The Role of Fluorescence in situ hybridization FISH

The Role of Fluorescence in situ hybridization (FISH) in Sequencing the Tomato Genome
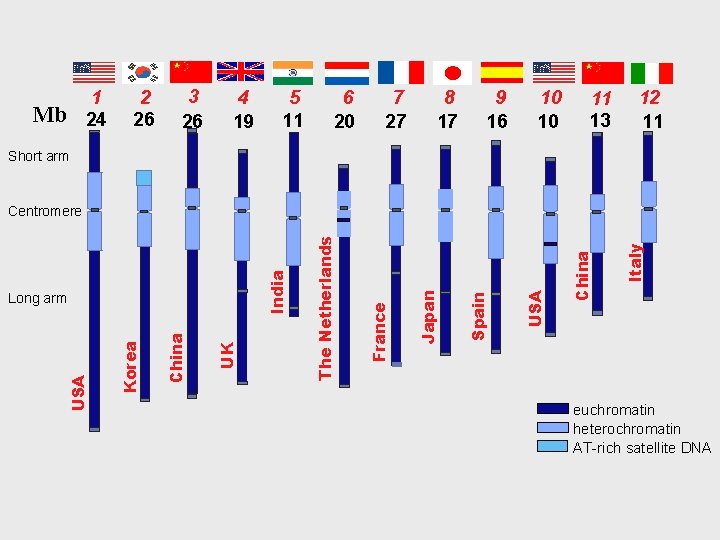
Mb 1 24 2 26 3 26 4 19 5 11 6 20 7 27 8 17 9 16 10 10 11 13 12 11 Short arm Italy China USA Spain Japan France UK China Korea USA Long arm The Netherlands India Centromere euchromatin heterochromatin AT-rich satellite DNA

Maize 1 C = 2634 Mb Tomato 1 C = 916 Mb Arabidopsis 1 C = 157 Mb


77% of the DNA is in heterochromatin 23% of DNA is in euchromatin

77% of the DNA is in heterochromatin 23% of the DNA is in euchromatin Thus, 0. 23 x 916 Mb = 211 Mb

77% of the DNA is in heterochromatin 23% of the DNA is in euchromatin Thus, 0. 23 x 916 Mb = 211 Mb Arabidopsis 1 C = 157 Mb

Hind. III library 15 tomato genome equivalents 129, 000 clones Averaging 117 kb
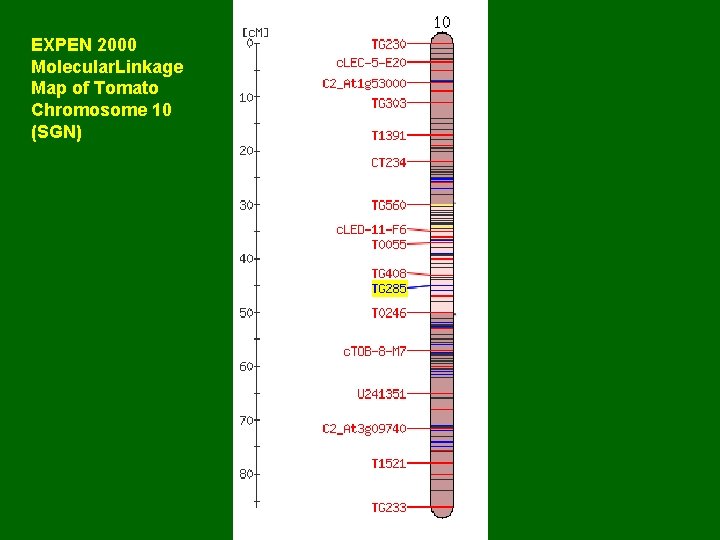
EXPEN 2000 Molecular. Linkage Map of Tomato Chromosome 10 (SGN)

Anchor (seed) BAC LE_HBa 0234 C 10 Probe TG 285
gaatcttggccatc aatgcctaggcat Anchor (seed) BAC LE_HBa 0234 C 10 Probe TG 285
gctacgcttagaa gaatcttggccatc aatgcctaggcat Anchor (seed) BAC LE_HBa 0234 C 10 Probe TG 285

Seed BAC LE_HBa 0234 C 10 Probe TG 285

Seed BAC LE_HBa 0234 C 10 Probe TG 285
Seed BAC LE_HBa 0234 C 10 Probe TG 285
Contig Seed BAC LE_HBa 0234 C 10 Probe TG 285

Fluorescence in situ Hybridization (FISH) China France The Netherlands Korea Japan USA

Fluorescence in situ Hybridization (FISH) China France The Netherlands Korea Japan USA



BAC DNA Nick translation biotindigoxygenindinitrophenol-labeled nucleotides FISH Probes

Hybridization mixture 50 -100 -fold excess of unlabeled tomato Cot 100 DNA Chromosomal in situ Suppression (CISS) Hybridization



Functions of FISH in Sequencing the Tomato Genome 1. To determine the locations of anchor BACs 2. To define eu-heterochromatin borders 3. To determine distances in Mb 4. To locate problem BACs
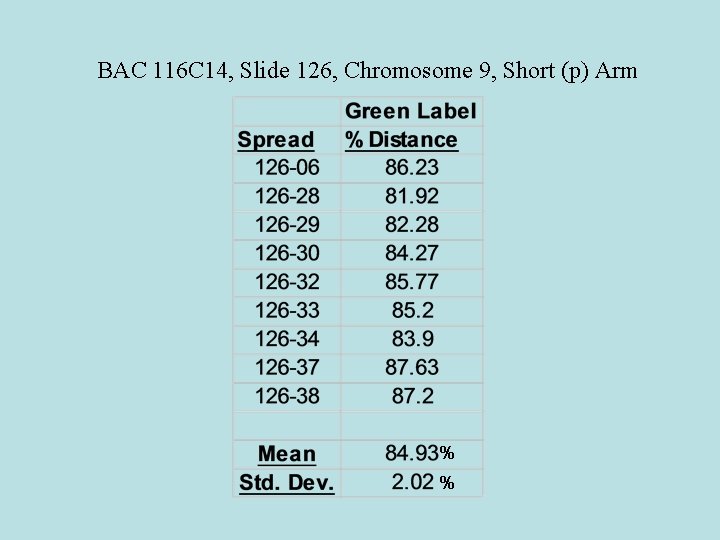
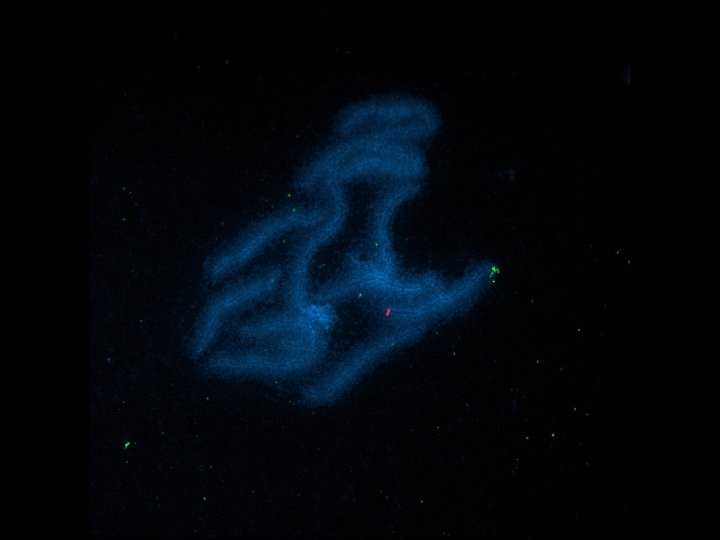
BAC 116 C 14, Slide 126, Chromosome 9, Short (p) Arm % %
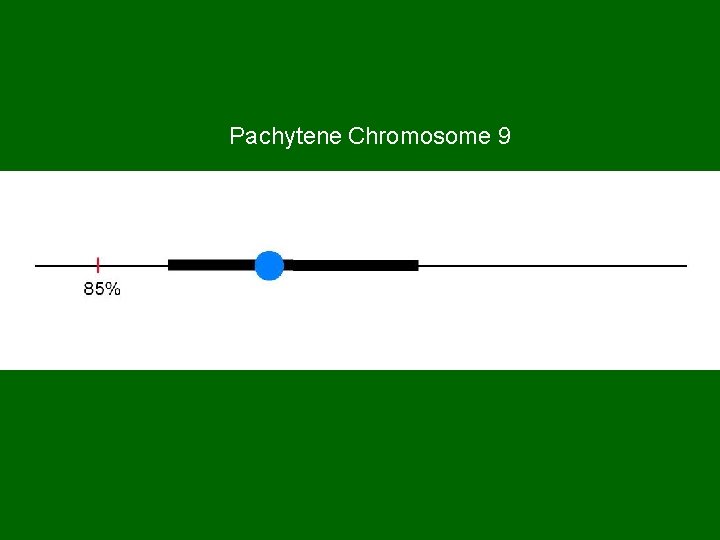
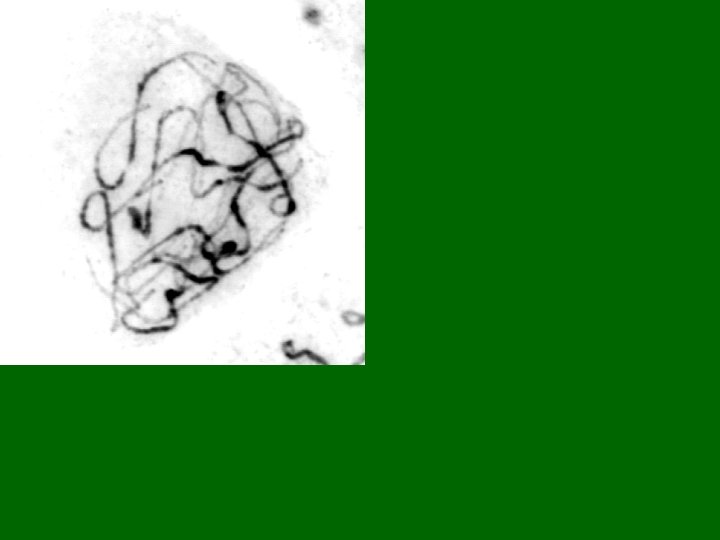
Pachytene Chromosome 9
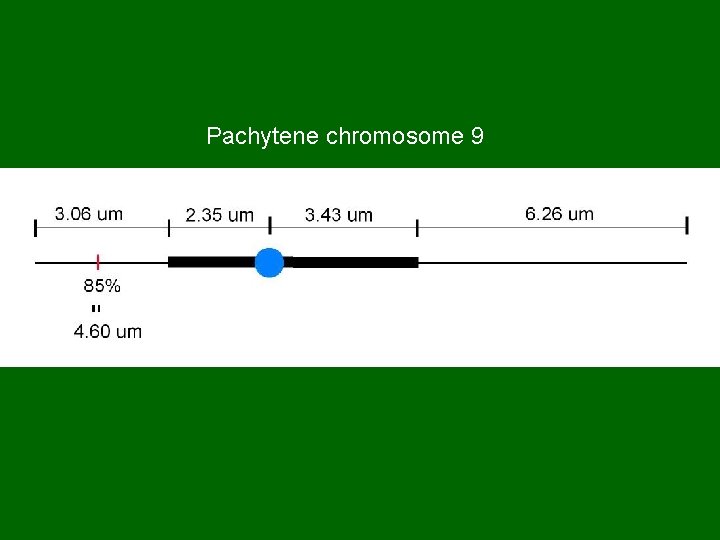
Pachytene chromosome 9

Functions of FISH in Sequencing the Tomato Genome 1. To determine the locations of anchor BACs 2. To define eu-heterochromatin borders 3. To determine distances in Mb 4. To locate problem BACs
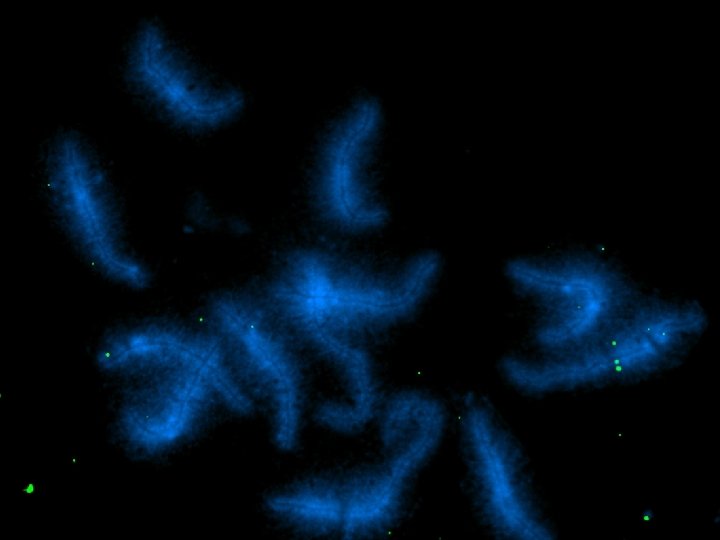

Functions of FISH in Sequencing the Tomato Genome 1. To determine the locations of anchor BACs 2. To define eu-heterochromatin borders 3. To determine distances in Mb 4. To locate problem BACs
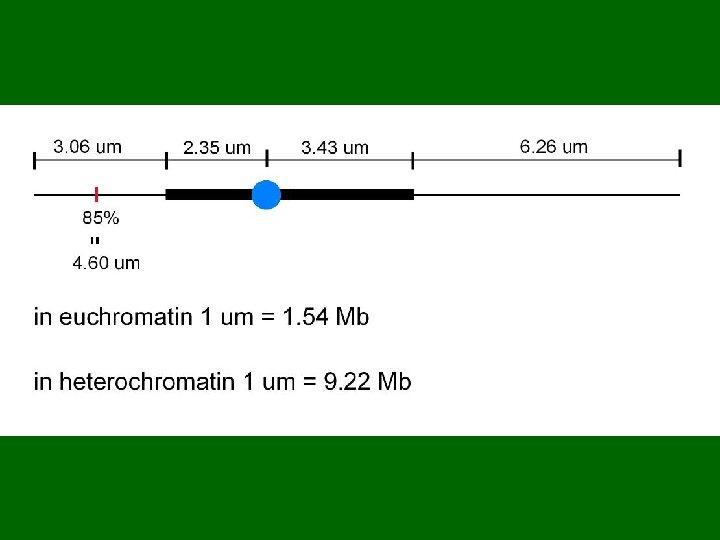

Functions of FISH in Sequencing the Tomato Genome 1. To determine the locations of anchor BACs 2. To define eu-heterochromatin borders 3. To determine distances in Mb 4. To locate problem BACs

RESULTS SO FAR 177 BACs FISHED

RESULTS SO FAR 177 BACs FISHED 126 BACS (74%) successfully localized.

RESULTS SO FAR 177 BACs FISHED 126 BACS (74%) successfully localized. 51 failed to localize because they either gave no FISH signal or there was more than one signal in spite of CISS hybridization.

Of 126 BACs localized 22 (17. 5%) FISHed to wrong chromosomes

Of 126 BACs localized 22 (17. 5%) FISHed to wrong chromosomes Of these, 11 have been checked by sequencing: 7 were overgo false positives 1 was due to a picking error 1 was due to a typographical error 2 were due to mapping errors

Of 126 BACs localized 22 (17. 5%) FISHed to wrong chromosomes Of these 11 have been checked by sequencing: 7 were overgo false positives 1 were due to a picking error 1 were due to a typographical error 2 were due to mapping errors Suggesting ≤ 3% mapping errors on the EXPEN 2000 linkage map

FUTURE ACTIVITIES an increasing emphasis on defining the size of gaps in sequencing 1) within BACs, 2) between contigs, 3) between contigs and euchromatinheterochromatin borders

The Role of BAC Fluorescence in situ hybridization (FISH) in Sequencing the Tomato Genome Senior Personnel Lorrie Anderson Song-Bin Chang Suzanne Royer Lindsay Shearer Steve Stack Undergraduates Madeline Fujishiro Lauren Larsen Dylan Westfall Jessica Wu
- Slides: 44