The Uni Prot knowledgebase www uniprot org a

The Uni. Prot knowledgebase www. uniprot. org a hub of integrated protein data Marie-Claude. Blatter@isb-sib. ch Swiss-Prot group, Geneva SIB Swiss Institute of Bioinformatics

Science cover, february 2011

data protein sequence knowledge functional information

Uni. Prot consortium EBI : European Bioinformatics Institute (UK) SIB : Swiss Institute of Bioinformatics (CH) PIR : Protein information resource (US)

www. uniprot. org
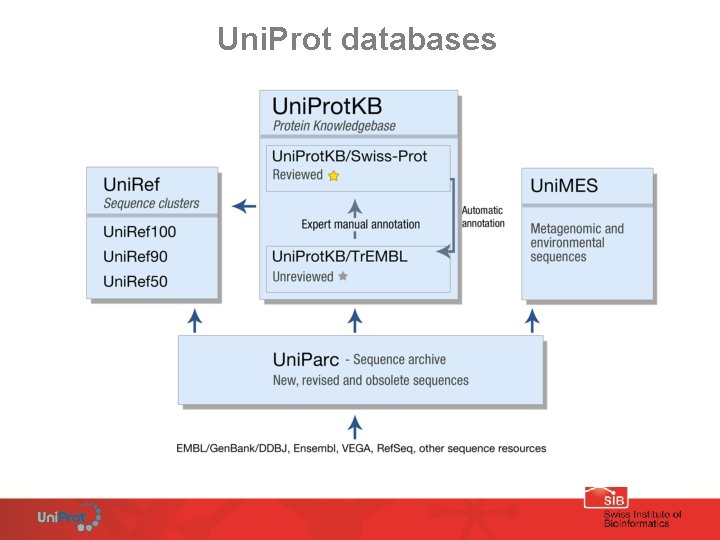
Uni. Prot databases

Uni. Prot. KB: Uni. Prot. KB protein sequence knowledgebase, 2 sections Uni. Prot. KB/Swiss-Prot and Uni. Prot. KB/Tr. EMBL (query, Blast, download) (~15 mo entries) Uni. Parc: Uni. Parc protein sequence archive (ENA equivalent at the protein level). Each entry contains a protein sequence with cross-links to other databases where you find the sequence (active or not). Not annotated (query, Blast, download) (~25 mo entries) Uni. Ref: Uni. Ref 3 clusters of protein sequences with 100, 90 and 50 % identity; useful to speed up sequence similarity search (BLAST) (query, Blast, download) (Uni. Ref 100 10 mo entries; Uni. Ref 90 7 mo entries; Uni. Ref 50 3. 3 mo entries) Uni. MES: Uni. MES protein sequences derived from metagenomic projects (mostly Global Ocean Sampling (GOS)) (download) (8 mo entries, included in Uni. Parc)
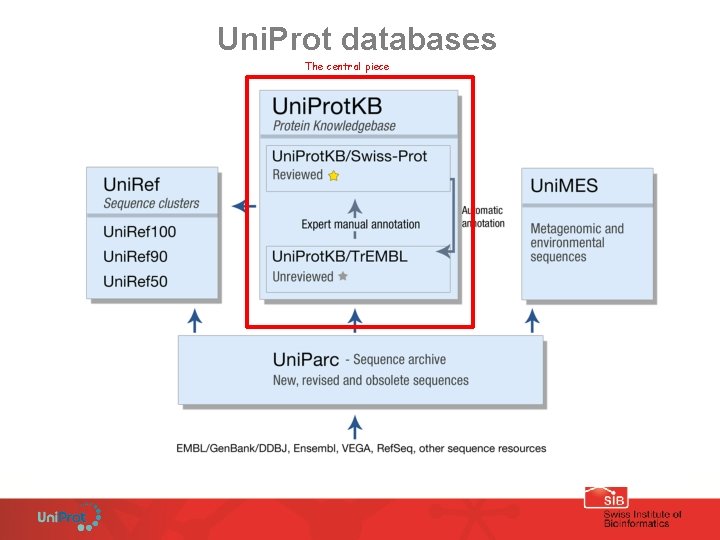
Uni. Prot databases The central piece

Uni. Prot. KB an encyclopedia on proteins composed of 2 sections Uni. Prot. KB/Tr. EMBL and Uni. Prot. KB/Swiss-Prot unreviewed and reviewed automatically annotated and manually annotated released every 4 weeks

Uni. Prot. KB Origin of protein sequences Uni. Prot. KB protein sequences are mainly derived from - INSDC (translated submitted coding sequences - CDS) Ensembl (gene prediction ) and Ref. Seq sequences Sequences of PDB structures Direct submission or sequences scanned from literature 85 % 15 % Notes: - Uni. Prot is not doing any gene prediction - Most non-germline immunoglobulins, T-cell receptors , most patent sequences, highly over-represented data (e. g. viral antigens), pseudogenes sequences are excluded from Uni. Prot. KB, - but stored in Uni. Parc - Data from the PIR database have been integrated in Uni. Prot. KB since 2003.

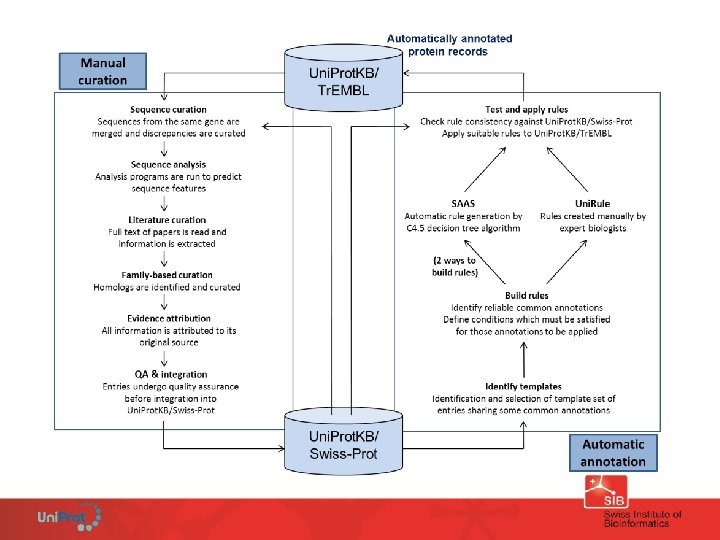
EMBL Manual annotation of the sequence and associated biological information Tr. EMBL Automated extraction of protein sequence (translated CDS), gene name and references. Automated annotation Swiss-Prot

Uni. Prot. KB/Tr. EMBL unreviewed Automatic annotation released every 4 weeks
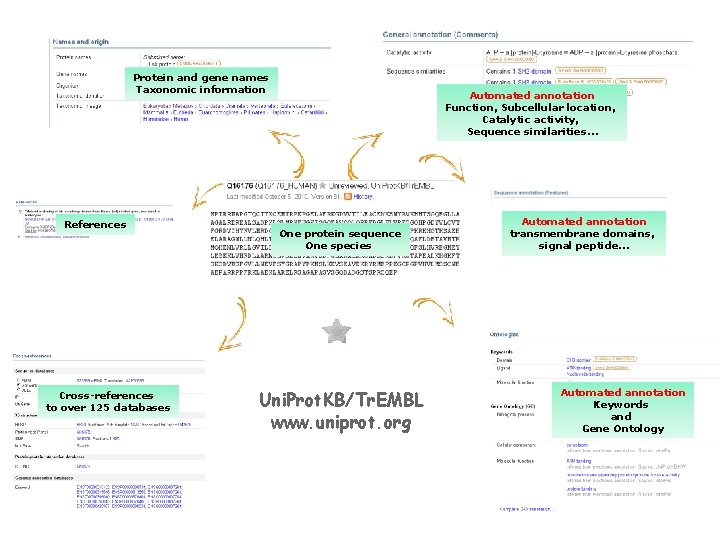
Protein and gene names Taxonomic information References Cross-references to over 125 databases Automated annotation Function, Subcellular location, Catalytic activity, Sequence similarities… One protein sequence One species Uni. Prot. KB/Tr. EMBL www. uniprot. org Automated annotation transmembrane domains, signal peptide… Automated annotation Keywords and Gene Ontology

Uni. Prot. KB/Tr. EMBL Automatic annotation Protein sequence - The quality of the protein sequences is dependent on the information provided by the submitter of the original nucleotide entry (CDS) or of the gene prediction pipeline (i. e. Ensembl). - 100% identical sequences (same length, same organism are merged automatically). Biological information Sources of annotation - Provided by the submitter (EMBL, PDB, TAIR…) - From automated annotation (automated generated annotation rules (i. e. SAAS) and/or manually generated annotation rules (i. e. Uni. Rule))
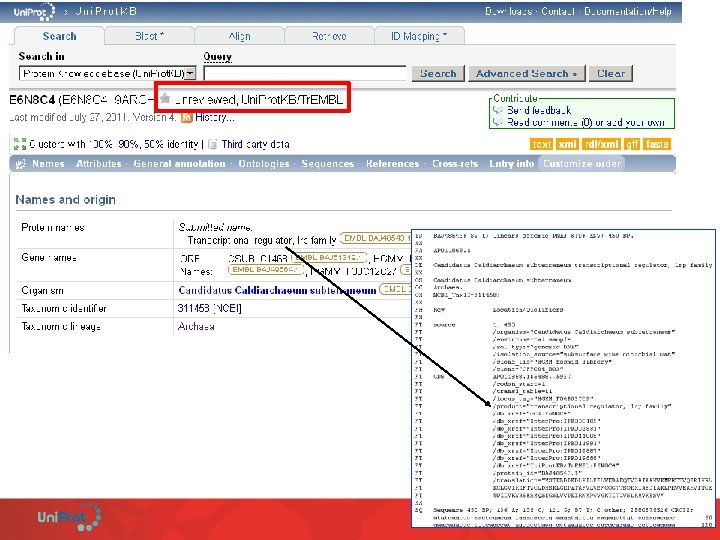
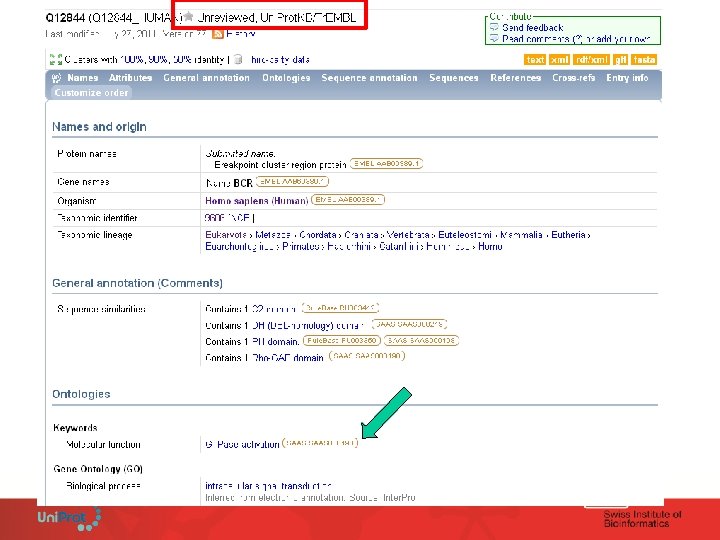

Uni. Prot. KB/Tr. EMBL Example of fully automatic annotation: SAAS • Rules are derived from the Uni. Prot. KB/Swiss-Prot manual annotation. • Fully automated rule generation based on C 4. 5 decision tree algorithm. • One annotation, one rule. • High stringency – require 99% or greater estimated precision to generate annotation (test on Uni. Prot. KB/Swiss-Prot) • Rules are produced, updated and validated at each release.

Uni. Prot. KB/Swiss-Prot reviewed manually annotated released every 4 weeks
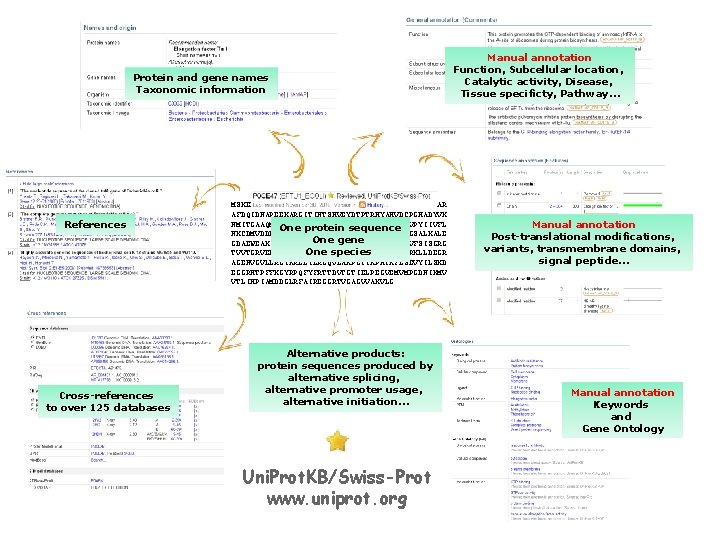
Protein and gene names Taxonomic information References Cross-references to over 125 databases MSKEKFERTKPHVNVGTIGHVDHGKTTLTAAITTVLAKTYGGAAR AFDQIDNAPEEKARGITINTSHVEYDTPTRHYAHVDCPGHADYVK NMITGAAQMDGAILVVAATDGPMPQTREHILLGRQVGVPYIIVFL One protein sequence NKCDMVDDEELLELVEMEVRELLSQYDFPGDDTPIVRGSALKALE One gene GDAEWEAKILELAGFLDSYIPEPERAIDKPFLLPIEDVFSISGRG One species TVVTGRVERGIIKVGEEVEIVGIKETQKSTCTGVEMFRKLLDEGR AGENVGVLLRGIKREEIERGQVLAKPGTIKPHTKFESEVYILSKD EGGRHTPFFKGYRPQFYFRTTDVTGTIELPEGVEMVMPGDNIKMV VTLIHPIAMDDGLRFAIREGGRTVGAGVVAKVLG Alternative products: protein sequences produced by alternative splicing, alternative promoter usage, alternative initiation… Uni. Prot. KB/Swiss-Prot www. uniprot. org Manual annotation Function, Subcellular location, Catalytic activity, Disease, Tissue specificty, Pathway… Manual annotation Post-translational modifications, variants, transmembrane domains, signal peptide… Manual annotation Keywords and Gene Ontology

Uni. Prot. KB/Swiss-Prot Manual annotation 1. Protein sequence (merge available CDS, annotate sequence discrepancies, report sequencing mistakes…) 2. Biological information (sequence analysis, extract literature information, ortholog data propagation, …)

Uni. Prot. KB/Swiss-Prot 1 - Protein sequence curation

Uni. Prot. KB/Swiss-Prot a gene-centric view of the protein space 1 entry <-> 1 gene (1 species) The displayed protein sequence: …canonical, representative, consensus… + alternative sequences (described within the entry)

What is the current status? • At least 20% of Swiss-Prot entries required a minimal amount of curation effort so as to obtain the “correct” sequence. • Typical problems – unsolved conflicts – uncorrected initiation sites – frameshifts – wrong gene prediction – other ‘problems’
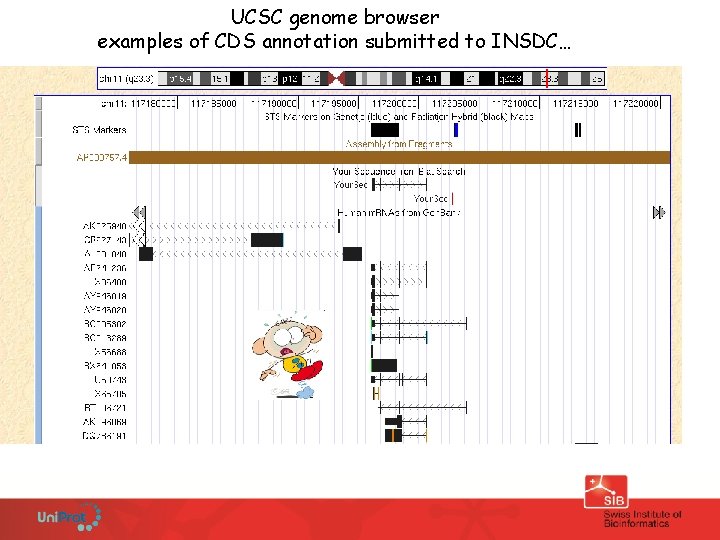
UCSC genome browser examples of CDS annotation submitted to INSDC…

Uni. Prot. KB/Swiss-Prot 2 - Biological data curation

Extract literature information and protein sequence analysis maximum usage of controlled vocabulary Uni. Prot. KB/Swiss-Prot gathers data form multiple sources: - publications (literature/Pubmed) - prediction programs (Prosite, TMHMM, …) - contacts with experts - other databases - nomenclature committees An evidence attribution system allows to easily trace the source of each annotation

Protein and gene names

General annotation (Comments) …enable researchers to obtain a summary of what is known about a protein… www. uniprot. org
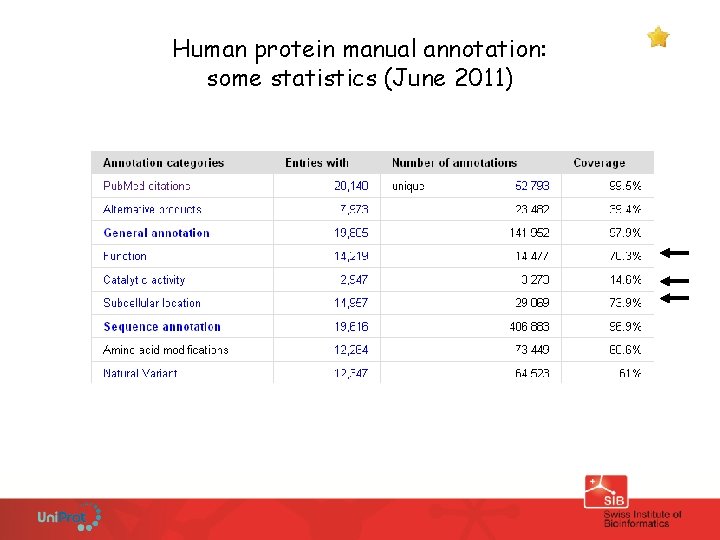
Human protein manual annotation: some statistics (June 2011)

Sequence annotation (Features) …enable researchers to obtain a summary of what is known about a protein… www. uniprot. org

Non-experimental qualifiers Uni. Prot. KB/Swiss-Prot considers both experimental and predicted data and makes a clear distinction between both Type of evidence Qualifier Strong experimental evidence None or Ref. X Light experimental evidence Probable Inferred by similarity with homologous protein By similarity Inferred by prediction Potential
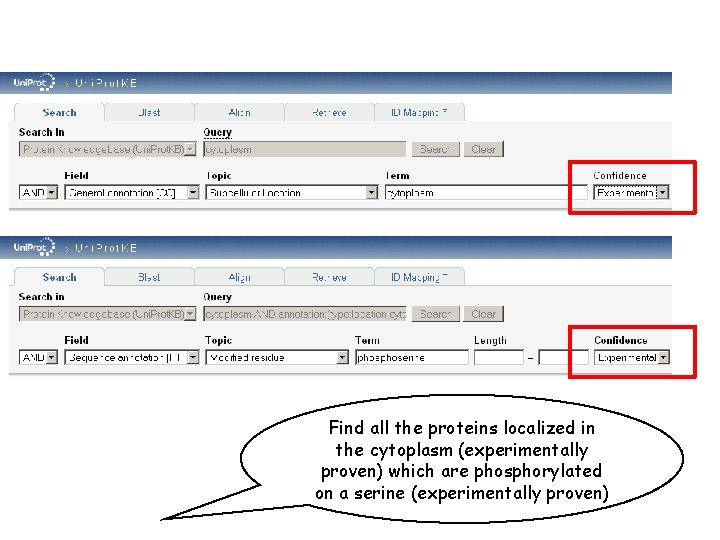
Find all the proteins localized in the cytoplasm (experimentally proven) which are phosphorylated on a serine (experimentally proven)

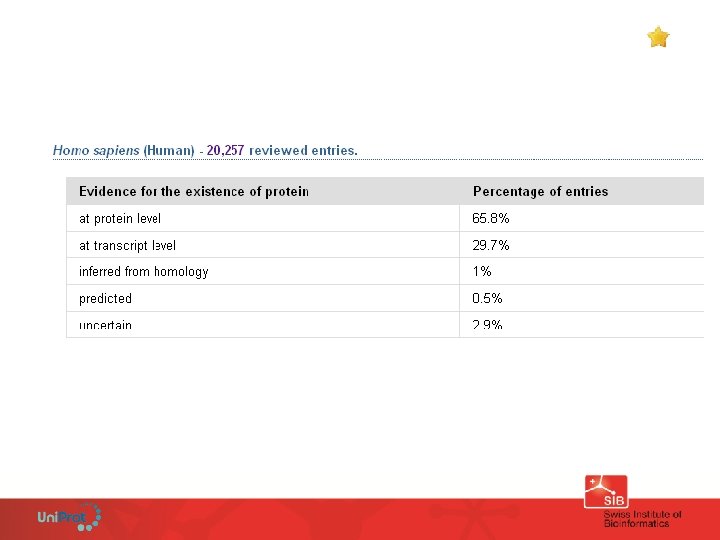
‘Protein existence’ tag • The ‘Protein existence’ tag indicates what is the evidence for the existence of a given protein; • Different qualifiers: 1. Evidence at protein level (~18%) (MS, western blot (tissue specificity), immuno (subcellular location), …) 2. Evidence at transcript level (~19%) 3. Inferred from homology (~58 %) 4. Predicted (~5%) 5. Uncertain (mainly in Tr. EMBL) http: //www. uniprot. org/docs/pe_criteria

Uni. Prot. KB Additional information can be found in the cross-references (to more than 140 databases)
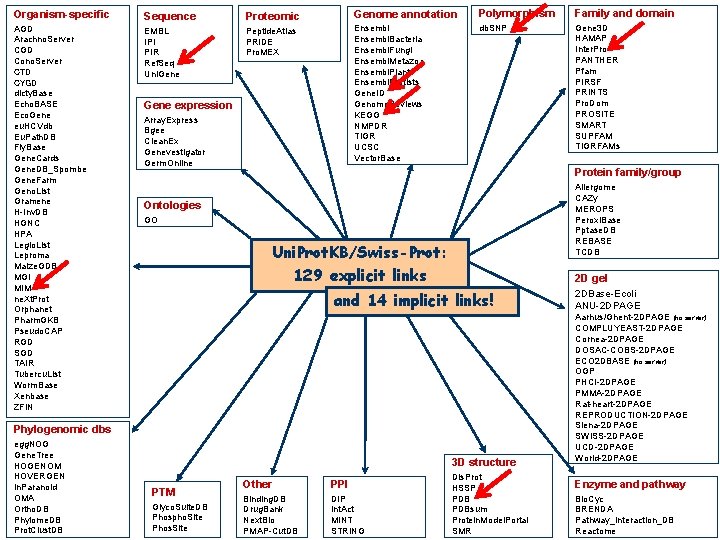
Organism-specific AGD Arachno. Server CGD Cono. Server CTD CYGD dicty. Base Echo. BASE Eco. Gene eu. HCVdb Eu. Path. DB Fly. Base Gene. Cards Gene. DB_Spombe Gene. Farm Geno. List Gramene H-Inv. DB HGNC HPA Legio. List Leproma Maize. GDB MGI MIM ne. Xt. Prot Orphanet Pharm. GKB Pseudo. CAP RGD SGD TAIR Tubercu. List Worm. Base Xenbase ZFIN Sequence EMBL IPI PIR Ref. Seq Uni. Gene Proteomic Genome annotation Polymorphism Family and domain Peptide. Atlas PRIDE Pro. MEX Ensembl. Bacteria Ensembl. Fungi Ensembl. Metazoa Ensembl. Plants Ensembl. Protists Gene. ID Genome. Reviews KEGG NMPDR TIGR UCSC Vector. Base db. SNP Gene 3 D HAMAP Inter. Pro PANTHER Pfam PIRSF PRINTS Pro. Dom PROSITE SMART SUPFAM TIGRFAMs Gene expression Array. Express Bgee Clean. Ex Genevestigator Germ. Online Protein family/group Allergome CAZy MEROPS Peroxi. Base Pptase. DB REBASE TCDB Ontologies GO Uni. Prot. KB/Swiss-Prot: 129 explicit links 2 D gel and 14 implicit links! Phylogenomic dbs egg. NOG Gene. Tree HOGENOM HOVERGEN In. Paranoid OMA Ortho. DB Phylome. DB Prot. Clust. DB 3 D structure PTM Glyco. Suite. DB Phospho. Site Phos. Site Other PPI Binding. DB Drug. Bank Next. Bio PMAP-Cut. DB DIP Int. Act MINT STRING Dis. Prot HSSP PDBsum Protein. Model. Portal SMR 2 DBase-Ecoli ANU-2 DPAGE Aarhus/Ghent-2 DPAGE (no server) COMPLUYEAST-2 DPAGE Cornea-2 DPAGE DOSAC-COBS-2 DPAGE ECO 2 DBASE (no server) OGP PHCI-2 DPAGE PMMA-2 DPAGE Rat-heart-2 DPAGE REPRODUCTION-2 DPAGE Siena-2 DPAGE SWISS-2 DPAGE UCD-2 DPAGE World-2 DPAGE Enzyme and pathway Bio. Cyc BRENDA Pathway_Interaction_DB Reactome

The Uni. Prot web site www. uniprot. org • Powerful search engine, google-like and easy-to-use, but also supports very directed field searches • Scoring mechanism presenting relevant matches first • Entry views, search result views and downloads are customizable • The URL of a result page reflects the query; all pages and queries are bookmarkable, supporting programmatic access • Search, Blast, Align, Retrieve, ID mapping
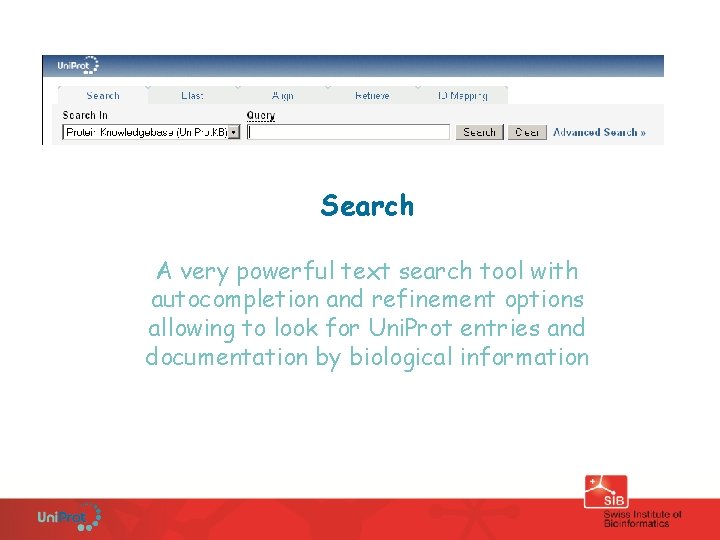
Search A very powerful text search tool with autocompletion and refinement options allowing to look for Uni. Prot entries and documentation by biological information

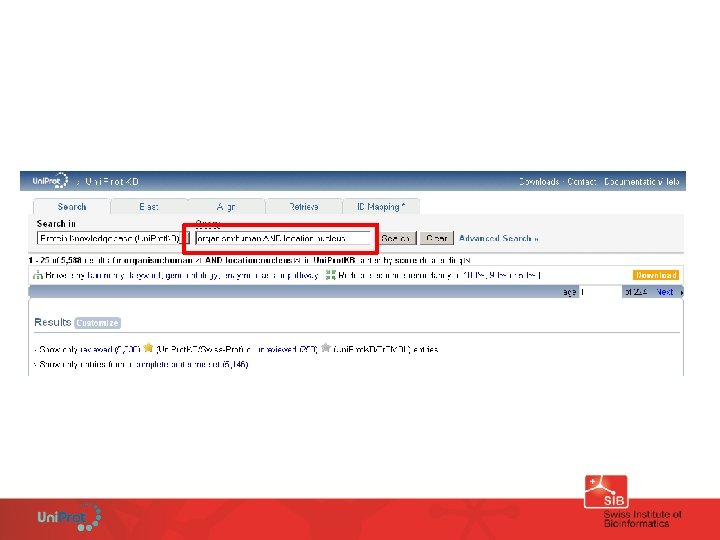
Find all human proteins located in the nucleus
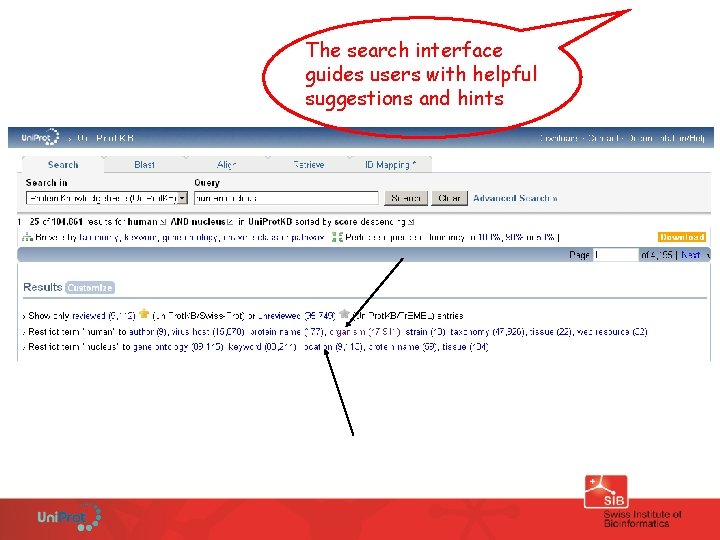
The search interface guides users with helpful suggestions and hints
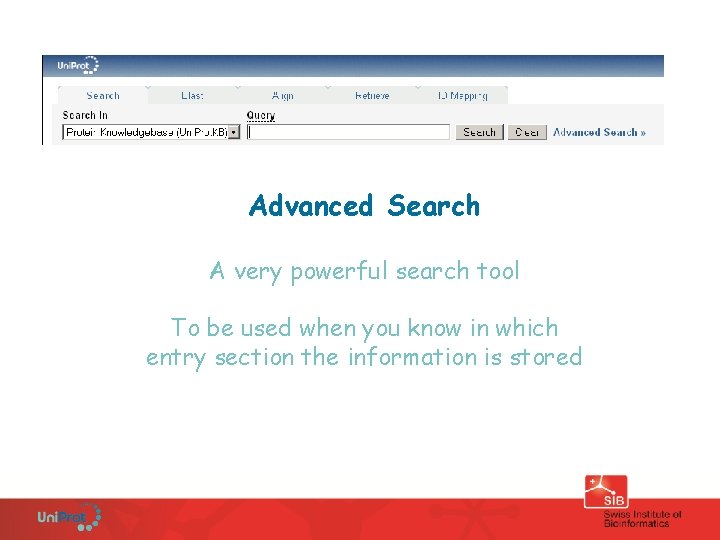
Advanced Search A very powerful search tool To be used when you know in which entry section the information is stored
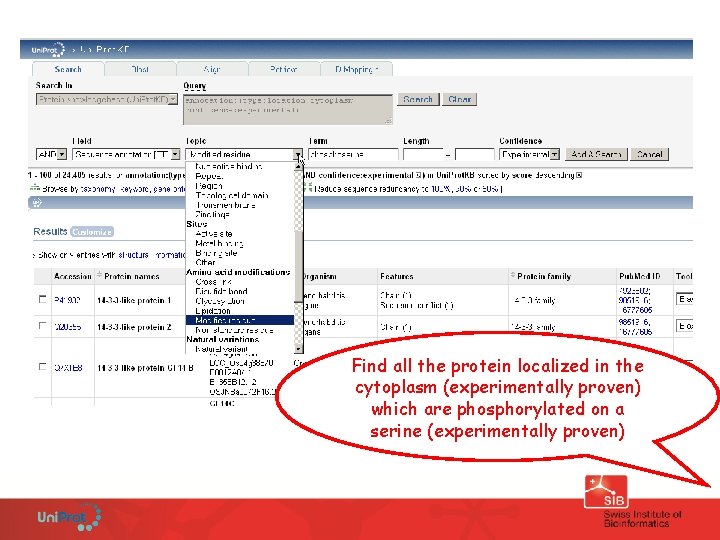
Find all the protein localized in the cytoplasm (experimentally proven) which are phosphorylated on a serine (experimentally proven)
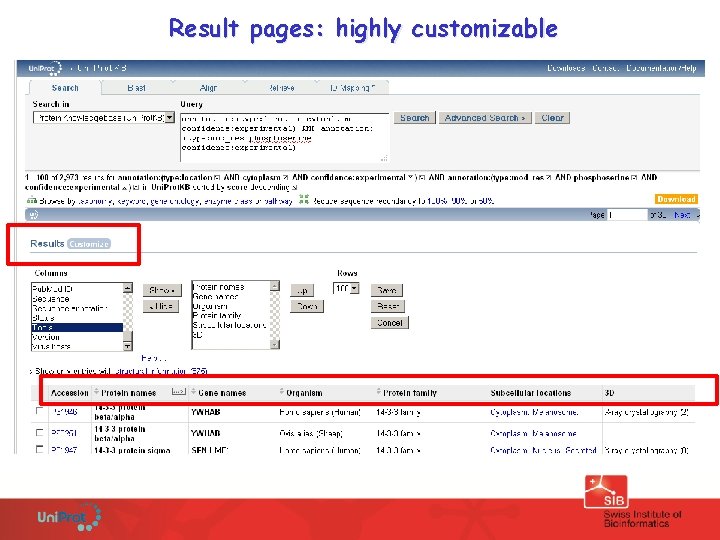
Result pages: highly customizable
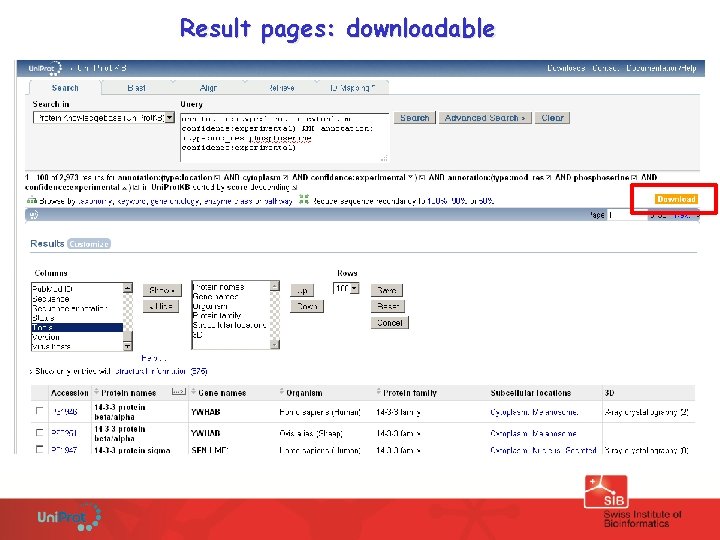
Result pages: downloadable
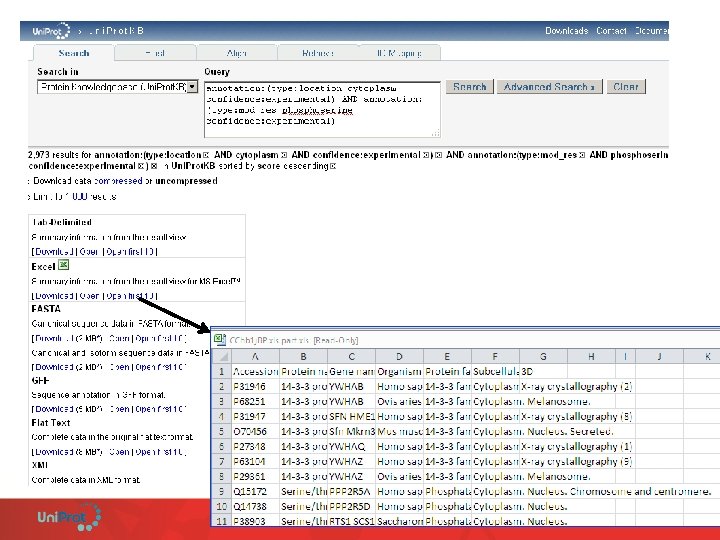
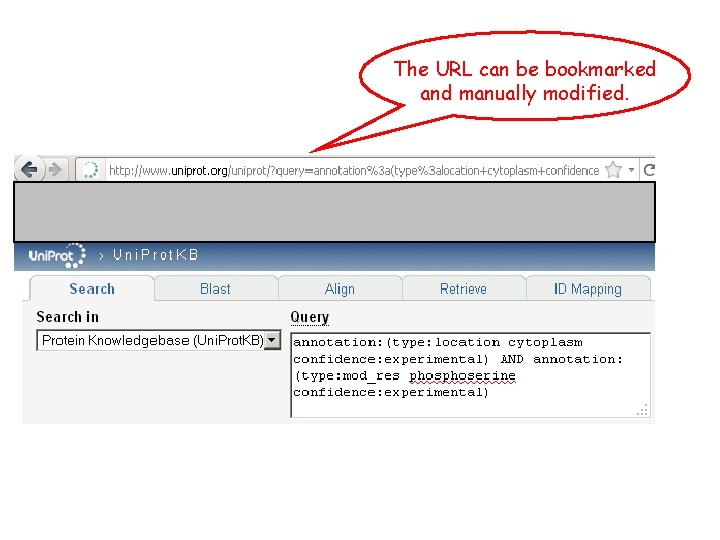
The URL can be bookmarked and manually modified.
Blast A tool associated with the standard options to search sequences in different Uni. Prot databases and data sets

Blast: customize the result display
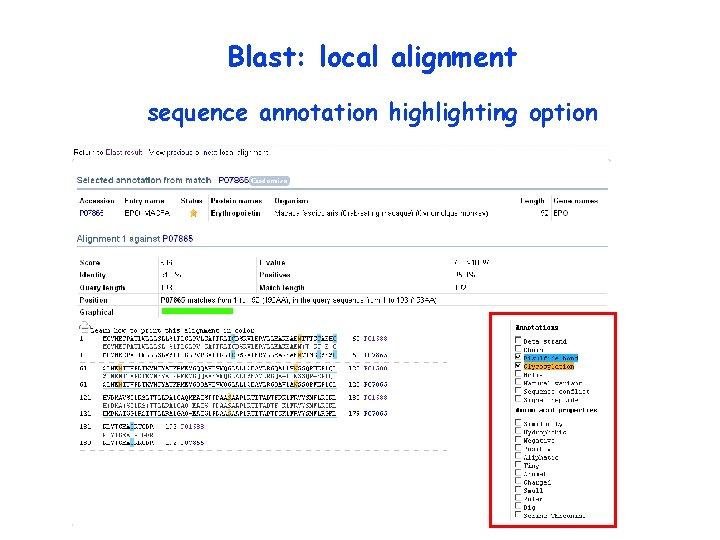
Blast: local alignment sequence annotation highlighting option
Align A Clustal. W multiple alignment tool with sequence annotation highlighting option
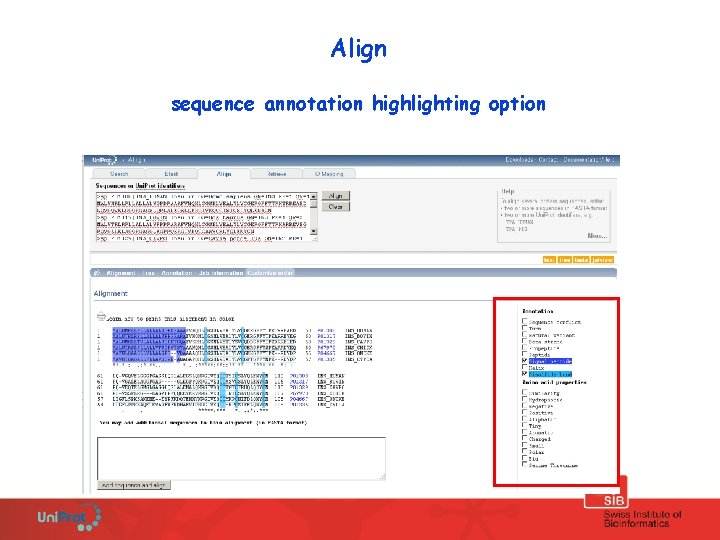
Align sequence annotation highlighting option
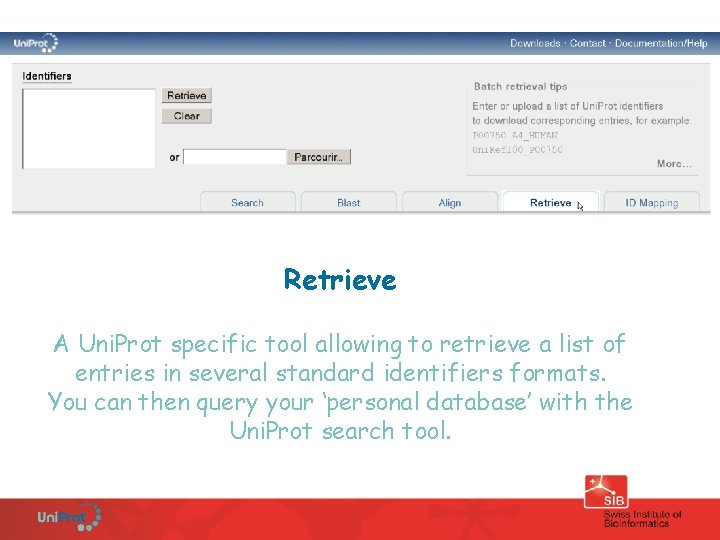
Retrieve A Uni. Prot specific tool allowing to retrieve a list of entries in several standard identifiers formats. You can then query your ‘personal database’ with the Uni. Prot search tool.
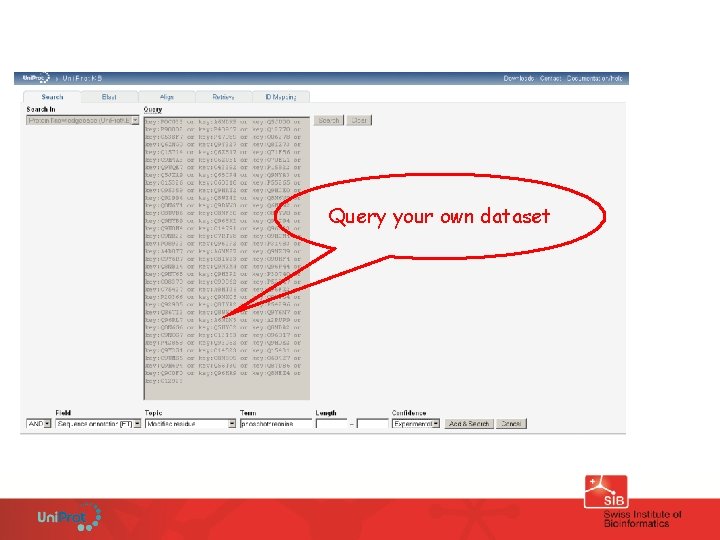
Query your own dataset
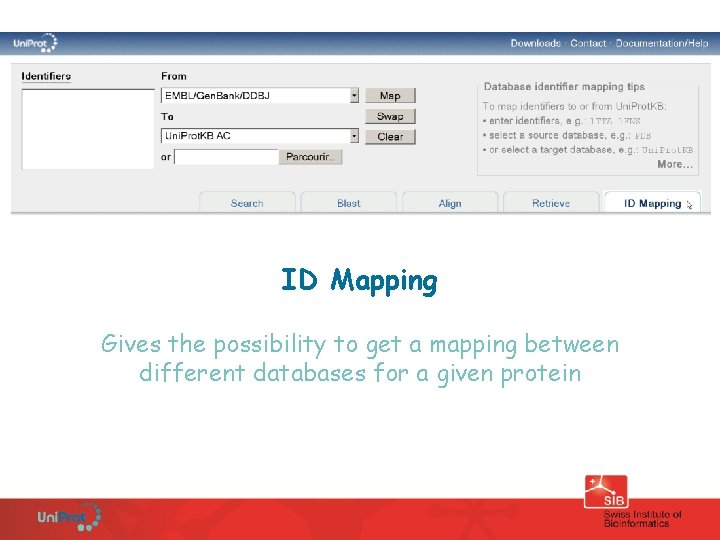
ID Mapping Gives the possibility to get a mapping between different databases for a given protein

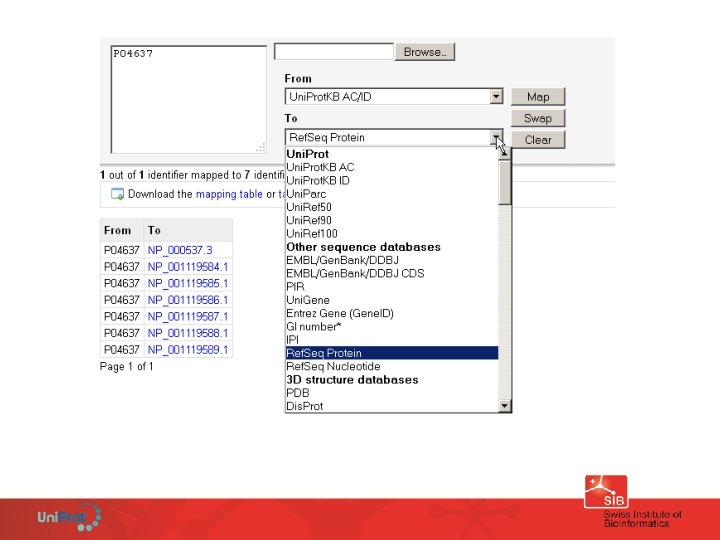
These identifiers are all pointing to a TP 53 (p 53) protein sequence ! P 04637, NP_000537, NP_001119584. 1, NP_001119585. 1, NP_001119584. 1, ENSG 00000141510, CCDS 11118, UPI 000002 ED 67, IPI 00025087, etc.

Download

Download Uni. Prot http: //www. uniprot. org/downloads
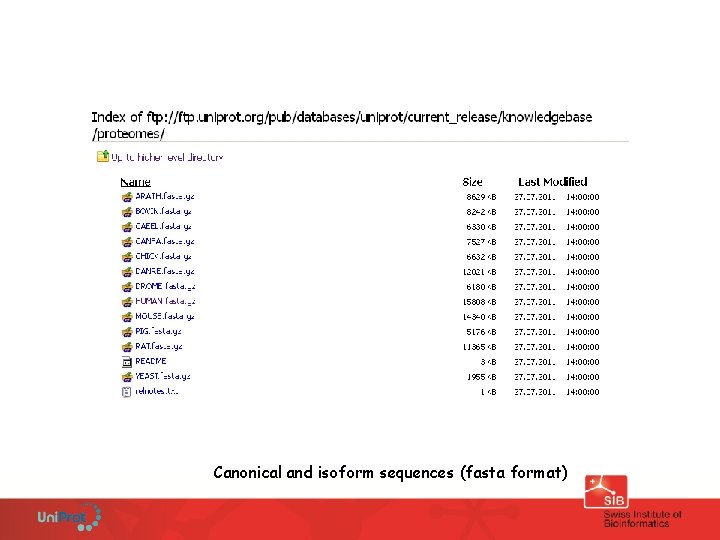
Canonical and isoform sequences (fasta format)

A few words on the Uni. Prot ‘complete proteome’ sequence sets…

Uni. Prot. KB - complete proteomes 2’ 747 complete proteomes ü Genome completely sequenced ü Proteins mapped to the genome ü Entries tagged with the KW ‘Complete proteome’ ü Uni. Prot. KB/Swiss-Prot isoform sequences are available in FASTA format only Fully manually reviewed (e. g. S. cerevisiae) Partially manually reviewed (e. g. Homo sapiens) Unreviewed (e. g. Acinetobacter baumannii (strain 1656 -2))

Uni. Prot. KB - complete proteomes Can be downloaded: ü From our complete proteome page www. uniprot. org/taxonomy/complete-proteomes ü From the ‘ftp download ‘ page ü By querying Uni. Prot. KB + download Query: organism: 93062 AND keyword: "complete proteome" Additional information: www. uniprot. org/faq/15
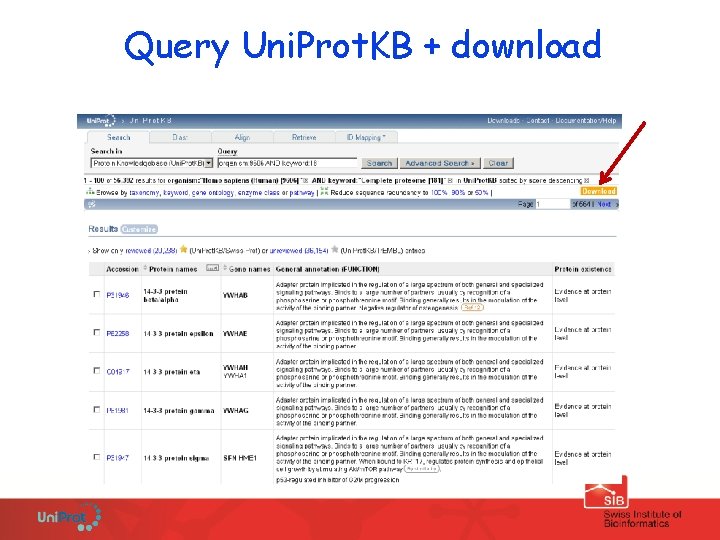
Query Uni. Prot. KB + download
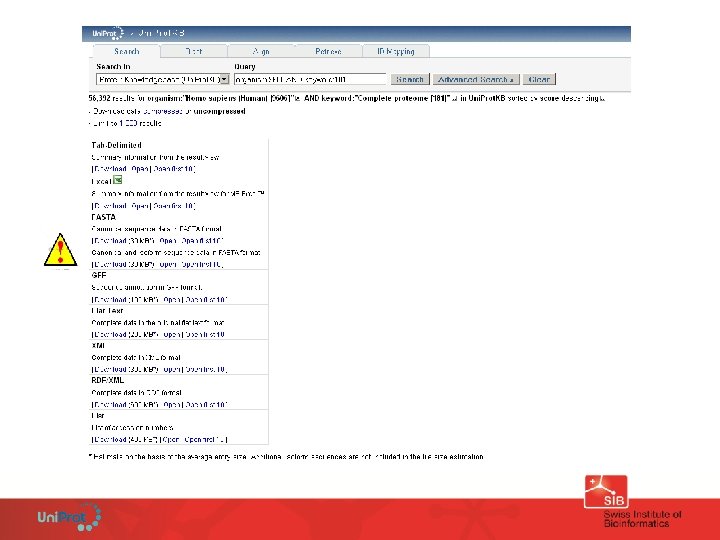

Human proteome ~ 20’ 200 genes Query for ‘homo sapiens’ (August 2011) • Uni. Prot. KB: 110, 056 entries + alt sequences (~ 15’ 435) = 125’ 491 • Uni. Prot. KB/Swiss-Prot: 20’ 244 entries + alt sequences (~ 15’ 435) = 35’ 679 • Uni. Prot. KB/Tr. EMBL: 89, 834 entries • Ref. Seq: 32’ 898 sequences • Ensembl: 90’ 720 sequences Query for ‘homo sapiens’ + Complete proteome (KW-181) • Uni. Prot. KB: 56’ 392 + alt sequences (15’ 435) = 71’ 827 • Uni. Prot. KB/Swiss-Prot: 20’ 238 + alt sequences (15’ 435) = 35’ 673 • Uni. Prot. KB/Tr. EMBL: 36’ 154 92% of human entries are linked with at least one Ref. Seq entry…

Summary
Do not hesitate to contact us ! help@uniprot. org

The Uni. Prot Consortium SIB Ioannis Xenarios, Lydie Bougueleret, Andrea Auchincloss, Kristian Axelsen, Delphine Baratin, Marie. Claude Blatter, Brigitte Boeckmann, Jerven Bolleman, Laurent Bollondi, Emmanuel Boutet, Lionel Breuza, Alan Bridge, Edouard de Castro, Lorenzo Cerutti, Elisabeth Coudert, Béatrice Cuche, Mikael Doche, Dolnide Dornevil, Severine Duvaud, Anne Estreicher, Livia Famiglietti, Marc Feuermann, Sebastien Gehant, Elisabeth Gasteiger, Alain Gateau, Vivienne Gerritsen, Arnaud Gos, Nadine Gruaz. Gumowski, Ursula Hinz, Chantal Hulo, Nicolas Hulo, Janet James, Florence Jungo, Guillaume Keller, Vicente Lara, Philippe Lemercier, Damien Lieberherr, Xavier Martin, Patrick Masson, Anne Morgat, Salvo Paesano, Ivo Pedruzzi, Sandrine Pilbout, Sylvain Poux, Monica Pozzato, Manuela Pruess, Nicole Redaschi, Catherine Rivoire, Bernd Roechert, Michel Schneider, Christian Sigrist, Karin Sonesson, Sylvie Staehli, Eleanor Stanley, André Stutz, Shyamala Sundaram, Michael Tognolli, Laure Verbregue, Anne-Lise Veuthey EBI Rolf Apweiler, Maria Jesus Martin, Claire O'Donovan, Michele Magrane, Yasmin Alam-Faruque, Ricardo Antunes, Benoit Bely, Mark Bingley, David Binns, Lawrence Bower, Wei Mun Chan, Emily Dimmer, Francesco Fazzini, Alexander Fedotov, John Garavelli, Leyla Garcia Castro, Rachael Huntley, Julius Jacobsen, Michael Kleen, Duncan Legge, Wudong Liu, Jie Luo, Sandra Orchard, Samuel Patient, Klemens Pichler, Diego Poggioli, Nikolas Pontikos, Steven Rosanoff, Tony Sawford, Harminder Sehra, Edward Turner, Matt Corbett, Mike Donnelly and Pieter van Rensburg PIR Cathy H. Wu, Cecilia N. Arighi, Leslie Arminski, Winona C. Barker, Chuming Chen, Yongxing Chen, Pratibha Dubey, Hongzhan Huang, Kati Laiho, Raja Mazumder, Peter Mc. Garvey, Darren A. Natale, Thanemozhi G. Natarajan, Jules Nchoutmboube, Natalia V. Roberts, Baris E. Suzek, Uzoamaka Ugochukwu, C. R. Vinayaka, Qinghua Wang, Yuqi Wang, Lai-Su Yeh and Jian Zhang www. uniprot. org

Uni. Prot is mainly supported by the National Institutes of Health (NIH) grant 1 U 41 HG 006104 -01. Additional support for the EBI's involvement in Uni. Prot comes from the NIH grant 2 P 41 HG 02273 -07. Swiss-Prot activities at the SIB are supported by the Swiss Federal Government through the Federal Office of Education and Science and the European Commission contracts SLING (226073), Gen 2 Phen (200754) and MICROME (222886). PIR activities are also supported by the NIH grants 5 R 01 GM 080646 -04, 3 R 01 GM 080646 -04 S 2, 1 G 08 LM 010720 -01, and 3 P 20 RR 016472 -09 S 2, and NSF grant DBI-0850319.

www. isb-sib. ch

Thank you for your attention http: //education. expasy. org/cours/Prague 2011/
- Slides: 73