The use of SNP chips for selection of


















- Slides: 34

The use of SNP chips for selection of dairy cattle Curt Van Tassell Bovine Functional Genomics Laboratory Agricultural Research Service, USDA Beltsville, MD curtvt@ars. usda. gov Applied Genomics for Sustainable Livestock Breeding 1) (

A Little History… 1908 USDA organizes a national milk-recording program based on success in Michigan 1936 The first national sire evaluations are calculated from daughter-dam comparisons 1989 Animal model evaluations use the relationships among all cows and bulls 1997 Inter. BULL evaluations introduced to incorporate foreign daughter information 2009 First official genetic evaluations incorporating genomic data Applied Genomics for Sustainable Livestock Breeding 2) (

History of genomic evaluations Oct. 2007 Bovine. SNP 50 Bead. Chip available Apr. 2008 First unofficial evaluation released Jan. 2009 Genomic evaluations official for Holstein and Jersey Aug. 2009 Official for Brown Swiss Sept. 2010 Unofficial evaluations from 3 K chip released Dec. 2010 3 K genomic evaluations official Applied Genomics for Sustainable Livestock Breeding 3) (

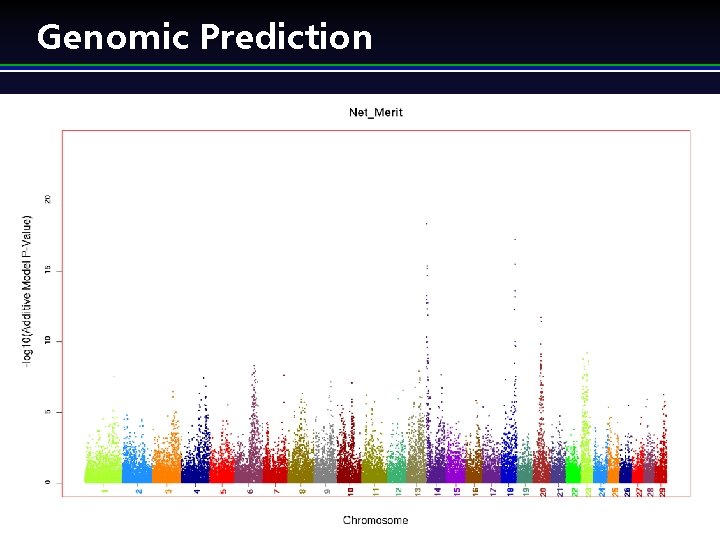
Genome Selection l “Train” system using phenotypic and genotypic data w l l Large regression system Predict genetic merit at birth by combining pedigree merit and merit predicted from SNP Final genetic predictions are transparent to technology Applied Genomics for Sustainable Livestock Breeding 4) (

Genomic Prediction Applied Genomics for Sustainable Livestock Breeding 5) (

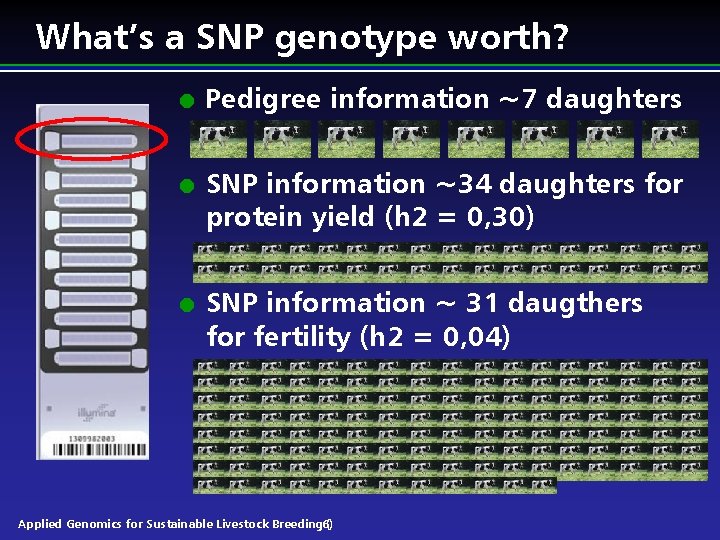
What’s a SNP genotype worth? l l l Pedigree information ~7 daughters SNP information ~34 daughters for protein yield (h 2 = 0, 30) SNP information ~ 31 daugthers for fertility (h 2 = 0, 04) Applied Genomics for Sustainable Livestock Breeding 6) (

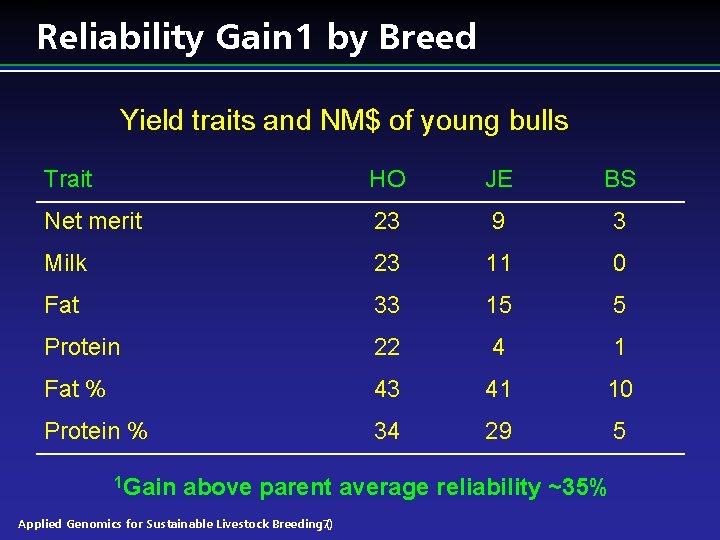
Reliability Gain 1 by Breed Yield traits and NM$ of young bulls Trait HO JE BS Net merit 23 9 3 Milk 23 11 0 Fat 33 15 5 Protein 22 4 1 Fat % 43 41 10 Protein % 34 29 5 1 Gain above parent average reliability ~35% Applied Genomics for Sustainable Livestock Breeding 7) (

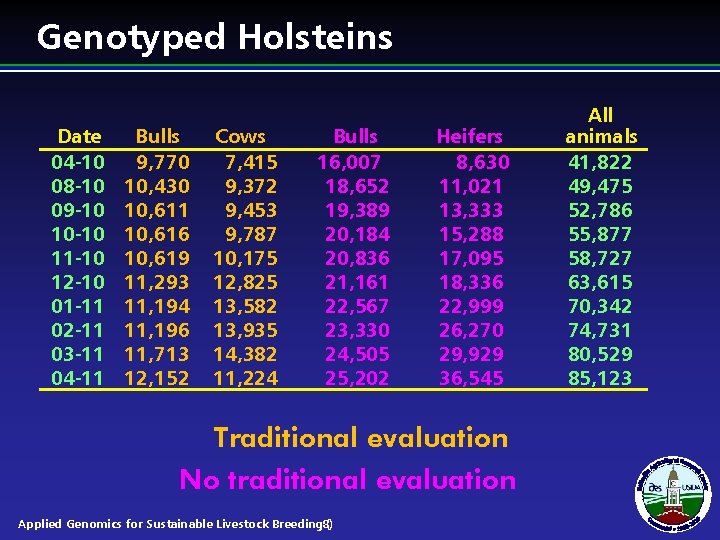
Genotyped Holsteins Date 04 -10 08 -10 09 -10 10 -10 11 -10 12 -10 01 -11 02 -11 03 -11 04 -11 Bulls 9, 770 10, 430 10, 611 10, 616 10, 619 11, 293 11, 194 11, 196 11, 713 12, 152 Cows 7, 415 9, 372 9, 453 9, 787 10, 175 12, 825 13, 582 13, 935 14, 382 11, 224 Bulls 16, 007 18, 652 19, 389 20, 184 20, 836 21, 161 22, 567 23, 330 24, 505 25, 202 Heifers 8, 630 11, 021 13, 333 15, 288 17, 095 18, 336 22, 999 26, 270 29, 929 36, 545 Traditional evaluation No traditional evaluation Applied Genomics for Sustainable Livestock Breeding 8) ( All animals 41, 822 49, 475 52, 786 55, 877 58, 727 63, 615 70, 342 74, 731 80, 529 85, 123

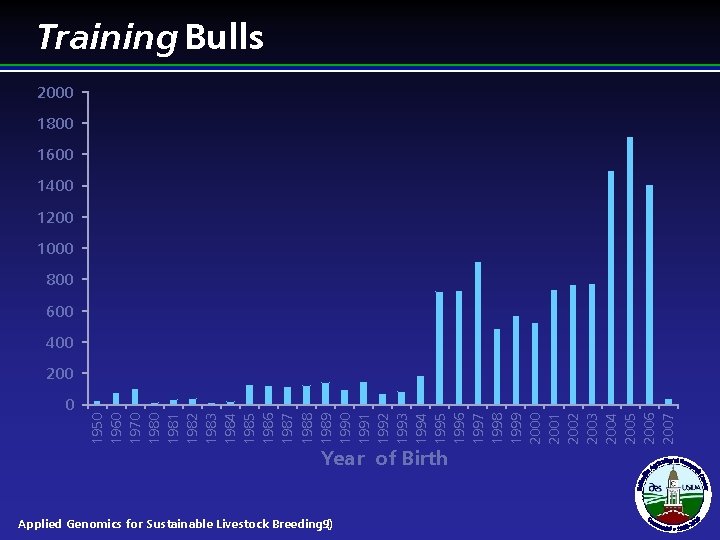
0 Year of Birth 1950 1960 1970 1981 1982 1983 1984 1985 1986 1987 1988 1989 1990 1991 1992 1993 1994 1995 1996 1997 1998 1999 2000 2001 2002 2003 2004 2005 2006 2007 Training Bulls 2000 1800 1600 1400 1200 1000 800 600 400 200 Applied Genomics for Sustainable Livestock Breeding 9) (

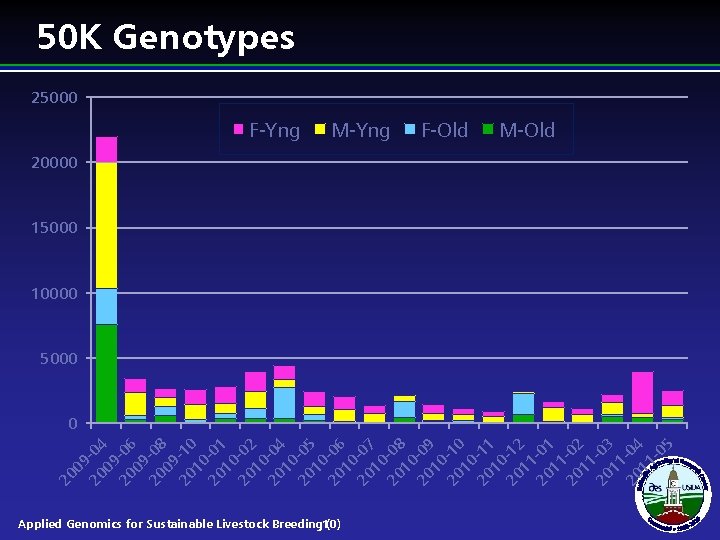
20 -04 09 20 -06 09 20 -08 09 20 -10 10 20 -01 10 20 -02 10 20 -04 10 20 -05 10 20 -06 10 20 -07 10 20 -08 10 20 -09 10 20 -10 10 20 -11 10 20 -12 11 20 -01 11 20 -02 11 20 -03 11 20 -04 11 -0 5 09 20 50 K Genotypes 25000 F-Yng M-Yng Applied Genomics for Sustainable Livestock Breeding 10) ( F-Old M-Old 20000 15000 10000 5000 0

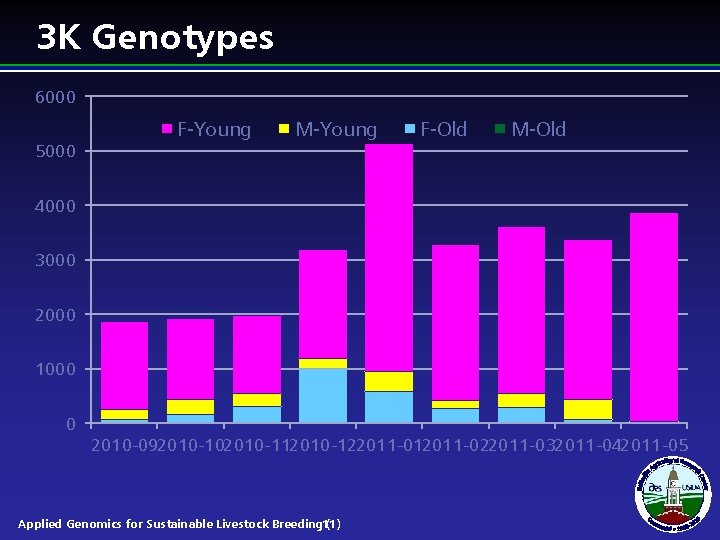
3 K Genotypes 6000 5000 F-Young M-Young F-Old M-Old 4000 3000 2000 1000 0 2010 -092010 -102010 -112010 -122011 -012011 -022011 -032011 -042011 -05 Applied Genomics for Sustainable Livestock Breeding 11) (

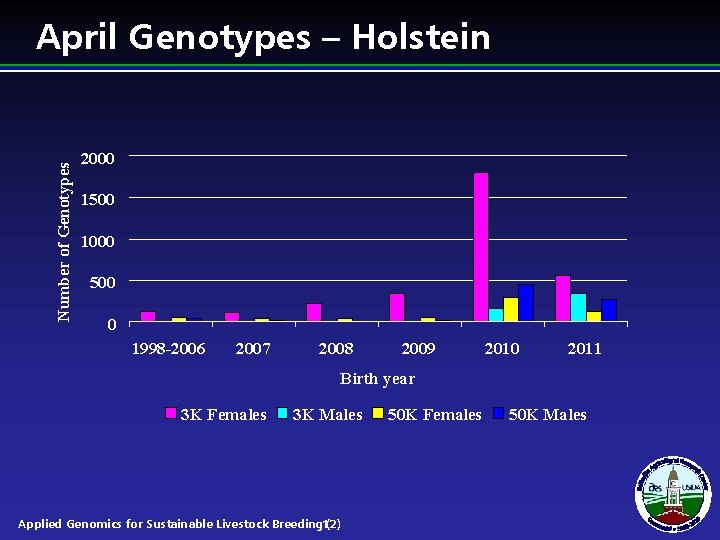
Number of Genotypes April Genotypes – Holstein 2000 1500 1000 500 0 1998 -2006 2007 2008 2009 2010 2011 Birth year 3 K Females 3 K Males Applied Genomics for Sustainable Livestock Breeding 12) ( 50 K Females 50 K Males

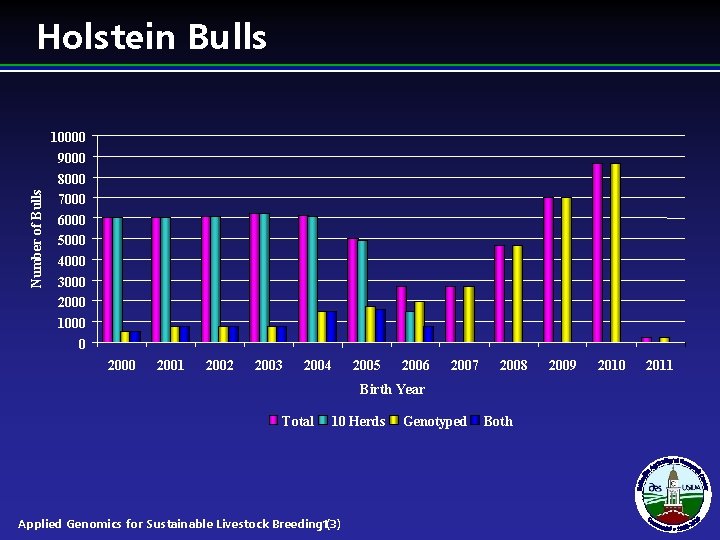
Number of Bulls Holstein Bulls 10000 9000 8000 7000 6000 5000 4000 3000 2000 1000 0 2001 2002 2003 2004 2005 2006 2007 2008 Birth Year Total 10 Herds Applied Genomics for Sustainable Livestock Breeding 13) ( Genotyped Both 2009 2010 2011

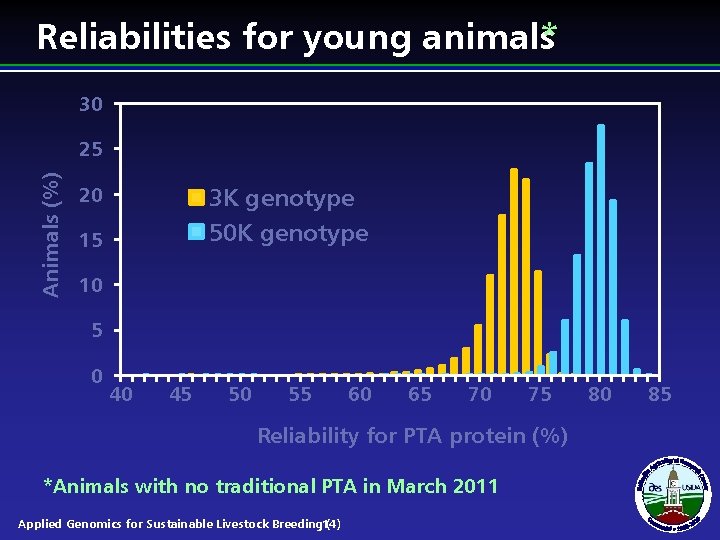
Reliabilities for young animals* 30 Animals (%) 25 20 3 K genotype 50 K genotype 15 10 5 0 40 45 50 55 60 65 70 75 Reliability for PTA protein (%) *Animals with no traditional PTA in March 2011 Applied Genomics for Sustainable Livestock Breeding 14) ( 80 85

Determination of Reliability l Sum of genomic relationships with the predictor animals weighted by reliability l Call rate l Expected imputation error rate l Reliability of parent average w Larger impact if dam not genotyped Applied Genomics for Sustainable Livestock Breeding 15) (

Use of genomic evaluations l Determine which young bulls to bring into AI service l Use to select mating sires l Pick bull dams l Market semen from 2 -year-old bulls Applied Genomics for Sustainable Livestock Breeding 16) (

Use of 3 K genomic evaluations l Sort heifers for breeding w Flush w Sexed semen w Beef bull l Confirm parentage to avoid inbreeding l Predict inbreeding depression better l Precision mating considering genomics(future) Applied Genomics for Sustainable Livestock Breeding 17) (

Results Applied Genomics for Sustainable Livestock Breeding 18) (

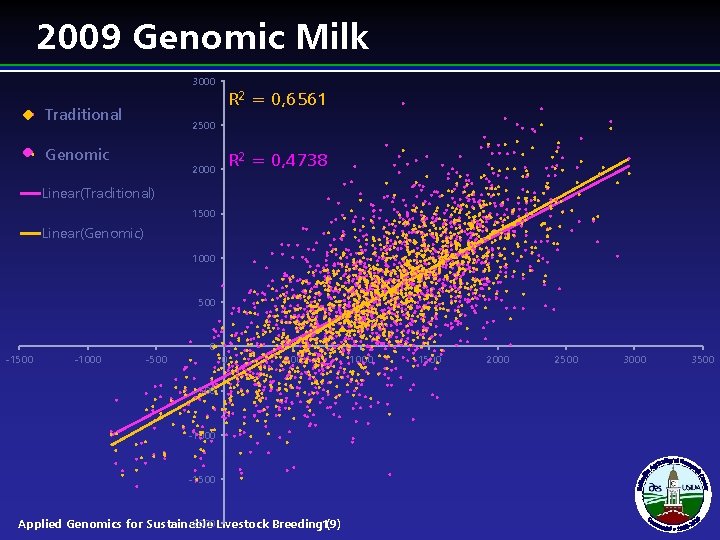
2009 Genomic Milk 3000 R 2 = 0, 6561 Traditional 2500 Genomic R 2 = 0, 4738 2000 Linear(Traditional) 1500 Linear(Genomic) 1000 500 0 -1500 -1000 -500 0 500 -1000 -1500 -2000 Livestock Breeding 19) Applied Genomics for Sustainable ( 1000 1500 2000 2500 3000 3500

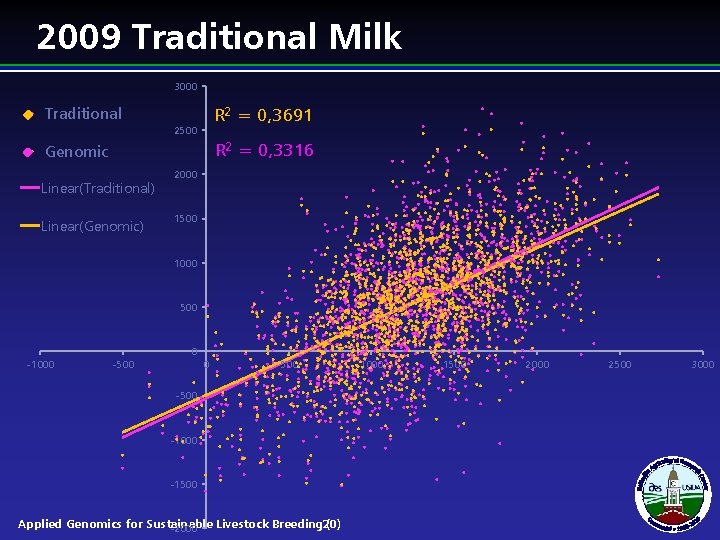
2009 Traditional Milk 3000 R 2 = 0, 3691 Traditional 2500 R 2 = 0, 3316 Genomic Linear(Traditional) Linear(Genomic) 2000 1500 1000 500 0 -1000 -500 0 500 -1000 -1500 Applied Genomics for Sustainable Livestock Breeding 20) ( -2000 1500 2000 2500 3000

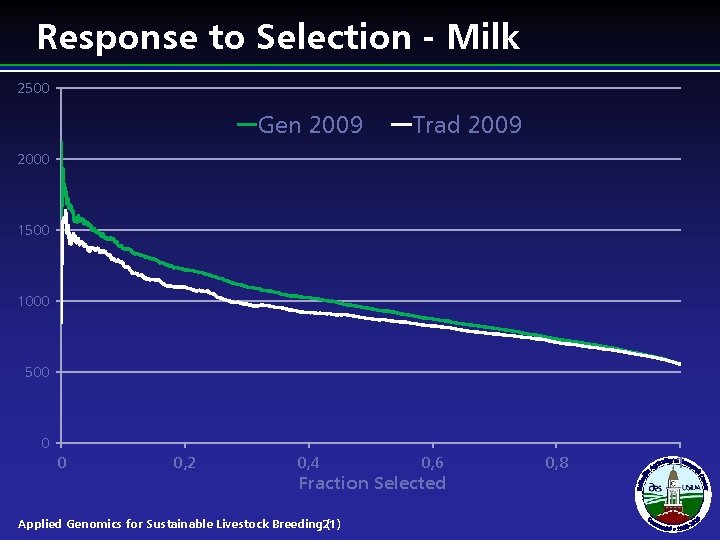
Response to Selection - Milk 2500 Gen 2009 Trad 2009 2000 1500 1000 500 0 0 0, 2 0, 4 0, 6 Fraction Selected Applied Genomics for Sustainable Livestock Breeding 21) ( 0, 8 1

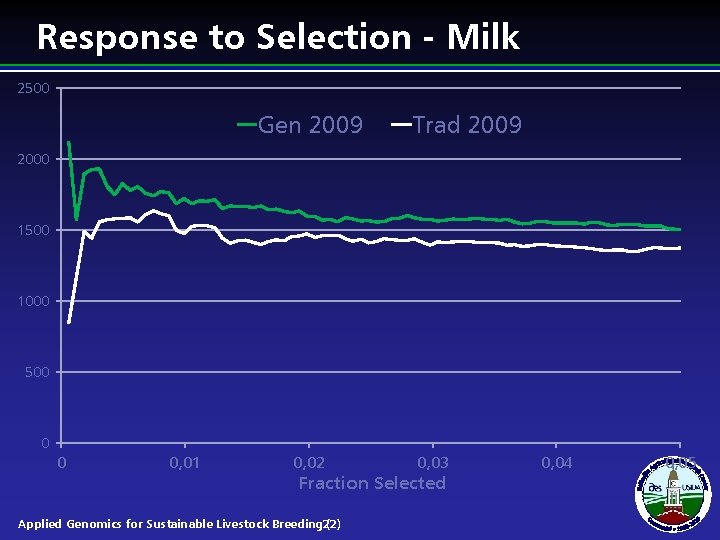
Response to Selection - Milk 2500 Gen 2009 Trad 2009 2000 1500 1000 500 0 0 0, 01 0, 02 0, 03 Fraction Selected Applied Genomics for Sustainable Livestock Breeding 22) ( 0, 04 0, 05

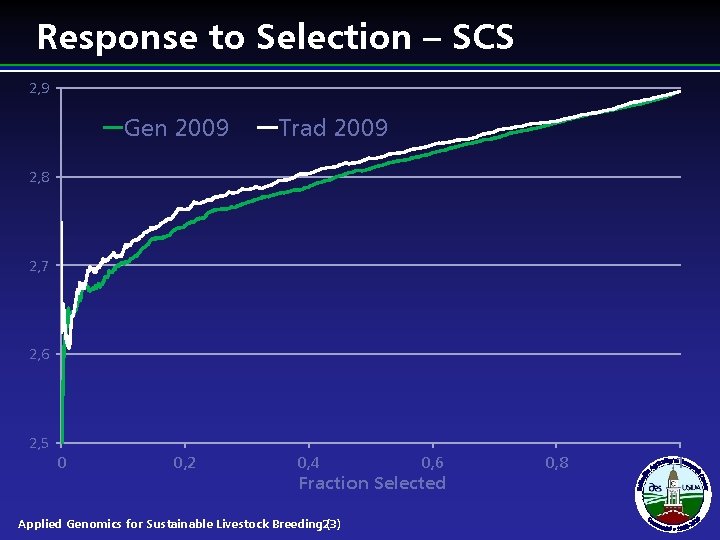
Response to Selection – SCS 2, 9 Gen 2009 Trad 2009 2, 8 2, 7 2, 6 2, 5 0 0, 2 0, 4 0, 6 Fraction Selected Applied Genomics for Sustainable Livestock Breeding 23) ( 0, 8 1

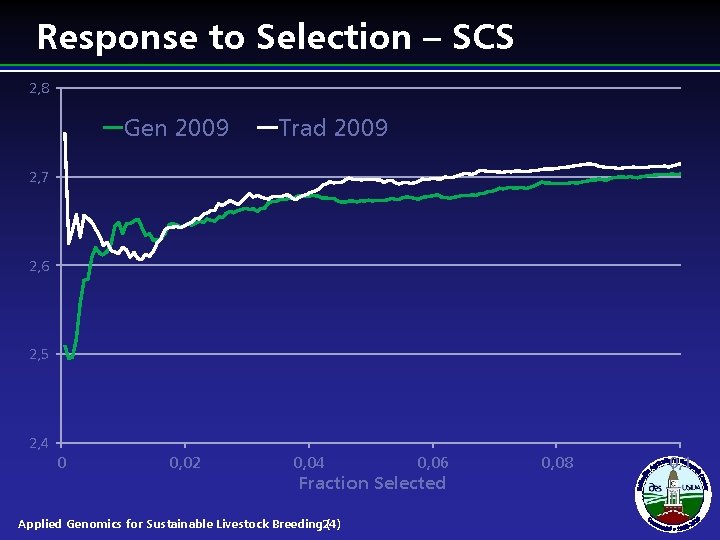
Response to Selection – SCS 2, 8 Gen 2009 Trad 2009 2, 7 2, 6 2, 5 2, 4 0 0, 02 0, 04 0, 06 Fraction Selected Applied Genomics for Sustainable Livestock Breeding 24) ( 0, 08 0, 1

Future Applied Genomics for Sustainable Livestock Breeding 25) (

Increase in accuracy l l Genotyped bulls get traditional evaluation when 5 years old Possible genotyping of 10, 000 bulls with semen in CDDR l Collaboration with more countries l Use of more SNP from HD chips l Full sequencing Applied Genomics for Sustainable Livestock Breeding 26) (

Application to more traits l l l Animal’s genotype is good for all traits Traditional evaluations required for accurate estimates of SNP effects Traditional evaluations not currently available for heat tolerance or feed efficiency Research populations could provide data for traits that are expensive to measure Will resulting evaluations work in target population? Applied Genomics for Sustainable Livestock Breeding 27) (

High density (HD) genotypes l Illumina Bovine HD Bead. Chip available 777, 962 SNP w Collaborations with GBR, ITA, & CAN to provide genotypes w Over 400 genotypes from research projects in database w l Affymetrics HD w 648, 855 SNP w Optimized for genetic coverage w NAAB negotiating plan for use Applied Genomics for Sustainable Livestock Breeding 28) (

Use of HD l l Some increase in accuracy from better tracking of QTL Potential for across breed evaluations Requires few new HD genotypes once adequate base for imputation developed Imputation improvements were particularly beneficial in imputing HD Applied Genomics for Sustainable Livestock Breeding 29) (

Low-Plex Genotyping l Need cost-effective genotyping platform for application to commercial cows w l <$5 / animal Applications: w Parentage, traceability w Relatedness w Targetted panels Applied Genomics for Sustainable Livestock Breeding 30) (

Genome Sequencing? l Whole-genome sequences on individuals will be available in the next few years w l l How will we store and use those data? Not feasible to calculate effects for 3, 000, 000 nucleotides or even 3, 000 SNP May be best to fine map then target genotype… Applied Genomics for Sustainable Livestock Breeding 31) (

Exome Sequencing? l Capture and sequence coding part of genome and regulatory regions ~50 -60 Mb l Higher likelihood of functional SNPs l Less data than complete sequence l Soon price difference irrelevant? Applied Genomics for Sustainable Livestock Breeding 32) (

Summary l l l Extraordinarily rapid implementation of genomic evaluations Young-bull acquisition and marketing now based on genomic evaluations Genotyping of many females because of 3 K chip Applied Genomics for Sustainable Livestock Breeding 33) (

Applied Genomics for Sustainable Livestock Breeding 34) (