The PSI KB Protein Model Portal Torsten Schwede

The PSI KB Protein Model Portal Torsten Schwede NIGMS PSI „Bottlenecks“ Workshop Bethesda, April 14, 2008 Swiss Institute of Bioinformatics

Overview: The KB Modeling Portal n Introduction n The KB Protein Model Portal n n Mission and Goals Version 1. 0: Content, Features & Technical Implementation Outlook: Next steps n New Features & Functions n Modeling Portal Community Workshop Questions & Discussion


gggtctctcttgttagaccagatctgagcctgggagctctctggctaactagggaacccactgcttaagcctcaataaagcttgccttgagtgcttcaagtagtgtgtgcccgtctgttgtgtgactctgatagctagagatcccttc agaccaaatttagtcagtgtgaaaaatctctagcagtggcgcctgaacagggacttgaaagcgaaagagaaaccagagaagctctctcgacgcaggactcggcttgctgaagcgcgcacggcaagaggcgaggggacggcgactggtg agtacgccaaaattttgactagcggaggctagaaggagatgggtgcgagagcgtcgatattaagcgggggaggattagatgggaaaaaattcggttaaggccagggggaaaaaatatagattaaaacatttagtat gggcaagcagggagctagaacgattcgcagtcaatcctggcctattagaaacatcagaaggttgtagacaaatactgggacaactacaaccagcccttcagacaggatcagaagaacttagatcattatataatacagtagcaaccct ctattgtgtgcatcaaaagatgtaaaagacaccaaggaagctttagatagaggaagagcaaaagtaagaaaaaagcacagcagcagctgacacaggaaatagcagccaggtcagccaaaattaccccata gtgcagaacatccaggggcaaatggtacatcaggccatatcacctagaactttaaatgcatgggtaaaagtagtagaaggctttcagcccagaagtaatacccatgttttcagcattatcagaaggagccaccccacaagatt taaacaccatgctaaacacagtggggggacatcaagcagccatgcaaatgttaaaagagaccatcaatgaggaagctgcagaatgggatagattgcatccagtgcagggcctcatccaccaggccagatgagagaaccaagggg aagtgacatagcaggaactactagtacccttcaggaacaaatagcatggatgacaaataatccacctatcccagtaggagaaatctataagagatggataatcctgggattaaaatagtaaggatgtatagccctaccagcatt ctggacataaaacaaggaccaaaggaaccctttagagactatgtagaccggttctataagactctaagagccgagcaagcttcacaggaggtaaaaaattggatgacagaaaccttgttggtccaaaatgcgaacccagattgtaaga ctattttaaaagcattgggaccagcagctacactagaagaaatgatgacagcatgtcagggagtgggaggacccggccataaagcaagagttttggcagaagcaatgagccaagtaacaaattcagctaccataatgatgcagaaagg caattttaggaaccaaagaaaaattgttaagtgtttcaattgtggcaaagaagggcacatagccaaaaattgcagggcccctaggaaaaggggctgttggaaatgtggaaaggagggacaccaaatgaaagattgtactgagagacag gctaattttttagggaaaatctggccttcccacaggggaaggccagggaattttcctcagaacagactagagccaacagccccaccagaagagagcttcaggtttggggaagagacaacaactccctctcagaagcagg agctgatagacaaggaactgtatccttcagcttccctcaaatcactctttggcaacgaccccttgtcacaataaagataggggggcaactaaaggaagctctattagatacaggagcagatgatacagtattagaagaaatttg ccaggaagatggaaaccaaaaatgatagggggaattggaggttttatcaaagtaagacagtatgatcaaatactcgtagaaatctgtggacataaagctataggtacagtattagtaggacctacacctgtcaacataattggaagaa atctgttgactcagattggttgcactttaaattttcccattagtcctattgaaactgtaccagtaaaattaaagccaggaatggcccaaaagttaaacaatggccattgacagaagaaaaaataaaagcattagtagaaatctg tacagaaatggaaaaggaaaaatttcaaaaatcgggcctgaaaatccatataatactccagtatttgccataaagaaaaaagacagtactaaatggagaaaattagtagatttcagagaacttaataagaaaactcaagacttc tgggaagttcaattaggaataccacatcccgcagggttaaaaaagaaaaaatcagtaacagtactggatgtgggtgatgcatatttttcagttcccttagataaagaattcaggaagtacactgcatttaccatacctagtataaaca atgagacaccagggattagatatcagtacaatgtgcttccacagggatggaaaggatcaccagcaatattccaaagcagcatgacaaaaatcttagagccttttagaaaacaaaatccagacatagttatcaatacatggacga tttgtaggatctgacttagaaatagggcagcatagaacaaaaatagaggaactgagacaacatctgttgaagtggggatttaccacaccagacaaaaaacatcagaacctccattcctttggatgggttatgaactccat cctgataaatggacagtacagcctatagtgctgccagaaaaggacagctggactgtcaatgacatacagaagttagtgggaaaattgggcaagtcagatttacccagggattaaagcaattatgtagactccttaggg gaaccaaggcactaacagaagtaataccactaacaaaagaagcagagctagaactggcagaaaacagggaaattctaaaagaaccagtacatggagtgtattatgacccatcaaaagacttaatagcggaaatacagaagcaggggca aggtcaatggacatatcaaatttatcaagagccatttaaaaatctgaaaacaggaaaatatgcaagaatgaggggtgcccacactaatgatgtaaaacaattaacagaggcagtgcaaaaaataaccacagaaagcatagtaatatgg ggaaagactcctaaatttaaactacccatacaaaaagaaacatggtggacagagtattggcaagccacctggattcctgagtgggagtttgtcaataccccttagtaaaattatggtaccagttagagaac ccataataggagcagaaactttctatgtagatggggcagctaacagggagactaaattaggaaaagcaggatatgttactaacaaagggagacaaaaagttgtctccataactgacacaacaaatcagaagactgagttacaagcaat tcttctagcattacaggattctggattagaagtaaacatagtaacagactcacaatatgcattaggaatcattcaagcacaaccagataaaagtgaatcagagatagtcaaataatagagcagttaataaaaaaaggtc tacctgacatgggtaccagcgcacaaaggaattggaggaaatgaacaagtagataaattagtcagtactggaatcaggaaagtactctttttagatggaatagataaagcccaagaagaacatgaaaaatatcacagtaattggaggg caatggctagtgattttaacctgccacctgtggtagcaaaagagatagtagccagctgtgataaatgtcagctaaaaggagaagccatggacaagtagactgtagtccaggaatatggcaactagattgtacacatttagaagg aaaaattatcctggtagcagttcatgtagccagtggatatatagaagcagaagttattccagcagaaacagggcaggaaacagcatactttctcttaaaattagcaggaagatggccagtaaaaacagtacagacaatggcagc aatttcaccagtactacagttaaggccgcctgttggtgggcaggaatcaagcaggaatttggcattccctacaatccccaaagtcaaggagtagtagaatctataaagaattaaagttataggacagataagagatcagg ctgaacatcttaagacagcagtacaaatggcagtattcatccacaattttaaaaggggggattggggggtacagtgcaggggaaagaatagtagacataatagcaacagacatacaaactaaagaactacaaaaacaaattac aaaaattcaaaattttcgggtttattacagggacagcagagatccactttggaaaggaccagcaaagcttctctggaaaggtgaaggggcagtagtaatacaagataatagtgacataaaagtagtgccaagaagaaaagcaaagatc attagggattatggaaaacagatggcaggtgatgattgtgtggcaagtagacaggatgaggattagaacatggaaaagtttagtaaaacaccatatgtttcaaggaaagctaagggatggttttatagacatcactatgaaagt actcatccgagaataagttcagaagtacacatcccactagggaatgcaaaattggtaataacaacatattggggtctacaggagaaagagactggcatttgggtcaaggagtctccatagaattgaggaaaaggagatatagca cacaattagaccctaacctagcagaccaactaattcatctgcattactttgattgtttttcagaatctgctataagaaatgccatattaggacatatagttagccctaggtgtgaatatcaagcaggacataacaaggtaggatctct acagtacttggcactaacagcattagtaagaccaagaaaaaagataaagccacctttgcctagtgttacaaaactgacagaggatagatggaacaagccccagaagaccaagggccacaaagggaaccatacaatggacactag aacttttagaggagctcaagaatgaagctgttagacattttcctaggatatggctccatagcttagggcaacatatctatgaaacttatggagatacttgggcaggagtggaagccataataagaattctgcaacaactgctgtttat tcatttcagaattgggtgtcaacatagcagaatagacattcttcgacgaaggagagcaagaaatggagccagtagatcctagagccctggaagcatccaggaagtcagcctaggactgcttgtaccaattgctattgtaaaaa gtgttgctttcattgccaagtttcataacaaaaggcttaggcatctcctatggcaggaagaagcggagacagcgacgaagagctcctcaagacagtcagactcatcaagtttctctatcaaagcagtagtacatgtaatg caatctttacaaatattagcagtagtagcattagtagtagcagcaataatagcaatagttgtgtggtccatagtattcatagaatataggaaaataagaagacaaaatagaaaggttgatagaataatagaaagagcag aagacagtggcaatgagagtgacggagatcaggaagaattatcagcacttgtggaaatggggcacgatgctccttgggatgttaatgatctgtaaagctgcagaaaatttgtgggtcacagtttattatggggtacctgtgtggaaag aagcaaccaccactctattttgtgcctcagatgctaaagcgtatgatacagaggtacataatgtttgggccacacatgcctgtgtacccacagaccccaacccacaagaagtagaactgaagaatgtgacagaaaattttaacatgtg gaaaaataacatggtagaccaaatgcatgaggatataattagtttatgggatcaaagcctaaagccatgtgtaaaattaaccccactctgtgttactttaaattgcactgattatgggaatgatactaacaccaataatagtagtgct actaaccccactagtagtagcgggggaatggaggggagaaataaaaaattgctctttcaatatcaccagaagcataagagataaagtgaagaatatgcacttttttatagtcttgatgtaataccaataaaagatgata atactagctataggttgagaagttgtaacacctcagtcattacacaggcctgtccaaaggtatcctttgaaccaattcccatacattattgtgccccggctggttttgcgattctaaagtgtaatgataaaaagttcaatggaaaagg accatgtacaaatgtcagcacagtacaatgtacacatggaattaggccagtagtatcaactgctgttaaatggcagtctagcagaagaagaggtagtaattagatcagacaatttctcggacaatgctaaagtcataatagta catctgaatctgtagaaattgtacaagactcaacaacattacaaggagaagtatacatgtaggaccaggcagagcaatttatacaacaggaataataggaaaaataagacaagcacattgtaacattagta gagcaaaatggaataacactttaaaacagatagttacaaaattaagagaacaatttaagaataaaacaatagtctttaatcctcaggaggggacccagaaattgtaatgcacagttttaattgtggaggggaatttttctactg taattcaacacaactgtttaacagtacttggaatggtactgcatggtcaaataacactgaaggaaatgacacaatcacactcccatgcagaataaaacaaattataaacatgtggcaggaagtaggaaaagcaatgtatgca cctcccatcagaggacaaattagatgttcatcaaatattacagggctgatattaacaagagatggtggtattaaccagaccaacaccaccgagattttcaggcctggaggaggagatatgaaggacaattggagaagtgaattatata aatataaagtagtaaaaattgaaccattaggagtagcaccaaggcaaagagtggtgcaaagagaaaaaagagcagtgggaataataggagctatgctccttgggttcttgggagcagcaggaagcactatgggcgcagc gtcaatgacgctgacggtacaggccagacaattattgtctggtatagtgcaacagcagaacaatttgctgagggctattgaggcgcaacagcatctgttgcacctcacagtctggggcatcaagcagctccaagagtcctggct gtggaaagatacctaagggatcaacagctcctggggttttggggttgctctggaaaactcatttgcaccactgctgtgccttggaatactagttggagtaataaatctctgagtcagatttgggataacatgacctggatgcagtggg aaagggaaattgataattacacaagcttaatatacaacttaattgaagaatcgcaaaaccaacaagaatgaacaagagttattggaattagataactgggcaagtttgtggaattggtttagcataacaaattggctgtggta tataaaaatattcataatgatagtaggaggcttggtaggtttaagaatagtttttactgtactttctatagtaaatagagttaggcagggatactcaccattgtcgtttcagacgcgcctcccaggaggggacccgacaggccc gaaggaatcgaagaagaaggtggagagacagatccggtcaattagtggattcttagcaattatctgggtcgacctgcggagcctgtgcctcttcagctaccaccgcttgagagacttactcttgattgtaacga ggattgtggaacttctgggacgcagggggtgggaagccctcaaatattggtggaatctcctacaatattggattcaggaactaaagaatagtgctgttagcttgctcaacgccacagccatagcagtagctgagggaactgatagggt
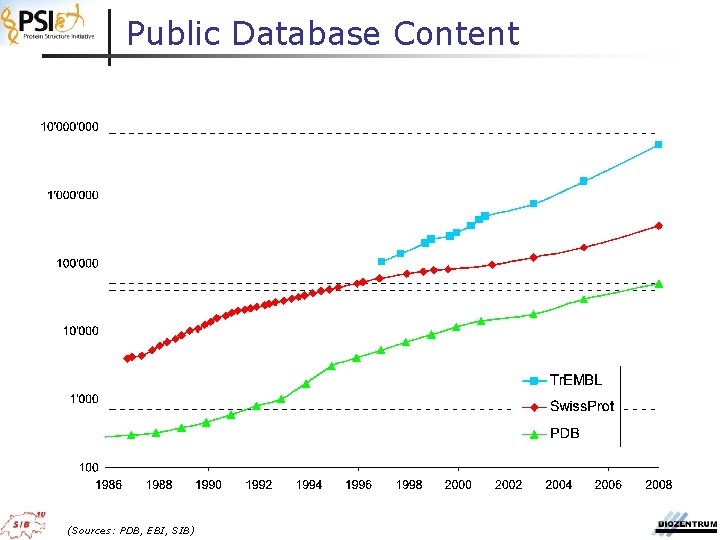
Public Database Content (Sources: PDB, EBI, SIB)

Overview: The KB Modeling Portal How can we find out if there is a model available for a given protein sequence? Well. . .

The KB Protein Model Portal n n The goal of the KB Protein Model Portal is to give access to the all models that can be leveraged from PSI targets and other experimental protein structures. The Protein Model Portal aims to provide a single interface to query simultaneously the existing pre-computed models at various sites, gives access to interactive services for template selection, target-template alignment, model building, and quality assessment.

The KB Protein Model Portal n The KB Protein Model Portal does NOT: n n n build models or develop modeling methods, but provides an interface to the participating expert groups and services; store all models in a database, but provides a query interface to the models provided by the partner sites; judge the quality of the models provided, but provides an interface to services for structure comparison and evaluation;
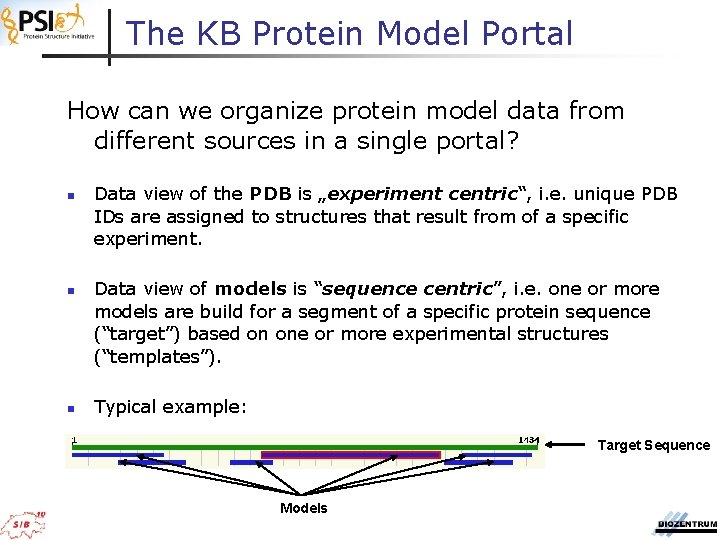
The KB Protein Model Portal How can we organize protein model data from different sources in a single portal? n n n Data view of the PDB is „experiment centric“, i. e. unique PDB IDs are assigned to structures that result from of a specific experiment. Data view of models is “sequence centric”, i. e. one or more models are build for a segment of a specific protein sequence (“target”) based on one or more experimental structures (“templates”). Typical example: Target Sequence Models

The KB Protein Model Portal Bottlenecks: n There is no common registry scheme for protein models. n For each target protein sequence, an ensemble of models will be available based on different templates, alternative alignments, and modeling methods. n Models will generally cover only fractions of the target sequence; parts of the target sequence might be missing, e. g. non modeled loops. n Protein models must update frequently to reflect updates of target sequence databases (Uni. Prot), template structure databases (PDB), and algorithmic improvements.

The KB Protein Model Portal Bottlenecks: n Sequence database accession codes are neither unique, nor stable. n Some groups do not store pre-computed models, but calculate models “on the fly”. n Some models will be based on outdated target sequences. Target protein sequences will be uniquely identified by hash values (UTSI) of their full length sequences as reference space.

The KB Protein Model Portal Data mapping of target sequences and structure models Unique (full length) target protein sequences are used as reference space to group models for identical targets. Target protein sequences will be uniquely identified by Hash function values (UTSI). >P 68399|CSK 21_BOVIN Casein kinase II subunit alpha - Bos taurus MSGPVPSRARVYTDVNTHRPREYWDYESHVVEWGNQDDYQLVRKLGRGKYSEVFEAINIT NNEKVVVKILKPVKKKKIKREIKILENLRGGPNIITLADIVKDPVSRTPALVFEHVNNTD FKQLYQTLTDYDIRFYMYEILKALDYCHSMGIMHRDVKPHNVMIDHEHRKLRLIDWGLAE FYHPGQEYNVRVASRYFKGPELLVDYQMYDYSLDMWSLGCMLASMIFRKEPFFHGHDNYD QLVRIAKVLGTEDLYDYIDKYNIELDPRFNDILGRHSRKRWERFVHSENQHLVSPEALDF LDKLLRYDHQSRLTAREAMEHPYFYTVVKDQARMGSSSMPGGSTPVSSANMMSGISSVPT PSPLGPLAGSPVIAAANPLGMPVPAAAGAQQ >P 68400|CSK 21_HUMAN Casein kinase II subunit alpha - Homo sapiens MSGPVPSRARVYTDVNTHRPREYWDYESHVVEWGNQDDYQLVRKLGRGKYSEVFEAINIT NNEKVVVKILKPVKKKKIKREIKILENLRGGPNIITLADIVKDPVSRTPALVFEHVNNTD FKQLYQTLTDYDIRFYMYEILKALDYCHSMGIMHRDVKPHNVMIDHEHRKLRLIDWGLAE Database Entry FYHPGQEYNVRVASRYFKGPELLVDYQMYDYSLDMWSLGCMLASMIFRKEPFFHGHDNYD QLVRIAKVLGTEDLYDYIDKYNIELDPRFNDILGRHSRKRWERFVHSENQHLVSPEALDF Uni. Prot. KB/Swiss-Prot P 68399 LDKLLRYDHQSRLTAREAMEHPYFYTVVKDQARMGSSSMPGGSTPVSSANMMSGISSVPT Uni. Prot. KB/Swiss-Prot P 68400 PSPLGPLAGSPVIAAANPLGMPVPAAAGAQQ Uni. Prot. KB/Swiss-Prot P 19138 >Q 5 U 065|Q 5 U 065_HUMAN Casein kinase 2, alpha 1 polypeptide - Homo sapiens Uni. Prot. KB/Tr. EMBL Q 5 U 065 MSGPVPSRARVYTDVNTHRPREYWDYESHVVEWGNQDDYQLVRKLGRGKYSEVFEAINIT Tr. EMBLnew AAH 53532 NNEKVVVKILKPVKKKKIKREIKILENLRGGPNIITLADIVKDPVSRTPALVFEHVNNTD Tr. EMBLnew AAH 11668 FKQLYQTLTDYDIRFYMYEILKALDYCHSMGIMHRDVKPHNVMIDHEHRKLRLIDWGLAE FYHPGQEYNVRVASRYFKGPELLVDYQMYDYSLDMWSLGCMLASMIFRKEPFFHGHDNYD International Protein Index (IPI) IPI 00016613 QLVRIAKVLGTEDLYDYIDKYNIELDPRFNDILGRHSRKRWERFVHSENQHLVSPEALDF International Protein Index (IPI) IPI 00744507 LDKLLRYDHQSRLTAREAMEHPYFYTVVKDQARMGSSSMPGGSTPVSSANMMSGISSVPT International Protein Index (IPI) IPI 00707334 PSPLGPLAGSPVIAAANPLGMPVPAAAGAQQ Ref. Seq >Q 9 D 9 I 4|TBC 20_MOUSE TBC 1 domain family member 20 NP_808227 - Mus musculus Ref. Seq NP_001886 MALRPSKGDGSAGRWDRGAGKADFNAKRKKKVAEIHQALNSDPIDLAALRRMAISEGGLL Ref. Seq NP_777060 TDEIRCQVWPKLLNVNTSEPPPVSRKDLRDMSKDYQQVLLDVRRSLRRFPPGMPDEQREG LQEELIDIILLVLDRNPQLHYYQGYHDIVVTFLLVVGERLATSLVEKLSTHHLRDFMDPT Ref. Seq XP_850579 MDNTKHILNYLMPIIDQVSPELHDFMQSAEVGTIFALSWLITWFGHVLMDFRHVVRLYDF Ref. Seq XP_001112324 FLACHPLMPIYFAAVIVLYREQEVLDCDCDMASVHHLLSQIPQDLPYETLISRAGDLFVQ Ref. Seq XP_001112363 FPPSELAREAAAQQEAERTAASTFKDFELASTQQRPDMVLRQRFRGLLRPEARTKDVLTK Ensembl ENSCAFP 00000010339 PRTNRFVKLAVMGLTVALGAAALAVVKSALEWAPKFQLQLFP Ensembl >ipi|IPI 00707334. 1 CASEIN KINASE II ENSP 00000217244 SUBUNIT ALPHA. Ensembl ENSP 00000339247 MSGPVPSRARVYTDVNTHRPREYWDYESHVVEWGNQDDYQLVRKLGRGKYSEVFEAINIT NNEKVVVKILKPVKKKKIKREIKILENLRGGPNIITLADIVKDPVSRTPALVFEHVNNTD Ensembl ENSP 00000341595 FKQLYQTLTDYDIRFYMYEILKALDYCHSMGIMHRDVKPHNVMIDHEHRKLRLIDWGLAE Ensembl ENSMMUP 00000037659 FYHPGQEYNVRVASRYFKGPELLVDYQMYDYSLDMWSLGCMLASMIFRKEPFFHGHDNYD EMBL Annotated CONs EAX 10665 QLVRIAKVLGTEDLYDYIDKYNIELDPRFNDILGRHSRKRWERFVHSENQHLVSPEALDF EMBL CDS AAA 35503 LDKLLRYDHQSRLTAREAMEHPYFYTVVKDQARMGSSSMPGGSTPVSSANMMSGISSVPT PIR-PSD Archive A 30319 PSPLGPLAGSPVIAAANPLGMPVPAAAGAQQ European Patent Office (EPO) US Patent Office (USPO) Japan Patent Office (JPO) TROME H-Invitational Database (H- Inv. DB) CS 458506 AAE 81305 BD 879267 NT_011387_19_8 HIT 000053902 CRC 64: D 3 B 6 F 5 D 13 FF 7422 D MD 5: b 6 f 2 c 321 d 42 d 50 b 985186307434 b 5166 UPI: UPI 0000000 CB 5 Real time annotation Version 1 1 1 1 Organism Bos taurus (Bovine) Homo sapiens (Human) Bos taurus (Bovine) Macaca mulatta (Rhesus macaque) Homo sapiens (Human) 1 1 3 Last Seen 2007 -07 -24 2004 -11 -09 2007 -07 -24 2003 -08 -30 2003 -06 -14 2004 -11 -15 2007 -06 -29 2007 -05 -08 2006 -02 -19 2007 -05 -08 2007 -06 -04 2007 -05 -31 2006 -09 -27 2007 -06 -04 2007 -06 -12 2007 -06 -20 2003 -04 -04 2007 -06 -09 2007 -06 -01 2004 -08 -28 2006 -07 -02 Active Yes No No No Yes Yes Yes No Yes Yes No No CRC 64: A 3 B 6 F 5 D 13 DF 7422 E MD 5: 605 f 4802 e 88 ec 1443 d 36520 ac 05 df 3 b 9 UPI: UPI 0000044948 familiaris (Dog) Canis Homo sapiens (Human) 1 1 First Seen 2004 -11 -23 1990 -11 -01 2005 -05 -10 2003 -06 -14 2003 -03 -29 2003 -03 -14 2006 -05 -16 2006 -01 -24 2004 -07 -08 2004 -09 -24 2004 -09 -14 2005 -08 -31 2006 -06 -15 2004 -12 -09 2003 -04 -01 2004 -05 -12 2006 -04 -03 2006 -08 -01 2007 -01 -15 2003 -03 -12 2003 -03 -31 2007 -03 -20 2003 -03 -26 2007 -01 -11 2004 -08 -23 2006 -07 -02 Homo sapiens (Human)
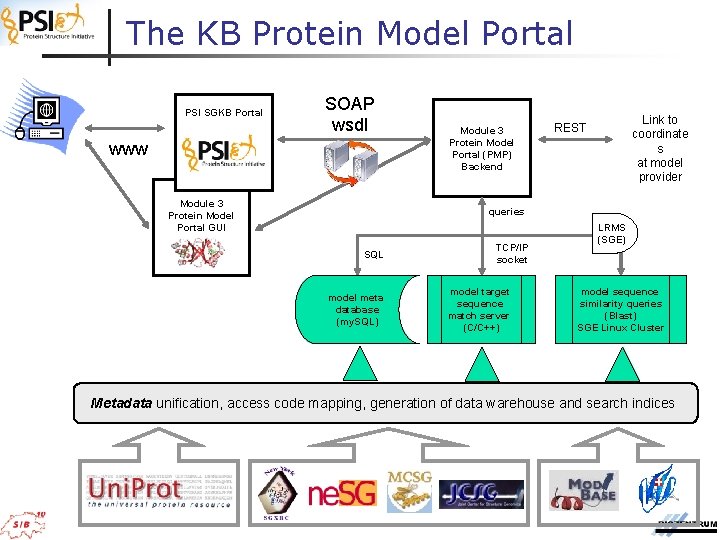
The KB Protein Model Portal PSI SGKB Portal SOAP wsdl www Module 3 Protein Model Portal GUI Module 3 Protein Model Portal (PMP) Backend Link to coordinate s at model provider REST queries SQL model meta database (my. SQL) TCP/IP socket model target sequence match server (C/C++) LRMS (SGE) model sequence similarity queries (Blast) SGE Linux Cluster Metadata unification, access code mapping, generation of data warehouse and search indices


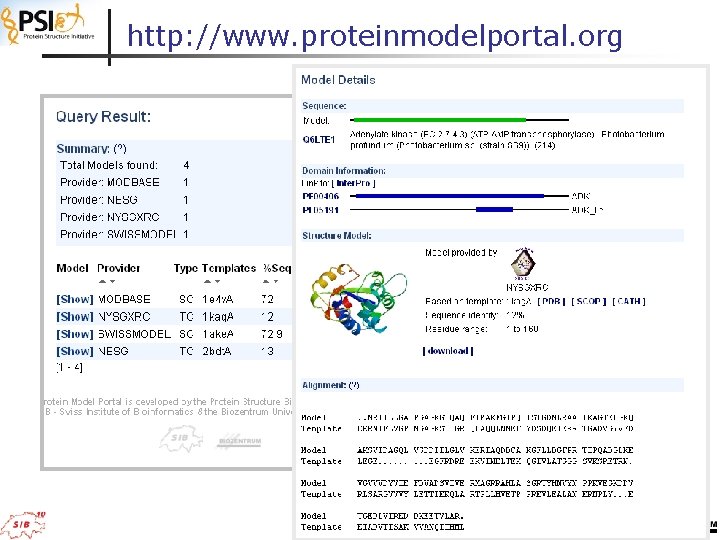
http: //www. proteinmodelportal. org
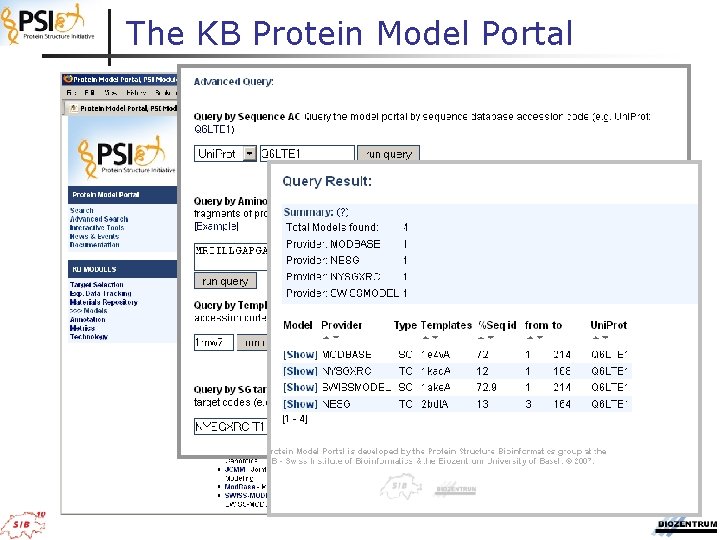
The KB Protein Model Portal
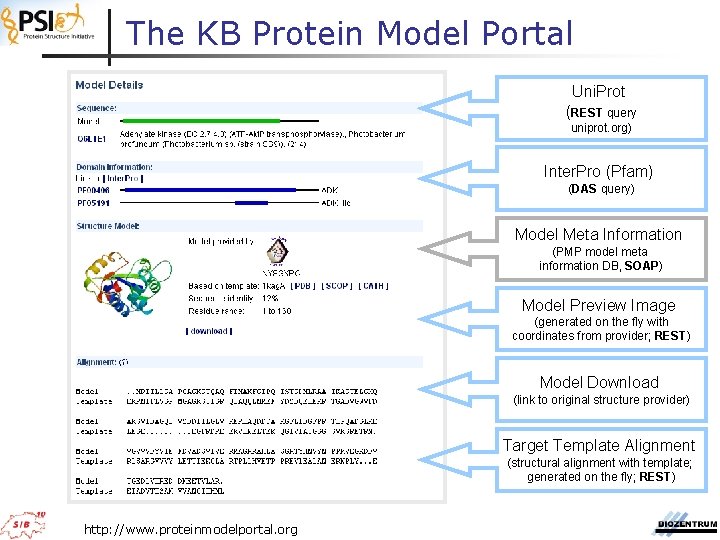
The KB Protein Model Portal Uni. Prot (REST query uniprot. org) Inter. Pro (Pfam) (DAS query) Model Meta Information (PMP model meta information DB, SOAP) Model Preview Image (generated on the fly with coordinates from provider; REST) Model Download (link to original structure provider) Target Template Alignment (structural alignment with template; generated on the fly; REST) http: //www. proteinmodelportal. org

Overview: The KB Model Portal n Introduction n Protein Model Portal n n n Mission and Goals n PMP Version 1. 0: Content and Features n Technical Implementation Outlook: Next steps n New Features & Functions n Modeling Portal Community Workshop Questions & Discussion

Outlook: Next steps n New Features & Functions n Better visualization of query results

Outlook: Next steps n New Features & Functions n Better visualization of query results n n Interactive structure / model comparison Visualization of mapped properties (sequence conservation; quality assessment results; Uni. Prot annotation) Residue-level annotations (Uni. Prot, Inter. Pro) Model quality assessment tools
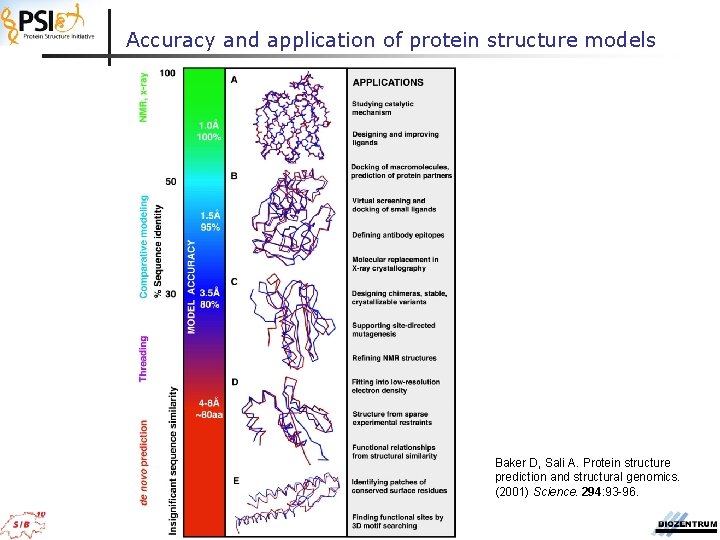
Accuracy and application of protein structure models Baker D, Sali A. Protein structure prediction and structural genomics. (2001) Science. 294: 93 -96.

Community Workshop on Applications of Protein Models in Biomedical Research University of California, San Francisco July 11 &12, 2008 How are protein structure models used in biomedical research projects today? n n Which requirements and limitations exist for the different applications? n Structure based drug design n Analysis of SNPs and disease related mutations n Phasing X-ray crystallography data by molecular replacement n Interpretation of low-resolution experimental data n Protein engineering and design n Functional characterization of novel proteins

Community Workshop on Applications of Protein Models in Biomedical Research University of California, San Francisco July 11 &12, 2008 We need your input! Please. . . n participate in the workshop; n send us examples of successful use of models in your work, and negative examples when models did not do what you expected; n let us know, what you expect from proteins models, and which aspects of modeling techniques need improvement to make models more useful to your research. n modeling_workshop@psi-structuralgenomics. org

Acknowledgements Biozentrum & SIB, University of Basel Michael Podvinec Jürgen Kopp Lorenza Bordoli Rainer Pöhlmann Konstantin Arnold James Battey Pascal Benkert Florian Kiefer SIB Geneva Eric Jain RCSB-PDB MCSG Helen Berman Christine Orengo John Westbrook David Lee Wendy Tao JCMM FCCC/NMHRCM Adam Godzik Roland Dunbrack Jr. NESG UCSF/NYSGXRC Diana Murray Andrej Sali Ursula Pieper Funding: NIH – National Institutes of Health SIB – Swiss Institue of Bioinformatics
- Slides: 25