Tools Enzymes Vectors Host DNA to be cloned

Tools Ø Enzymes Ø Vectors Ø Host Ø DNA to be cloned

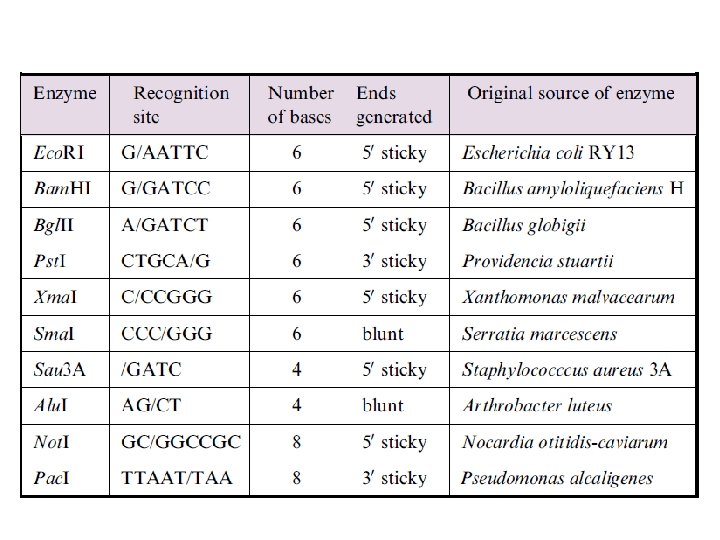
Enzymes §Nucleases §Polymerases §Ligases, §Modifying enzymes §Topoisomerase,

Exonucleases • Unit Enzyme 1 ug DNA at 37 C in 60 min in 50 ul reaction* Exonuclease. III Exonuclease. VII ss. DNA ds. DNA ss. DNA 3 -5 3 -5 and 5 -3 https: //www. neb. com/protocols/2012/12/07/optimizing-restriction-endonuclease-reactions

How they cut Exo III

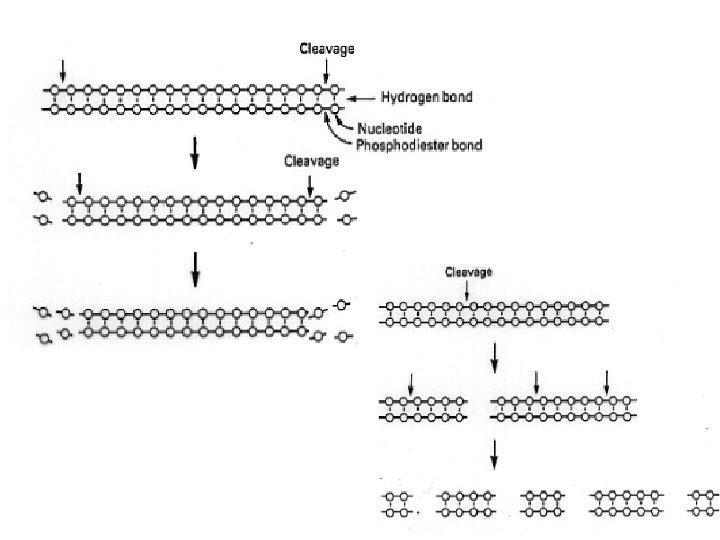
Endonucleases • Dnase I Ø Ø ss. DNA, ds. DNA template degradation in transcription reactions Removal of genomic DNA from RNA samples DNase I footprinting Nick Translation • Mung bean nucleases ss. DNA Ø Removal of single-stranded extensions (3' and 5') to leave ligat able blunt ends Ø Transcriptional mapping Ø Cleavage of hairpin loops

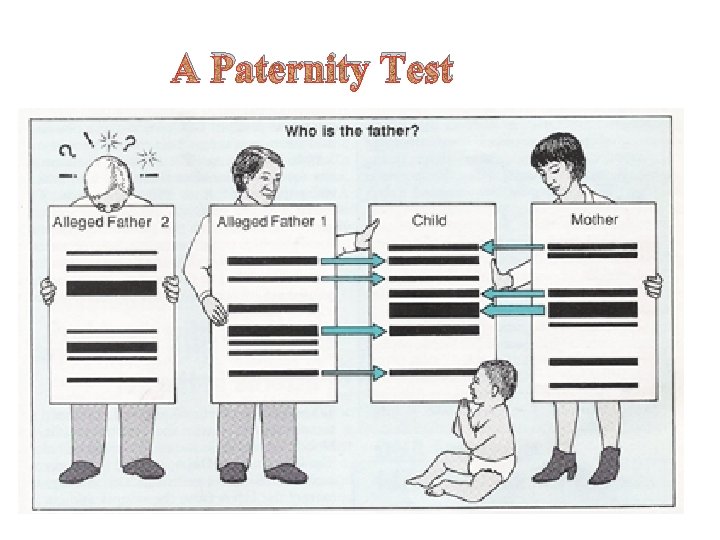
A Paternity Test
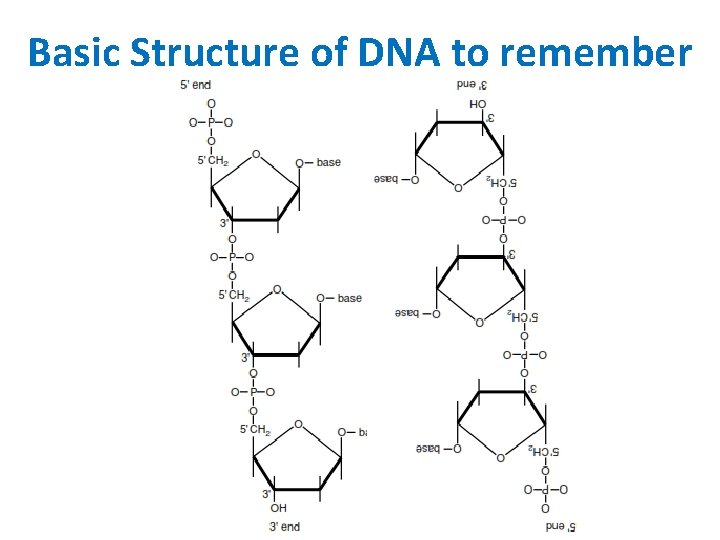
Basic Structure of DNA to remember

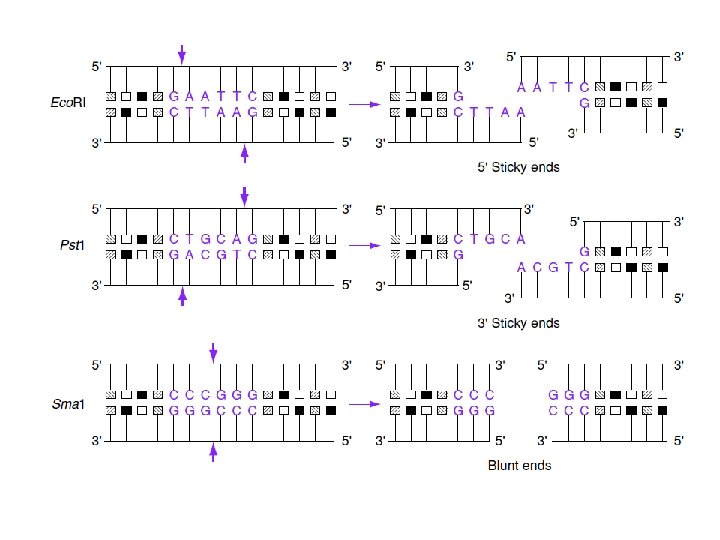
Iso schizomer Sph I (CGTAC^G) and Bbu I (CGTAC^G) Neo schizomer Sma I (CCC/GGG) and Xma I (C/CCGGG) Iso caudomer Nhe. I G*CTAG C and Avr. II C*CTAG G C GATC*G G GATC*C
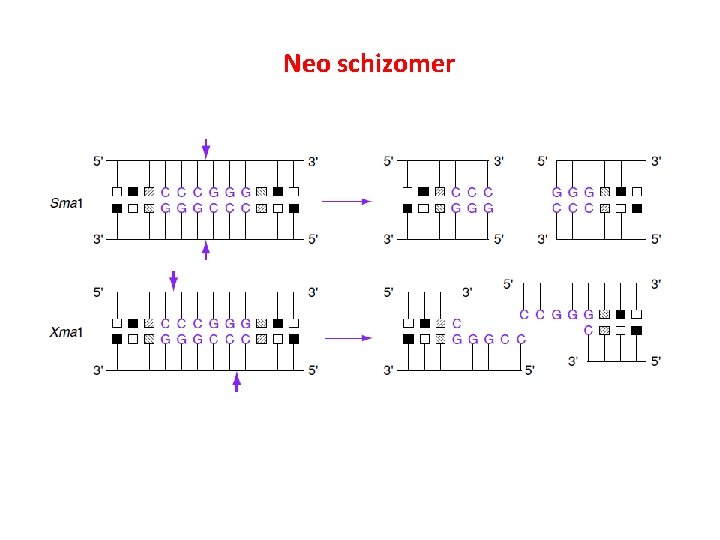
Neo schizomer
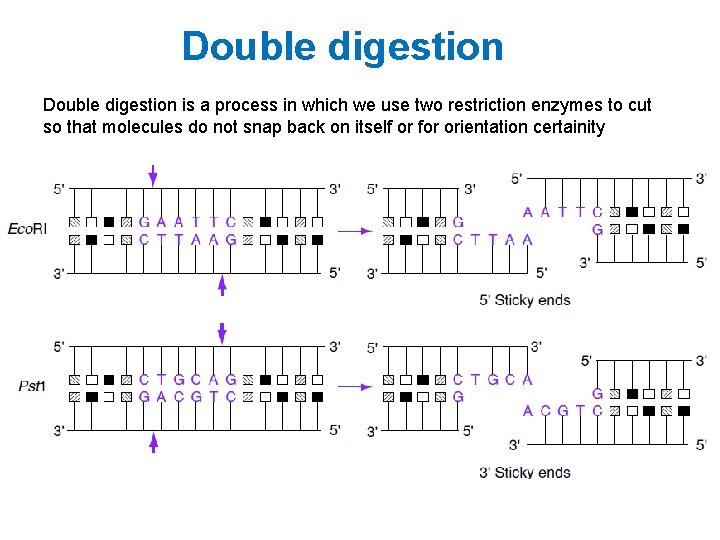
Double digestion is a process in which we use two restriction enzymes to cut so that molecules do not snap back on itself or for orientation certainity

RIBONUCLEASES RNase A Bovine pancreatic RNase A, Rana pipiens RNase. H RNase H family can be found in nearly all organisms, from archaea to bacteria and eukaryota. Rnase III Double stranded RNA degradation RNAi, micro. RNA
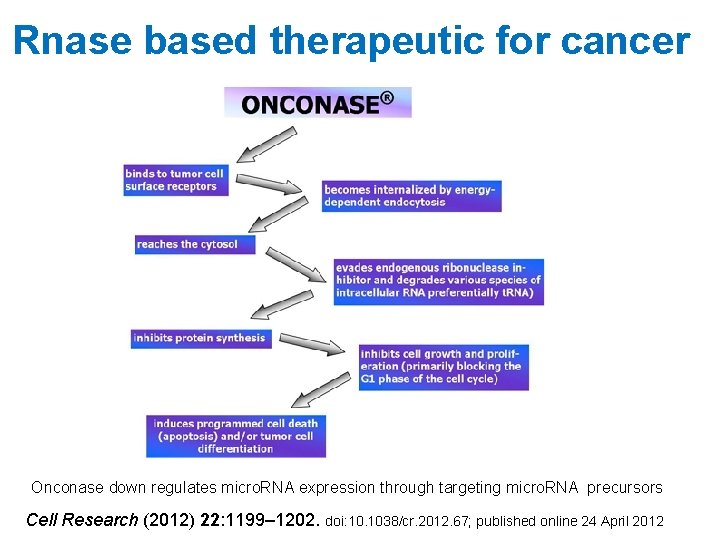
Rnase based therapeutic for cancer Onconase down regulates micro. RNA expression through targeting micro. RNA precursors Cell Research (2012) 22: 1199– 1202. doi: 10. 1038/cr. 2012. 67; published online 24 April 2012

Rnase H of HIV and HBV as an example 1 - Structural Basis for the Inhibition of RNase H Activity of HIV-1 Reverse Transcriptase by RNase H Active Site-Directed Inhibitors. 2010 Journal of Virology
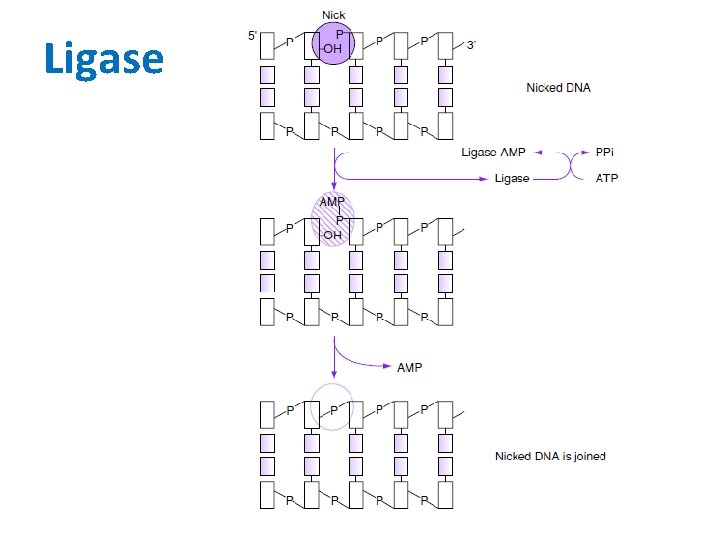
Ligase
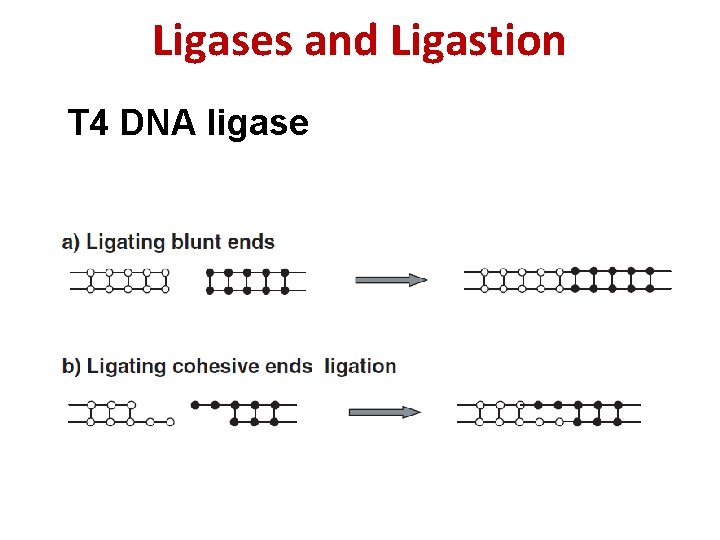
Ligases and Ligastion T 4 DNA ligase
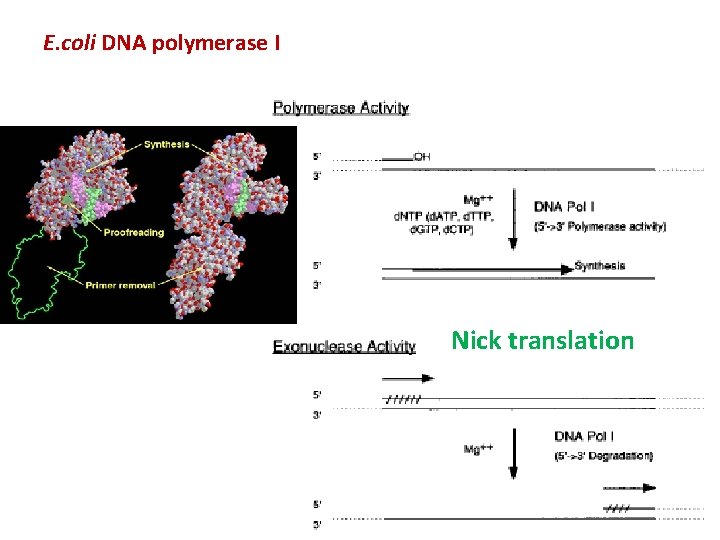
E. coli DNA polymerase I Nick translation
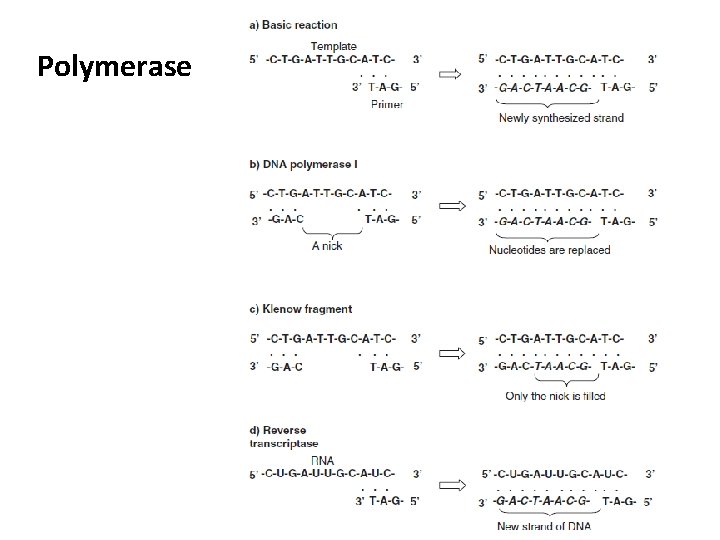
Polymerase
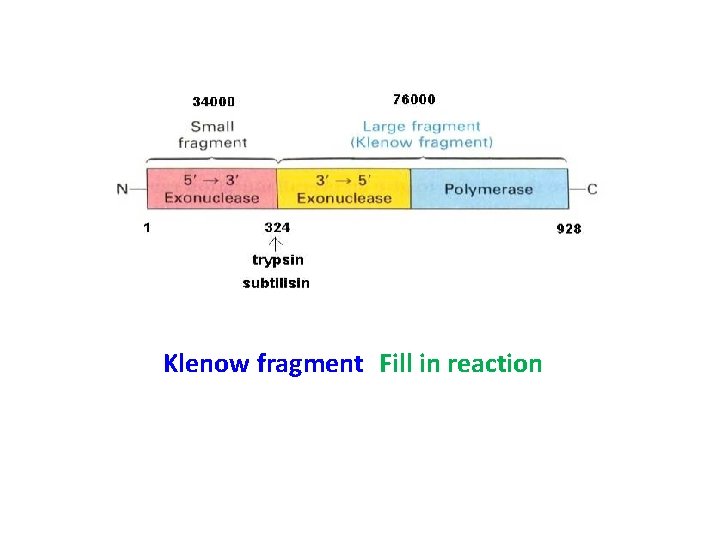
Klenow fragment Fill in reaction

Bacteriophage T 4 and T 7 polymerase Bacteriophage T 4 Active single-stranded 3'->5' exonuclease (ss DNA) (stronger than that of the Klenow fragment) Fill in Trimming back

The T 7 polymerase §Enzyme has very high proof reading and polymerization §The enzyme chemically or genetically modified §High processivity, and fast polymerase rate Used in DNA sequencing

Taq DNA polymerase Thermus aquaticus PCR optimization, Pfu DNA polymerase Pyrococcus furiosus PCR (if DNA has to use in cloning)

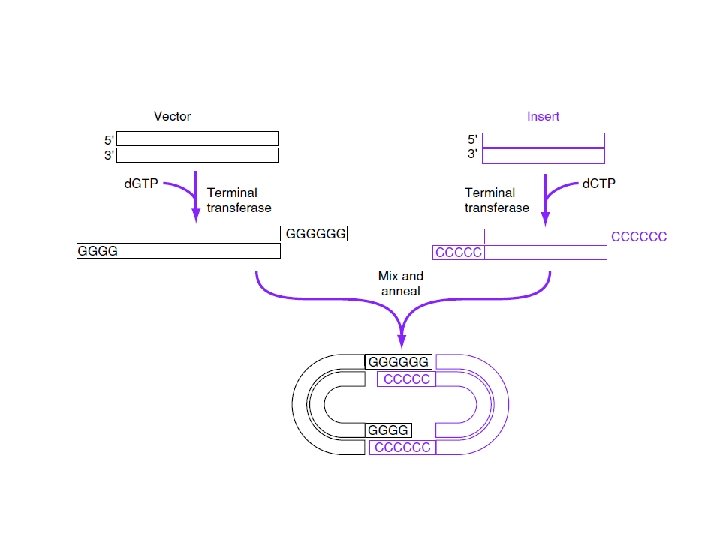
Terminal Deoxynuclotidyl Transferase Probe preparation tailing method

AMV reverse transcriptase • HIV-1 reverse transcriptase from human immunodeficiency virus • M-MLV reverse transcriptase from the Moloney murine leukemia virus • AMV reverse transcriptase from the avian myeloblastosis virus

RNA polymerases S 6 RNA Polymerases T 7 RNA Polymerases

Uses of different enzymes Primer removal from PCR mixtures: Exo 1 thermo prior to PCR product sequencing (see Reference 2) for one-tube "megaprimer" PCR mutagenesis (see Reference 3) Removal of single-stranded DNA containing a 3'-hydroxyl terminus from nucleic acid mixtures Assay for the presence of single-stranded DNA with a 3'-hydroxyl terminus (see Reference 4) http: //www. thermoscientificbio. com/dna-and-rna-modifying-enzymes/exonuclease-i/
- Slides: 33