The tangled web of epigenetics Bryan Turner Advances

The tangled web of epigenetics Bryan Turner Advances in Genetics: a Second Teachers’ Conference Nowgen Centre, Manchester 28 June 2017

What is epigenetics? Epigenetics is the study of the processes by which genotype gives rise to phenotype. Waddington 1942 “The changes in gene activity during development are generally referred to as epigenetic”. Holliday 1987 The boundary between genetics and epigenetics is blurred

The power of epigenetics Epigenetics defines the processes by which organisms access and use the information stored in DNA Epigenetics can tell us • How we interact with our environment • How to escape the tyranny of DNA • How to cure cancer

The Power of Epigenetics Epigenetic processes generate multiple phenotypes from the same genotype “…endless forms most beautiful and most wonderful…”

Epigenetics is underpinned by molecular mechanisms that regulate how, when and where genes are expressed Genome function (gene expression) is regulated by the interaction between DNA sequence, histones and non-histone proteins (eg. transcription factors).

Epigenetics is underpinned by molecular mechanisms that regulate how, when and where genes are expressed Genome function (gene expression) is regulated by the interaction between DNA sequence, histones and non-histone proteins (eg. transcription factors). Chromatin and the nucleosome are key players in epigenetic processes
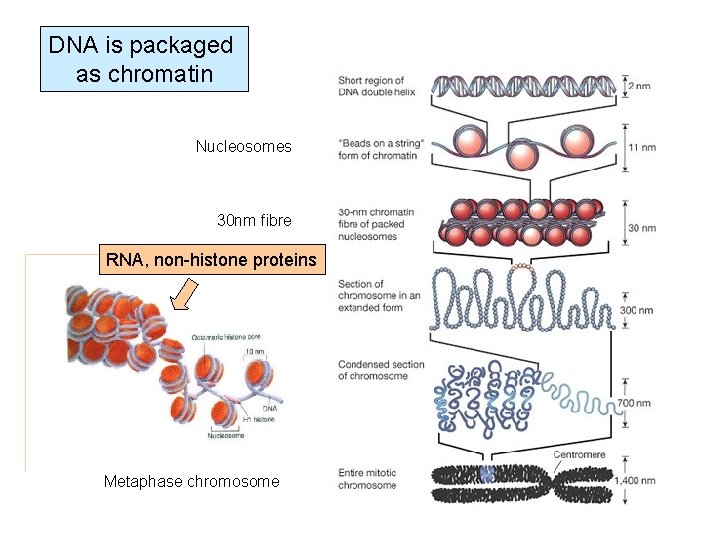
DNA is packaged as chromatin Nucleosomes 30 nm fibre RNA, non-histone proteins Metaphase chromosome
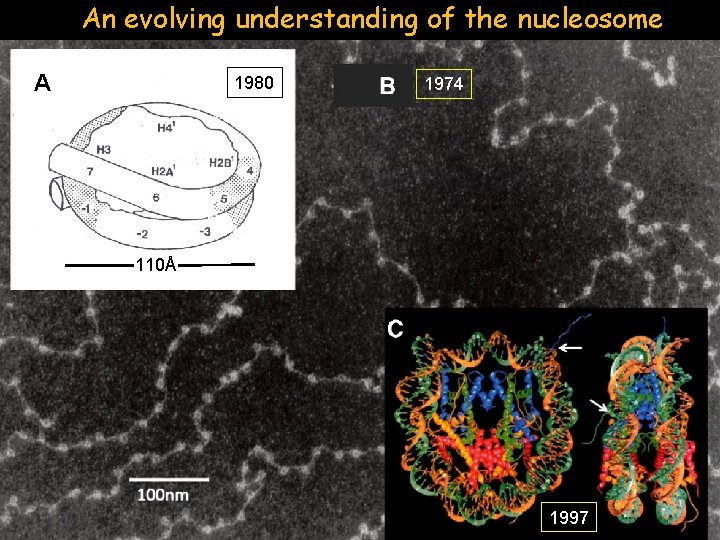
An evolving understanding of the nucleosome A 1980 1974 110Å 1997
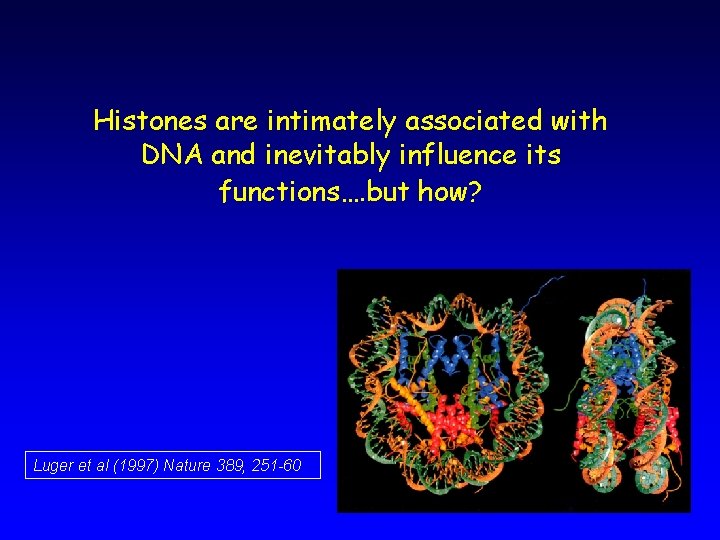
Histones are intimately associated with DNA and inevitably influence its functions…. but how? Luger et al (1997) Nature 389, 251 -60
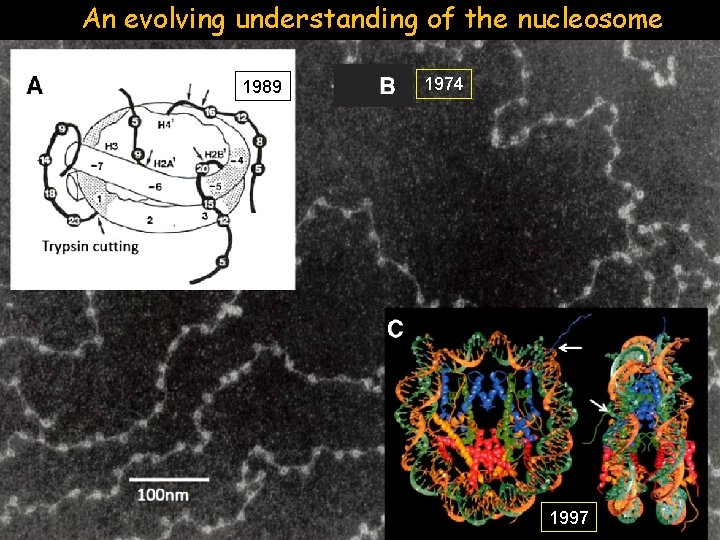
An evolving understanding of the nucleosome 1989 1974 1997
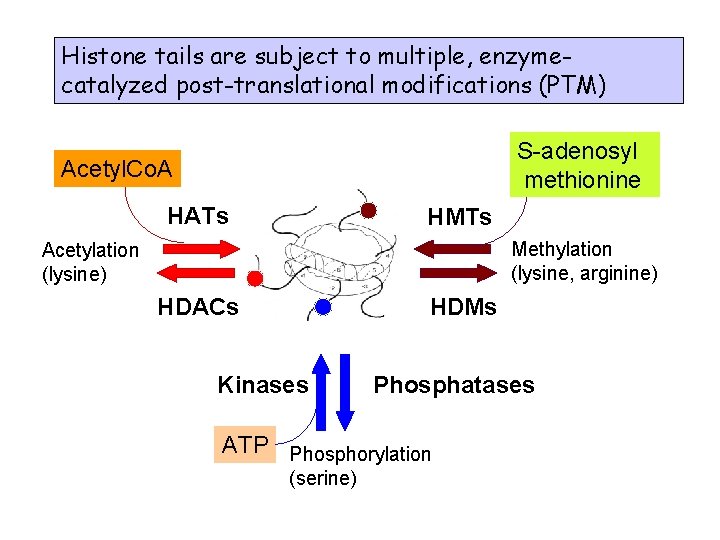
Histone tails are subject to multiple, enzymecatalyzed post-translational modifications (PTM) S-adenosyl methionine Acetyl. Co. A HATs HMTs Methylation (lysine, arginine) Acetylation (lysine) HDACs HDMs Kinases Phosphatases ATP Phosphorylation (serine)
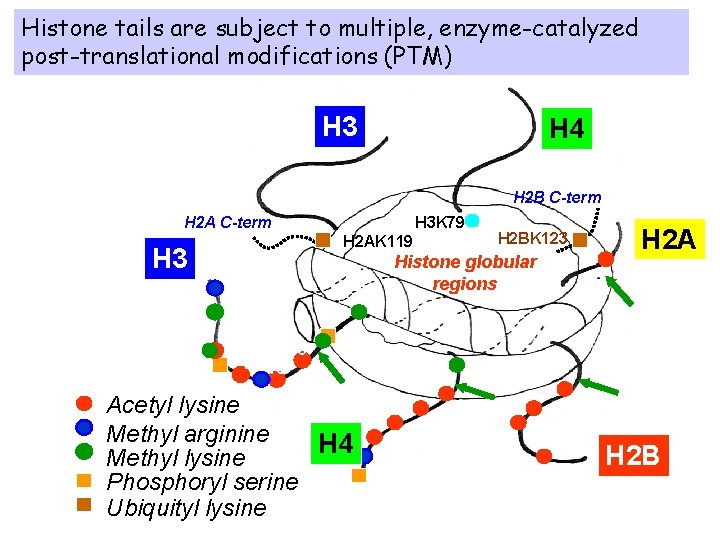
Histone tails are subject to multiple, enzyme-catalyzed post-translational modifications (PTM) H 3 H 4 H 2 B C-term H 2 A C-term H 3 K 79 H 2 BK 123 H 2 AK 119 H 3 Histone globular regions 2 4 9 5 H 2 A 36 14 18 23 Acetyl lysine Methyl arginine H 4 Methyl lysine Phosphoryl serine Ubiquityl lysine 20 3 1 5 8 12 16 15 12 5 20 H 2 B

Proteins with specific binding domains recognise specific modifications and thereby influence chromatin structure and function
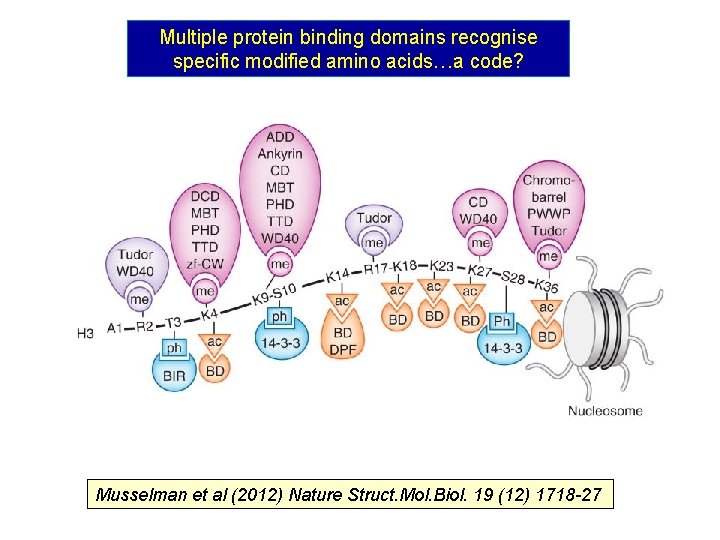
Multiple protein binding domains recognise specific modified amino acids…a code? Musselman et al (2012) Nature Struct. Mol. Biol. 19 (12) 1718 -27

Nucleosome signalling (A beautiful paradigm) Changes in nucleosome structure through incorporation of histone variants or the actions of modifying enzymes, can influence genome function. The nucleosome core particle carries on its surface, a variety of chemical signals in the form of modifications of specific histone amino acids. Modifications are put in place and removed by families of enzymes. Histones carrying specific modifications can be bound by proteins carrying defined binding domains. These proteins regulate the function of genomic regions to which they are bound.

Roles of chromatin modification histone modifications Short-term effects Long-term effects Ongoing processes Heritable changes Transcription, replication, repair Cellular memory
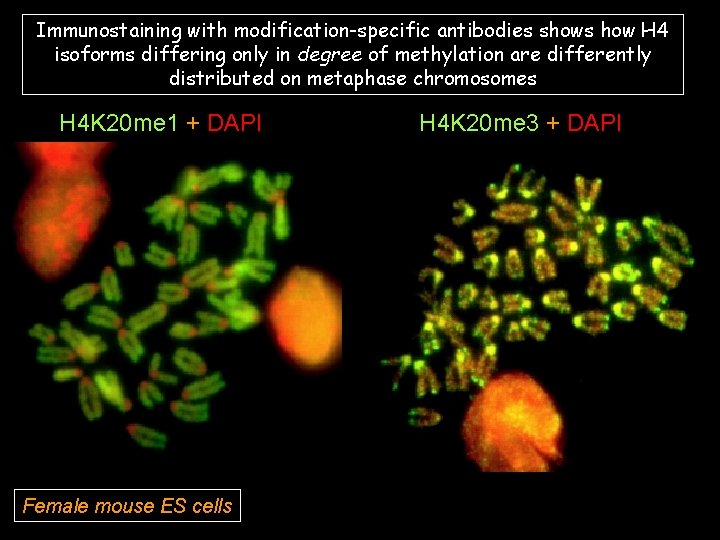
Immunostaining with modification-specific antibodies shows how H 4 isoforms differing only in degree of methylation are differently distributed on metaphase chromosomes H 4 K 20 me 1 + DAPI Female mouse ES cells H 4 K 20 me 3 + DAPI
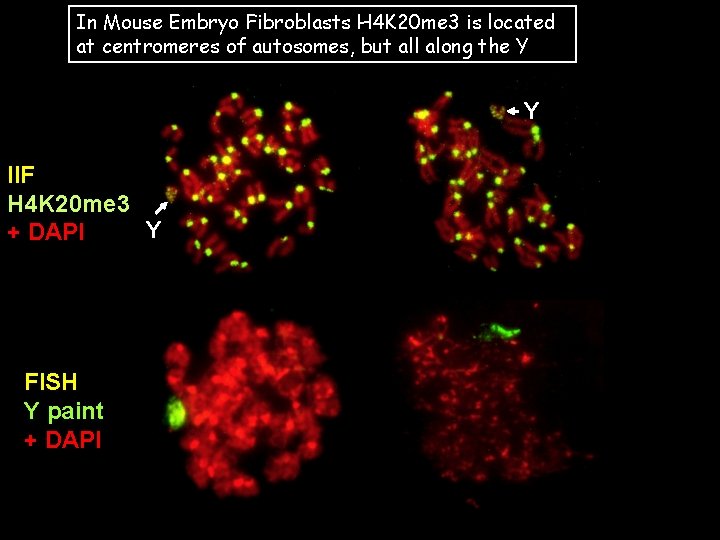
In Mouse Embryo Fibroblasts H 4 K 20 me 3 is located at centromeres of autosomes, but all along the Y Y IIF H 4 K 20 me 3 Y + DAPI FISH Y paint + DAPI
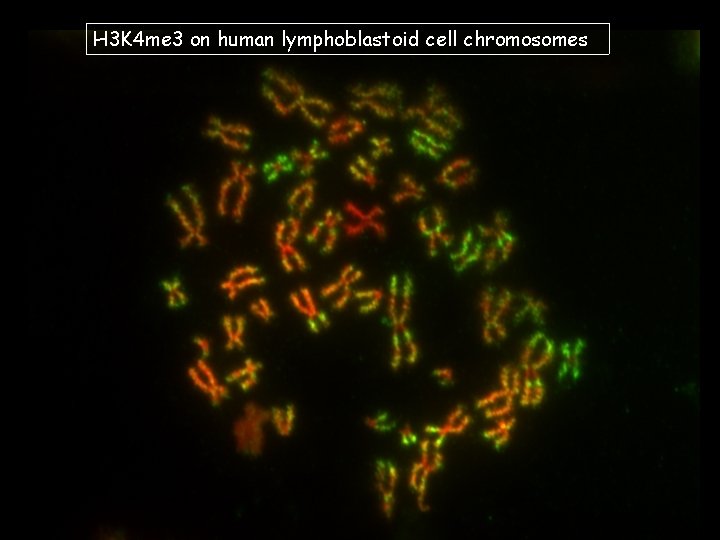
H 3 K 4 me 3 on human lymphoblastoid cell chromosomes

The nucleosome is more than just a DNA packaging device: It is a sophisticated signalling system that regulates gene expression It also, inevitably, links genome function to environmental agents
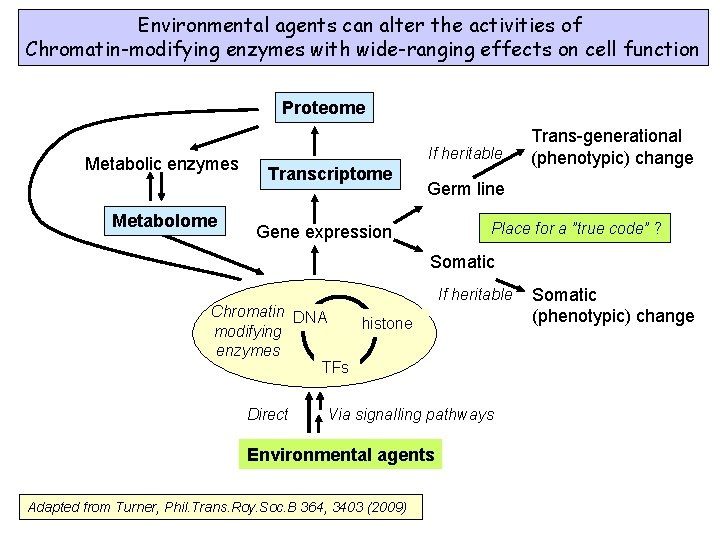
Environmental agents can alter the activities of Chromatin-modifying enzymes with wide-ranging effects on cell function Proteome Metabolic enzymes Metabolome If heritable Transcriptome Trans-generational (phenotypic) change Germ line Place for a ”true code” ? Gene expression Somatic Chromatin DNA histone modifying enzymes TFs Direct If heritable Via signalling pathways Environmental agents Adapted from Turner, Phil. Trans. Roy. Soc. B 364, 3403 (2009) Somatic (phenotypic) change

More than just neglect…. . Jean-Baptiste Lamarck (1744 -1829) Lamarck is the only major figure in the history of biology whose name has become…a term of abuse. Most scientists’ contributions are fated to be outgrown, but very few authors have written works which, two centuries later, are still rejected with an indignation so intense that the sceptic may suspect something akin to an uneasy conscience. C. H. Waddington, in Evolution after Darwin, Chicago University Press, 1960 Philosophie Zoologique, 1809 The environment drives heritable phenotypic change

How we escape the tyranny of DNA

Paradigm shifts in genetics 1850 -1900 : Proto-genetics Mendel, inheritance Darwin, natural selection 1900– 1950 : Age of genetics gene concept, mutation, genotype-phenotype 1950– 2000 : Age of DNA structure, genetic code, Selfish Gene, genomics Has DNA become the all-powerful deity of a new religion…

with its prophets…… …. who ask us to wonder at the marvels created by DNA in a deterministic universe

…and of course a patron saint Charles Darwin (1809 -1882) The combination of Natural Selection and Mendelian Genetics provides an almost complete explanation for the earth’s biodiversity “The Newton of Biology” and a major beneficiary of the Age of DNA The Origin of Species by Means of Natural Selection, 1859

But DNA is just an inert coding device, useless without epigenetic processes to interpret its messages.

But DNA is just an inert coding device, useless without epigenetic processes to interpret its messages. Epigenetic processes can be manipulated and changed by environment, lifestyle, diet etc Philosophically, epigenetics is an empowering science!

Paradigm shifts in genetics 1850 -1900 : Proto-genetics Mendelian inheritance Darwin, natural selection 1900 – 1950 : Age of genetics gene concept, mutation, genotype-phenotype 1950– 2000 : Age of DNA structure, genetic code, genome sequence 2000 - epigenetic code, epigenome, epigenetic medicine : Age of epigenetics

Jean-Baptiste Lamarck (1744 -1829) Epigenetics will free us from the tyranny of DNA Vive la révolution Philosophie Zoologique, 1809 The environment drives heritable phenotypic change

Epigenetic drugs to treat cancer
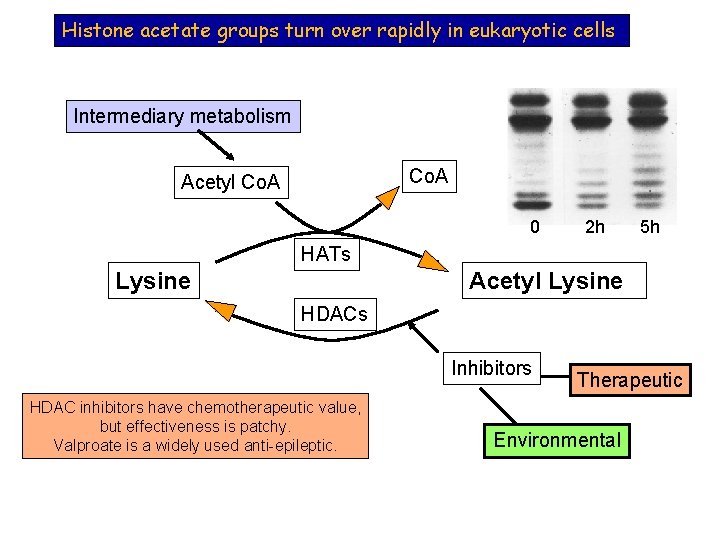
Histone acetate groups turn over rapidly in eukaryotic cells Intermediary metabolism Co. A Acetyl Co. A 0 2 h 5 h HATs Lysine Acetyl Lysine HDACs Inhibitors HDAC inhibitors have chemotherapeutic value, but effectiveness is patchy. Valproate is a widely used anti-epileptic. Therapeutic Environmental
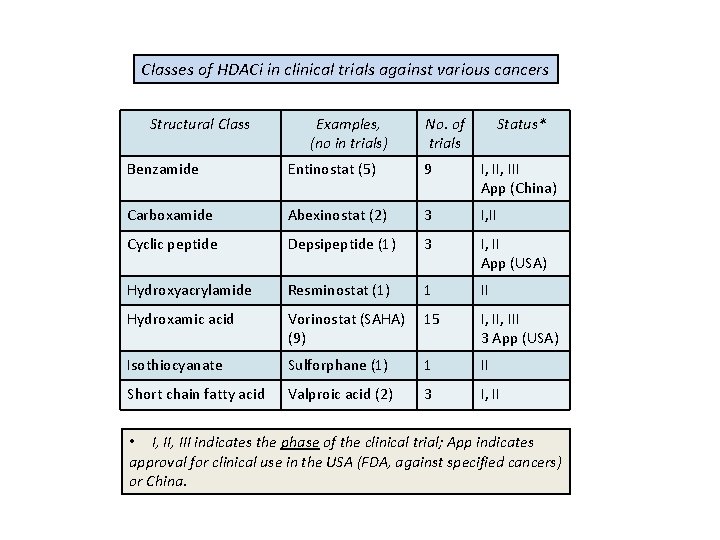
Classes of HDACi in clinical trials against various cancers Structural Class Examples, (no in trials) No. of trials Status* Benzamide Entinostat (5) 9 I, III App (China) Carboxamide Abexinostat (2) 3 I, II Cyclic peptide Depsipeptide (1) 3 I, II App (USA) Hydroxyacrylamide Resminostat (1) 1 II Hydroxamic acid Vorinostat (SAHA) (9) 15 I, III 3 App (USA) Isothiocyanate Sulforphane (1) 1 II Short chain fatty acid Valproic acid (2) 3 I, II • I, III indicates the phase of the clinical trial; App indicates approval for clinical use in the USA (FDA, against specified cancers) or China.

First interesting question Histone acetylation would be expected to cause massive (lethal) disruption of epigenetic systems Why are many normal cells so tolerant? And why are some cancers sensitive?

First interesting question Histone acetylation would be expected to cause massive (lethal) disruption of epigenetic systems Why are many normal cells so tolerant? And why are some cancers sensitive?

Second interesting question Why should human cells mount such a complex resistance response to a particular class of anti-cancer drugs?

HDAC inhibitors are naturally occurring (environmental) agents Short chain fatty acids (eg. butyric) are products of gut microflora and found at m. M concentrations in the colon Trichostatin A and other hydroxamic acid derivatives are secreted by certain fungi and are anti-fungal antibiotics. Depsipeptide (FDA approved) is a fermentation product of Chromobacterium violaceum

Second interesting question Why should human cells mount such a complex resistance response to a particular class of anti-cancer drugs? The answer may lie in our evolutionary history
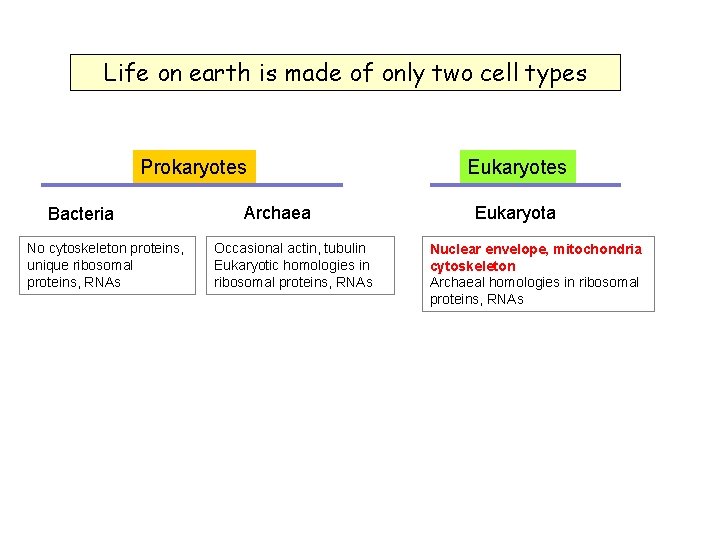
Life on earth is made of only two cell types Prokaryotes Bacteria No cytoskeleton proteins, unique ribosomal proteins, RNAs Archaea Occasional actin, tubulin Eukaryotic homologies in ribosomal proteins, RNAs Eukaryotes Eukaryota Nuclear envelope, mitochondria cytoskeleton Archaeal homologies in ribosomal proteins, RNAs
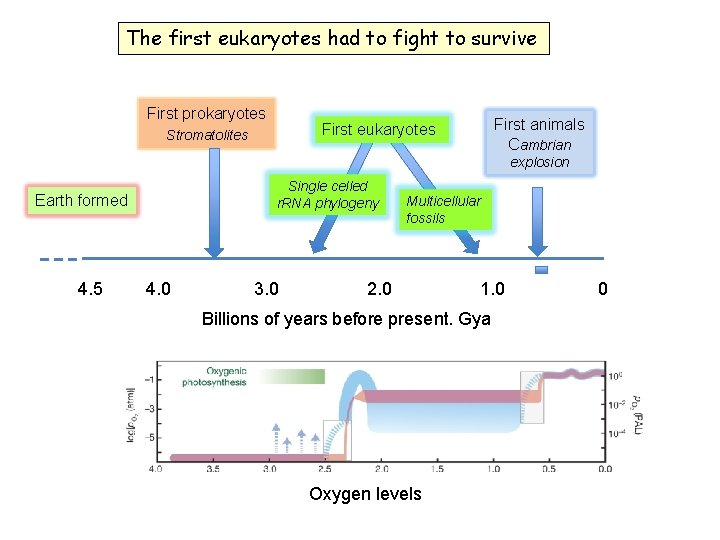
The first eukaryotes had to fight to survive First prokaryotes First animals Cambrian First eukaryotes Stromatolites explosion Single celled r. RNA phylogeny Earth formed 4. 5 4. 0 3. 0 Multicellular fossils 2. 0 1. 0 Billions of years before present. Gya Oxygen levels 0

A microbial arms race Prokaryotes attacked (or defended themselves against) the earliest eukaryotic interlopers (predators? ) by secreting toxins to disrupt their unique epigenetic systems.

A microbial arms race Prokaryotes defended themselves against the earliest eukaryotic predators by secreting toxins to disrupt their unique epigenetic systems. We see these today as bacterial anti-fungals (TSA, depsipeptide) T

A microbial arms race Prokaryotes defended themselves against the earliest eukaryotic predators by secreting toxins to disrupt their unique epigenetic systems. We see these today as bacterial anti-fungals (TSA, depsipeptide) Eukaryotes fought back, and we see this today as fungal antibiotics such as penicillin

A microbial arms race Prokaryotes defended themselves against the earliest eukaryotic predators by secreting toxins to disrupt their unique epigenetic systems. We see these today as bacterial anti-fungals (TSA, depsipeptide) Eukaryotes fought back, and we see this today as fungal antibiotics such as penicillin They also developed ways of defending their epigenetic systems against chemical attack, which we have detected as a survival response against HDACi
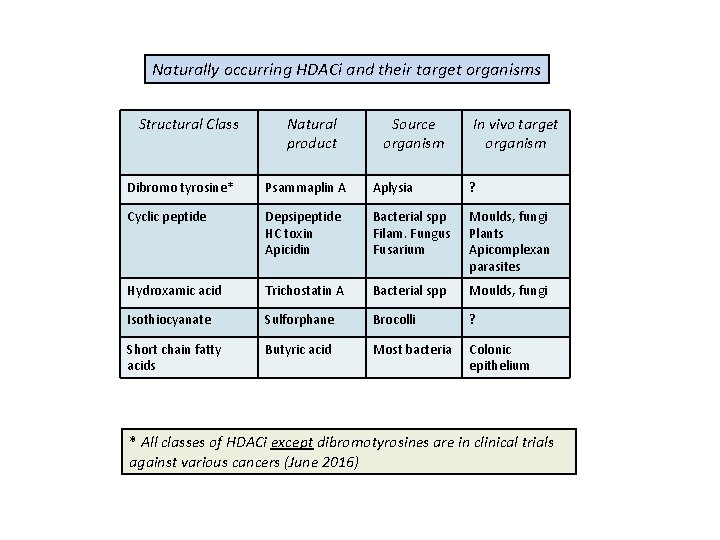
Naturally occurring HDACi and their target organisms Structural Class Natural product Source organism In vivo target organism Dibromo tyrosine* Psammaplin A Aplysia ? Cyclic peptide Depsipeptide HC toxin Apicidin Bacterial spp Filam. Fungus Fusarium Moulds, fungi Plants Apicomplexan parasites Hydroxamic acid Trichostatin A Bacterial spp Moulds, fungi Isothiocyanate Sulforphane Brocolli ? Short chain fatty acids Butyric acid Most bacteria Colonic epithelium * All classes of HDACi except dibromotyrosines are in clinical trials against various cancers (June 2016)
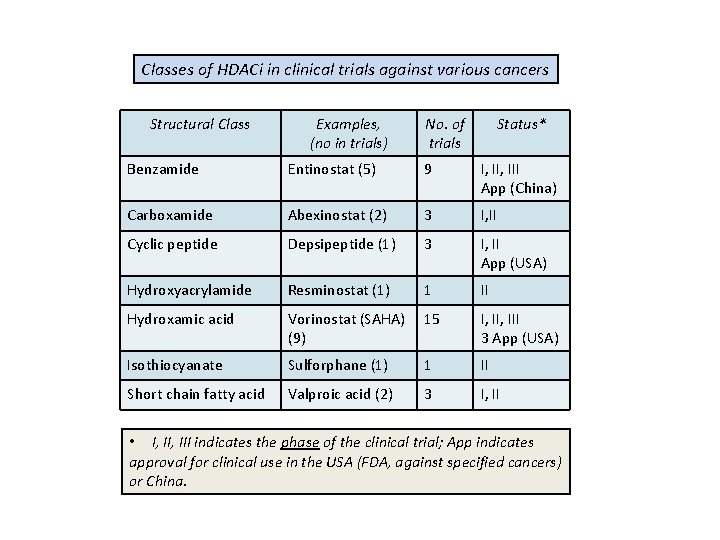
Classes of HDACi in clinical trials against various cancers Structural Class Examples, (no in trials) No. of trials Status* Benzamide Entinostat (5) 9 I, III App (China) Carboxamide Abexinostat (2) 3 I, II Cyclic peptide Depsipeptide (1) 3 I, II App (USA) Hydroxyacrylamide Resminostat (1) 1 II Hydroxamic acid Vorinostat (SAHA) (9) 15 I, III 3 App (USA) Isothiocyanate Sulforphane (1) 1 II Short chain fatty acid Valproic acid (2) 3 I, II • I, III indicates the phase of the clinical trial; App indicates approval for clinical use in the USA (FDA, against specified cancers) or China.

Human cells can mount a chromatin-based response that limits the toxic effects of HDAC inhibitors Are cancers susceptible to HDACi those in which key components of the resistance response have been lost through mutation? Can we develop drugs to target these components in cancer cells? “Seen in the light of evolution, biology is perhaps the most satisfying and inspiring science. Without that light, it becomes a pile of sundry facts…” Theodosius Dobzhansky Turner B (2011) FEBS Lett 585, 2032 -40 (Epigenetics special edition)
- Slides: 47