Third m RNA base 3 end of codon
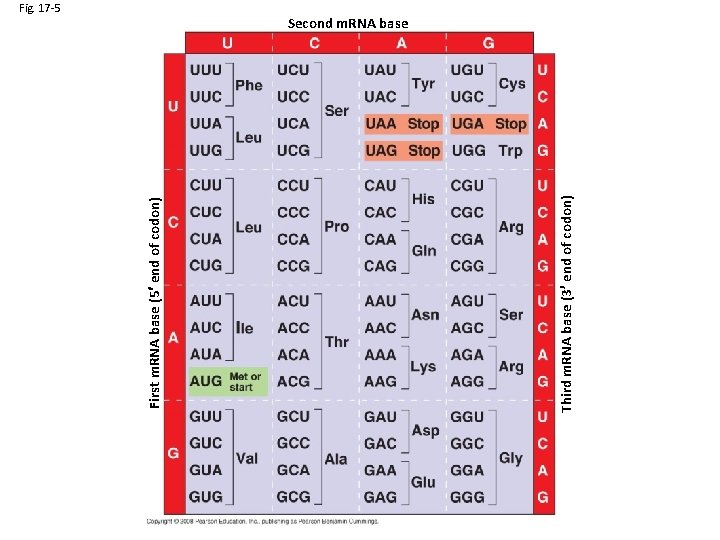
Third m. RNA base (3 end of codon) First m. RNA base (5 end of codon) Fig. 17 -5 Second m. RNA base
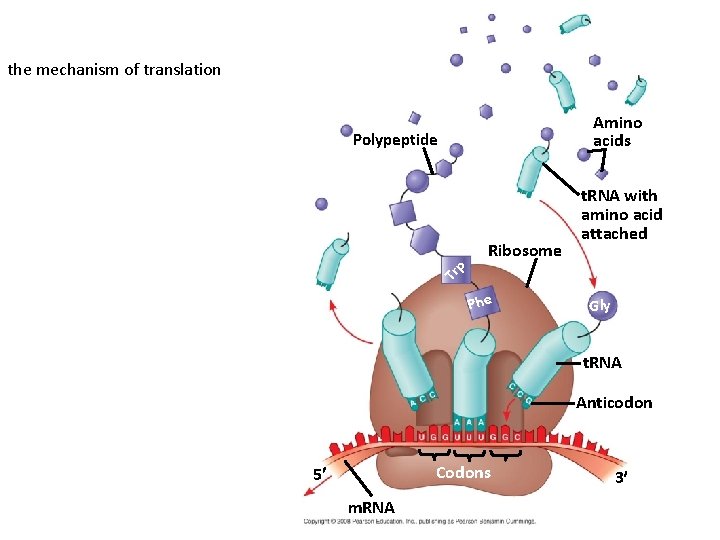
the mechanism of translation Amino acids Polypeptide Tr p Ribosome t. RNA with amino acid attached Phe Gly t. RNA Anticodon Codons 5 m. RNA 3
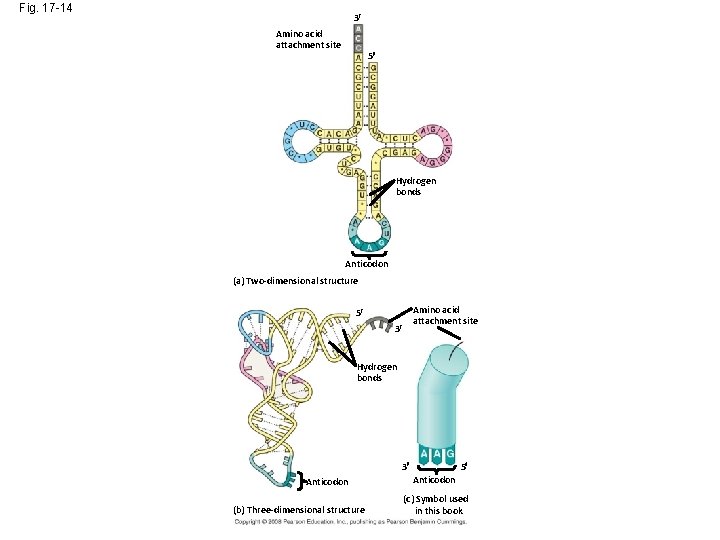
Fig. 17 -14 3 Amino acid attachment site 5 Hydrogen bonds Anticodon (a) Two-dimensional structure Amino acid attachment site 5 3 Hydrogen bonds 3 Anticodon (b) Three-dimensional structure 5 Anticodon (c) Symbol used in this book
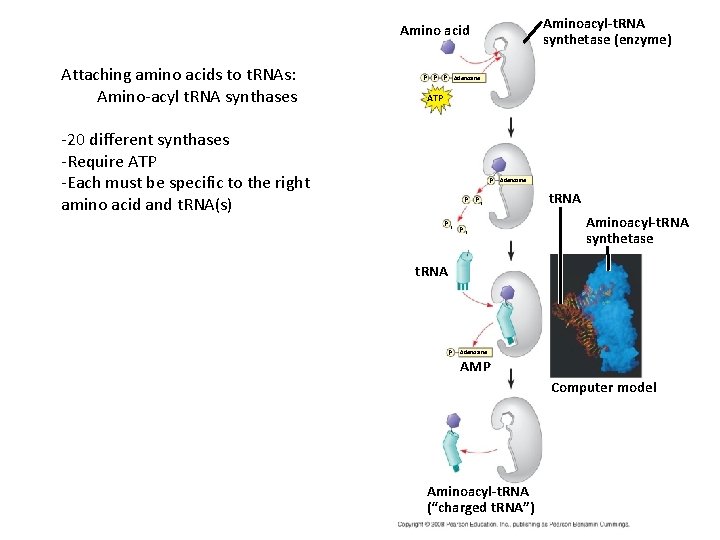
Aminoacyl-t. RNA synthetase (enzyme) Amino acid Attaching amino acids to t. RNAs: Amino-acyl t. RNA synthases P P P Adenosine ATP -20 different synthases -Require ATP -Each must be specific to the right amino acid and t. RNA(s) P Adenosine P Pi Pi P i t. RNA Aminoacyl-t. RNA synthetase t. RNA P Adenosine AMP Computer model Aminoacyl-t. RNA (“charged t. RNA”)
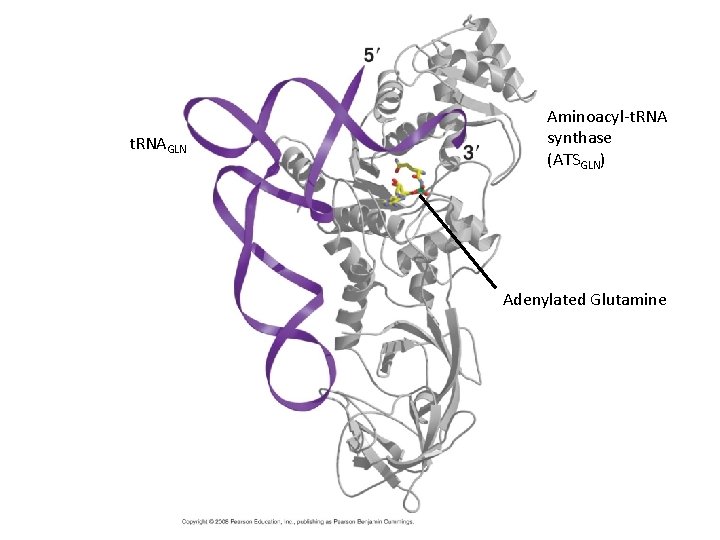
t. RNAGLN Aminoacyl-t. RNA synthase (ATSGLN) Adenylated Glutamine
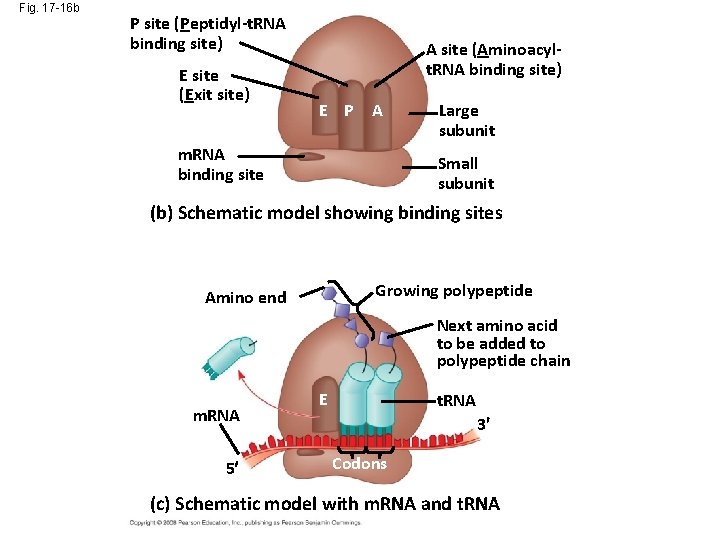
Fig. 17 -16 b P site (Peptidyl-t. RNA binding site) E site (Exit site) A site (Aminoacylt. RNA binding site) E P A m. RNA binding site Large subunit Small subunit (b) Schematic model showing binding sites Growing polypeptide Amino end Next amino acid to be added to polypeptide chain m. RNA 5 E t. RNA 3 Codons (c) Schematic model with m. RNA and t. RNA
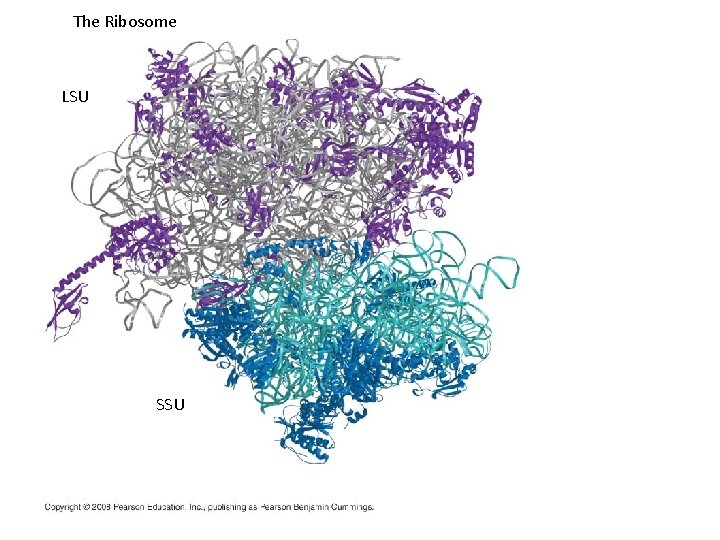
The Ribosome LSU SSU
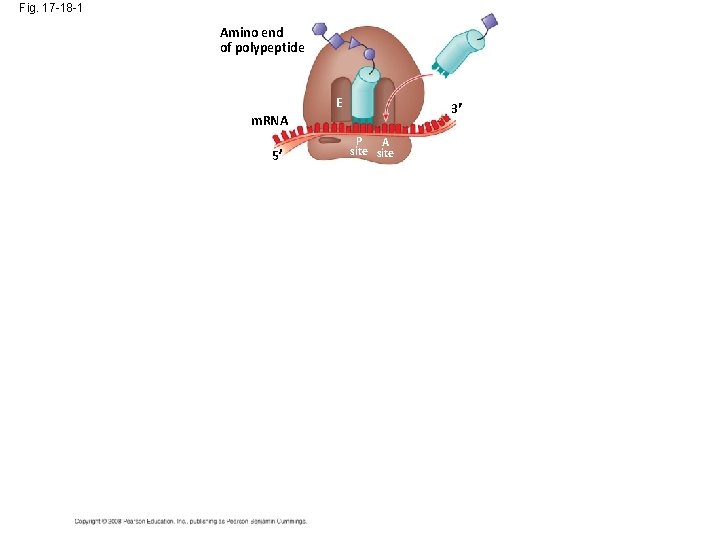
Fig. 17 -18 -1 Amino end of polypeptide E 3 m. RNA 5 P A site
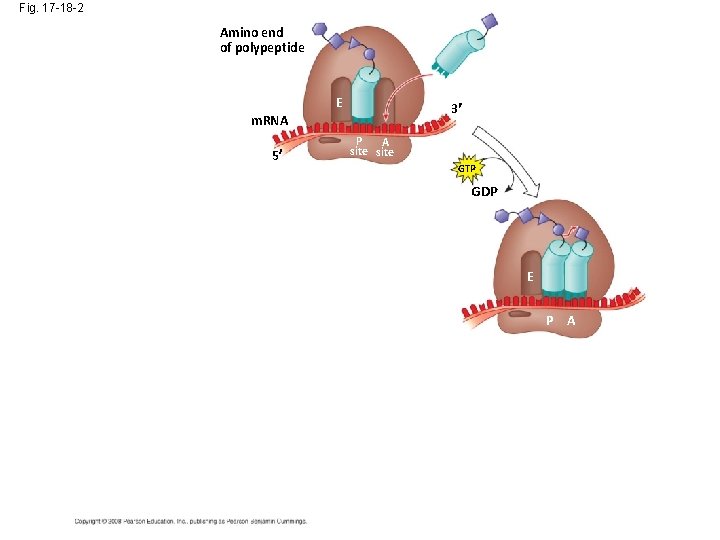
Fig. 17 -18 -2 Amino end of polypeptide E 3 m. RNA 5 P A site GTP GDP E P A
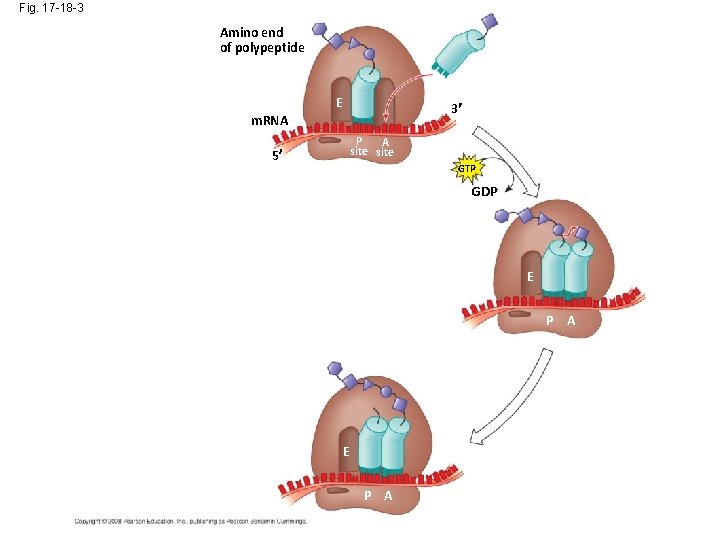
Fig. 17 -18 -3 Amino end of polypeptide E 3 m. RNA P A site 5 GTP GDP E P A
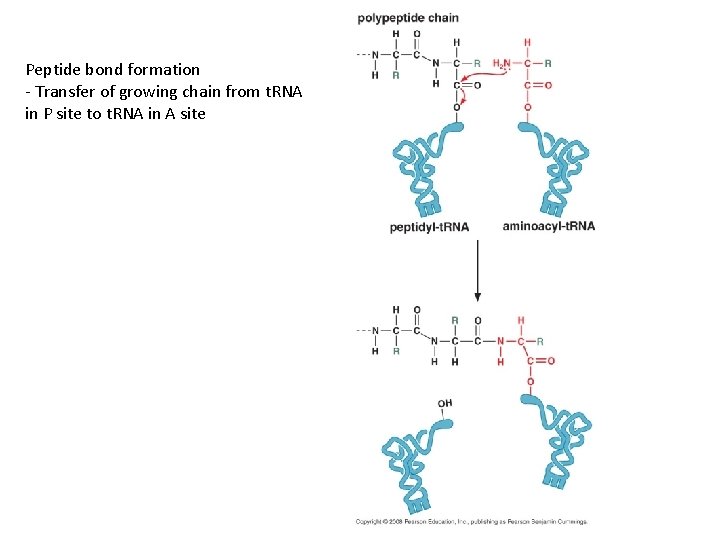
Peptide bond formation - Transfer of growing chain from t. RNA in P site to t. RNA in A site
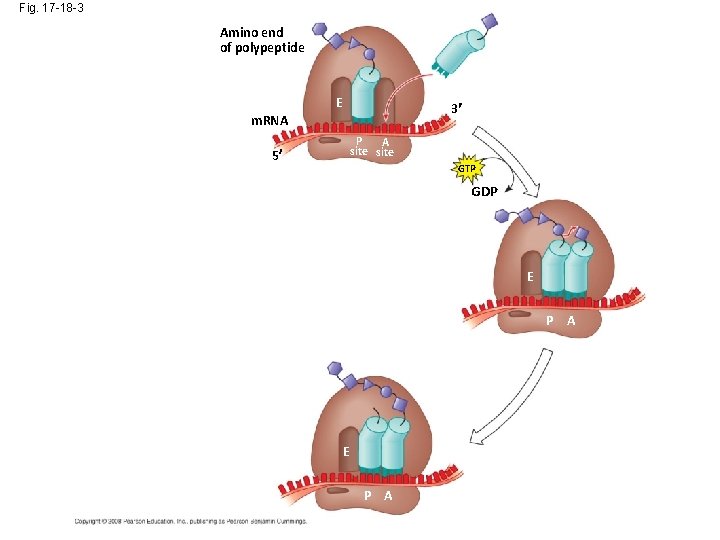
Fig. 17 -18 -3 Amino end of polypeptide E 3 m. RNA P A site 5 GTP GDP E P A
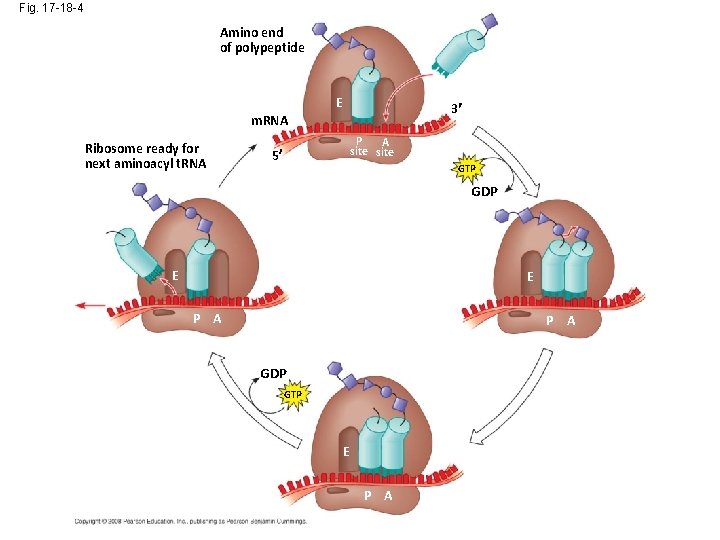
Fig. 17 -18 -4 Amino end of polypeptide E 3 m. RNA Ribosome ready for next aminoacyl t. RNA P A site 5 GTP GDP E E P A GDP GTP E P A
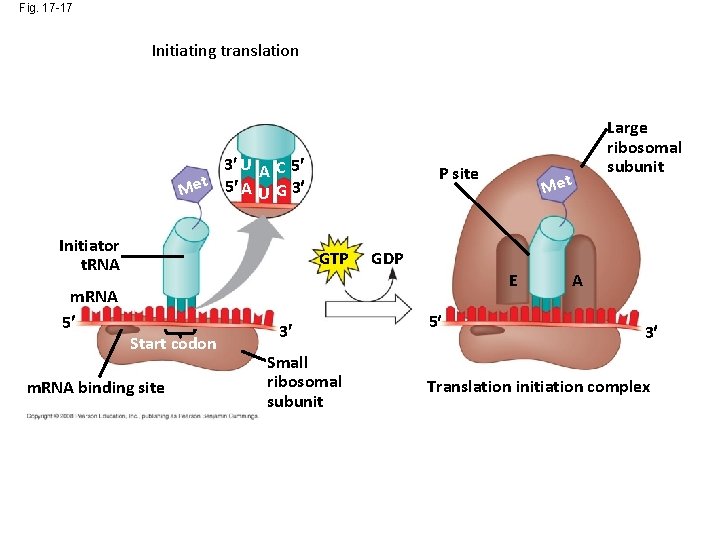
Fig. 17 -17 Initiating translation 3 U A C 5 Met 5 A U G 3 Initiator t. RNA m. RNA 5 P site GTP Start codon m. RNA binding site 3 Small ribosomal subunit Met Large ribosomal subunit GDP E 5 A 3 Translation initiation complex
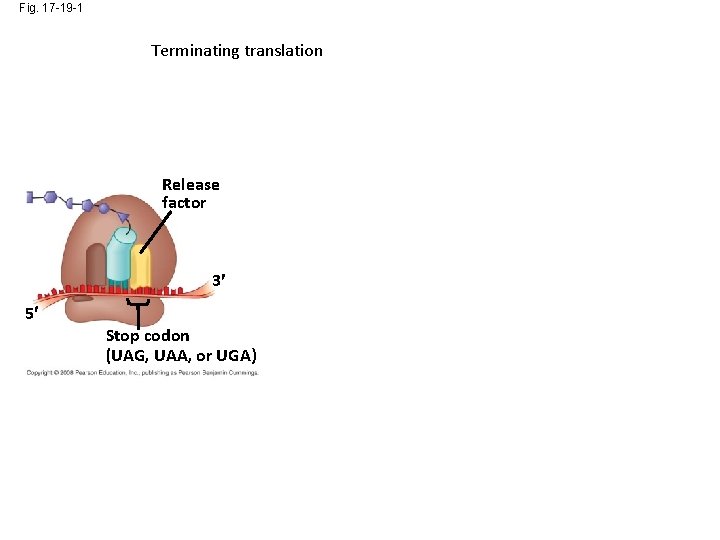
Fig. 17 -19 -1 Terminating translation Release factor 3 5 Stop codon (UAG, UAA, or UGA)
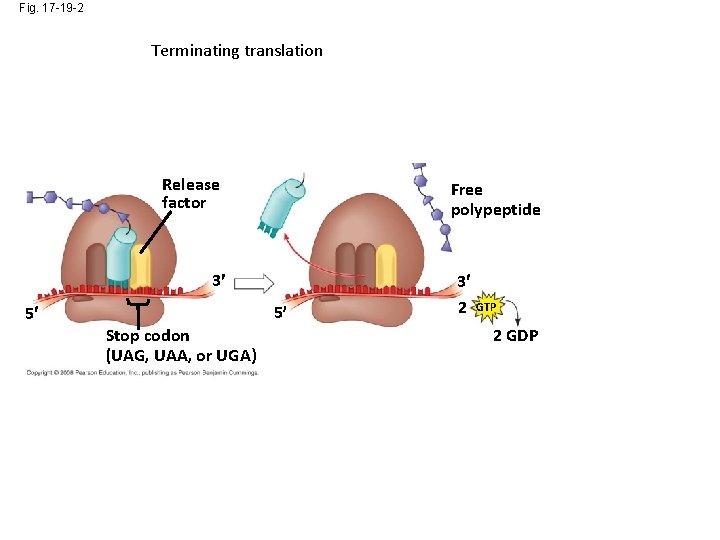
Fig. 17 -19 -2 Terminating translation Release factor Free polypeptide 3 5 5 Stop codon (UAG, UAA, or UGA) 3 2 GTP 2 GDP
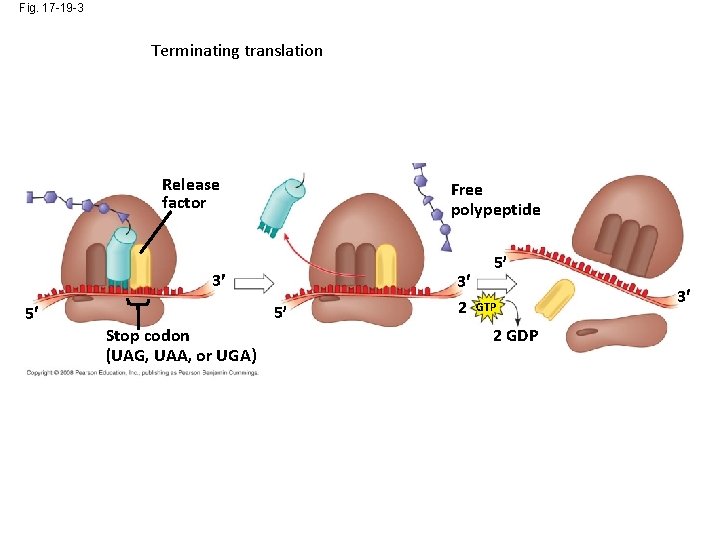
Fig. 17 -19 -3 Terminating translation Release factor Free polypeptide 3 5 5 Stop codon (UAG, UAA, or UGA) 3 2 5 GTP 2 GDP 3
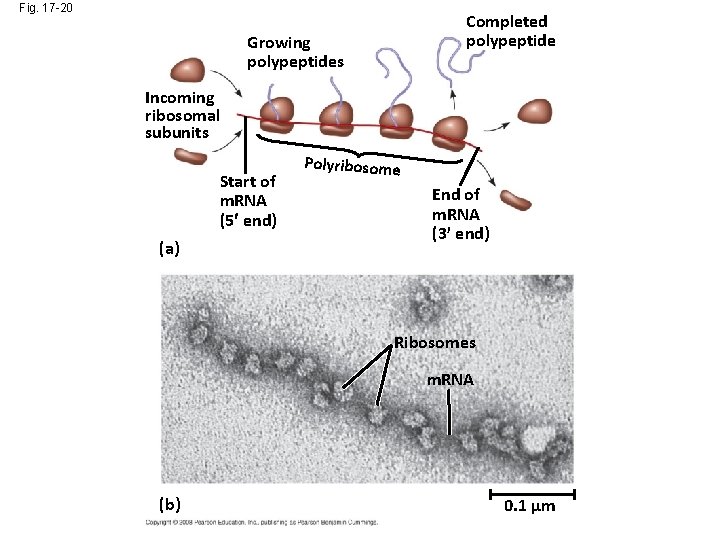
Fig. 17 -20 Completed polypeptide Growing polypeptides Incoming ribosomal subunits Start of m. RNA (5 end) (a) Polyribosome End of m. RNA (3 end) Ribosomes m. RNA (b) 0. 1 µm
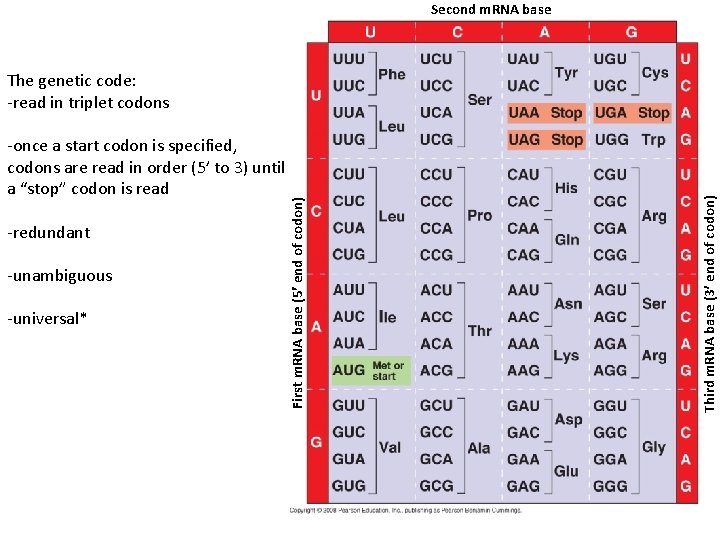
Second m. RNA base -redundant -unambiguous -universal* Third m. RNA base (3 end of codon) -once a start codon is specified, codons are read in order (5’ to 3) until a “stop” codon is read First m. RNA base (5 end of codon) The genetic code: -read in triplet codons
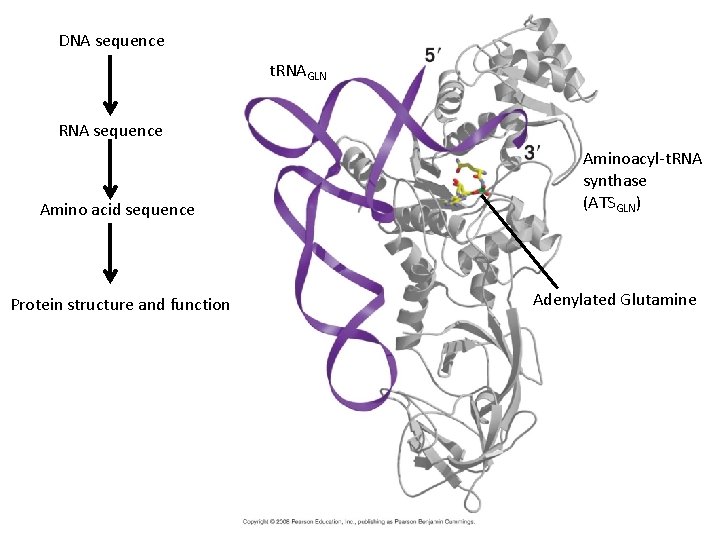
DNA sequence t. RNAGLN RNA sequence Amino acid sequence Protein structure and function Aminoacyl-t. RNA synthase (ATSGLN) Adenylated Glutamine
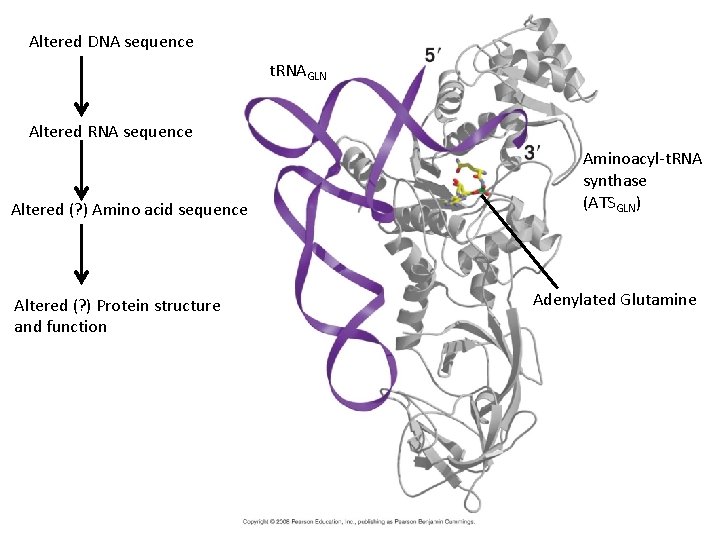
Altered DNA sequence t. RNAGLN Altered RNA sequence Altered (? ) Amino acid sequence Altered (? ) Protein structure and function Aminoacyl-t. RNA synthase (ATSGLN) Adenylated Glutamine
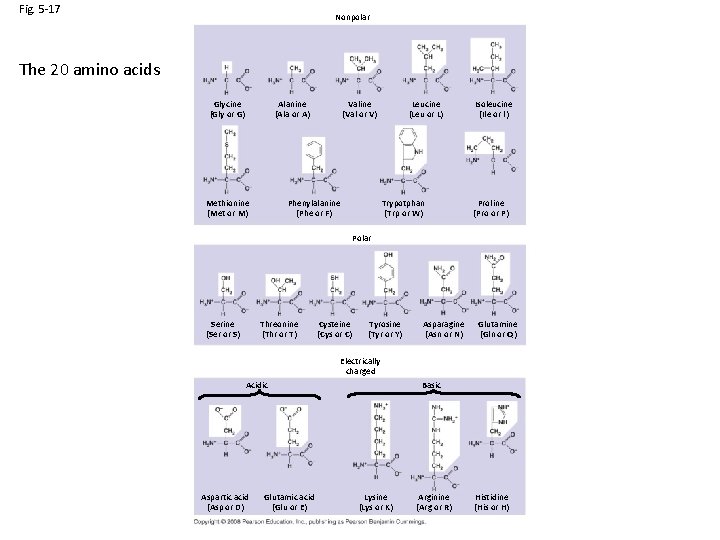
Fig. 5 -17 Nonpolar The 20 amino acids Glycine (Gly or G) Valine (Val or V) Alanine (Ala or A) Methionine (Met or M) Leucine (Leu or L) Trypotphan (Trp or W) Phenylalanine (Phe or F) Isoleucine (Ile or I) Proline (Pro or P) Polar Serine (Ser or S) Threonine (Thr or T) Cysteine (Cys or C) Tyrosine (Tyr or Y) Asparagine (Asn or N) Glutamine (Gln or Q) Electrically charged Acidic Aspartic acid (Asp or D) Glutamic acid (Glu or E) Basic Lysine (Lys or K) Arginine (Arg or R) Histidine (His or H)
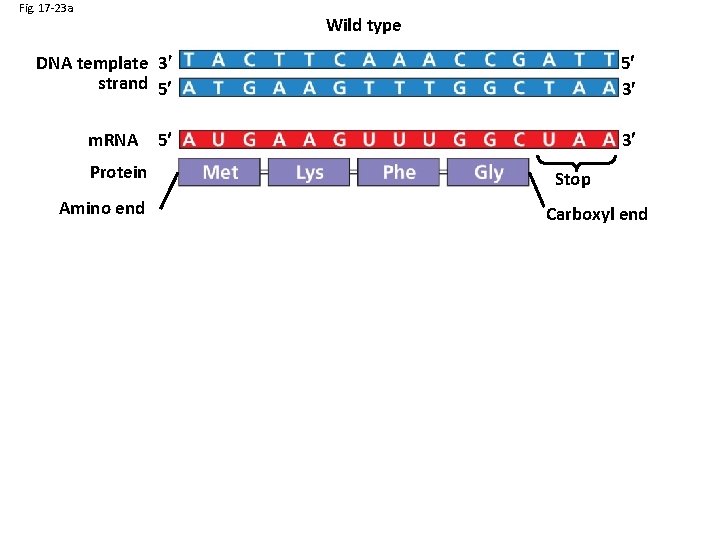
Fig. 17 -23 a Wild type DNA template 3 strand 5 5 3 5 3 m. RNA Protein Stop Amino end Carboxyl end A instead of G 5 3 3 5 U instead of C 5 3 Stop Silent (no effect on amino acid sequence)
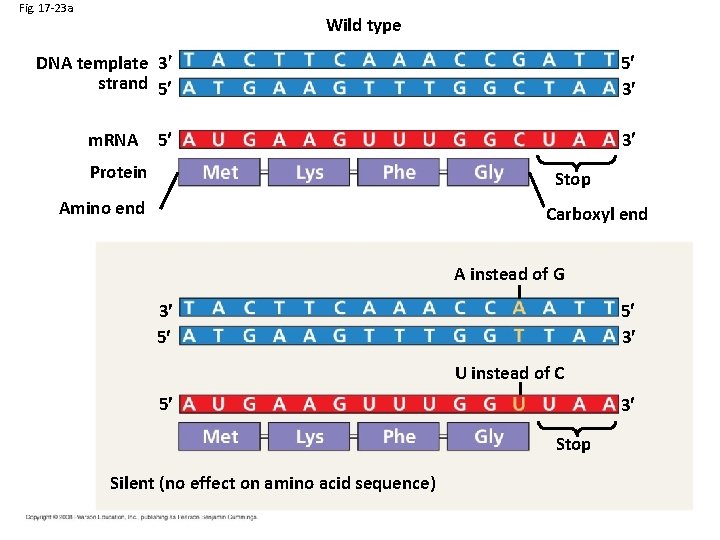
Fig. 17 -23 a Wild type DNA template 3 strand 5 5 3 5 3 m. RNA Protein Stop Amino end Carboxyl end A instead of G 5 3 3 5 U instead of C 5 3 Stop Silent (no effect on amino acid sequence)
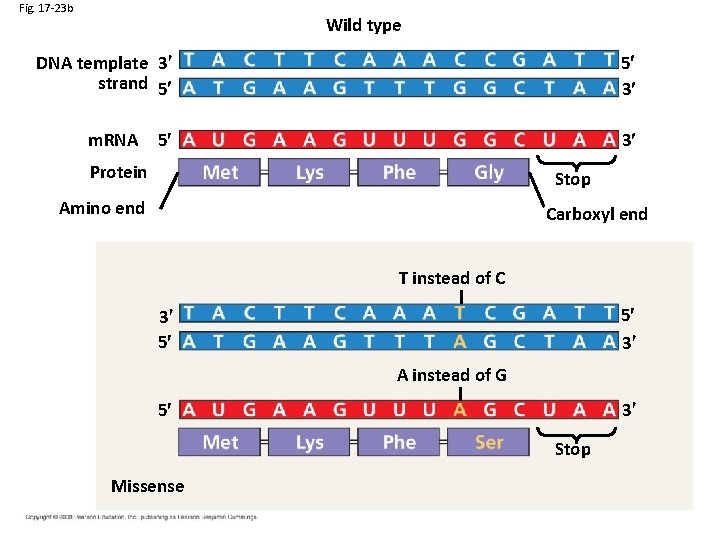
Fig. 17 -23 b Wild type DNA template 3 strand 5 5 3 5 3 m. RNA Protein Stop Amino end Carboxyl end T instead of C 5 3 3 5 A instead of G 3 5 Stop Missense
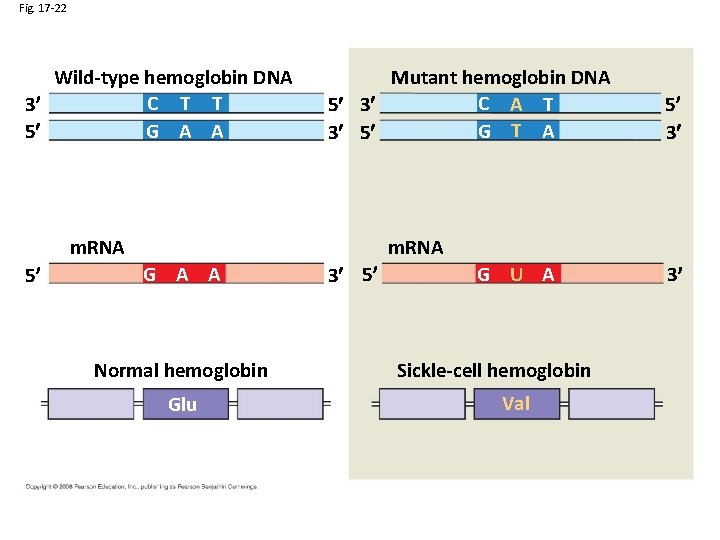
Fig. 17 -22 Wild-type hemoglobin DNA C T T 3 5 G A A Mutant hemoglobin DNA C A T 5 3 G T A 3 5 m. RNA 5 5 3 m. RNA G A A Normal hemoglobin Glu 3 5 G U A Sickle-cell hemoglobin Val 3
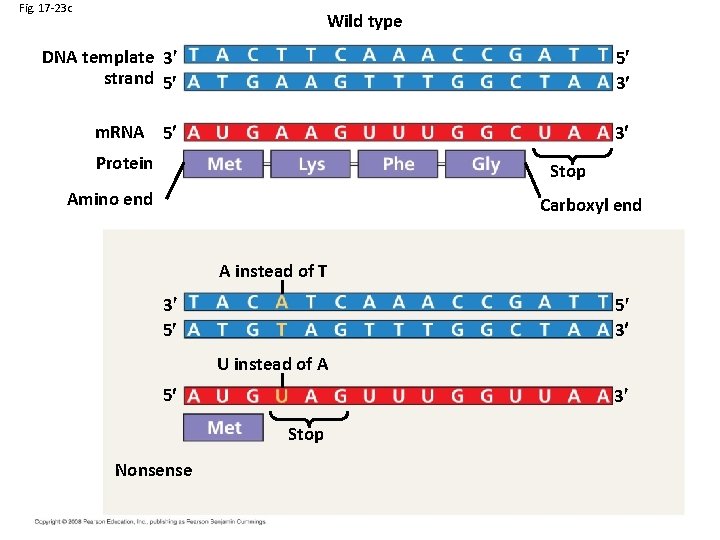
Fig. 17 -23 c Wild type DNA template 3 strand 5 5 3 m. RNA 5 3 Protein Stop Amino end Carboxyl end A instead of T 3 5 5 3 U instead of A 5 3 Stop Nonsense
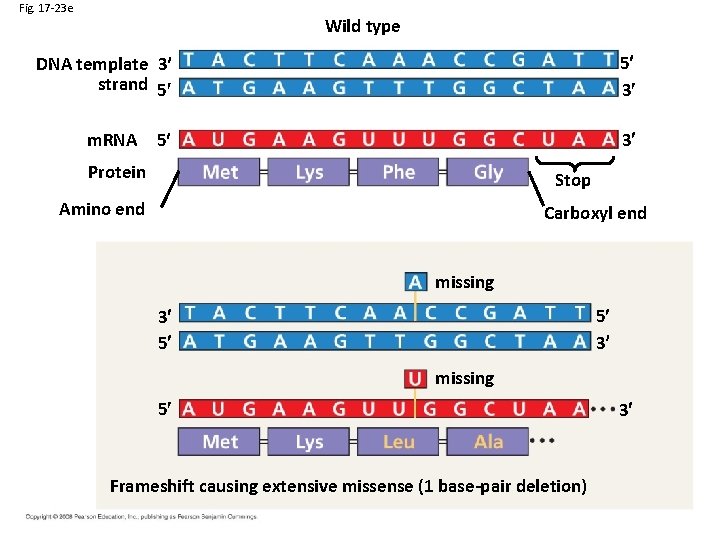
Fig. 17 -23 e Wild type 5 3 DNA template 3 strand 5 m. RNA 3 5 Protein Stop Amino end Carboxyl end missing 5 3 3 5 missing 5 Frameshift causing extensive missense (1 base-pair deletion) 3
- Slides: 28