Tom Kepler Santa Fe Institute Normalization and Analysis



![Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA](https://slidetodoc.com/presentation_image_h2/8e7788919ad719f62778b237ae060cae/image-6.jpg)
![Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA](https://slidetodoc.com/presentation_image_h2/8e7788919ad719f62778b237ae060cae/image-7.jpg)
![Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA](https://slidetodoc.com/presentation_image_h2/8e7788919ad719f62778b237ae060cae/image-8.jpg)












- Slides: 36

Tom Kepler Santa Fe Institute Normalization and Analysis of DNA Microarray Data by Self-Consistency and Local Regression kepler@santafe. edu


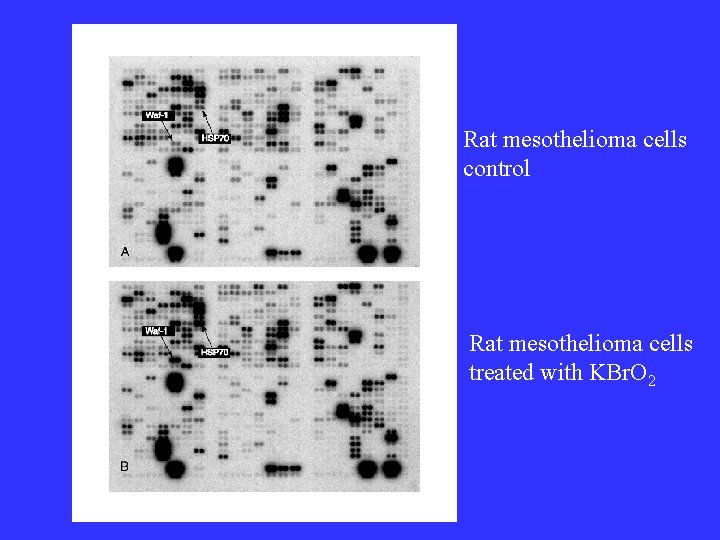
Rat mesothelioma cells control Rat mesothelioma cells treated with KBr. O 2

Normalization Method to be improved: 1. Assume that some genes will not change under the treatment under investigation. 2. Identify these core genes in advance of the experiment. 3. Normalize all genes against these genes assuming they do not change

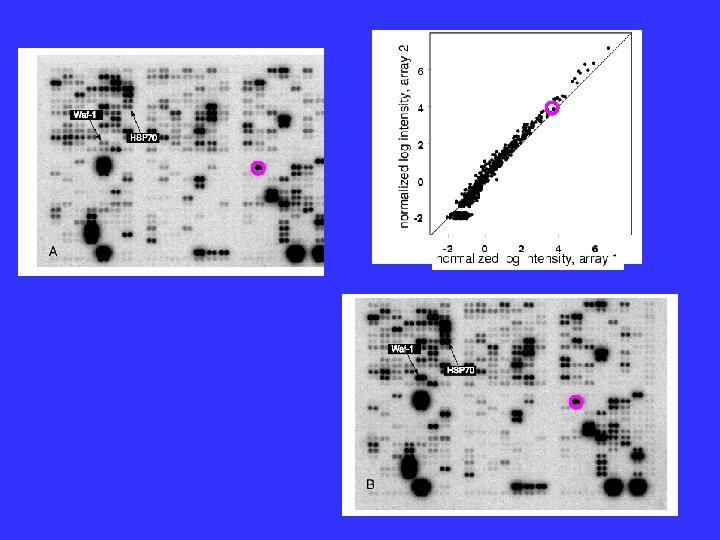
Normalization New Method: 1. Assume that some genes will not change under the treatment under investigation. 2. Choose these core genes arbitrarily. 3. Normalize (provisionally) all genes against these genes assuming they do not change. 4. Determine which genes do not change under this normalization. 5. Make this set the new core. If this core differs from the previous core, go to 3. Else, done.
![Error Model I spot intensity m RNA concentration of specific m RNA Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA](https://slidetodoc.com/presentation_image_h2/8e7788919ad719f62778b237ae060cae/image-6.jpg)
Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA c = normalization constant
![Error Model I spot intensity m RNA concentration of specific m RNA Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA](https://slidetodoc.com/presentation_image_h2/8e7788919ad719f62778b237ae060cae/image-7.jpg)
Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA c = normalization constant = lognormal multiplicative error
![Error Model I spot intensity m RNA concentration of specific m RNA Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA](https://slidetodoc.com/presentation_image_h2/8e7788919ad719f62778b237ae060cae/image-8.jpg)
Error Model I = spot intensity [m. RNA] = concentration of specific m. RNA c = normalization constant = lognormal multiplicative error index 1, i: treatment group index 2, j: replicate within treatment index 3, k: spot (gene)

Y = log spot intensity = mean log concentration of specific m. RNA = treatment effect (conc. specific m. RNA) = normalization constant = normal additive error index 1, i: treatment group index 2, j: replicate within treatment index 3, k: spot (gene)

Model: Identifiability constraints: Estimate by ordinary least squares:

Model: Identifiability constraints: But note: cannot identify between a and d

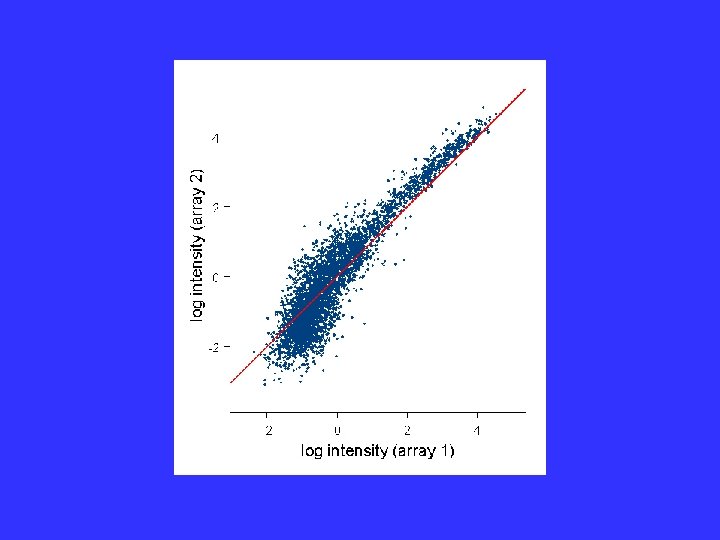
Self-consistency: The weight wk( ) is small if the kth gene is judged to be changed; close to one if it is judged to be unchanged. Procedure is iterative.




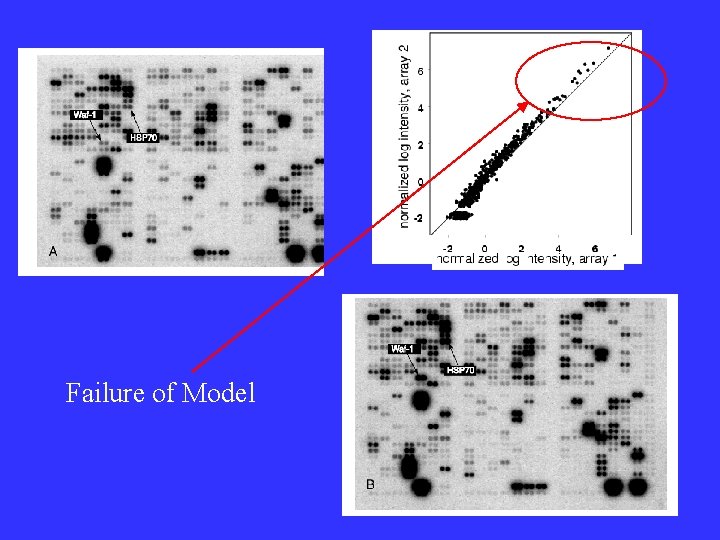
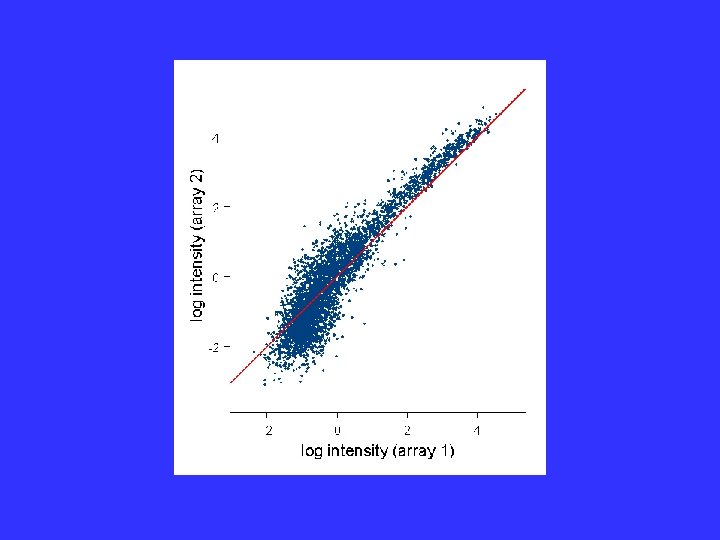
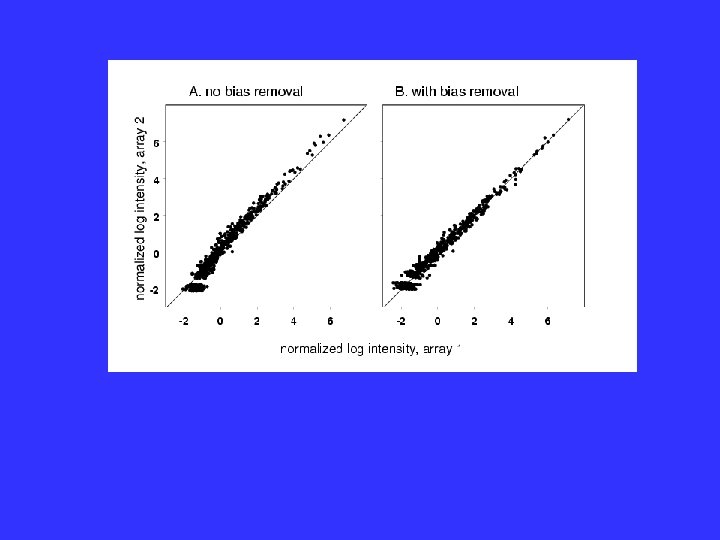
Failure of Model


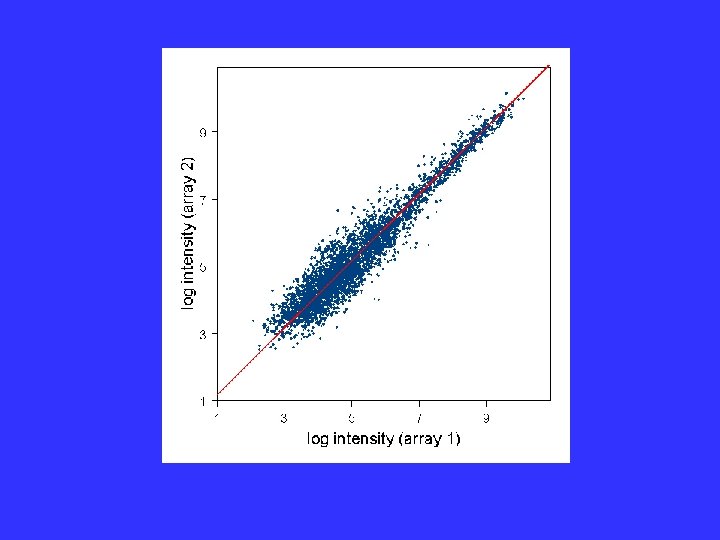
Generalized Model The normalization ij( k) and the heteroscedasticity function gij( k) are slowly varying functions of the intensity, . Estimate by Local Regression

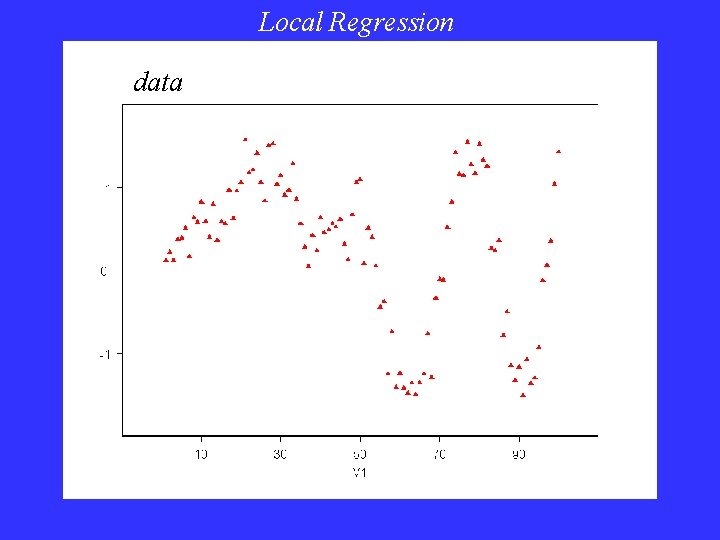
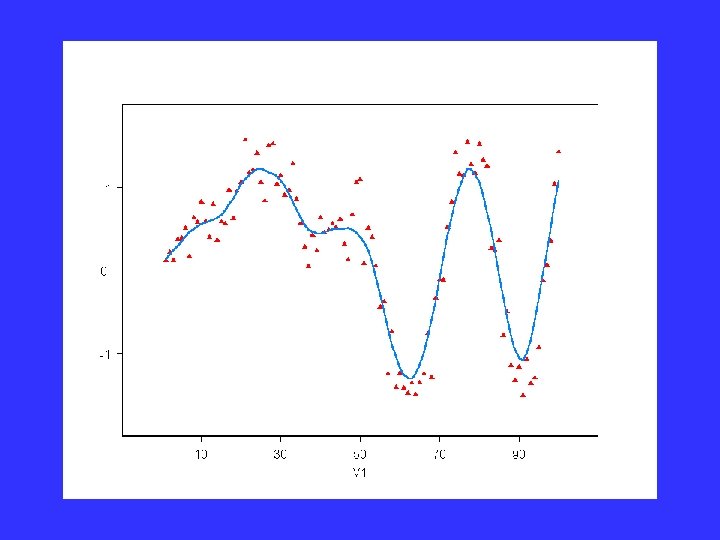
Local Regression data

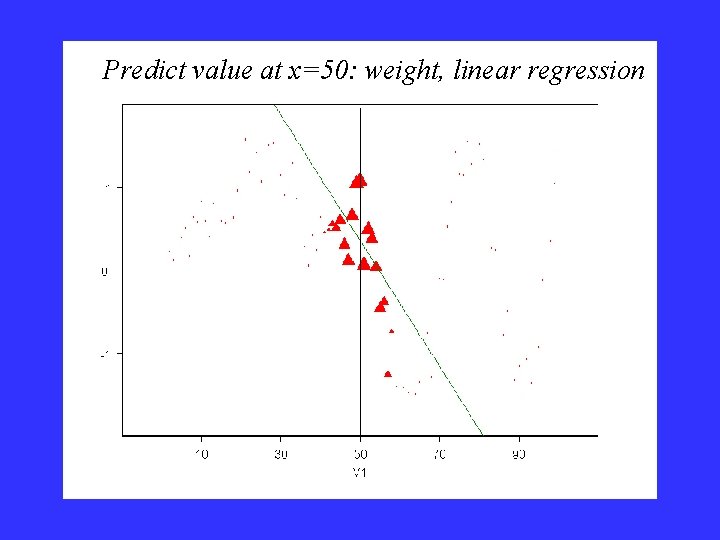
Predict value at x=50: weight, linear regression

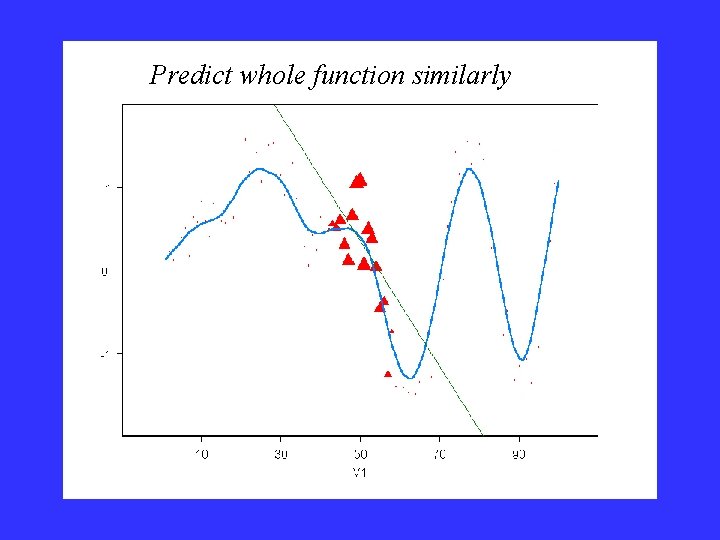
Predict whole function similarly


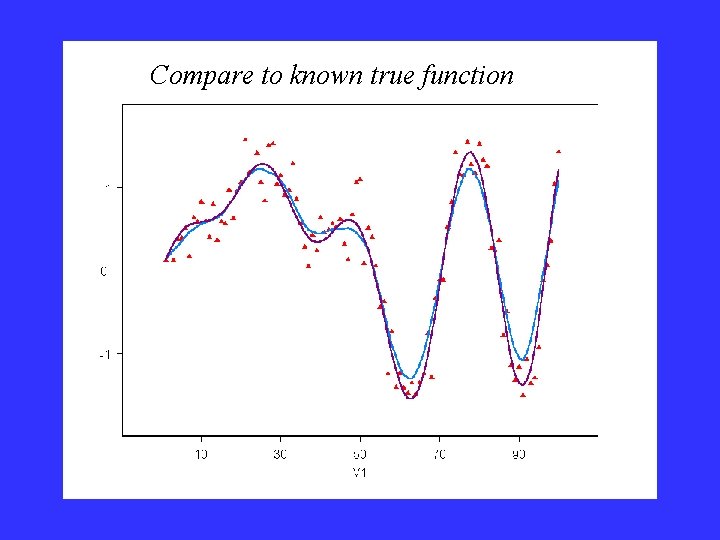
Compare to known true function




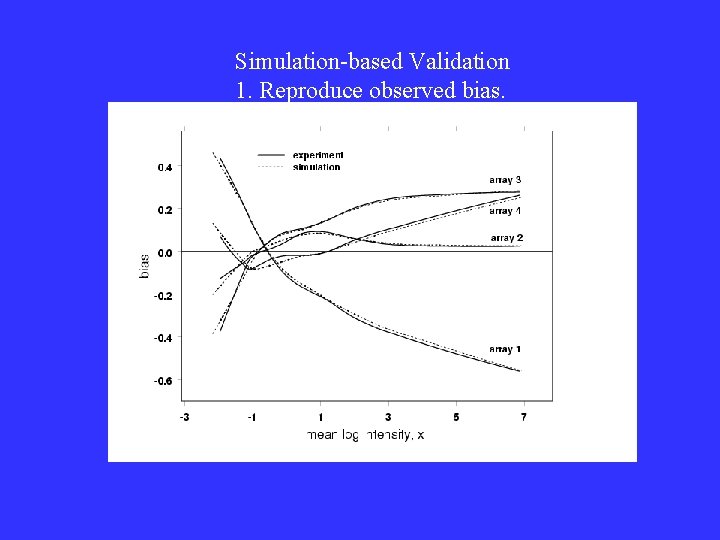
Simulation-based Validation 1. Reproduce observed bias.

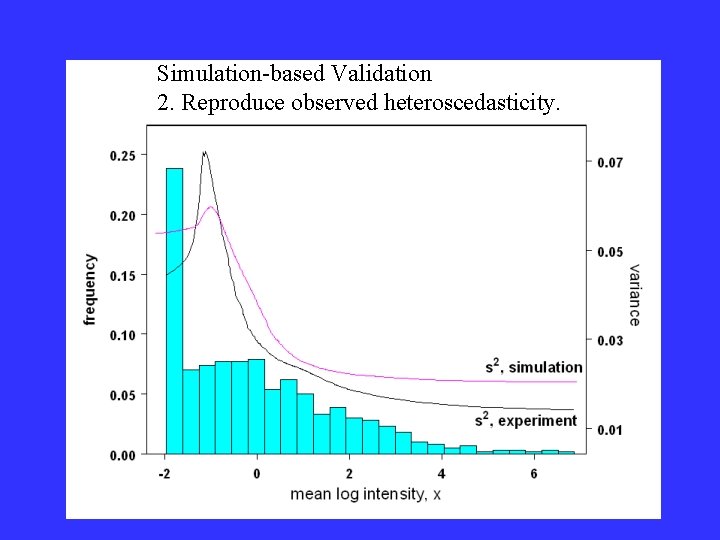
Simulation-based Validation 2. Reproduce observed heteroscedasticity.

Test based on z statistic:

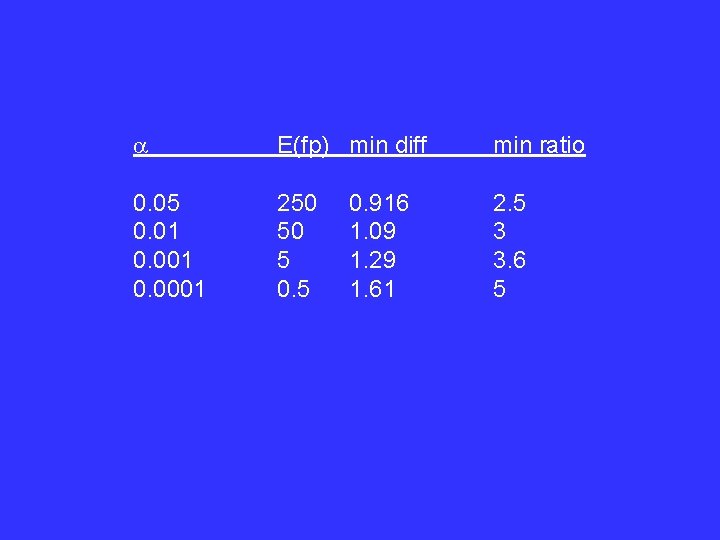
Choice of significance level: expected number of false positives: E(false positives) = a N But minimum detectable difference increases as a gets smaller

E(fp) min diff min ratio 0. 05 0. 01 0. 0001 250 50 5 0. 5 2. 5 3 3. 6 5 0. 916 1. 09 1. 29 1. 61

bias “-fold change” Proportion changed spots Validation of method against simulated data 3. Hypothesis testing: Simulated from stated model “rate false pos. ” = mean observed / expected

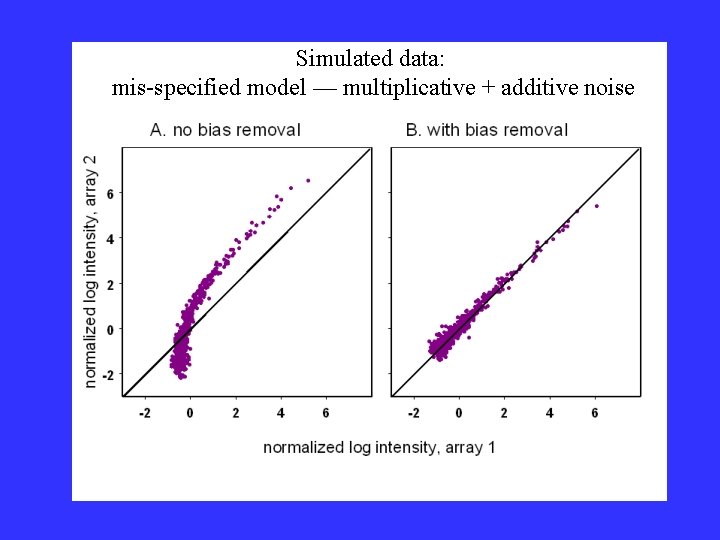
Simulated data: mis-specified model — multiplicative + additive noise

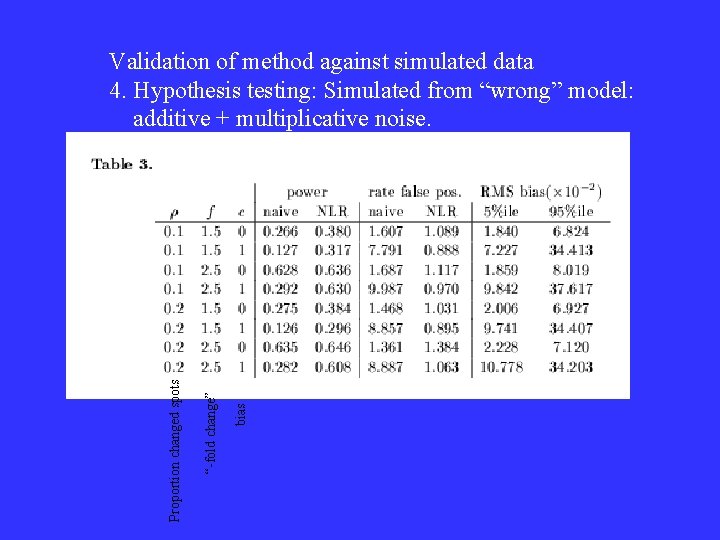
bias “-fold change” Proportion changed spots Validation of method against simulated data 4. Hypothesis testing: Simulated from “wrong” model: additive + multiplicative noise.

Acknowledgments Lynn Crosby North Carolina State University Kevin Morgan Strategic Toxicological Sciences Glaxo. Wellcome

Santa Fe Institute www. santafe. edu postdoctoral fellowships available (apply before the end of the year) kepler@santafe. edu