Todays Announcements 1 Read ECB Will assign later

Today’s Announcements 1. Read ECB: Will assign later today. 2. Homework assigned (later) today Today’s take-home lessons (i. e. what you should be able to answer at end of lecture) 1. Molecular motors: What are they? (3 families, 2 which walk on microtubules; one family which walks on actin) 2. Super-Accuracy: FIONA (nanometer accuracy, << l/2) 3. Confocal microscopy (Can discriminate according to z-axis) 4. Super-resolution microscopy—STED, STORM, (PALM next time) microscopy (gets resolution << l/2)

Fluorescence Imaging with One Nanometer Accuracy Very good accuracy: 1. 5 nm, 1 -500 msec
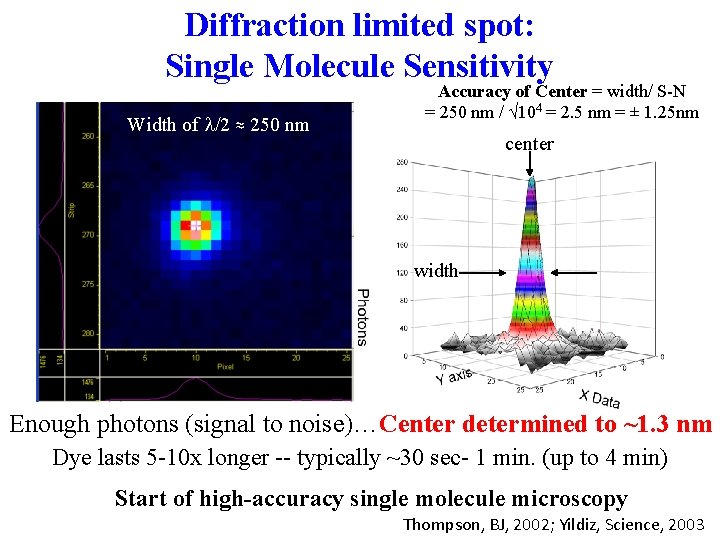
Diffraction limited spot: Single Molecule Sensitivity Width of l/2 ≈ 250 nm Accuracy of Center = width/ S-N = 250 nm / √ 104 = 2. 5 nm = ± 1. 25 nm center width Enough photons (signal to noise)…Center determined to ~1. 3 nm Dye lasts 5 -10 x longer -- typically ~30 sec- 1 min. (up to 4 min) Start of high-accuracy single molecule microscopy Thompson, BJ, 2002; Yildiz, Science, 2003
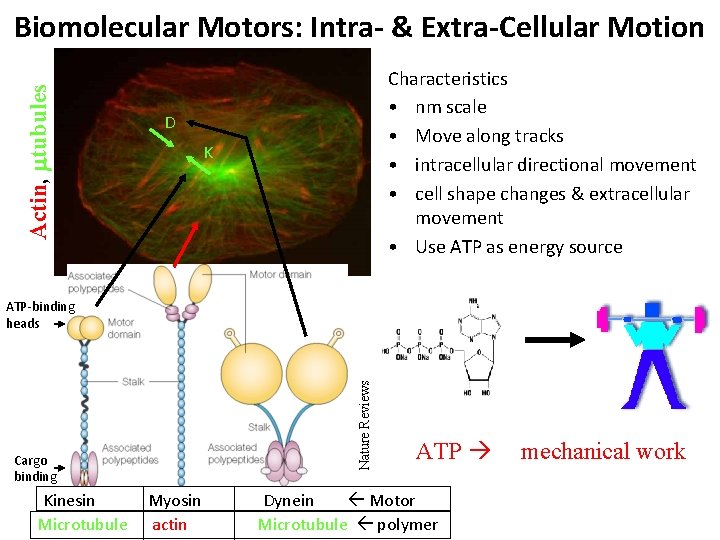
Actin, mtubules Biomolecular Motors: Intra- & Extra-Cellular Motion Characteristics • nm scale • Move along tracks • intracellular directional movement • cell shape changes & extracellular movement • Use ATP as energy source D K Nature Reviews ATP-binding heads Cargo binding Kinesin Microtubule Myosin actin ATP Dynein Motor Microtubule polymer mechanical work
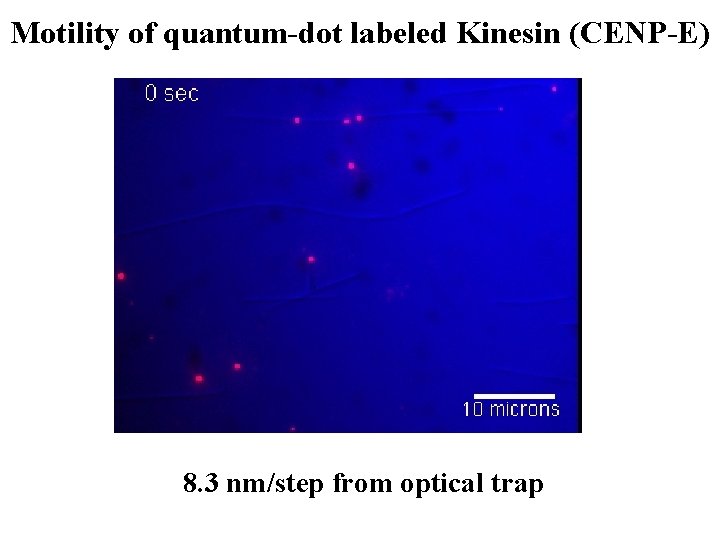
Motility of quantum-dot labeled Kinesin (CENP-E) Streptavidin Quantum Dot Streptavidin conjugate Six-histidine tag Axoneme or microtubule Biotinylated Anti-Pentahis antibody Leucine zippered CENP-E dimer w/ six histidine-tag - 8. 3 nm/step from optical trap +
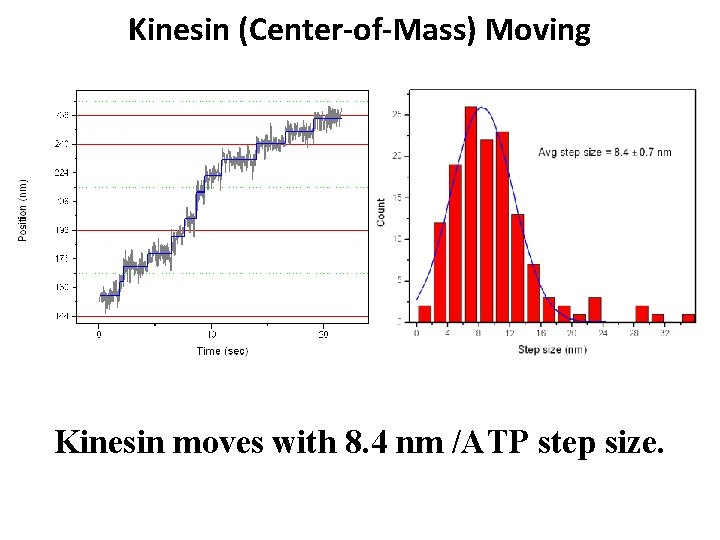
Kinesin (Center-of-Mass) Moving Kinesin moves with 8. 4 nm /ATP step size.
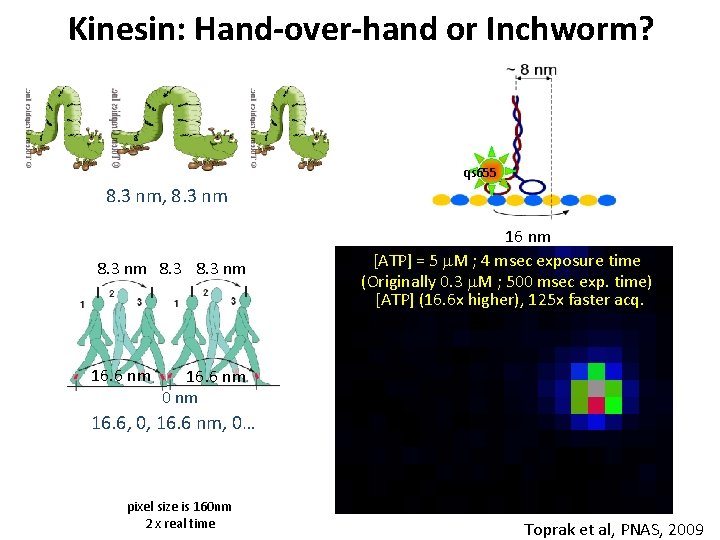
Kinesin: Hand-over-hand or Inchworm? qs 655 8. 3 nm, 8. 3 nm 16. 6 nm 16 nm [ATP] = 5 m. M ; 4 msec exposure time (Originally 0. 3 m. M ; 500 msec exp. time) [ATP] (16. 6 x higher), 125 x faster acq. 16. 6 nm 0 nm 16. 6, 0, 16. 6 nm, 0… pixel size is 160 nm 2 x real time Toprak et al, PNAS, 2009
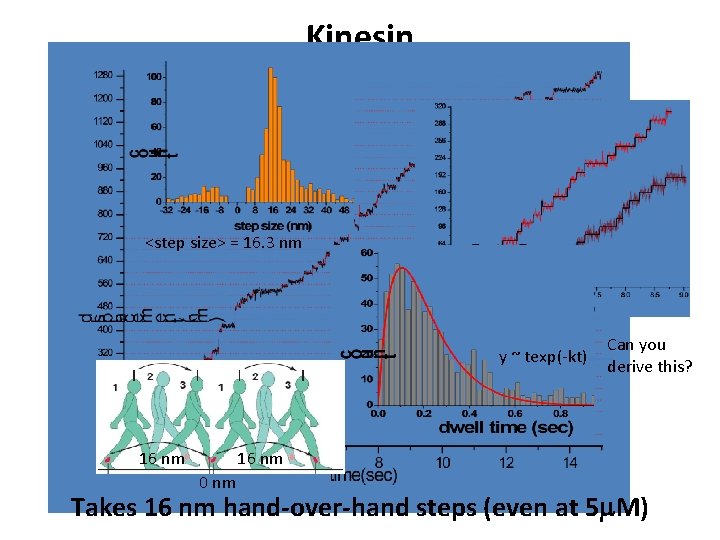
Kinesin <step size> = 16. 3 nm y ~ texp(-kt) 16 nm Can you derive this? 16 nm 0 nm Takes 16 nm hand-over-hand steps (even at 5 m. M)
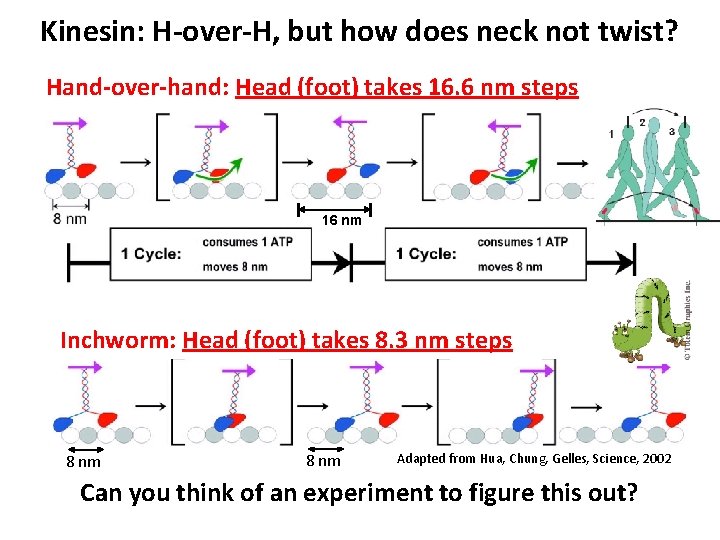
Kinesin: H-over-H, but how does neck not twist? Hand-over-hand: Head (foot) takes 16. 6 nm steps 16 nm Inchworm: Head (foot) takes 8. 3 nm steps 8 nm Adapted from Hua, Chung, Gelles, Science, 2002 Can you think of an experiment to figure this out?
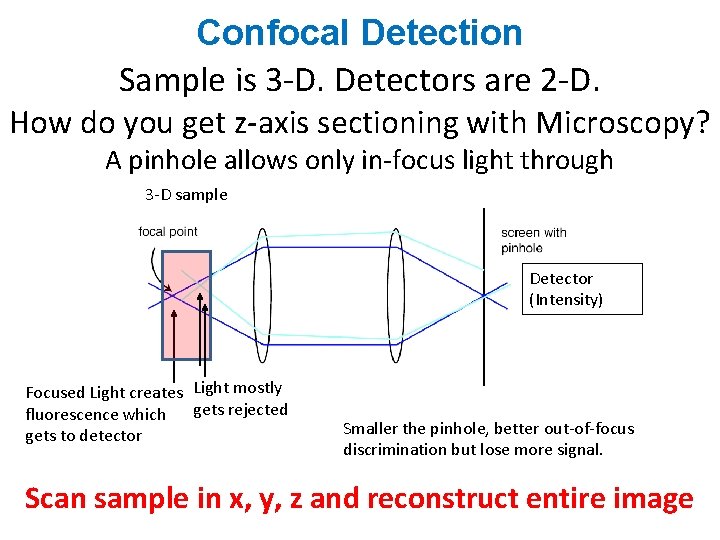
Confocal Detection Sample is 3 -D. Detectors are 2 -D. How do you get z-axis sectioning with Microscopy? A pinhole allows only in-focus light through 3 -D sample Detector (Intensity) Focused Light creates Light mostly gets rejected fluorescence which gets to detector Smaller the pinhole, better out-of-focus discrimination but lose more signal. Scan sample in x, y, z and reconstruct entire image
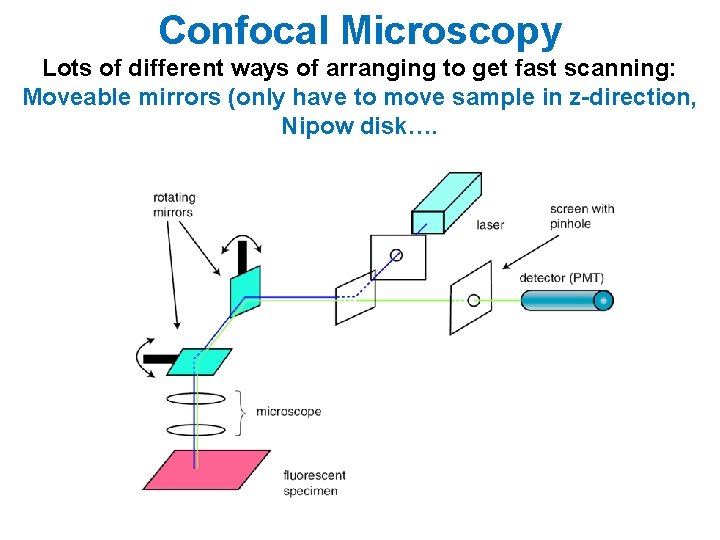
Confocal Microscopy Lots of different ways of arranging to get fast scanning: Moveable mirrors (only have to move sample in z-direction, Nipow disk….
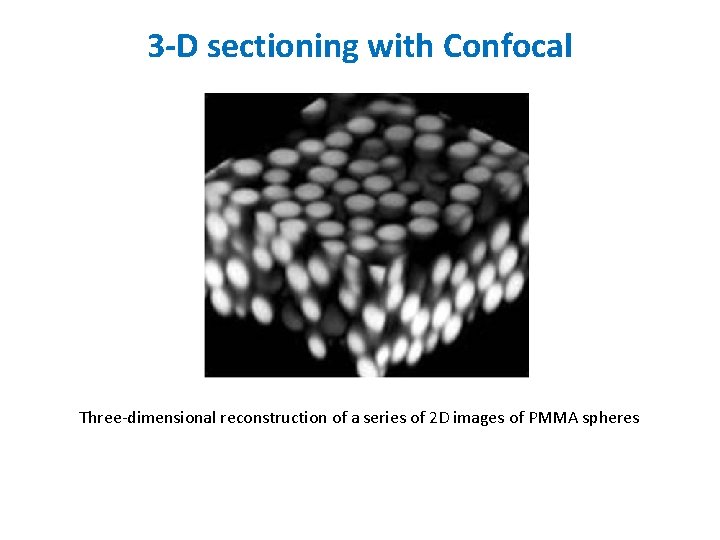
3 -D sectioning with Confocal Three-dimensional reconstruction of a series of 2 D images of PMMA spheres

Super-resolution Breaking the classical diffraction limit Can we achieve nanometer resolution? i. e. resolve two point objects separated by d << l/2? Idea: 1) Make Point-Spread Function smaller << l/2 2) Make one temporally or permanently disappear, find center (via FIONA) and then reconstruct image.
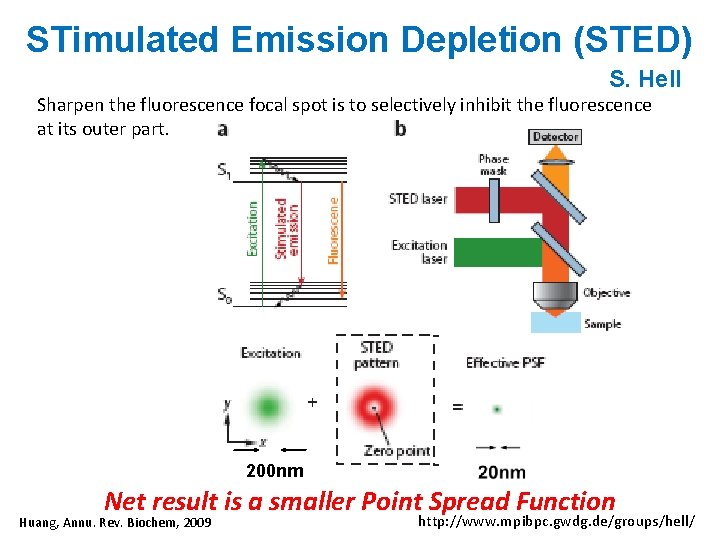
STimulated Emission Depletion (STED) S. Hell Sharpen the fluorescence focal spot is to selectively inhibit the fluorescence at its outer part. 200 nm Net result is a smaller Point Spread Function Huang, Annu. Rev. Biochem, 2009 http: //www. mpibpc. gwdg. de/groups/hell/
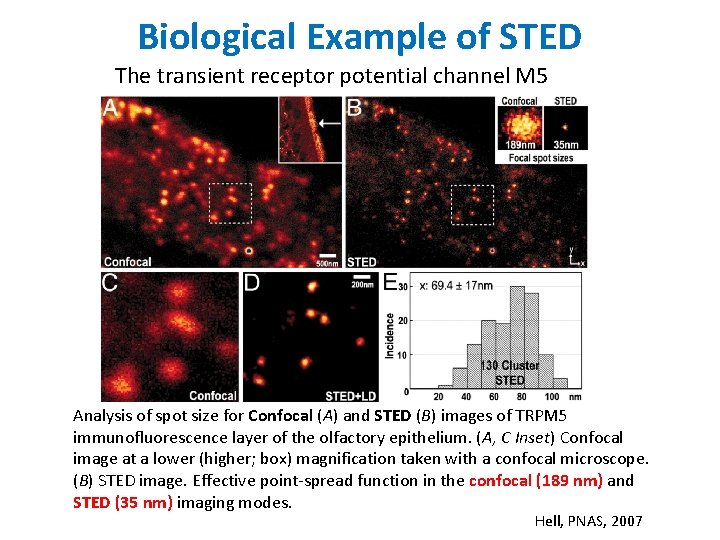
Biological Example of STED The transient receptor potential channel M 5 Analysis of spot size for Confocal (A) and STED (B) images of TRPM 5 immunofluorescence layer of the olfactory epithelium. (A, C Inset) Confocal image at a lower (higher; box) magnification taken with a confocal microscope. (B) STED image. Effective point-spread function in the confocal (189 nm) and STED (35 nm) imaging modes. Hell, PNAS, 2007
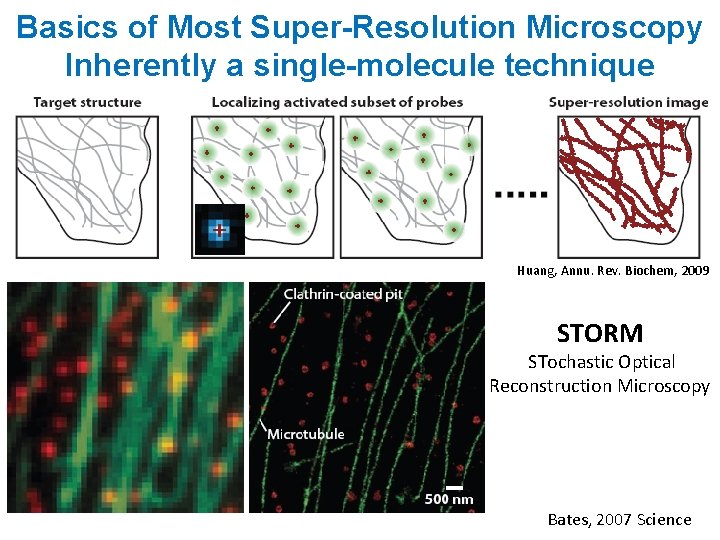
Basics of Most Super-Resolution Microscopy Inherently a single-molecule technique Huang, Annu. Rev. Biochem, 2009 STORM STochastic Optical Reconstruction Microscopy Bates, 2007 Science

Class evaluation 1. What was the most interesting thing you learned in class today? 2. What are you confused about? 3. Related to today’s subject, what would you like to know more about? 4. Any helpful comments. Answer, and turn in at the end of class.
- Slides: 17