Tools in Bioinformatics Genome Browsers Retrieving genomic information

Tools in Bioinformatics Genome Browsers

Retrieving genomic information n Previous lesson(s): annotation-based perspective of search/data n Today: genomic-based perspective: look at all the data from the prism of a specific chromosome location n Next: sequence-based searches

Genome browsers n NCBI Map Viewer n http: //www. ncbi. nih. gov/mapview n Ensembl n http: //www. ensembl. org/ n UCSC Genome Browser n http: //genome. ucsc. edu/
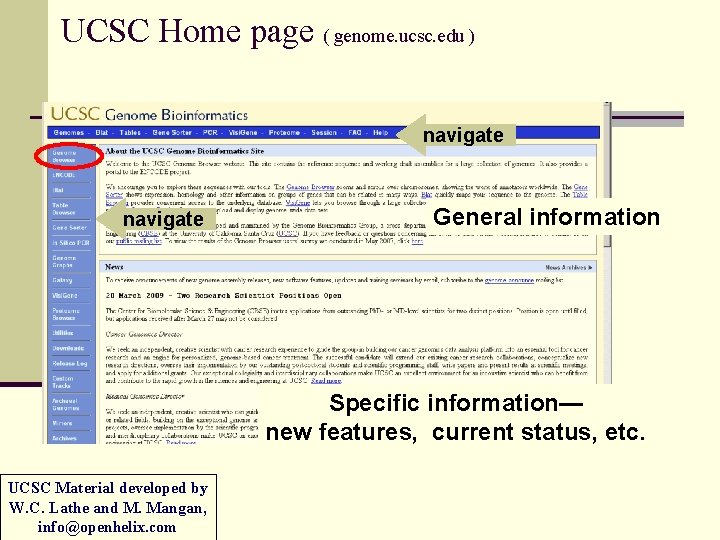
UCSC Home page ( genome. ucsc. edu ) navigate General information Specific information— new features, current status, etc. UCSC Material developed by W. C. Lathe and M. Mangan, info@openhelix. com

The Genome Browser Gateway start page, basic search Start all text/ID searches here! s ple xamow e l h rc s be a e l s ion fu gest p l He sug Use this Gateway page to search by: gene names, gene symbols, chromosome number, region, keyword, identifiers (NP, NM, OMIM, Locus. Link), and more. See examples on page format ,

The Genome Browser Gateway start page choices, sample search for Human BRCA 1 Perform a sample search with the May 2004 data assembly, and examine the results. We will use human BRCA 1 as our example. breast and ovarian cancer susceptibility gene, Science. 1994 Oct 7; 266(5182): 66 -71 Use the Submit button to send the query to the database.
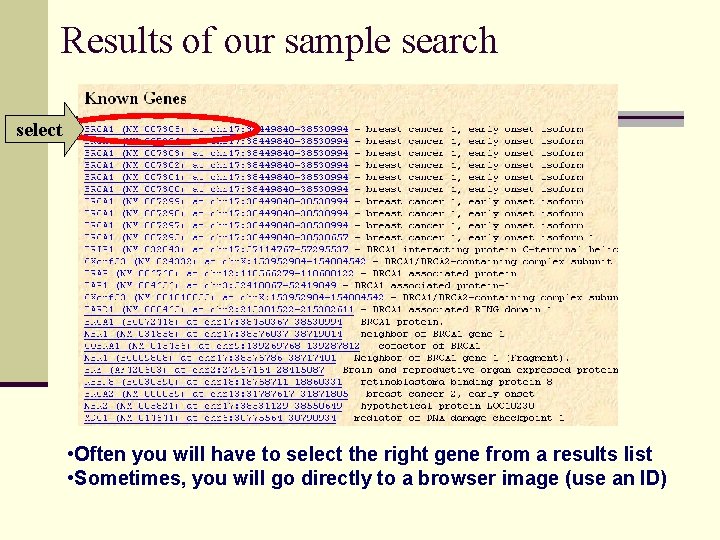
Results of our sample search select • Often you will have to select the right gene from a results list • Sometimes, you will go directly to a browser image (use an ID)
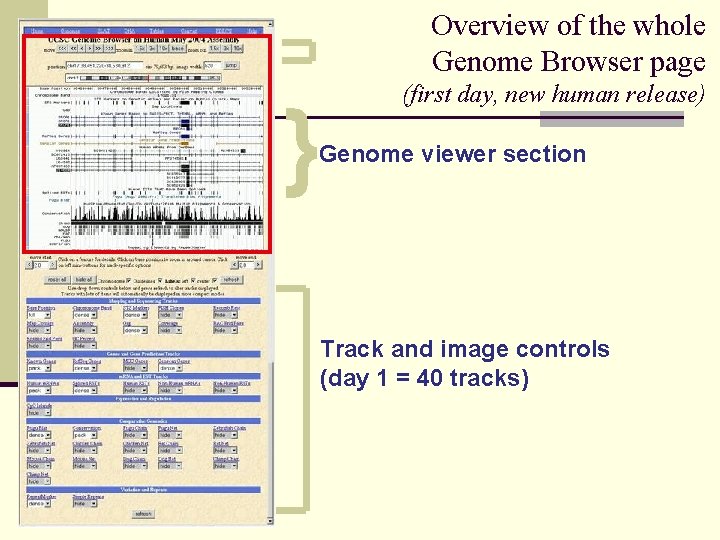
Overview of the whole Genome Browser page } (first day, new human release) Genome viewer section Track and image controls (day 1 = 40 tracks)

} Overview of the whole Genome Browser page (mature release) Genome viewer section Track and image controls (72 tracks = mature assembly) Groups of data Mapping and Sequencing Tracks Genes and Gene Prediction Tracks m. RNA and EST Tracks Expression and Regulation Comparative Genomics ENCODE Tracks Variation and Repeats

First sample Genome Viewer image Genome backbone STS markers Known genes Ref. Seq genes Gene predictions Full-length clones if available GB human ESTs, m. RNAs Microarray data if available species comparisons repeats

Different species, different tracks
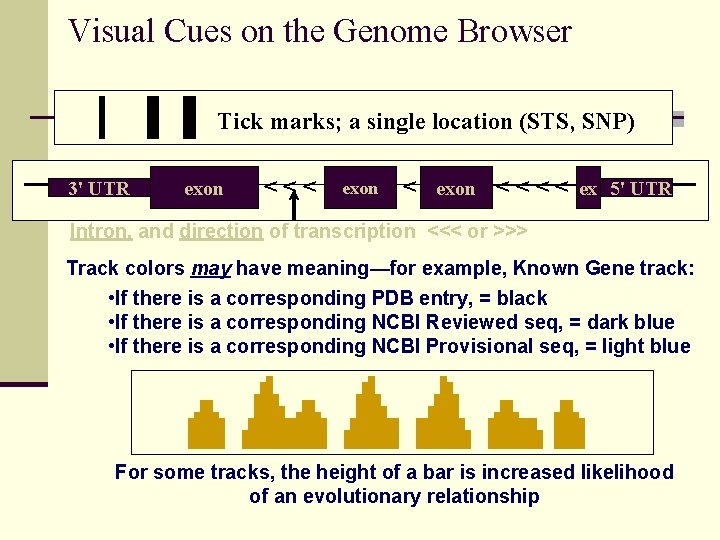
Visual Cues on the Genome Browser Tick marks; a single location (STS, SNP) 3' UTR exon <<< exon < < ex 5' UTR Intron, and direction of transcription <<< or >>> Track colors may have meaning—for example, Known Gene track: • If there is a corresponding PDB entry, = black • If there is a corresponding NCBI Reviewed seq, = dark blue • If there is a corresponding NCBI Provisional seq, = light blue For some tracks, the height of a bar is increased likelihood of an evolutionary relationship

Options for changing the images: Walk left or right Specify a position Zoom in Zoom out How wide on screen • Change your overview from the controls at the top • Change viewer window size • Use “base” to get right down to the nucleotides
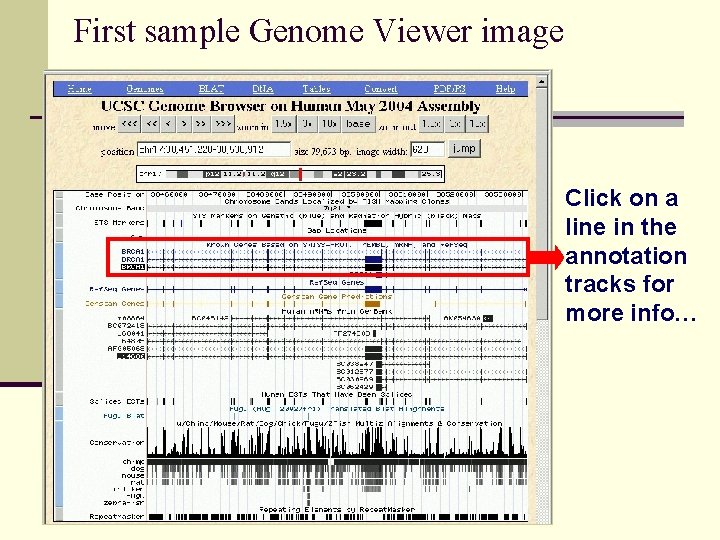
First sample Genome Viewer image Click on a line in the annotation tracks for more info…

Clicking an annotation line, new page of detailed information You will get detail for that single item Example: click on the BRCA 1 Black Known Genes line Click the line New web page opens Many links to more data about BRCA 1

informative description other resource links to sequences “Known gene” BRCA 1 sample page microarray data m. RNA secondary structure protein domains and structure homologs in other species Gene Ontology™ descriptions m. RNA descriptions pathways Not all genes have this much detail. Different annotation tracks carry different detail data.
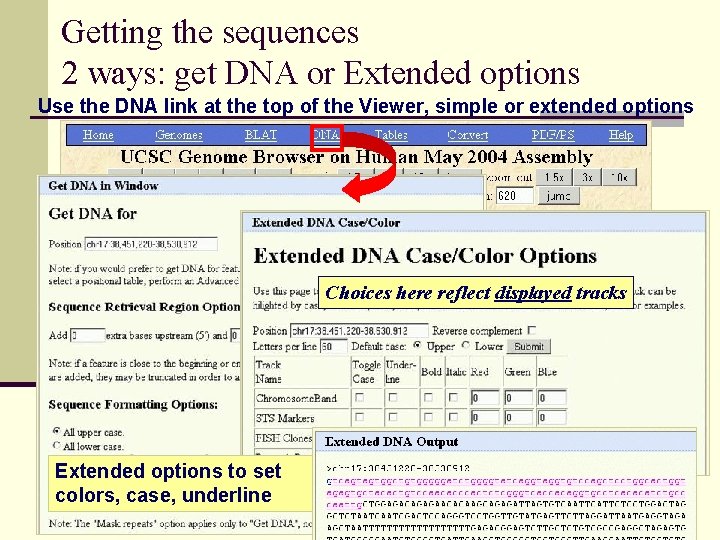
Getting the sequences 2 ways: get DNA or Extended options Use the DNA link at the top of the Viewer, simple or extended options Choices here reflect displayed tracks Extended options to set colors, case, underline
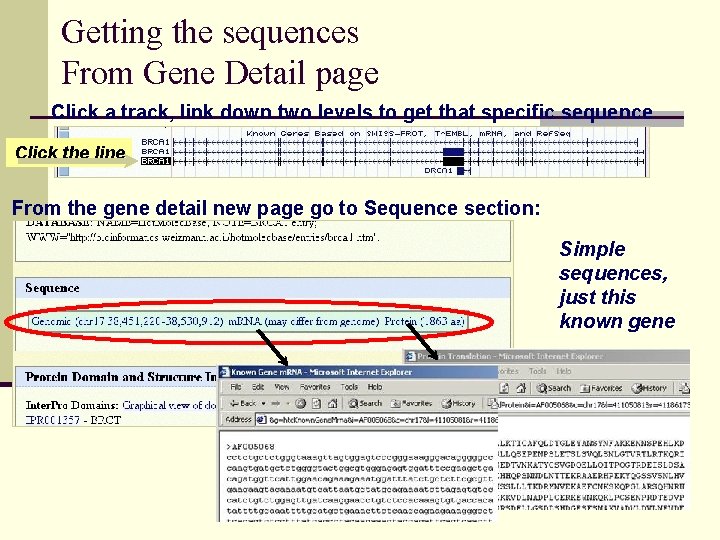
Getting the sequences From Gene Detail page Click a track, link down two levels to get that specific sequence Click the line From the gene detail new page go to Sequence section: Simple sequences, just this known gene

Annotation Track display options hide, dense, squish, pack, full…? Links to details and/or filters refresh to make changes Change track view Some data is ON or OFF by default Menu links to info about tracks—content and methods. You change the view with pulldown menus. After changing a pulldown, REFRESH to commit the change
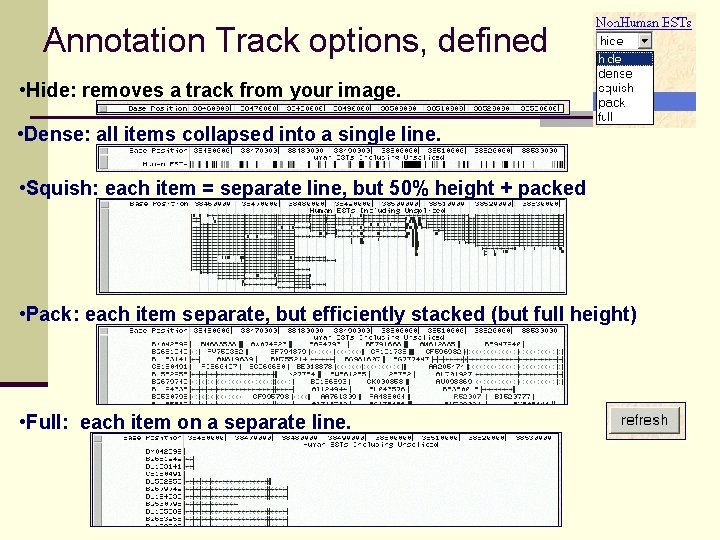
Annotation Track options, defined • Hide: removes a track from your image. • Dense: all items collapsed into a single line. • Squish: each item = separate line, but 50% height + packed • Pack: each item separate, but efficiently stacked (but full height) • Full: each item on a separate line.
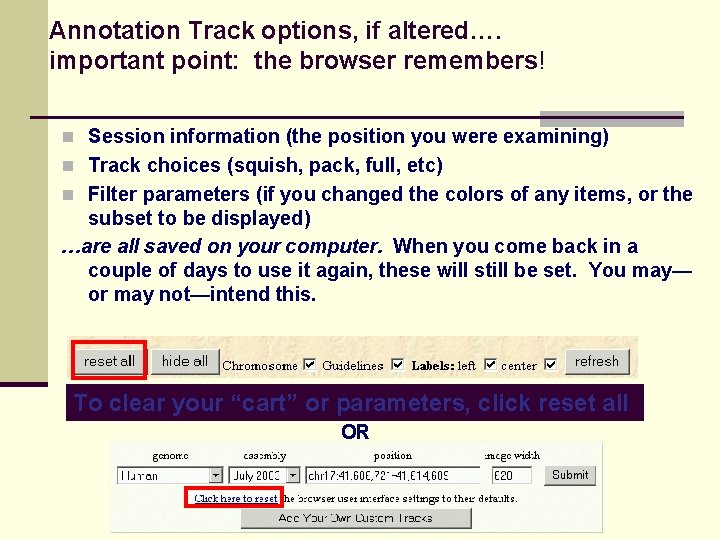
Annotation Track options, if altered…. important point: the browser remembers! n Session information (the position you were examining) n Track choices (squish, pack, full, etc) n Filter parameters (if you changed the colors of any items, or the subset to be displayed) …are all saved on your computer. When you come back in a couple of days to use it again, these will still be set. You may— or may not—intend this. To clear your “cart” or parameters, click reset all OR
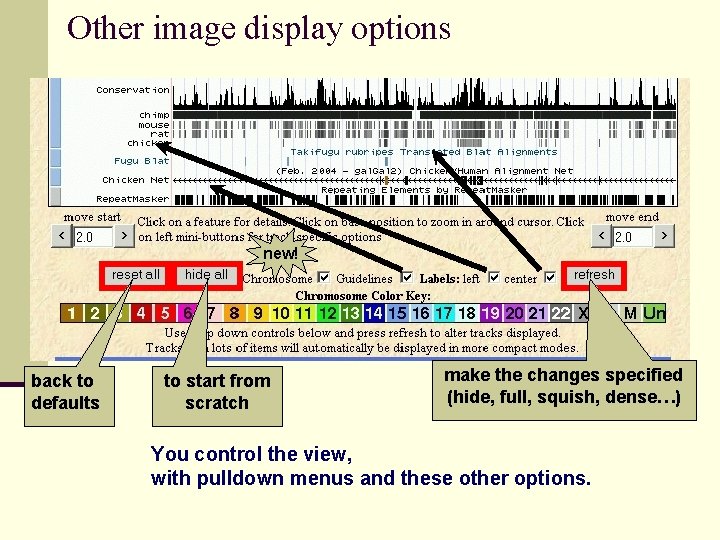
Other image display options new! back to defaults to start from scratch make the changes specified (hide, full, squish, dense…) You control the view, with pulldown menus and these other options.
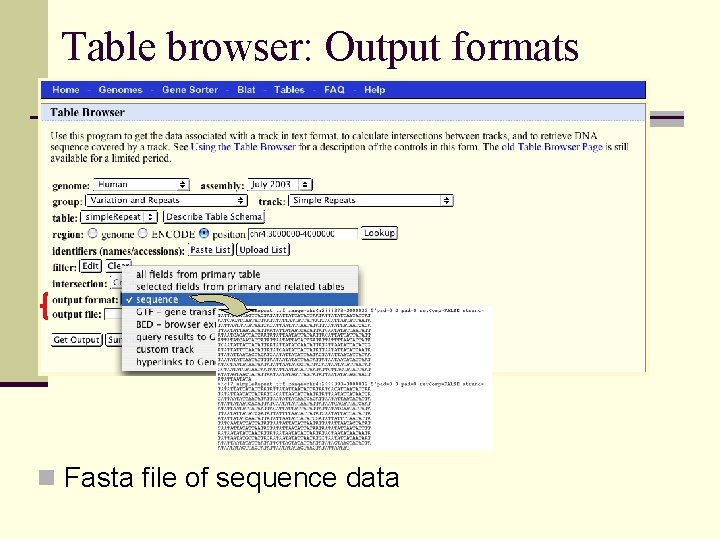
Table browser: Output formats n Fasta file of sequence data

SFRS 5 examples
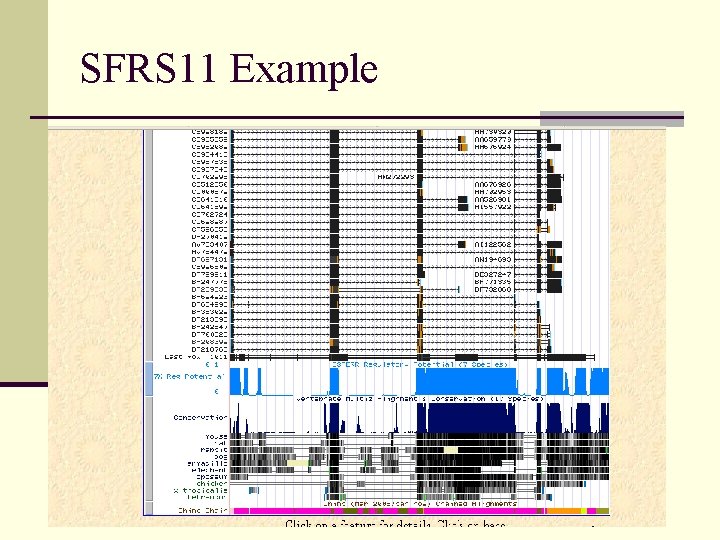
SFRS 11 Example
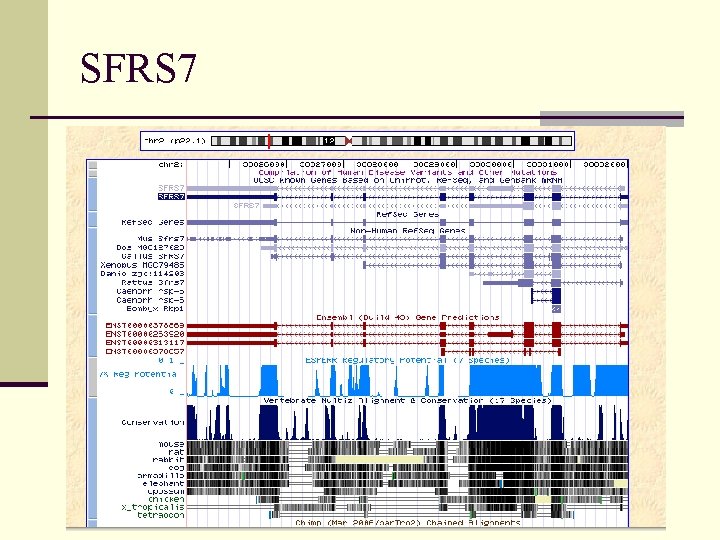
SFRS 7
- Slides: 26