Thomas Bayes 1702 1761 0 58 0 91

Thomas Bayes 1702 -1761 0. 58 0. 91 0. 99 1. 00 0. 76 0. 96 0. 99 0. 58 Leptothorax gredosi Leptothorax racovitzae Camponotus herculeanus

Computational phylogenetics CSC 10. -12. 2006 Mikko Kolkkala Bayesian inference
How to read a tree?

Bayesian inference Only very recently phylogenetical applications (”Why”? We’ll return to that…) Controversial philosophy Subjective probability concept; degrees of belief measured as probabilities A learning process Prior and posterior probabilities Spam filters Subjective! Quack!

Conditional probability: ”|” p = probability D = Data Θ = model/hypothesis/parameters | = read: ”provided that"

An example Suppose we have ten identical looking dice, nine ordinary, one die loaded so that a six appears with probability 1/2. Let’s pick one die randomly. The probability of it being loaded is (of course) 1/10 (= prior) Next, we roll the die once - and get a six: What is the probability that we have picked the loaded die now? p( loaded die | a six ) = p( a six | loaded die ) • p( loaded die ) p(a six) = 1/2 • 1/10 + 1/6 • 9/10 = 1/4 (= posterior)

An exercise A reliable test? Test for a rare disease (prevalence 0. 1 %): Disease - positive result with probability 0. 99 No disease - positive result with probability 0. 05. What is the probability that the test is positive but the individual tested has not the disease? Answer: 0. 98 (http: //en. wikipedia. org/wiki/Bayesian_inference)

“loaded die" model “a six" data p(model | data) = p(data | model) p(data) • p(model)
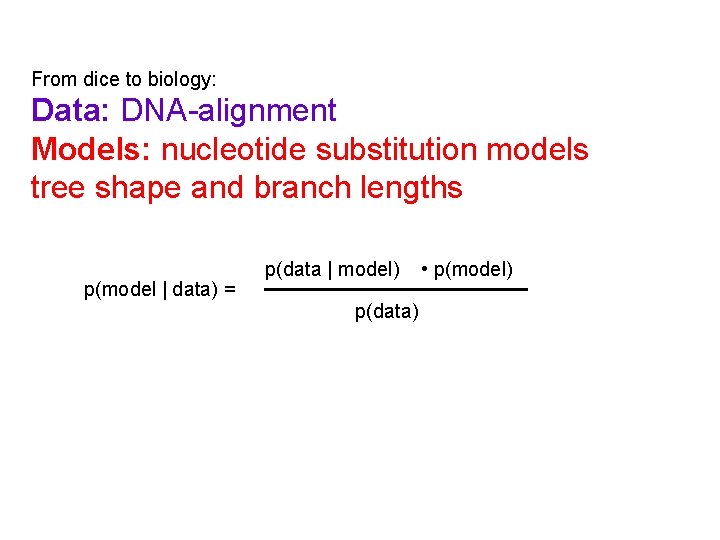
From dice to biology: Data: DNA-alignment Models: nucleotide substitution models tree shape and branch lengths p(model | data) = p(data | model) p(data) • p(model)

Posterior distribution Prior distribution Likelihood function

If this Bayesian thing is so excellent why hasn’t It been used in phylogenetic analyses? No-one can solve the equations! Numerical solutions possible - but only with powerful computers
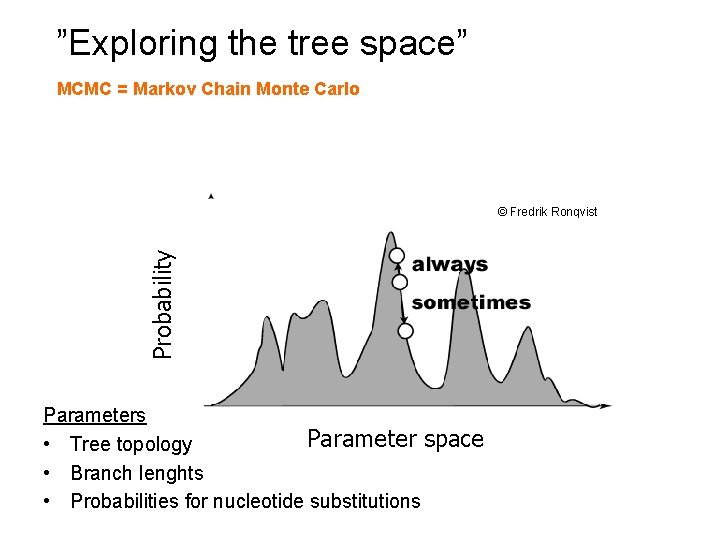
“”Exploring the tree space” MCMC = Markov Chain Monte Carlo Probability © Fredrik Ronqvist Parameters Parameter space • Tree topology • Branch lenghts • Probabilities for nucleotide substitutions

Metropolis-Coupled Markov Chain Monte Carlo MCMCMC = (MC)3 “Heated chains" “Flattened" parameter landscape

(MC)3 © John Huelsenbeck

(MC)3 © John Huelsenbeck
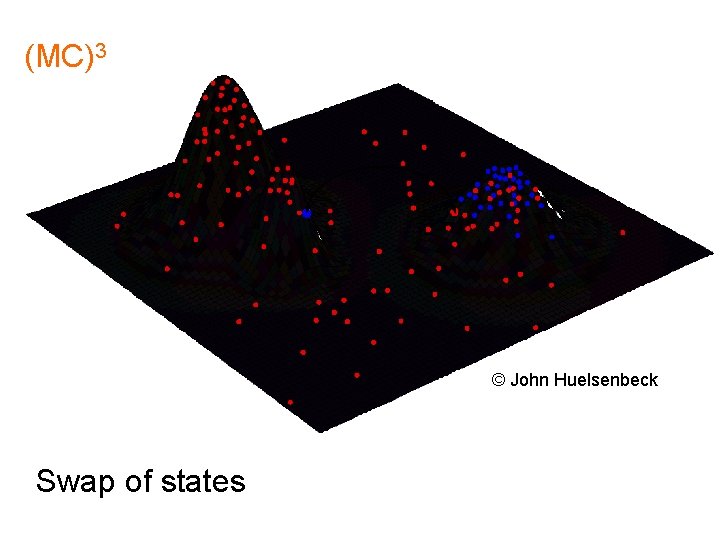
(MC)3 © John Huelsenbeck Swap of states

p-values directly No need for bootstrapping
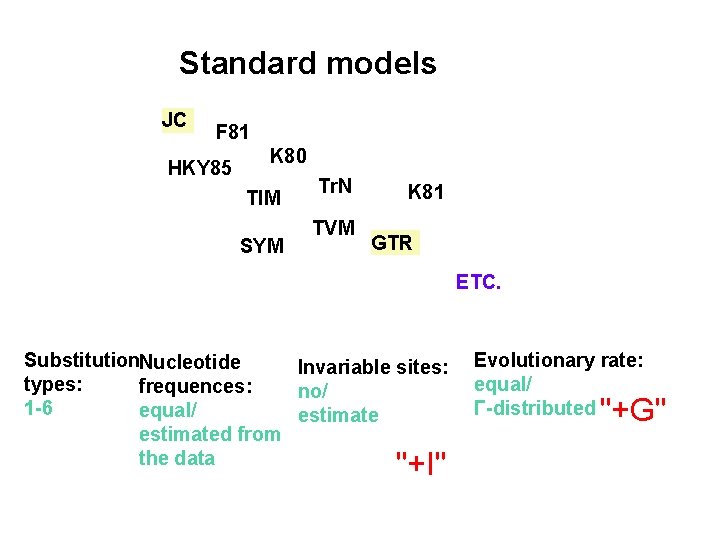
Standard models JC F 81 HKY 85 K 80 TIM SYM Tr. N TVM K 81 GTR ETC. Substitution. Nucleotide Invariable sites: types: frequences: no/ 1 -6 equal/ estimated from the data "+I" Evolutionary rate: equal/ Γ-distributed "+G"
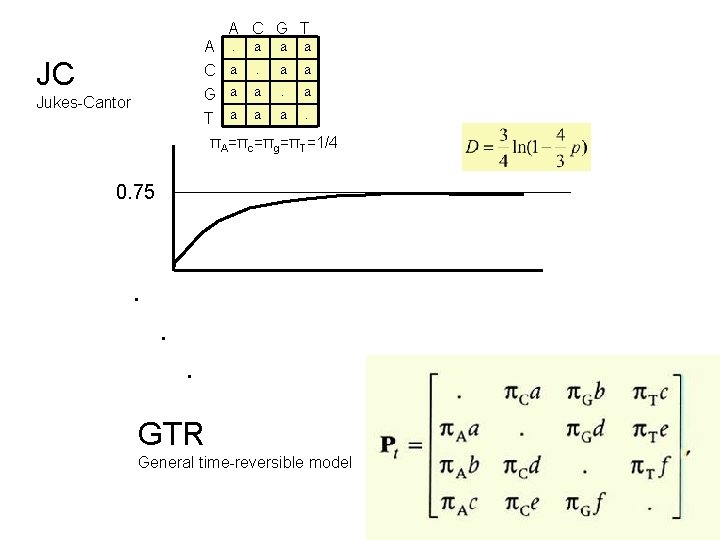
A. a a a C G T a a a. A C G T πA=πc=πg=πT=1/4 JC Jukes-Cantor 0. 75 . . . GTR General time-reversible model
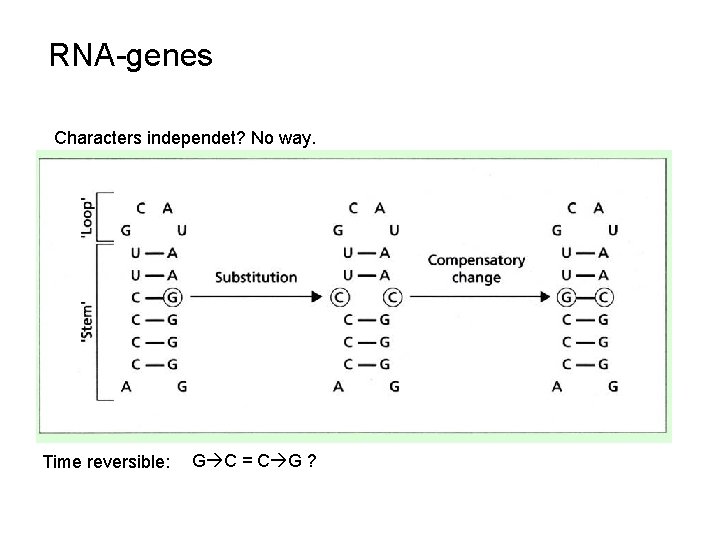
RNA-genes Characters independet? No way. Time reversible: G C = C G ?

Coding regions SSR-models (site-specific rates) Different evolutionary rate for 1. /2. /3. positions of codons Problematic (see: Buckley ym. 2001 Syst. Biol. 50: 67 -86)
But – how to chooce the model? Well, nobody said it would be easy. How many parameters Does it take to fit an elephant? 30

“What do you consider the largest map that would be really useful? " "About six inches to the mile. " "Only six inches! […] We actually made a map of the country, on the scale of a mile to the mile!" (Lewis Carroll 1893)

Choosing a model AIC (Akaike information criterion) AICc (Consistent Akaike information criterion) BIC (Bayesian information criterion) Programs: Modeltest (bad) Find. Model (plop!) Mr. Aic ?


Most commonly used program: Mr. Bayes Future? Alignment and phylogeny co-estimation BAli-Phy (Redeling & Suchard 2005) Beast (Lunter et al. 2005) Lunter, G. et al. 2005: Bayesian coestimation of phylogeny and sequence alignment. BMC Bioinformatics 6: 83 Redelings, B. D. & Suchard, M. A 2005: Joint Bayesian estimation of alignment and phylogeny. Syst. Biol. 54: 401 -418
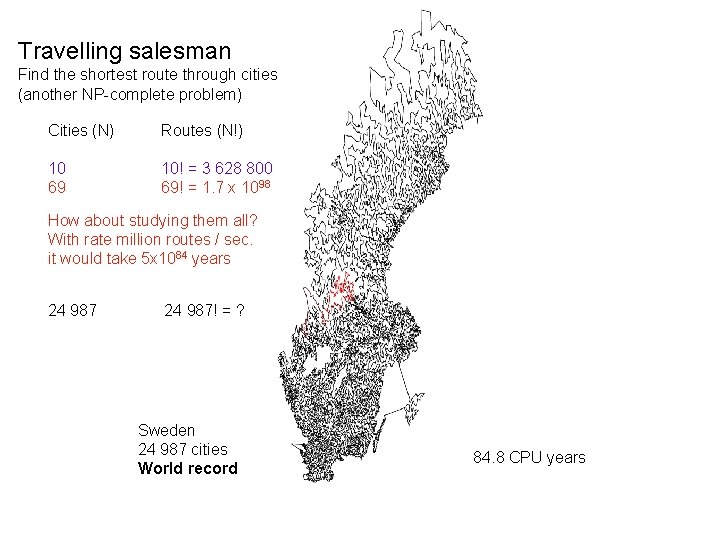
Travelling salesman Find the shortest route through cities (another NP-complete problem) Cities (N) Routes (N!) 10 69 10! = 3 628 800 69! = 1. 7 x 1098 How about studying them all? With rate million routes / sec. it would take 5 x 1084 years 24 987! = ? Sweden 24 987 cities World record 84. 8 CPU years

Acknowledgements Fredrik Ronqvist John Huelsenbeck Wife and Mom

Mr. Bayes Ronqvist, F. & Huelsenbeck, J. 2001: Bioinformatics 17: 754 -755 (2005: v. 3. 1. ) -Command-line interface -UNIX, Macintosh and PC platforms

Mr. Bayes Homepage Manual Wiki, FAQ Mailing list (archives)

Mr. Bayes Running the analysis All you have to do: Type execute filename. nex * at the Mr. Bayes > prompt and press enter * Replace filename. nex with your nexus-file containing Mr. Bayes commands (type full path if the file is not in the same folder as Mr. Bayes program).

Mr. Bayes – an example nexus file #nexus begin data; dimensions ntax=6 nchar=20; format datatype=dna; matrix Otus 1 aaaaaaaaaa Otus 2 aaaaaaaaaa Otus 3 aaaaaaaaaa Otus 4 cccccccccc Otus 5 gggggggggg Otus 6 tttttttttt ; end; begin mrbayes; mcmcp ngen= 100000 samplefreq=100; mcmc; end;

A real thing:

Mr. Bayes After the run Summarize the parameter values, type: sump burnin= Summarize the trees, type: sumt burnin= With a proper burnin value
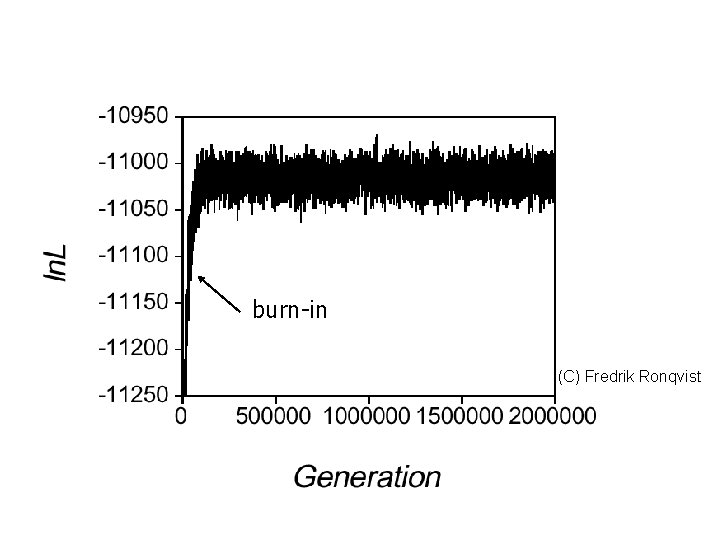
burn-in (C) Fredrik Ronqvist

Mr. Bayes After the run Burnin discards initial values before the analysis reached convergence (burnin=2500 if you have run a million generations, sampled every 100 th of them, and want to discard the first 25%) Note: you have to run “enough” generations -Check the plot generated by sump; there should be no obvious trends -The standard deviation of split frequencies should be less than 0. 01.

Mr. Bayes Models Restriction: Can handle only 24 substitution models Command for example: lset nst=6 rates=invgamma Confused? Try typing: help lset Priors, command: prset Defaults (try help prset) should work fine for most analysis

Cladistic parsimony Prefer the tree with the fewest number of evolutionary steps – only parsimony informative sites count Otus 1 Otus 2 Otus 3 Otus 4 Otus 5 Otus 6 aaaaaaaaaaaaaaaaaaaa cccccccccc gggggggggg tttttttttt Otus 1 Otus 2 Otus 3 Otus 4 Otus 5 Otus 6

Fain ja Houde 2004: Evolution 58: 2558 -2573

Mr. Bayes Exercises: 1. Study program defaults with help command (e. g. lset and prset) 2. Run program with a few arbitrary sequences (e. g. palikka. nex) -Try sump and sumt commands with different burnin values -Study the files made by the program – where is the tree? 3. Run program with some real data (e. g. your own or birds. txt) -Align sequences -Put them into a nexus file -Try to find out how to select JC, K 2 P and GTR model with gamma-distributed rate variation and without with correction for invariable sites and without -Try the model suggested by Find. Model (AIC-criterion) -
- Slides: 40