The Three Domains of Living Organisms Classification Vocabulary

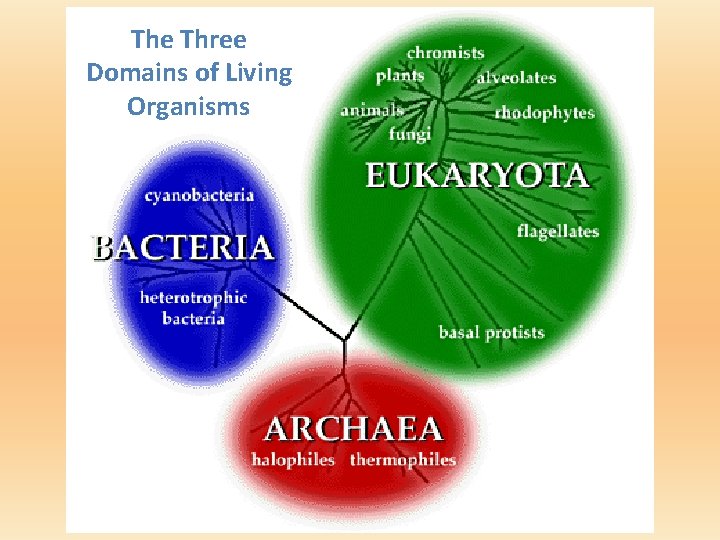
The Three Domains of Living Organisms

Classification Vocabulary • prokaryote: unicellular organism that does not have membrane-bound organelles (like a nucleus); e. g. , bacteria. • eukaryote: unicellular or multicellular organism that contains a true nucleus and other membrane-bound organelles; e. g. , yeast, paramecium, plant, animal. • organelle: membrane-bound structures with particular functions within eukaryotic cells; e. g. , nucleus, mitochondrion, chloroplast.
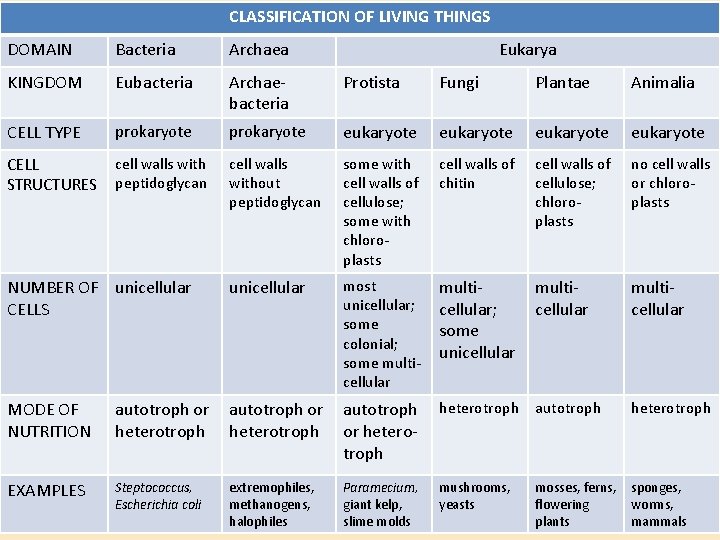
CLASSIFICATION OF LIVING THINGS DOMAIN Bacteria Archaea Eukarya KINGDOM Eubacteria Archaebacteria Protista Fungi Plantae Animalia CELL TYPE prokaryote eukaryote CELL STRUCTURES cell walls with peptidoglycan cell walls without peptidoglycan some with cell walls of cellulose; some with chloroplasts cell walls of chitin cell walls of cellulose; chloroplasts no cell walls or chloroplasts NUMBER OF unicellular CELLS unicellular most unicellular; some colonial; some multicellular; some unicellular multicellular MODE OF NUTRITION autotroph or heterotroph autotroph heterotroph EXAMPLES Steptococcus, Escherichia coli extremophiles, methanogens, halophiles Paramecium, giant kelp, slime molds mushrooms, yeasts mosses, ferns, flowering plants sponges, worms, mammals

Evolutionary Classification • Darwin’s ideas about descent with modification have given rise to the study of phylogeny. Øphylogeny: evolutionary relationships among organisms. • Biologists now group organisms into categories that represent lines of evolutionary descent, or phylogeny, not just physical similarities. • Grouping organisms together based on their evolutionary history is called evolutionary classification.

Evolutionary Classification • How are evolutionary relationships determined? • Evolutionary relationships are determined on the basis of similarities in structure, breeding behavior, geographical distribution, chromosomes, similarities in DNA and RNA, molecular clocks, and biochemistry. ØThese traits provide clues about how species evolved and probable evolutionary relationships of species.
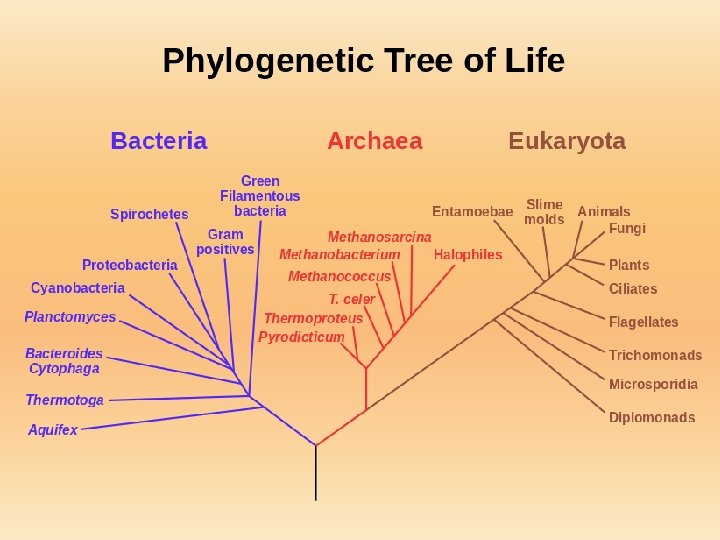

Phylogenetic (or Evolutionary) Classification • Species that share a common ancestor also share an evolutionary history or phylogeny. • A classification system that shows the evolutionary history of species is a phylogenetic classification. • Cladistics: a biological classification system based on phylogeny.

Cladistics • As groups of organisms evolve from a common ancestor, they retain some unique inherited characteristics called shared derived traits (or shared derived characters). • Derived traits are used to make a branching diagram called a cladogram that is a model of the phylogeny of a species. • Cladograms are similar to pedigrees with one important difference: pedigrees show direct ancestry of an organism from two parents, whereas cladograms show the probable evolution of a group of organisms from an ancestral group.
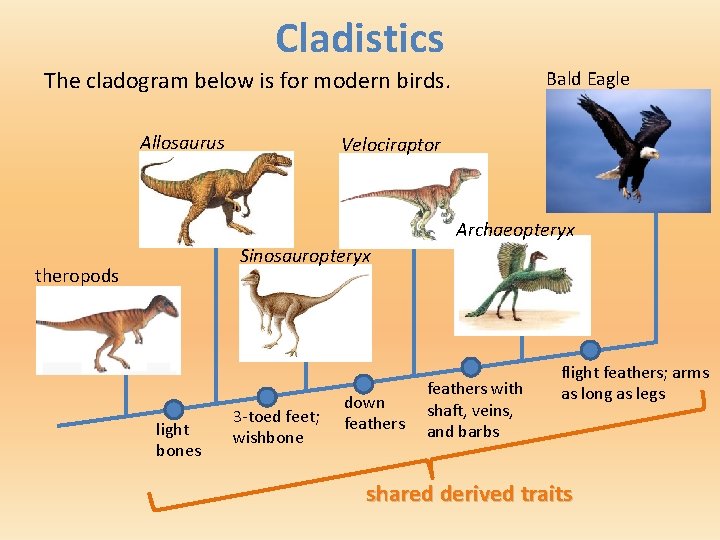
Cladistics The cladogram below is for modern birds. Allosaurus Bald Eagle Velociraptor Archaeopteryx Sinosauropteryx theropods light bones 3 -toed feet; wishbone down feathers with shaft, veins, and barbs flight feathers; arms as long as legs shared derived traits
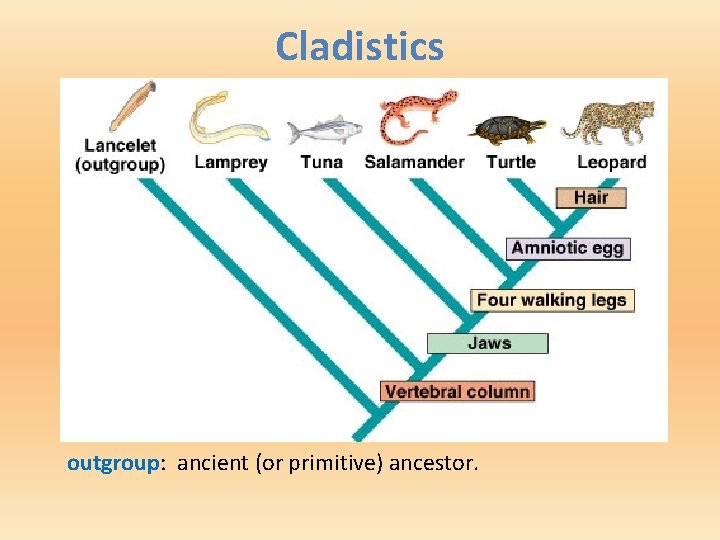
Cladistics outgroup: ancient (or primitive) ancestor.
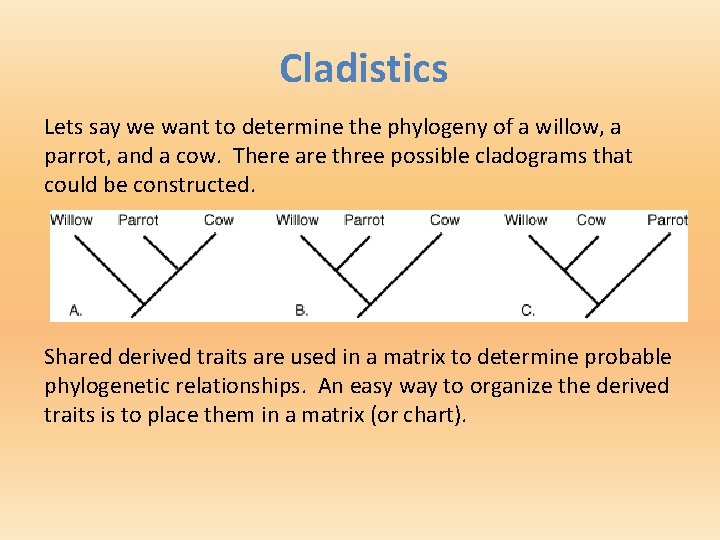
Cladistics Lets say we want to determine the phylogeny of a willow, a parrot, and a cow. There are three possible cladograms that could be constructed. Shared derived traits are used in a matrix to determine probable phylogenetic relationships. An easy way to organize the derived traits is to place them in a matrix (or chart).
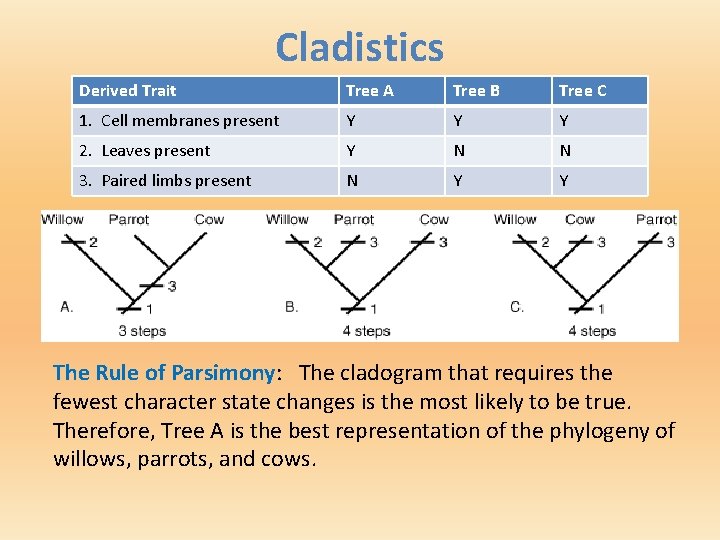
Cladistics Derived Trait Tree A Tree B Tree C 1. Cell membranes present Y Y Y 2. Leaves present Y N N 3. Paired limbs present N Y Y The Rule of Parsimony: The cladogram that requires the fewest character state changes is the most likely to be true. Therefore, Tree A is the best representation of the phylogeny of willows, parrots, and cows.

Cladistics • We you examine different character states such as “limbs present” or “limbs absent, ” we need to know which traits are ancestral and which are derived, because only shared derived characters tell us anything about evolutionary relationships of organisms. • To construct a cladogram, you need an outgroup that is distantly related to the organisms you want to classify, and use the outgroup as a standard for comparison. • You assume that the character states of the outgroup are ancestral.

Cladistics Imagine that we have landed on the Potato Head Planet. Natives tell us that Taxon A is distantly related to the others, so we choose it as our outgroup. We will score character traits as it is done by phylogenists, with 0 = ancestral and 1 = derived. Character A B (outgroup) C D 1. Eyes present 0 0 2. Feet present 0 0 0 1 3. Hat present 0 1 0 0 4. Lips present 0 0 1 1
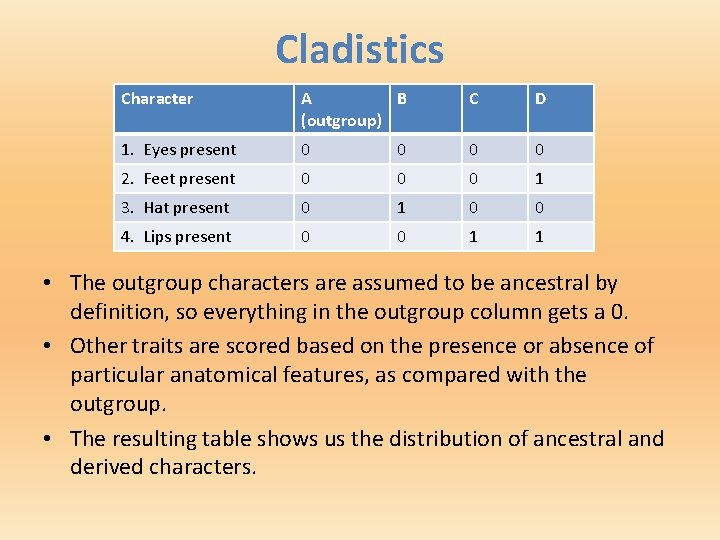
Cladistics Character A B (outgroup) C D 1. Eyes present 0 0 2. Feet present 0 0 0 1 3. Hat present 0 1 0 0 4. Lips present 0 0 1 1 • The outgroup characters are assumed to be ancestral by definition, so everything in the outgroup column gets a 0. • Other traits are scored based on the presence or absence of particular anatomical features, as compared with the outgroup. • The resulting table shows us the distribution of ancestral and derived characters.
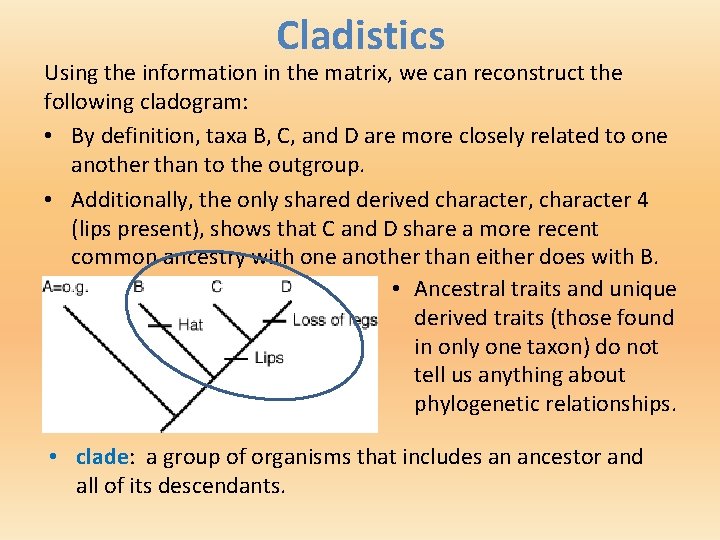
Cladistics Using the information in the matrix, we can reconstruct the following cladogram: • By definition, taxa B, C, and D are more closely related to one another than to the outgroup. • Additionally, the only shared derived character, character 4 (lips present), shows that C and D share a more recent common ancestry with one another than either does with B. • Ancestral traits and unique derived traits (those found in only one taxon) do not tell us anything about phylogenetic relationships. • clade: a group of organisms that includes an ancestor and all of its descendants.
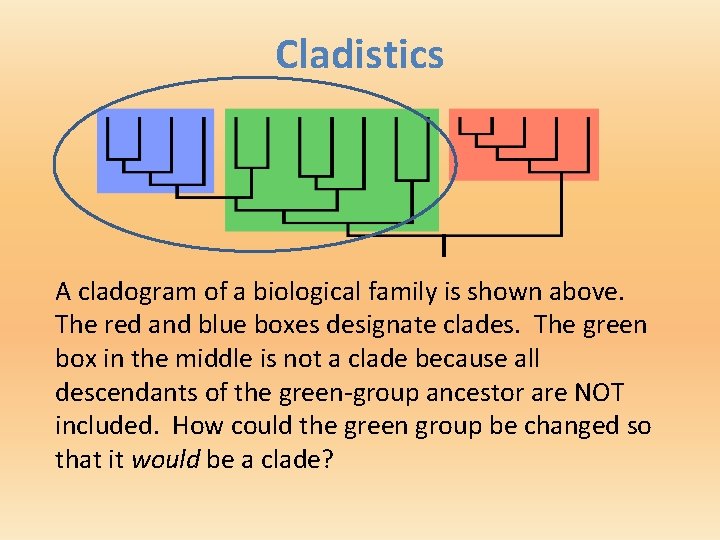
Cladistics A cladogram of a biological family is shown above. The red and blue boxes designate clades. The green box in the middle is not a clade because all descendants of the green-group ancestor are NOT included. How could the green group be changed so that it would be a clade?
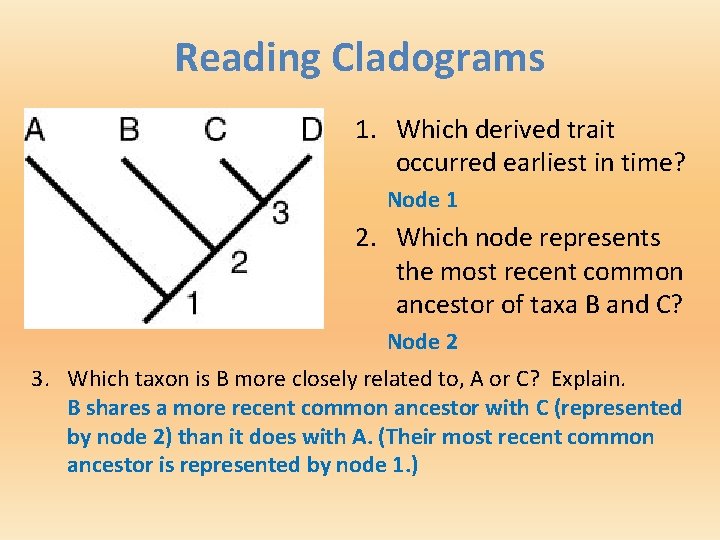
Reading Cladograms 1. Which derived trait occurred earliest in time? Node 1 2. Which node represents the most recent common ancestor of taxa B and C? Node 2 3. Which taxon is B more closely related to, A or C? Explain. B shares a more recent common ancestor with C (represented by node 2) than it does with A. (Their most recent common ancestor is represented by node 1. )
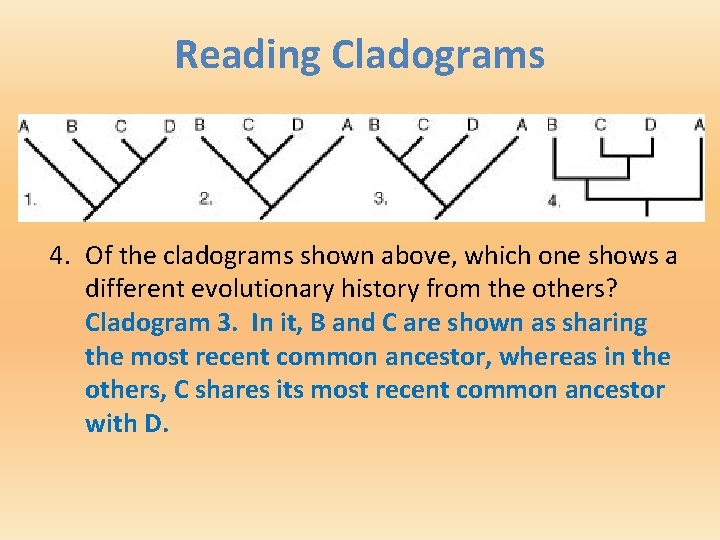
Reading Cladograms 4. Of the cladograms shown above, which one shows a different evolutionary history from the others? Cladogram 3. In it, B and C are shown as sharing the most recent common ancestor, whereas in the others, C shares its most recent common ancestor with D.
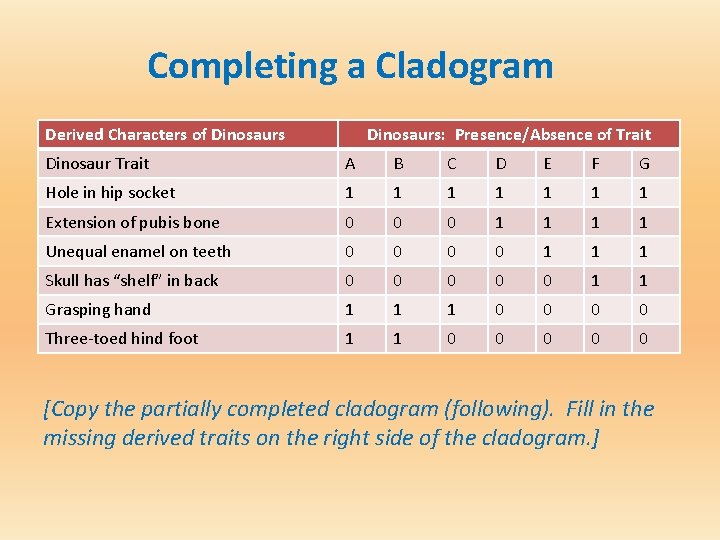
Completing a Cladogram Derived Characters of Dinosaurs: Presence/Absence of Trait Dinosaur Trait A B C D E F G Hole in hip socket 1 1 1 1 Extension of pubis bone 0 0 0 1 1 Unequal enamel on teeth 0 0 1 1 1 Skull has “shelf” in back 0 0 0 1 1 Grasping hand 1 1 1 0 0 Three-toed hind foot 1 1 0 0 0 [Copy the partially completed cladogram (following). Fill in the missing derived traits on the right side of the cladogram. ]
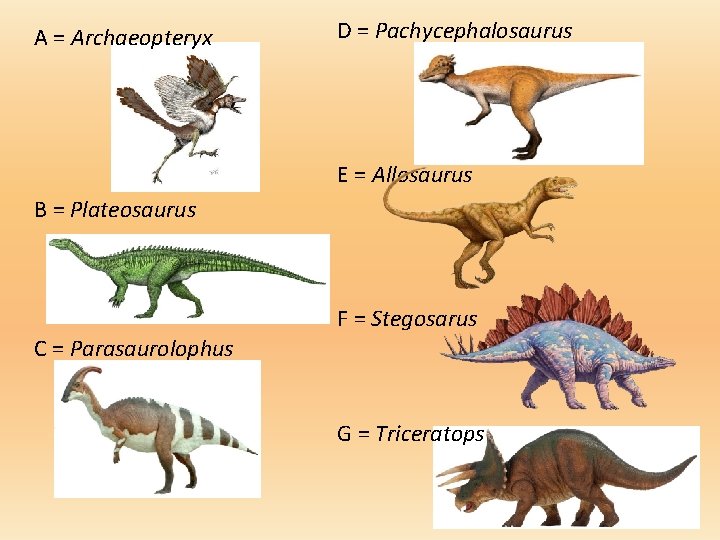
A = Archaeopteryx D = Pachycephalosaurus E = Allosaurus B = Plateosaurus F = Stegosarus C = Parasaurolophus G = Triceratops
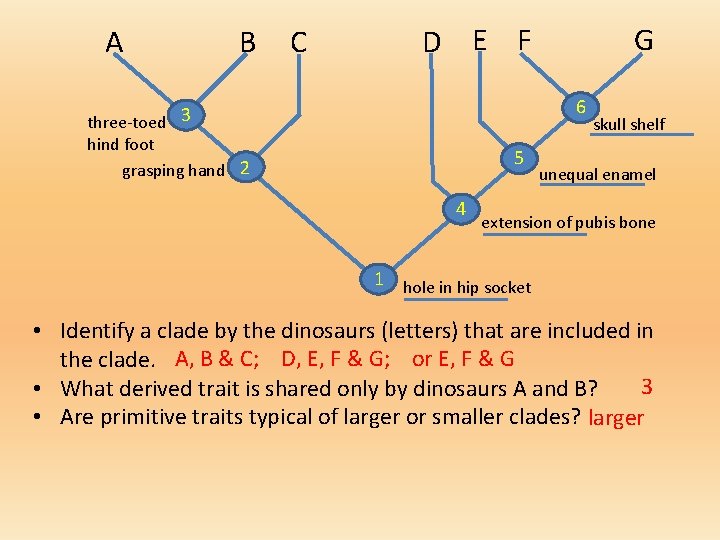
A B C D E F G 6 three-toed 3 hind foot grasping hand 2 5 4 skull shelf unequal enamel extension of pubis bone 1 hole in hip socket • Identify a clade by the dinosaurs (letters) that are included in the clade. A, B & C; D, E, F & G; or E, F & G 3 • What derived trait is shared only by dinosaurs A and B? • Are primitive traits typical of larger or smaller clades? larger
- Slides: 23