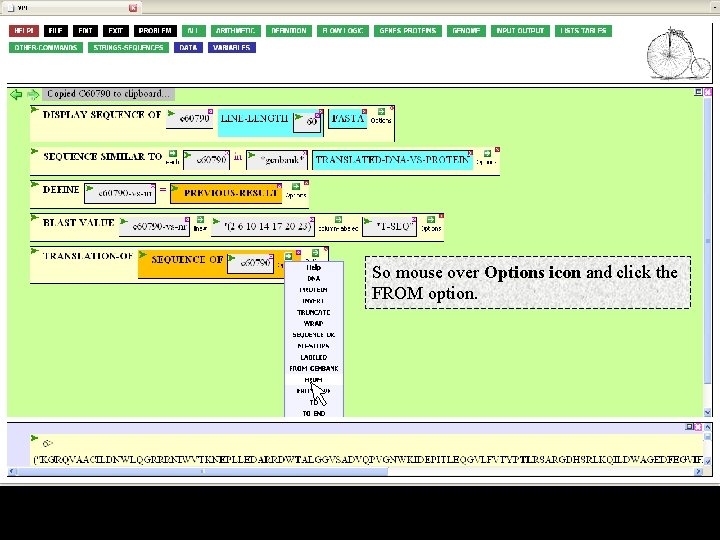
So mouse over Options icon and click the FROM option.

And do the same thing to get the TO option.
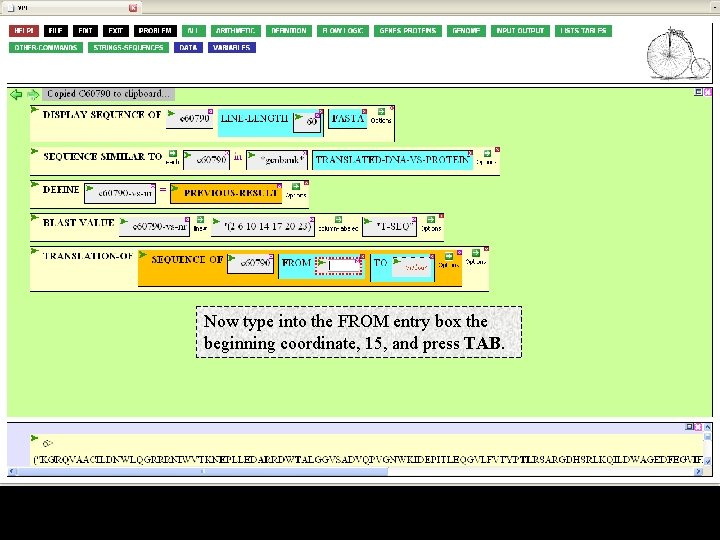
Now type into the FROM entry box the beginning coordinate, 15, and press TAB.
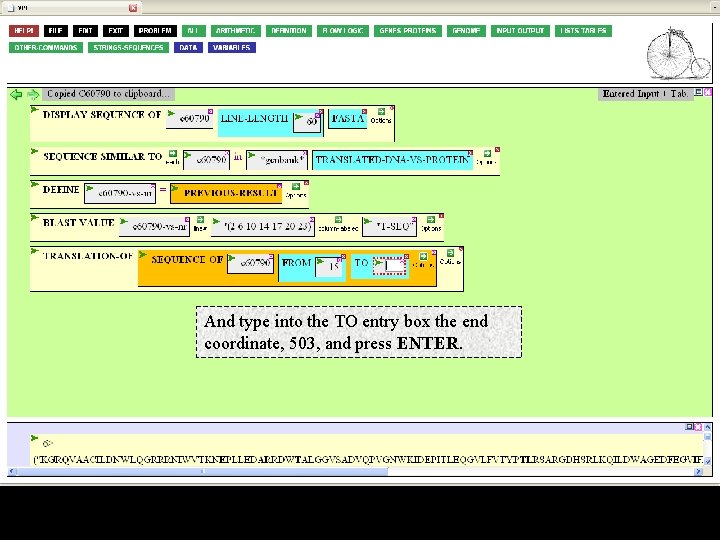
And type into the TO entry box the end coordinate, 503, and press ENTER.
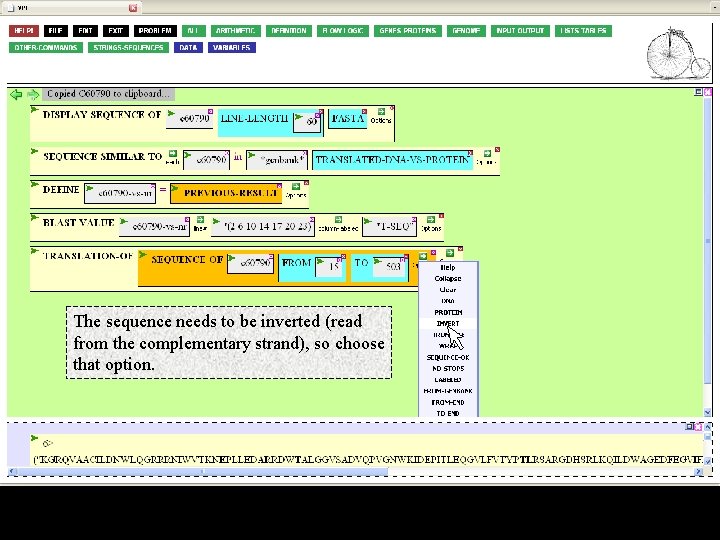
The sequence needs to be inverted (read from the complementary strand), so choose that option.
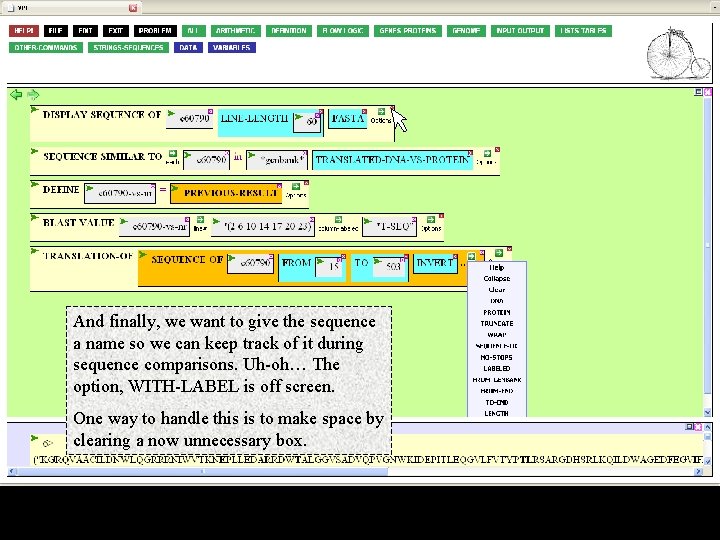
And finally, we want to give the sequence a name so we can keep track of it during sequence comparisons. Uh-oh… The option, WITH-LABEL is off screen. One way to handle this is to make space by clearing a now unnecessary box.
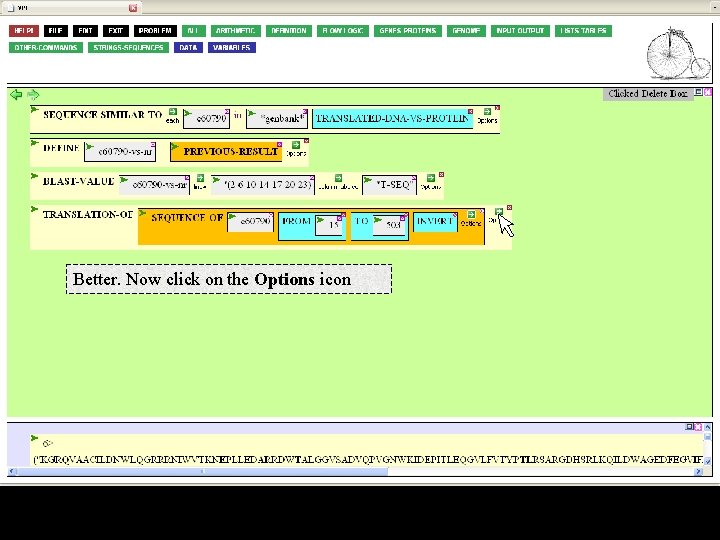
Better. Now click on the Options icon
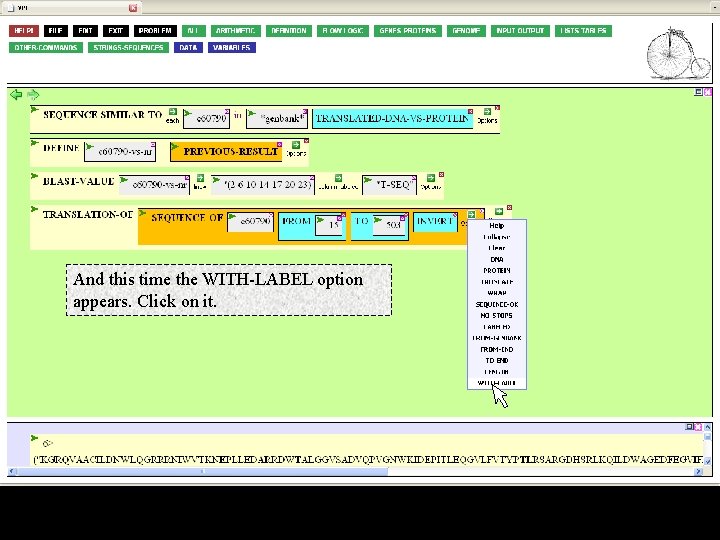
And this time the WITH-LABEL option appears. Click on it.
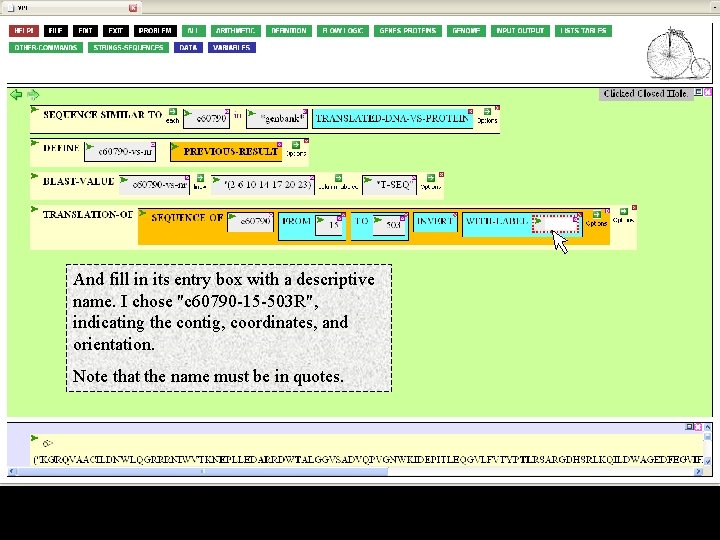
And fill in its entry box with a descriptive name. I chose "c 60790 -15 -503 R", indicating the contig, coordinates, and orientation. Note that the name must be in quotes.
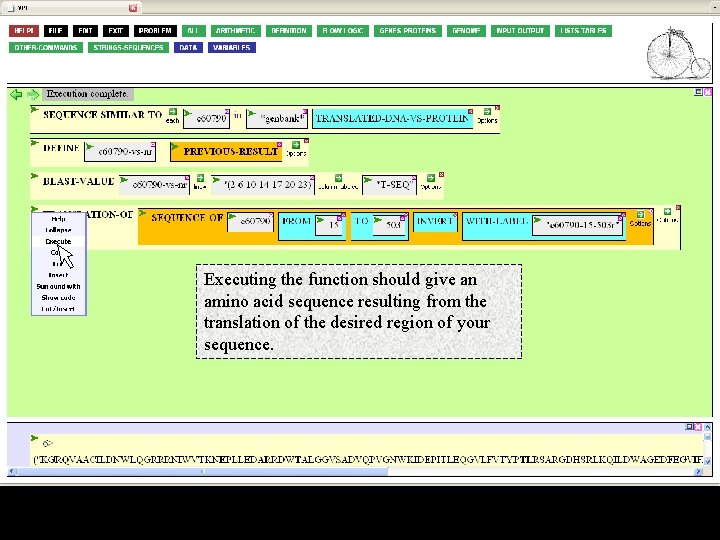
Executing the function should give an amino acid sequence resulting from the translation of the desired region of your sequence.
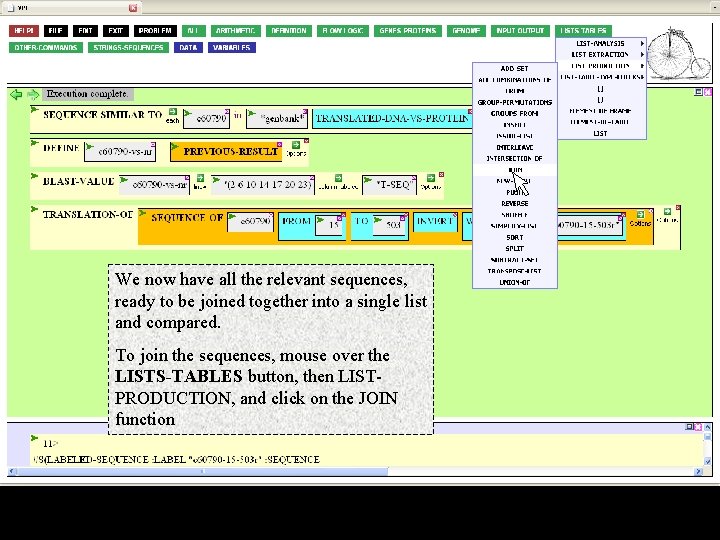
We now have all the relevant sequences, ready to be joined together into a single list and compared. To join the sequences, mouse over the LISTS-TABLES button, then LISTPRODUCTION, and click on the JOIN function
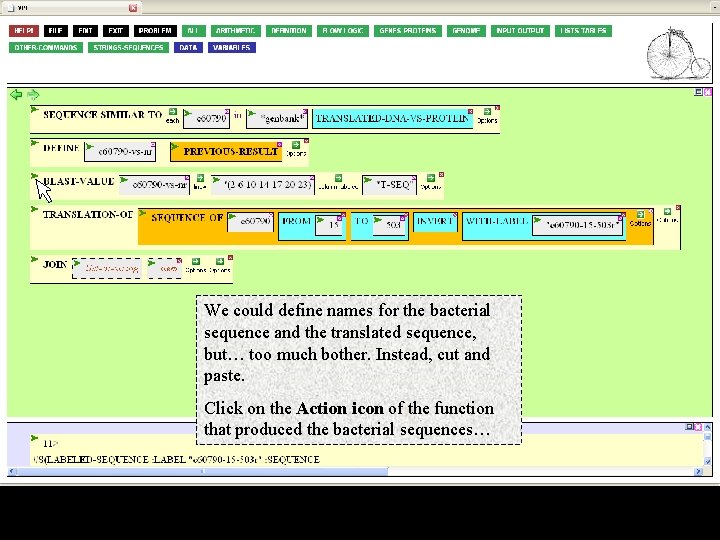
We could define names for the bacterial sequence and the translated sequence, but… too much bother. Instead, cut and paste. Click on the Action icon of the function that produced the bacterial sequences…

Cut the function box and paste it into the first argument box of JOIN.
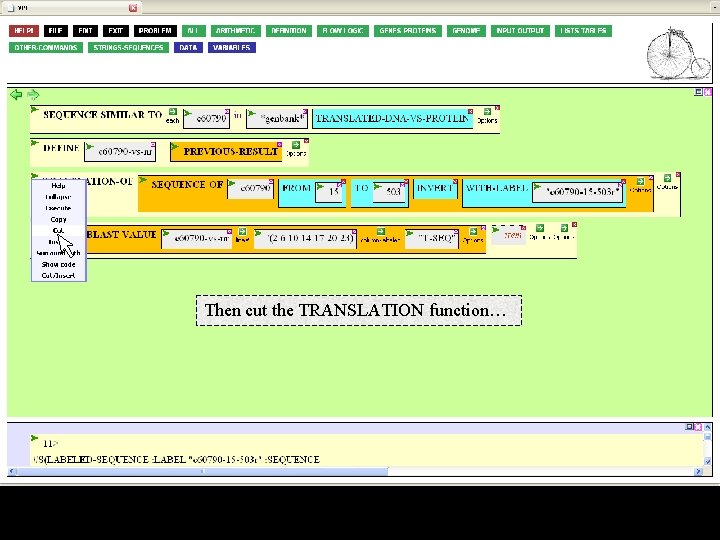
Then cut the TRANSLATION function…
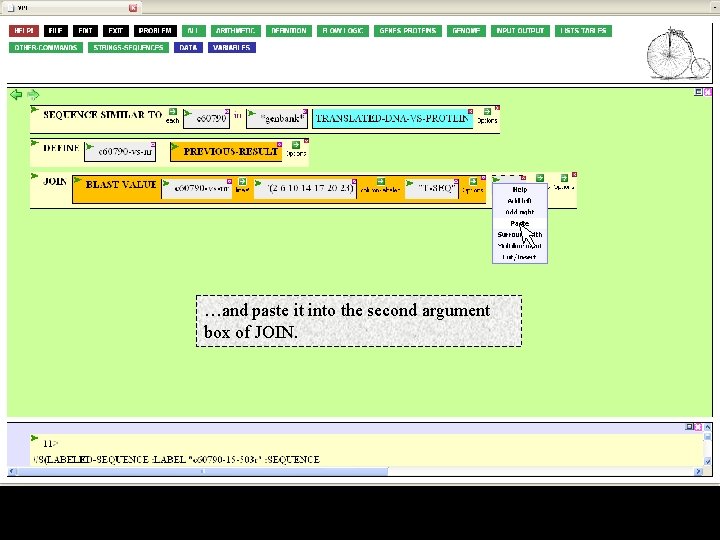
…and paste it into the second argument box of JOIN.
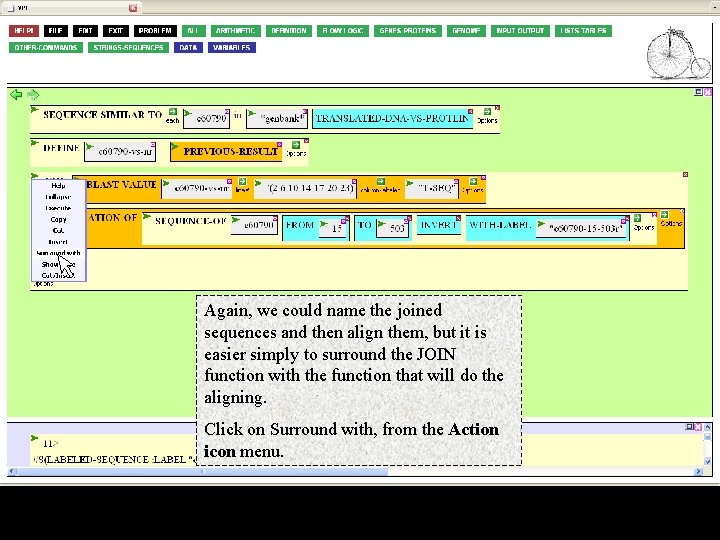
Again, we could name the joined sequences and then align them, but it is easier simply to surround the JOIN function with the function that will do the aligning. Click on Surround with, from the Action icon menu.

Then select ALIGNMENT-OF from the STRINGS-SEQUENCES menu, BIOINFORMATI-TOOLS submenu.
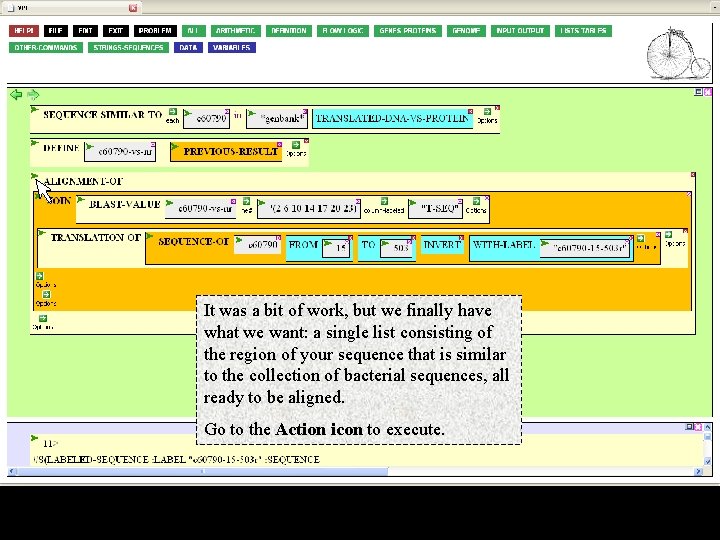
It was a bit of work, but we finally have what we want: a single list consisting of the region of your sequence that is similar to the collection of bacterial sequences, all ready to be aligned. Go to the Action icon to execute.
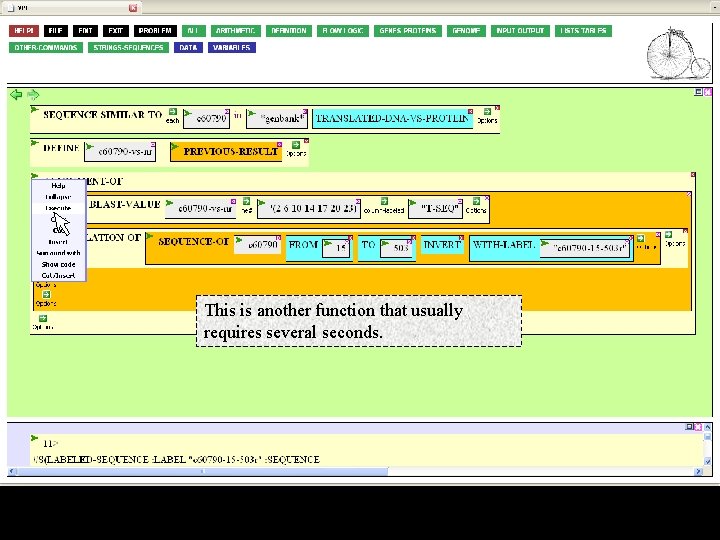
This is another function that usually requires several seconds.
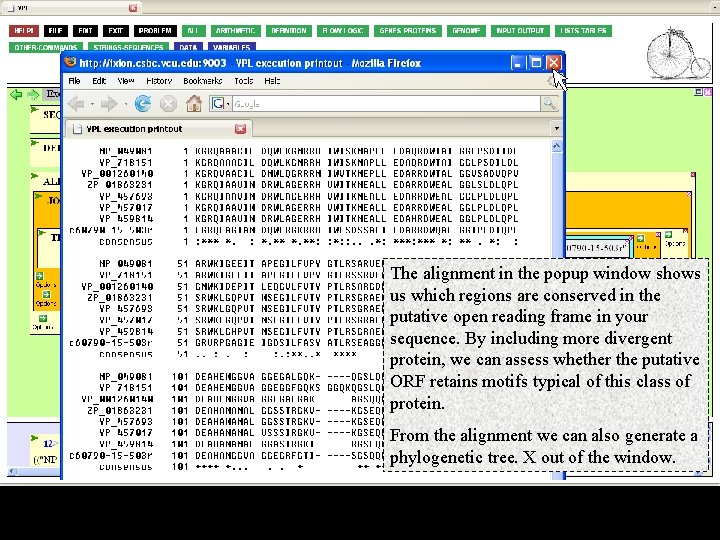
The alignment in the popup window shows us which regions are conserved in the putative open reading frame in your sequence. By including more divergent protein, we can assess whether the putative ORF retains motifs typical of this class of protein. From the alignment we can also generate a phylogenetic tree. X out of the window.
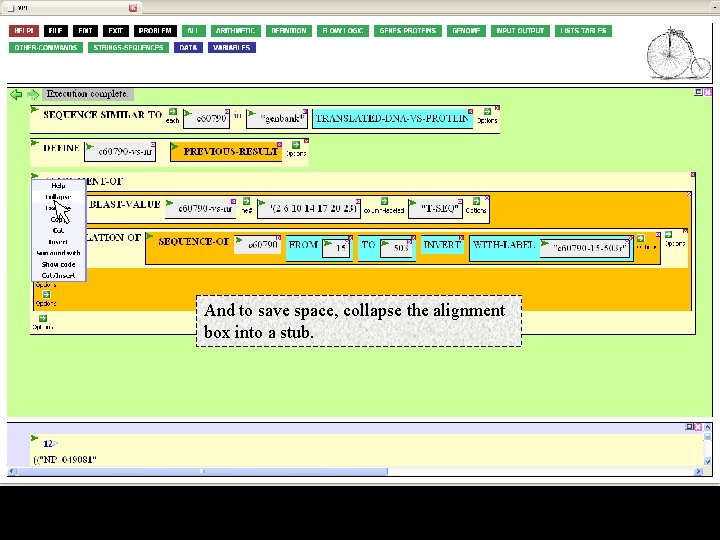
And to save space, collapse the alignment box into a stub.
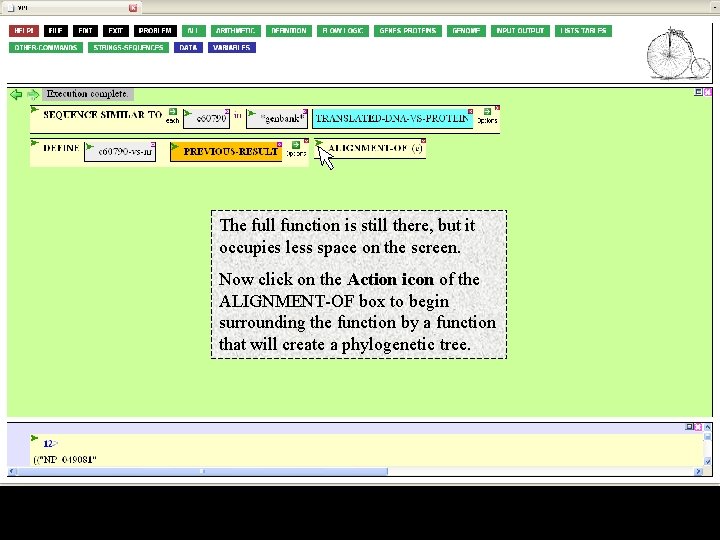
The full function is still there, but it occupies less space on the screen. Now click on the Action icon of the ALIGNMENT-OF box to begin surrounding the function by a function that will create a phylogenetic tree.
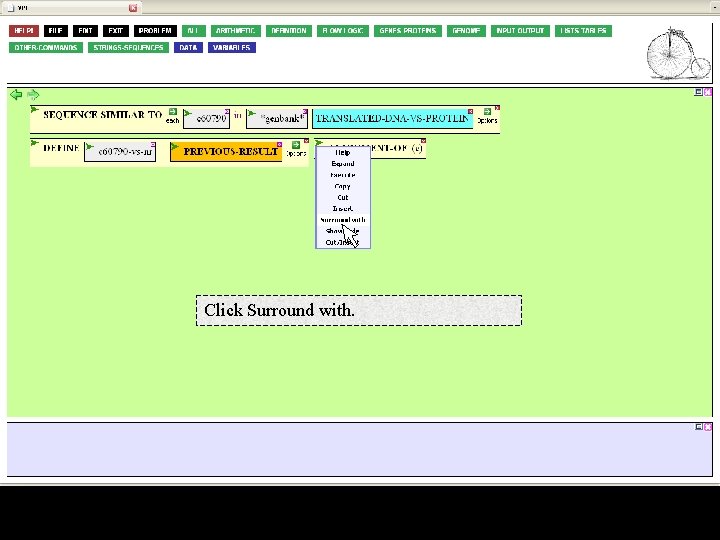
Click Surround with.
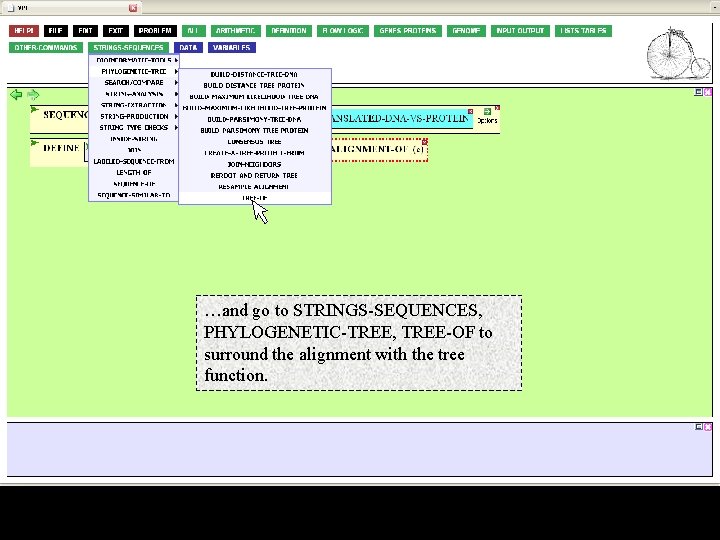
…and go to STRINGS-SEQUENCES, PHYLOGENETIC-TREE, TREE-OF to surround the alignment with the tree function.
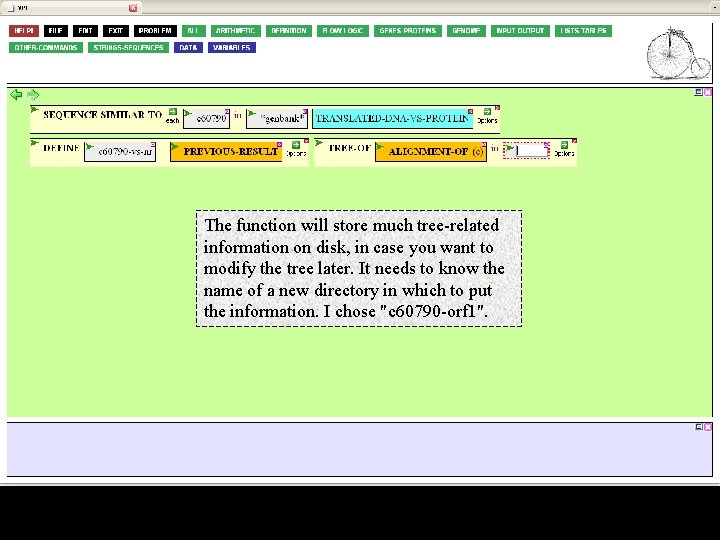
The function will store much tree-related information on disk, in case you want to modify the tree later. It needs to know the name of a new directory in which to put the information. I chose "c 60790 -orf 1".
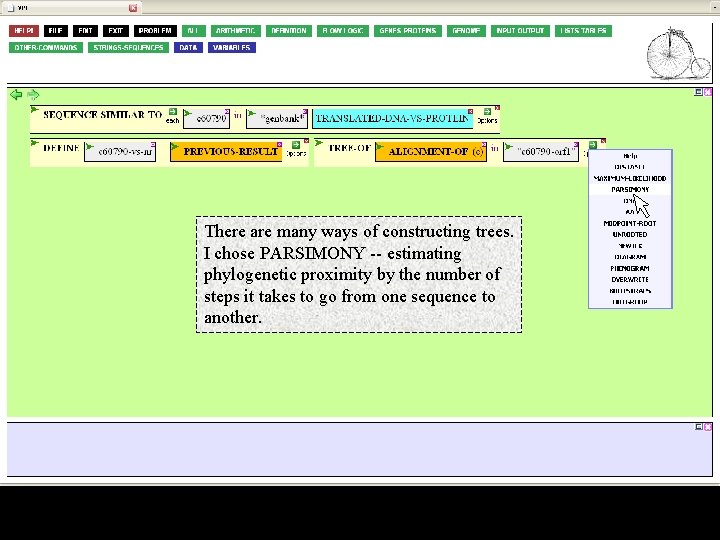
There are many ways of constructing trees. I chose PARSIMONY -- estimating phylogenetic proximity by the number of steps it takes to go from one sequence to another.
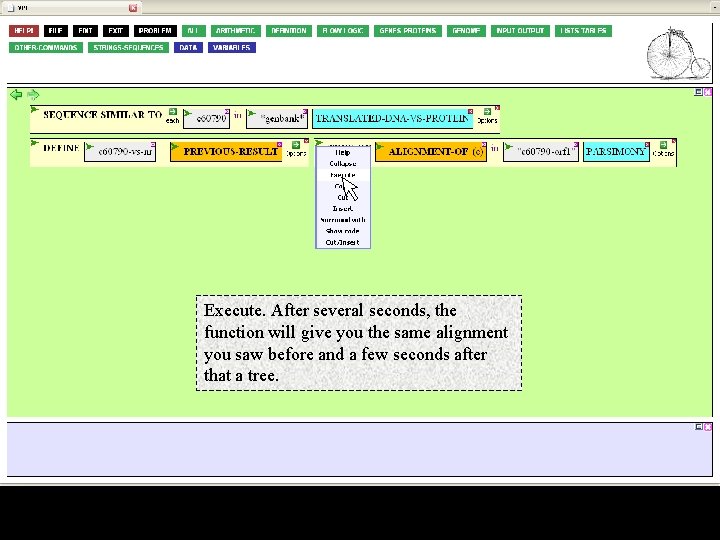
Execute. After several seconds, the function will give you the same alignment you saw before and a few seconds after that a tree.
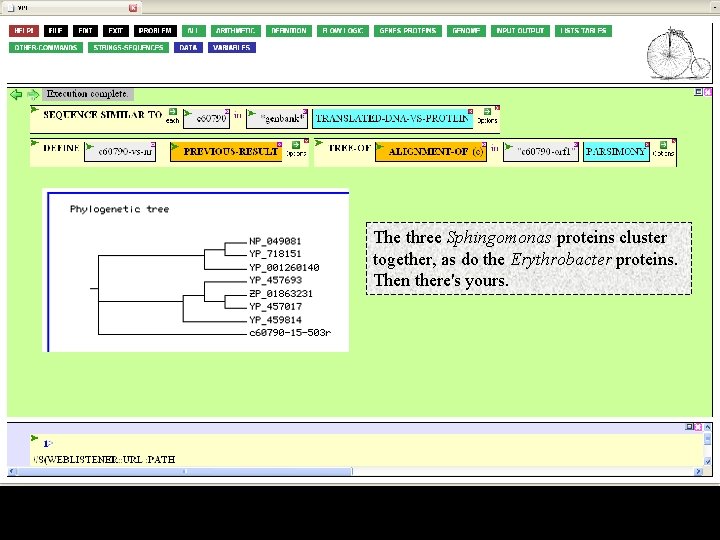
The three Sphingomonas proteins cluster together, as do the Erythrobacter proteins. Then there's yours.

If you want to return to this session or refer to it later, you can save it by mousing over the EDIT button and clicking Save user session.
- Slides: 92