Time Series Bitmap Experiments This file contains full

Time Series Bitmap Experiments • This file contains full color, large scale versions of the experiments shown in the paper, and additional experiments which were omitted because of space constraints • Note that in every case, all the data is freely available
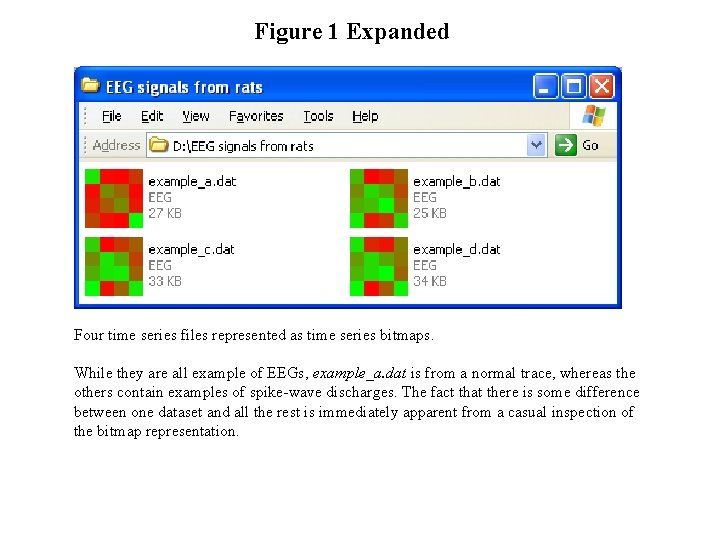
Figure 1 Expanded Four time series files represented as time series bitmaps. While they are all example of EEGs, example_a. dat is from a normal trace, whereas the others contain examples of spike-wave discharges. The fact that there is some difference between one dataset and all the rest is immediately apparent from a casual inspection of the bitmap representation.

Figure 3 Expanded The gene sequences of mitochondrial DNA of four animals, used to create their own file icons using a chaos game representation. Note that Pan troglodytes is the familiar Chimpanzee, and Loxodonta africana and Elephas maximus are the African and Indian Elephants, respectively. The file icons show that humans and chimpanzees have similar genomes, as do the African and Indian elephants.

Figure 6 Expanded A snapshot of a folder containing cardiograms when its files are arranged by “Cluster” option. Five cardiograms have been grouped into two different clusters based on their similarity. Cluster 1 (eeg 1 ~ 3): BIDMC Congestive Heart Failure Database (chfdb): record chf 02 Start times at 0, 82, 150, respectively Cluster 2 (eeg 6 ~ 7): BIDMC Congestive Heart Failure Database (chfdb): record chf 15 Start times at 0, 82 respectively

Figure 7 Expanded Data Key Cluster 1 (datasets 1 ~ 5): BIDMC Congestive Heart Failure Database (chfdb): record chf 02 Start times at 0, 82, 150, 200, 250, respectively Cluster 2 (datasets 6 ~ 10): BIDMC Congestive Heart Failure Database (chfdb): record chf 15 Start times at 0, 82, 150, 200, 250, respectively Cluster 3 (datasets 11 ~ 15): Long Term ST Database (ltstdb): record 20021 Start times at 0, 50, 100, 150, 200, respectively Cluster 4 (datasets 16 ~ 20): MIT-BIH Noise Stress Test Database (nstdb): record 118 e 6 Start times at 0, 50, 100, 150, 200, respectively
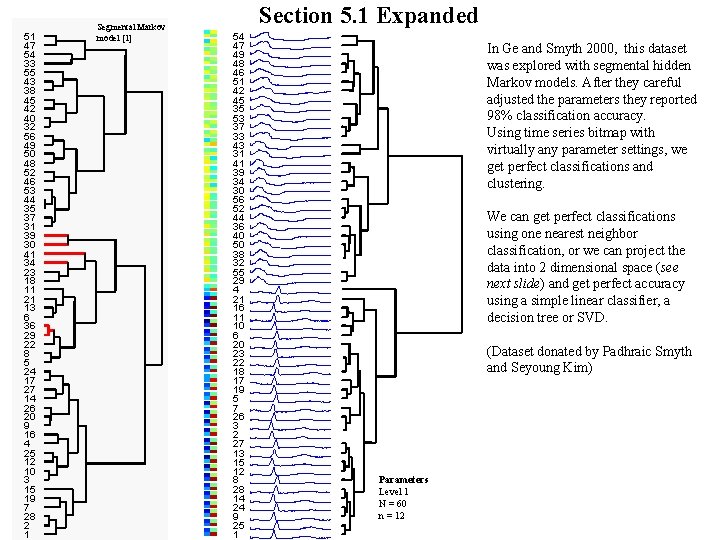
51 47 54 33 55 43 38 45 42 40 32 56 49 50 48 52 46 53 44 35 37 31 39 30 41 34 23 18 11 21 13 6 36 29 22 8 5 24 17 27 14 26 20 9 16 4 25 12 10 3 15 19 7 28 2 1 Segmental Markov model [1] Section 5. 1 Expanded 54 47 49 48 46 51 42 45 35 53 37 33 43 31 41 39 34 30 56 52 44 36 40 50 38 32 55 29 4 21 16 11 10 6 20 23 22 18 17 19 5 7 26 3 2 27 13 15 12 8 28 14 24 9 25 1 In Ge and Smyth 2000, this dataset was explored with segmental hidden Markov models. After they careful adjusted the parameters they reported 98% classification accuracy. Using time series bitmap with virtually any parameter settings, we get perfect classifications and clustering. We can get perfect classifications using one nearest neighbor classification, or we can project the data into 2 dimensional space (see next slide) and get perfect accuracy using a simple linear classifier, a decision tree or SVD. (Dataset donated by Padhraic Smyth and Seyoung Kim) Parameters Level 1 N = 60 n = 12
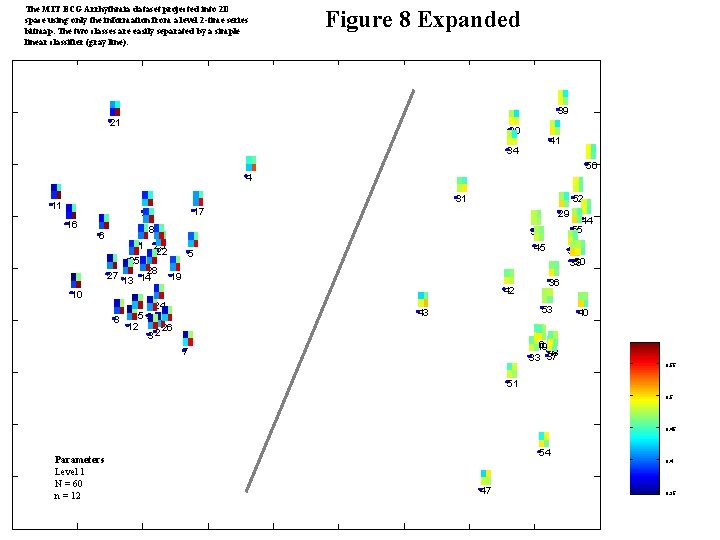
The MIT ECG Arrhythmia dataset projected into 2 D space using only the information from a level 2 -time series bitmap. The two classes are easily separated by a simple linear classifier (gray line). Figure 8 Expanded 39 21 30 41 34 56 4 31 11 16 52 17 20 29 18 6 1 25 27 35 23 22 28 13 14 45 5 19 42 10 8 24 15 9 12 26 32 32 50 38 36 53 43 46 49 48 33 37 7 44 55 40 0. 55 51 0. 5 0. 45 Parameters Level 1 N = 60 n = 12 54 47 0. 4 0. 35
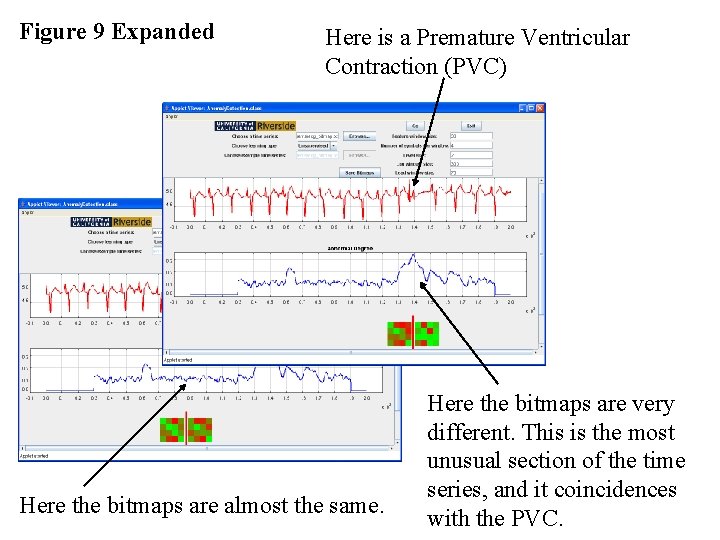
Figure 9 Expanded Here is a Premature Ventricular Contraction (PVC) Here the bitmaps are almost the same. Here the bitmaps are very different. This is the most unusual section of the time series, and it coincidences with the PVC.

Below are some more examples of anomaly detection in ECG with our bitmap approach. These examples did not make it into the paper because of space limitations
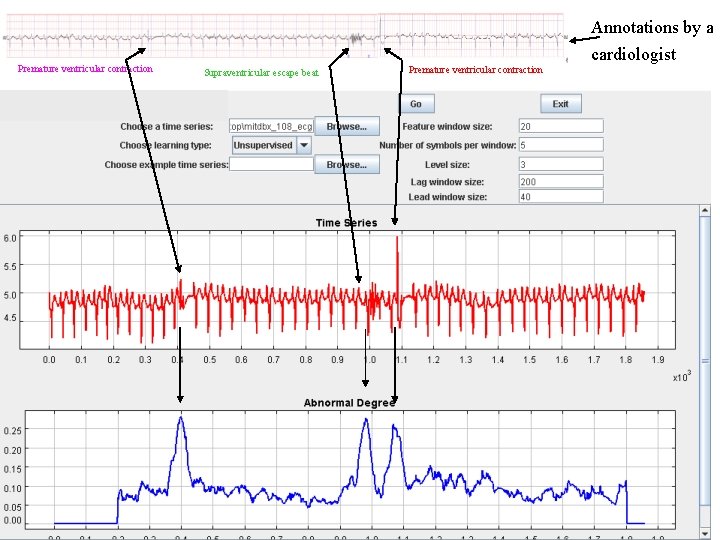
Annotations by a Premature ventricular contraction cardiologist Supraventricular escape beat Premature ventricular contraction
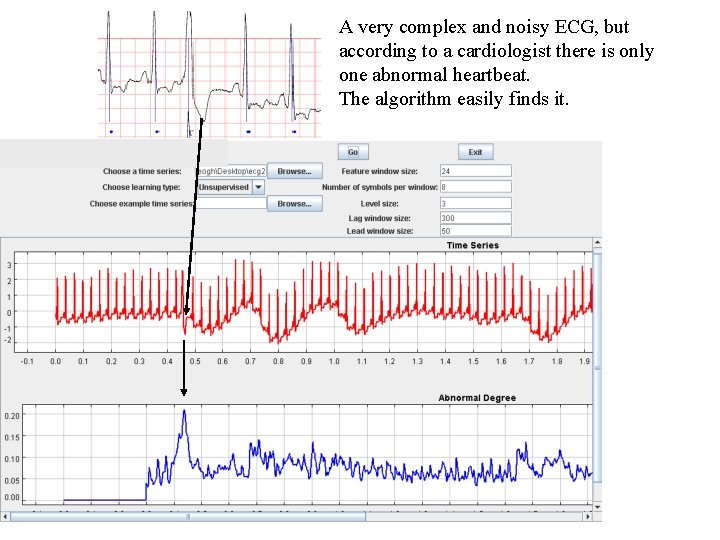
A very complex and noisy ECG, but according to a cardiologist there is only one abnormal heartbeat. The algorithm easily finds it.
- Slides: 11