Tools in Bioinformatics Genome Browsers Retrieving genomic information

Tools in Bioinformatics Genome Browsers

Retrieving genomic information n Previous lesson(s): annotation-based perspective of search/data Today: genomic-based perspective: look at all the data from the prism of a specific chromosome location Next: sequence-based searches
Genome browsers n NCBI Map Viewer n n Ensembl n n http: //www. ncbi. nih. gov/mapview http: //www. ensembl. org/ UCSC Genome Browser n http: //genome. ucsc. edu/
Important note to slide users: PC users n Mac users To maintain the color schemes/cues and the animations, if you import these slides into other slide sets please click the checkbox in the Power. Point Insert window that maintains slide format. Otherwise important information may be lost. Copyright Open. Helix. No use or reproduction without express written consent 5

The UCSC Genome Browser Introduction Materials prepared by Mary Mangan, Ph. D. www. openhelix. com Updated: Q 1 2009 Version 16 a_0209 Copyright Open. Helix. No use or reproduction without express written consent 6

UCSC Genome Browser Agenda n n n n Introduction and Credits Basic Searches Understanding Displays Get Details or Sequences Sequence Searches (BLAT) Summary Exercises UCSC Genome Browser: http: //genome. ucsc. edu Copyright Open. Helix. No use or reproduction without express written consent 7

Organization of Genomic Data Annotation Tracks sequence Genome backbone: base position number chromosome band sts sites gap locations known genes Links out to more data predicted genes microarray/expression data evolutionary conservation SNPs repeated regions more… Copyright Open. Helix. No use or reproduction without express written consent 8

A Sample of the UCSC Genome Browser gene details official sequence Annotation Tracks comparisons SNPs Copyright Open. Helix. No use or reproduction without express written consent 9

UCSC Genome Browser Agenda n n n n Introduction and Credits Basic Searches Understanding Displays Get Details or Sequences Sequence Searches (BLAT) Summary Exercises UCSC Genome Browser: http: //genome. ucsc. edu Copyright Open. Helix. No use or reproduction without express written consent 11

The UCSC Homepage: http: //genome. ucsc. edu navigate General information Specific information— new features, current status, etc. Copyright Open. Helix. No use or reproduction without express written consent 12
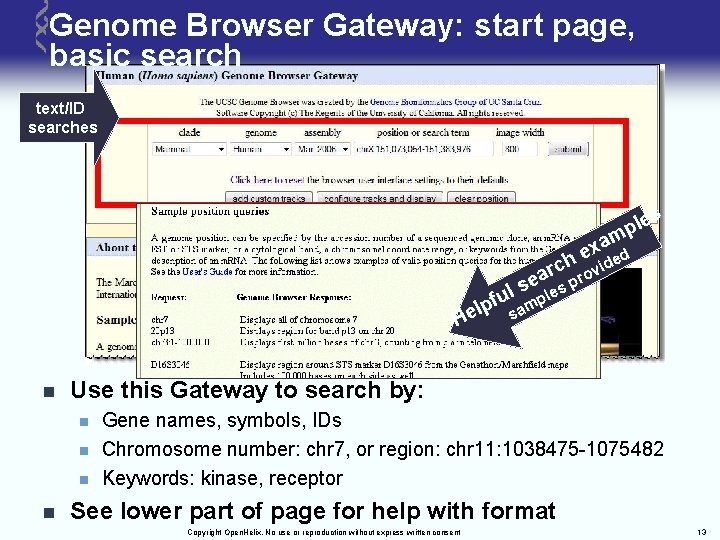
Genome Browser Gateway: start page, basic search text/ID searches le p m s xa d e ch ide ar prov e s les ul f amp p l e s H n Use this Gateway to search by: n n Gene names, symbols, IDs Chromosome number: chr 7, or region: chr 11: 1038475 -1075482 Keywords: kinase, receptor See lower part of page for help with format Copyright Open. Helix. No use or reproduction without express written consent 13
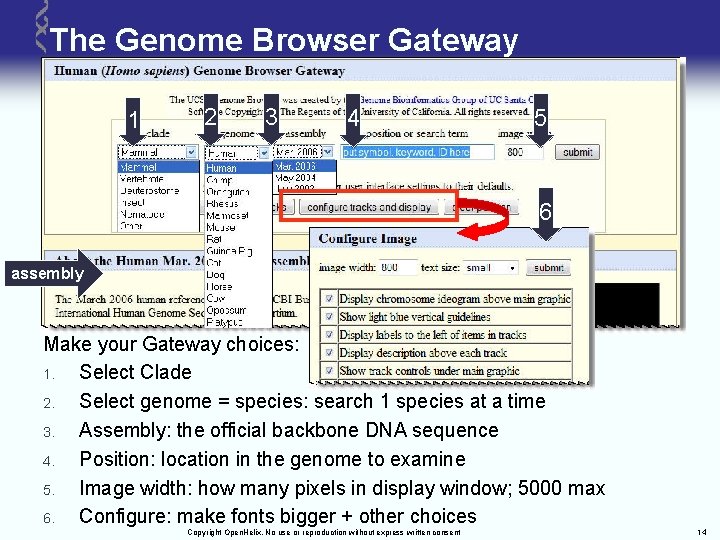
The Genome Browser Gateway 1 2 3 4 5 6 assembly Make your Gateway choices: 1. Select Clade 2. Select genome = species: search 1 species at a time 3. Assembly: the official backbone DNA sequence 4. Position: location in the genome to examine 5. Image width: how many pixels in display window; 5000 max 6. Configure: make fonts bigger + other choices Copyright Open. Helix. No use or reproduction without express written consent 14
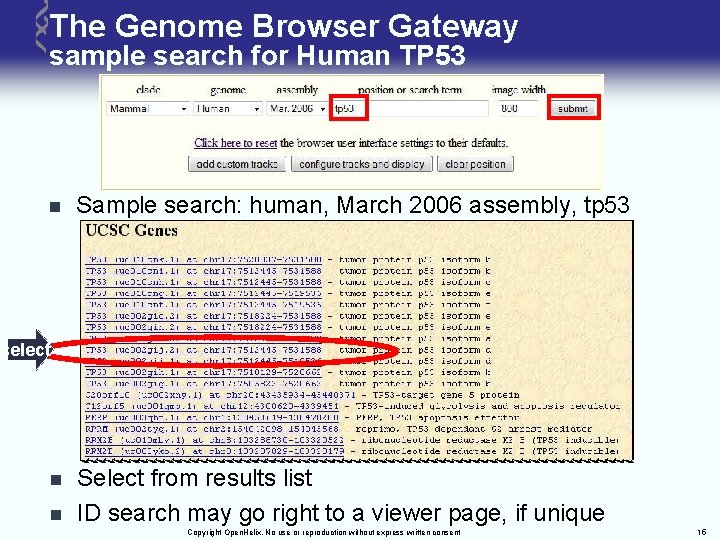
The Genome Browser Gateway sample search for Human TP 53 n Sample search: human, March 2006 assembly, tp 53 select n n Select from results list ID search may go right to a viewer page, if unique Copyright Open. Helix. No use or reproduction without express written consent 15

UCSC Genome Browser Agenda n n n n Introduction and Credits Basic Searches Understanding Displays Get Details or Sequences Sequence Searches (BLAT) Summary Exercises UCSC Genome Browser: http: //genome. ucsc. edu Copyright Open. Helix. No use or reproduction without express written consent 16

} Overview of the Whole Genome Browser Page Genome viewer section (mature release) Groups of data (Tracks) Mapping and Sequencing Tracks Phenotype and Disease Tracks Genes and Gene Prediction Tracks (including sno/mi. RNA data) m. RNA and EST Tracks Expression (such as microarray) Regulation (including TFBS) Comparative Genomics • As a group • Individual species Variation and Repeats (including SNPs, copy number variation) ENCODE Tracks Copyright Open. Helix. No use or reproduction without express written consent 17

Different Species, Different Tracks, Same Software n n Species may have different data tracks Layout, software, functions the same Copyright Open. Helix. No use or reproduction without express written consent 18

Sample Genome Viewer Image, TP 53 Region base position UCSC genes Ref. Seq genes MGC clones m. RNAs & ESTs many species compared single species compared SNPs repeats Copyright Open. Helix. No use or reproduction without express written consent 19

Visual Cues on the Genome Browser Tick marks; a single location (STS, SNP) 3' UTR exon <<< exon < < < <ex 5' UTR Intron and direction of transcription <<< or >>> Track colors may have meaning—for example, UCSC Gene track: • If there is a corresponding PDB entry = black • If there is a corresponding reviewed/validated seq = dark blue • If there is a non-Ref. Seq seq = lightest blue For some tracks, the height of a bar is increased likelihood of an evolutionary relationship (conservation track) Alignment indications (Conservation pairs: “chain” or “net” style) • Alignments = boxes, Gaps = lines Copyright Open. Helix. No use or reproduction without express written consent 20

Options for Changing Images: Upper Section Walk left or right Zoom in Specify a position Click to zoom 3 x and re-center n n n Zoom out Fonts, window, next item, more Change your view or location with controls at the top Use “base” to get right down to the nucleotides Configure: to change font, window size, more… n Next item, next exon navigation assistance can be turned on Copyright Open. Helix. No use or reproduction without express written consent 21
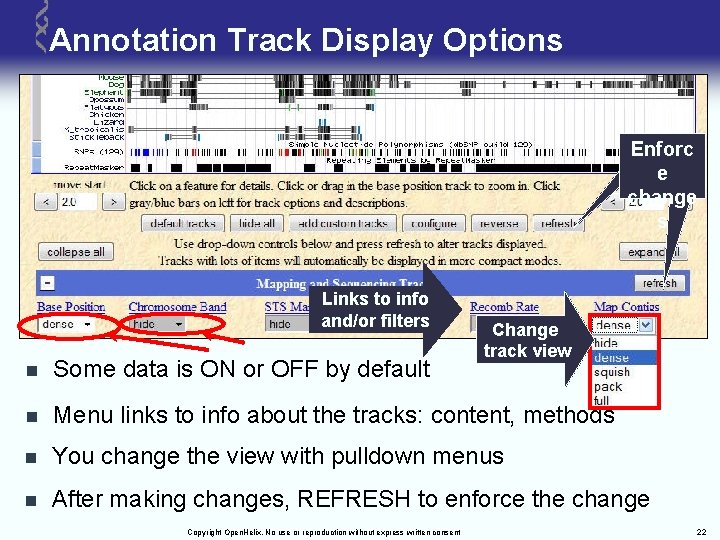
Annotation Track Display Options enforce Enforc change e change s s Links to info and/or filters Change track view n Some data is ON or OFF by default n Menu links to info about the tracks: content, methods n You change the view with pulldown menus n After making changes, REFRESH to enforce the change Copyright Open. Helix. No use or reproduction without express written consent 22

Annotation Track Options Defined n Hide: removes a track from view n Dense: all items collapsed into a single line n Squish: each item = separate line, but 50% height + packed n Pack: each item separate, but efficiently stacked (full height) n Full: each item on separate line Copyright Open. Helix. No use or reproduction without express written consent 23

Mid-page Options to Change Settings Flip display to Genomic 3’ 5’ Reset, back to defaults n n n Enforce any changes (hide, full, squish…) Start from scratch You control the views Use pulldown menus Configure options page Copyright Open. Helix. No use or reproduction without express written consent 24
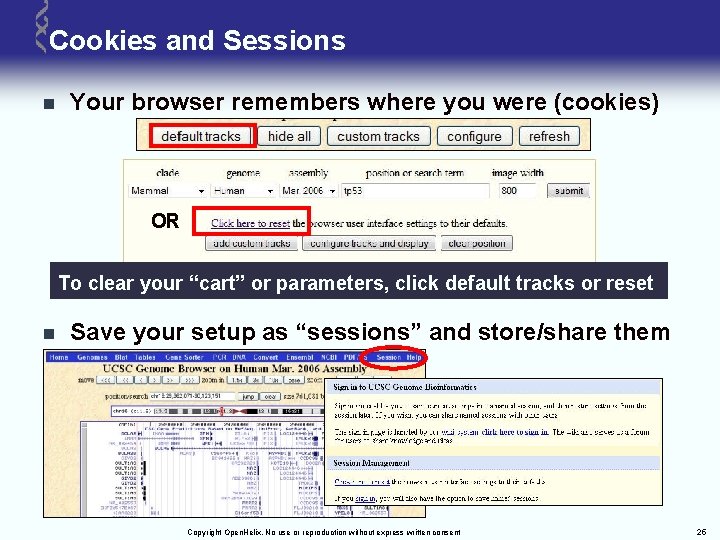
Cookies and Sessions n Your browser remembers where you were (cookies) OR To clear your “cart” or parameters, click default tracks or reset n Save your setup as “sessions” and store/share them Copyright Open. Helix. No use or reproduction without express written consent 25

UCSC Genome Browser Agenda n n n n Introduction and Credits Basic Searches Understanding Displays Get Details or Sequences Sequence Searches (BLAT) Summary Exercises UCSC Genome Browser: http: //genome. ucsc. edu Copyright Open. Helix. No use or reproduction without express written consent 26
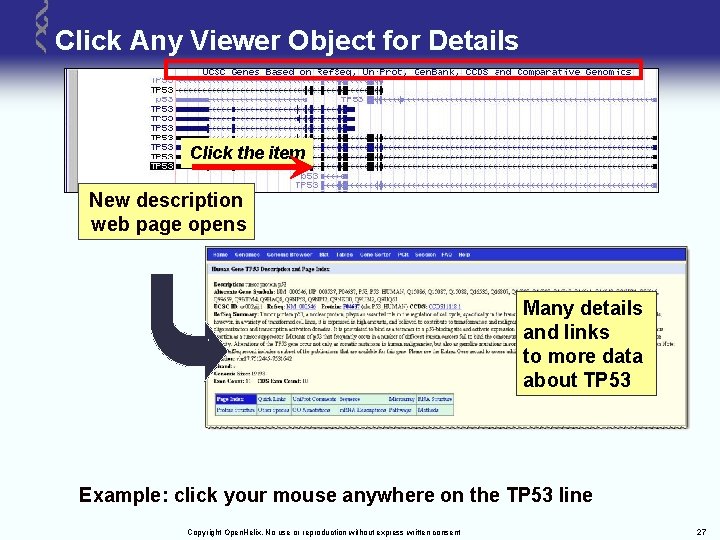
Click Any Viewer Object for Details Click the item New description web page opens Many details and links to more data about TP 53 Example: click your mouse anywhere on the TP 53 line Copyright Open. Helix. No use or reproduction without express written consent 27

informative Click description other resource links to sequences Annotation Track Item for Details Pages Not all genes have genetic association this much detail. studies comparative toxicology Different microarray data annotation tracks carry different data. m. RNA secondary structure protein domains/structure orthologs in other species Gene Ontology™ descriptions m. RNA descriptions pathways gene model Copyright Open. Helix. No use or reproduction without express written consent 28
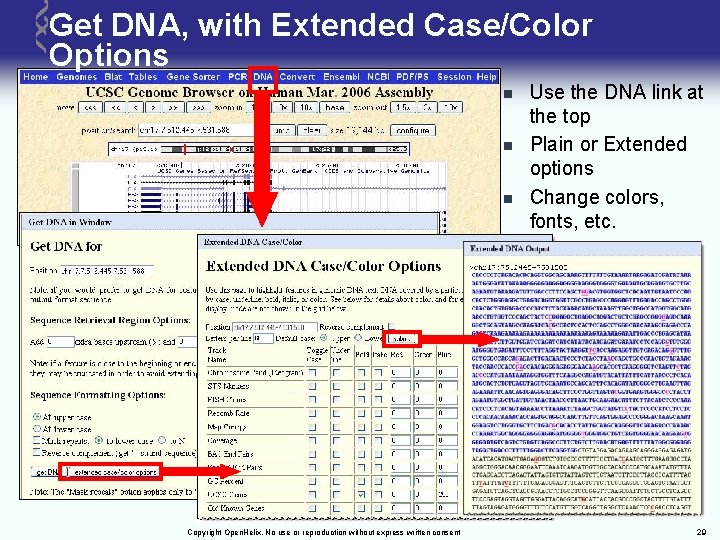
Get DNA, with Extended Case/Color Options n n n Copyright Open. Helix. No use or reproduction without express written consent Use the DNA link at the top Plain or Extended options Change colors, fonts, etc. 29
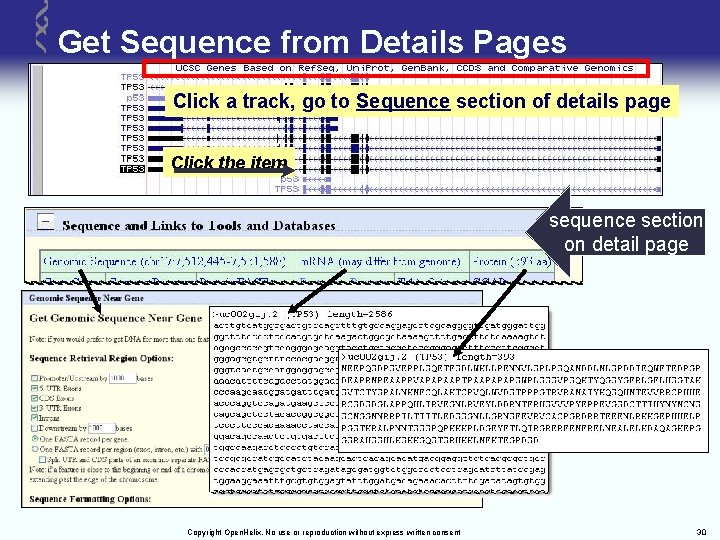
Get Sequence from Details Pages Click a track, go to Sequence section of details page Click the item sequence section on detail page Copyright Open. Helix. No use or reproduction without express written consent 30

UCSC Genome Browser Agenda n n n n Introduction and Credits Basic Searches Understanding Displays Get Details or Sequences Sequence Searches (BLAT) Summary Exercises UCSC Genome Browser: http: //genome. ucsc. edu Copyright Open. Helix. No use or reproduction without express written consent 31

Accessing the BLAT Tool BLAT = BLAST-like Alignment Tool n n n Rapid searches by INDEXING the entire genome Works best with high similarity matches See documentation and publication for details n Kent, WJ. Genome Res. 2002. 12: 656 Copyright Open. Helix. No use or reproduction without express written consent 32
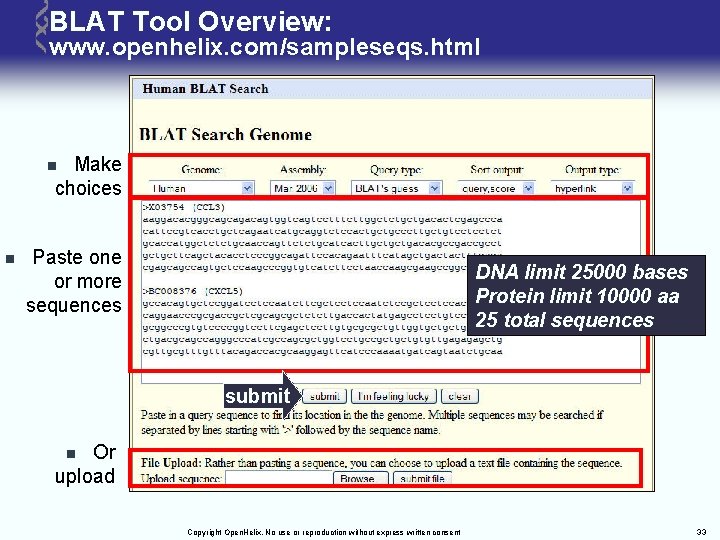
BLAT Tool Overview: www. openhelix. com/sampleseqs. html Make choices n n Paste one or more sequences DNA limit 25000 bases Protein limit 10000 aa 25 total sequences submit Or upload n Copyright Open. Helix. No use or reproduction without express written consent 33

n n sorting Results with demo sequences, settings default; sort = Query, Score n n go to alignment detail go to browser/viewer BLAT Results with Hyperlinks Score is a count of matches—higher number, better match Click browser to go to Genome Browser image location (next slide) Click details to see the alignment to genomic sequence (2 nd slide) Copyright Open. Helix. No use or reproduction without express written consent 34
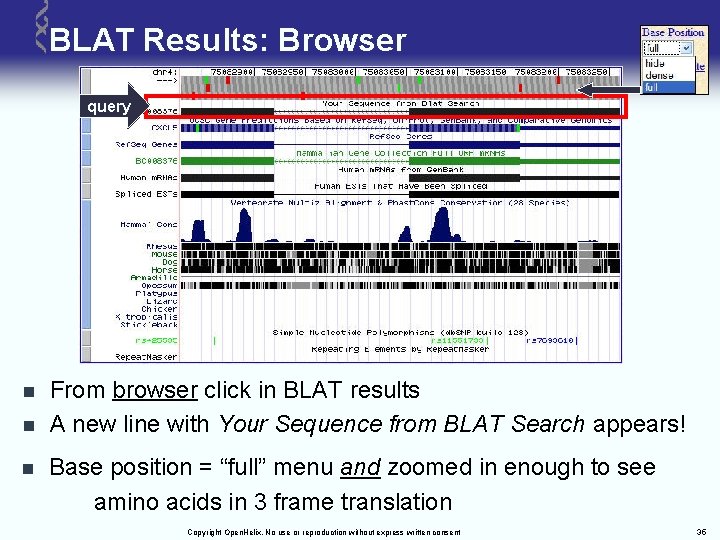
BLAT Results: Browser query n n n From browser click in BLAT results A new line with Your Sequence from BLAT Search appears! Base position = “full” menu and zoomed in enough to see amino acids in 3 frame translation Copyright Open. Helix. No use or reproduction without express written consent 35

BLAT Results, Alignment Details Your query Genomic match, color cues Side by Side Alignment yours genomic Copyright Open. Helix. No use or reproduction without express written consent 36

UCSC Genome Browser Agenda n n n n Introduction and Credits Basic Searches Understanding Displays Get Details or Sequences Sequence Searches (BLAT) Summary Exercises UCSC Genome Browser: http: //genome. ucsc. edu Copyright Open. Helix. No use or reproduction without express written consent 37
Introduction Summary n n n UCSC Genome Browser Visual cues and genomic context Many ways to alter your views Access to deeper data Access and use sequence data Copyright Open. Helix. No use or reproduction without express written consent 38
- Slides: 36