Tomato Chromosome 8 sequencing at Kazusa DNA Research








- Slides: 12

Tomato Chromosome 8 sequencing at Kazusa DNA Research Institute Erika Asamizu

Sequencing efforts in Japan Chiba & National Institute of Vegetable and Tea Science

Status of chr. 8 sequencing Status Nr. BACs Finished 21 (2, 485, 895 bp) Phase 2 2 Assembling 8 Sequencing 11 Total 42

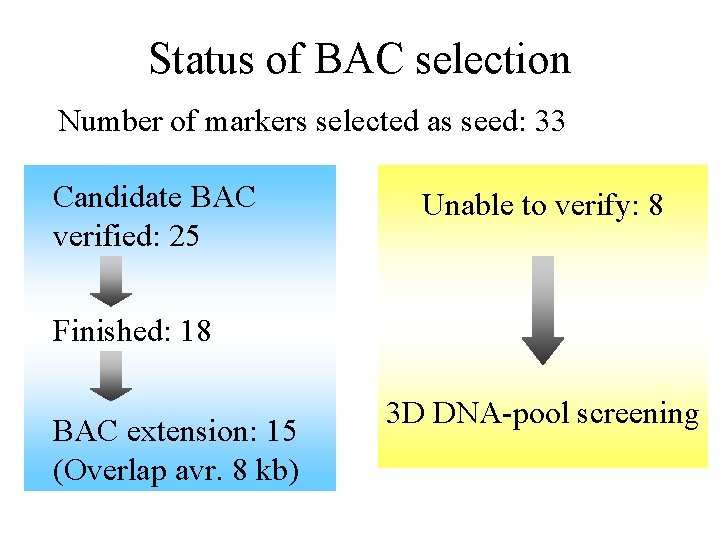
Status of BAC selection Number of markers selected as seed: 33 Candidate BAC verified: 25 Unable to verify: 8 Finished: 18 BAC extension: 15 (Overlap avr. 8 kb) 3 D DNA-pool screening

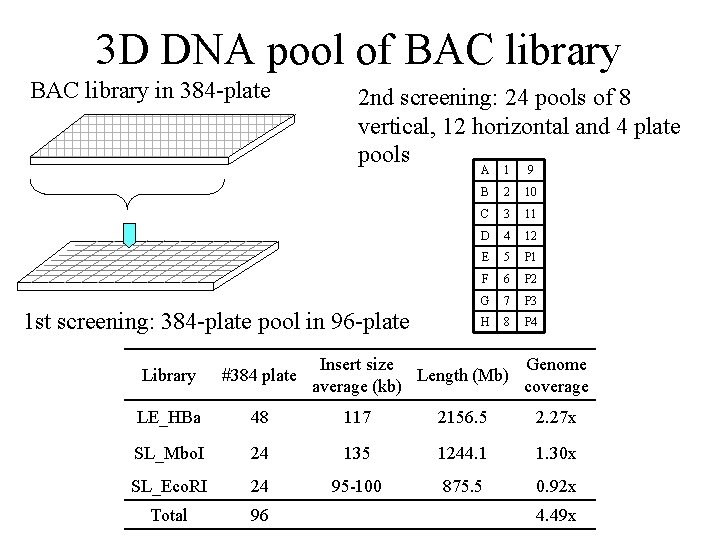
3 D DNA pool of BAC library in 384 -plate 2 nd screening: 24 pools of 8 vertical, 12 horizontal and 4 plate pools A 1 9 1 st screening: 384 -plate pool in 96 -plate B 2 10 C 3 11 D 4 12 E 5 P 1 F 6 P 2 G 7 P 3 H 8 P 4 Insert size Genome Length (Mb) average (kb) coverage Library #384 plate LE_HBa 48 117 2156. 5 2. 27 x SL_Mbo. I 24 135 1244. 1 1. 30 x SL_Eco. RI 24 95 -100 875. 5 0. 92 x Total 96 4. 49 x

FISH localization of ambiguously mapped BAC c. LET-5 -O 24 c. LEB-7 -J 7 HBa 0076 J 13 (58 kb) HBa 0059 L 14 (60 kb) HBa 0157 P 11 (124 kb) Marker Chr c. M Confidence Type Anchored BAC c. LET-5 -O 24 8 47 F(LOD 3) EST HBa 0076 J 13 c. LEB-7 -J 7 3 128 I RFLP HBa 0059 L 14 FISH of HBa 0157 P 11 is being performed in Colorado Sate U.

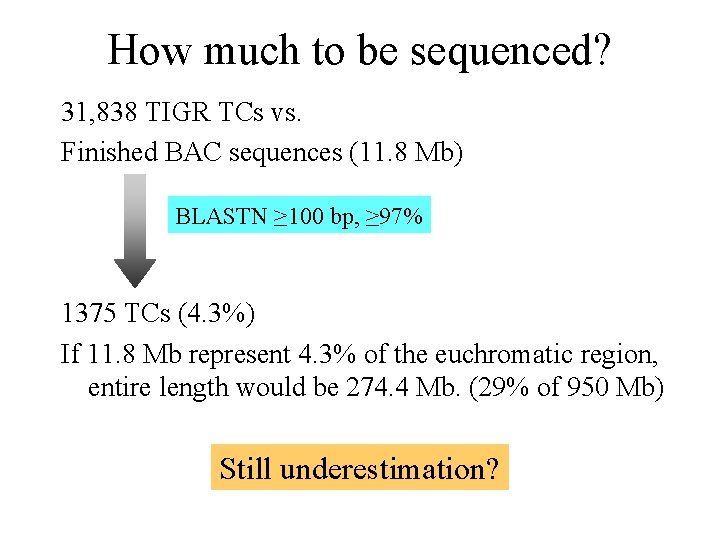
How much to be sequenced? 31, 838 TIGR TCs vs. Finished BAC sequences (11. 8 Mb) BLASTN ≥ 100 bp, ≥ 97% 1375 TCs (4. 3%) If 11. 8 Mb represent 4. 3% of the euchromatic region, entire length would be 274. 4 Mb. (29% of 950 Mb) Still underestimation?

Repeats in gene spaces 30, 288 BAC ends vs. 14, 229 repeat sequences (TIGR_Sol. Ath_repeat, mips_repeat_collection, SGN repeat collection) BLASTN E≤ 1 e-10 17, 412 BAC ends (57. 5%) hit repeat sequences. Euchromatin could be longer because of dispersed type of repeats?

Observed problem HBa 0127 P 15: Anchored to C 2_At 4 g 31130 (Verified) Finished length was only 39, 599 bp, and did NOT contain the marker sequence. Repeat 1 2 3 4 The digested pattern indicates that the length of sequenced BAC is ~40 kb. 1. λHind III 2. BAC/Not I 3. BAC/Xho I 4. 1 kb ladder Where did the marker go?

Possible problem caused by dispersed type of repeats We found the marker sequence in the shotgun library reads, but it was not incorporated into the finished assembly. Deleted clone was preferentially replicated in host cell? Deletion has occurred because of the repeat?

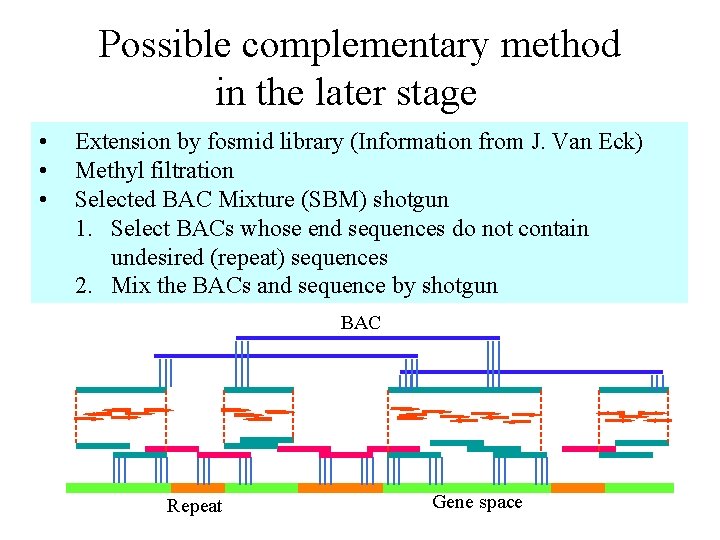
Possible complementary method in the later stage • • • Extension by fosmid library (Information from J. Van Eck) Methyl filtration Selected BAC Mixture (SBM) shotgun 1. Select BACs whose end sequences do not contain undesired (repeat) sequences 2. Mix the BACs and sequence by shotgun BAC Repeat Gene space

Plant genome sequencing team National Institute of Vegetable and Tea Science Satoshi Tabata Molecular Genetics and Physiology team Libraries Sequencing Shusei Sato Takakazu Kaneko Erika Asamizu Akiko Watanabe Shigemi Sasamoto Akiko Muraki Sayaka Shimpo Yoshie Kishida Naoko Torikai Naomi Nakazaki Informatics Midori Kato Yasukazu Nakamura Manabu Yamada Mitsuyo Kohara Shinobu Nakayama Tsunakazu Fujishiro Mayumi Iriguchi Kumiko Kawashima Yoshimi Shimizu Chiharu Minami Chika Takahashi Kyoko Hayashi Hiroyuki Fukuoka Satomi Negoro Yumika Kitamura