The Proteome Analysis Database http www ebi ac

The Proteome Analysis Database http: //www. ebi. ac. uk/proteome/ EMBL Outstation — The European Bioinformatics Institute

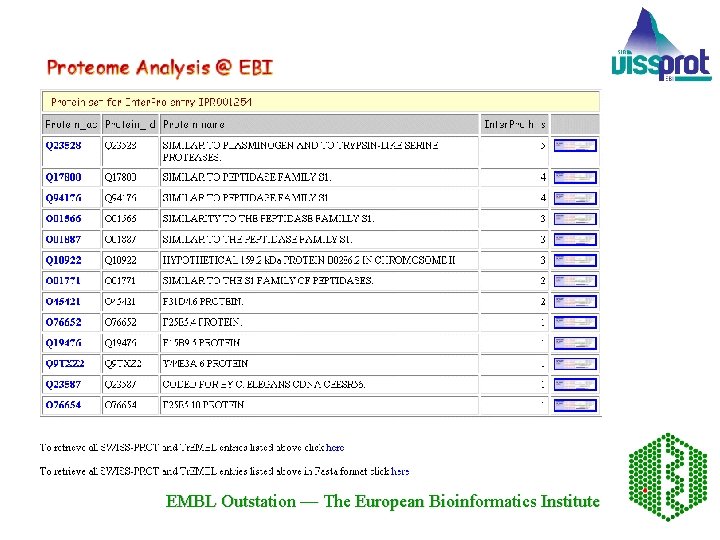
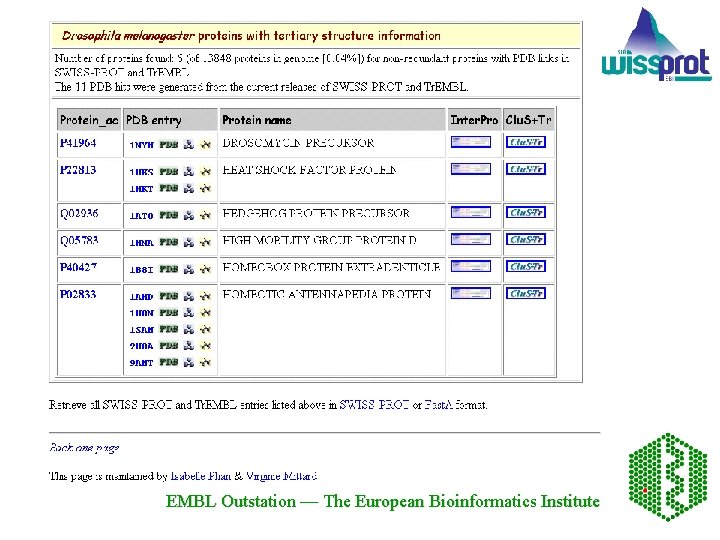
The Proteome Analysis Database - aims at integrating information from a variety of sources that will together facilitate the classification of the proteins in complete proteome sets. Structural information includes amino acid composition for each of the proteomes and links are provided to HSSP, the Homology derived Secondary Structure of Proteins, and PDB, the Protein Data Bank, for individual proteins from each of the proteomes. Functional classification using Gene Ontology (GO) is available. EMBL Outstation — The European Bioinformatics Institute

EMBL Outstation — The European Bioinformatics Institute

Complete proteome sets for each organism have been assembled from SPTR (SWISS-PROT + Tr. EMBLnew) database to be wholly non-redundant at the sequence level. Archaeal, bacterial and the A. thaliana and S. cerevisiae proteome sets: A standard procedure based on tracking protein identifiers from the nucleotide sequence database EMBL-Bank is used. D. melanogaster, C. elegans and H. sapiens proteome sets: There are no unique identifiers in EMBL-Bank that allow the identification of all genome-project sequences for these organisms. Each of these organisms is treated separately as a special case. EMBL Outstation — The European Bioinformatics Institute
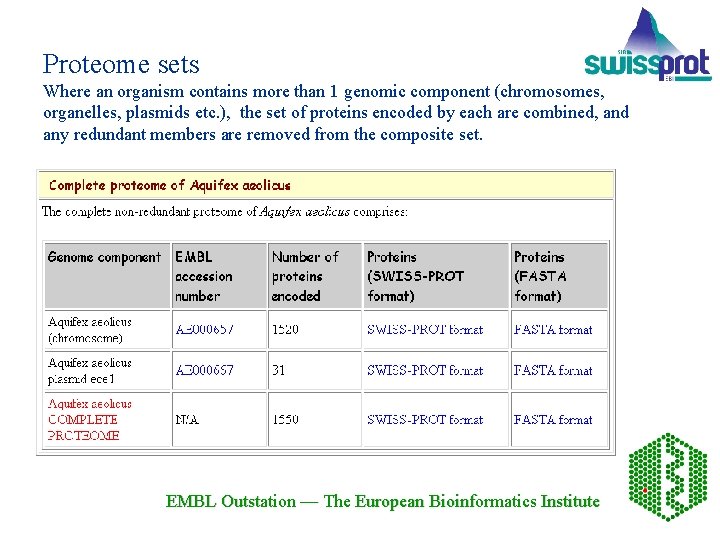
Proteome sets Where an organism contains more than 1 genomic component (chromosomes, organelles, plasmids etc. ), the set of proteins encoded by each are combined, and any redundant members are removed from the composite set. EMBL Outstation — The European Bioinformatics Institute

EMBL Outstation — The European Bioinformatics Institute

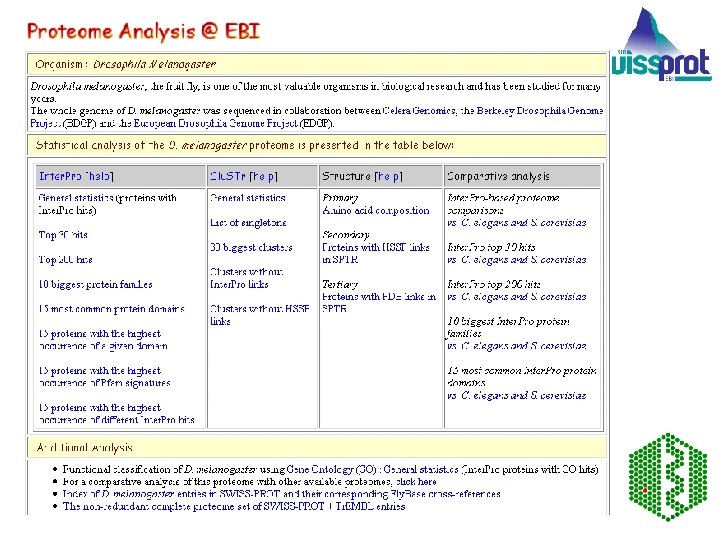
For D. melanogaster proteins, the complete set of are those predicted from the Celera genomic sequence. Each entry is tagged on entry into Tr. EMBL Outstation — The European Bioinformatics Institute

EMBL Outstation — The European Bioinformatics Institute

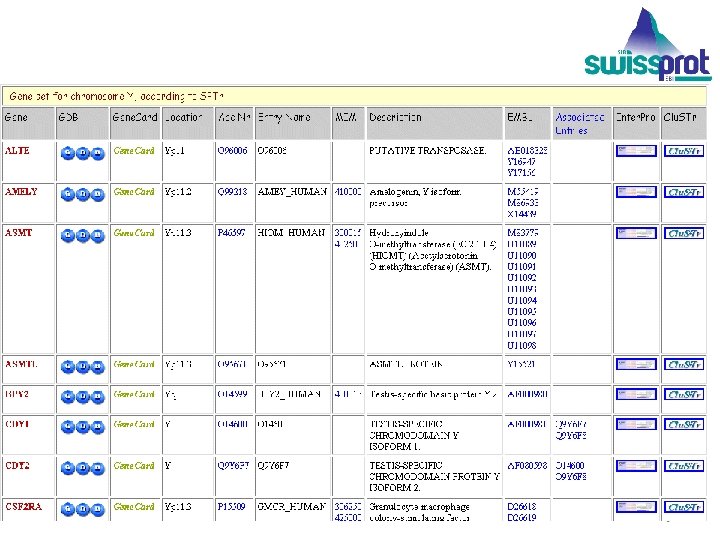
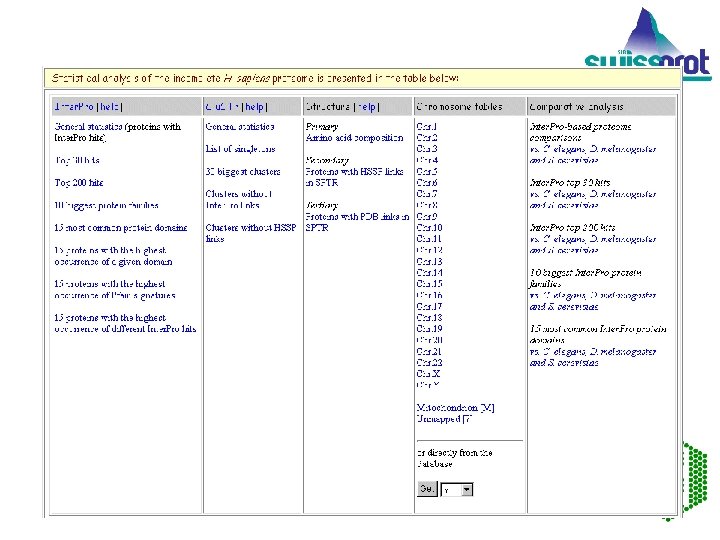
CHROMOSOME TABLES Map proteins to chromosomes for yeast and human. The information needed to make protein-chromosome mappings is distributed over several databases. Resources are pooled to make mappings. EMBL Outstation — The European Bioinformatics Institute
EMBL Outstation — The European Bioinformatics Institute
EMBL Outstation — The European Bioinformatics Institute

EMBL Outstation — The European Bioinformatics Institute
EMBL Outstation — The European Bioinformatics Institute
EMBL Outstation — The European Bioinformatics Institute
EMBL Outstation — The European Bioinformatics Institute
EMBL Outstation — The European Bioinformatics Institute
EMBL Outstation — The European Bioinformatics Institute

EMBL Outstation — The European Bioinformatics Institute

30 biggest clusters EMBL Outstation — The European Bioinformatics Institute
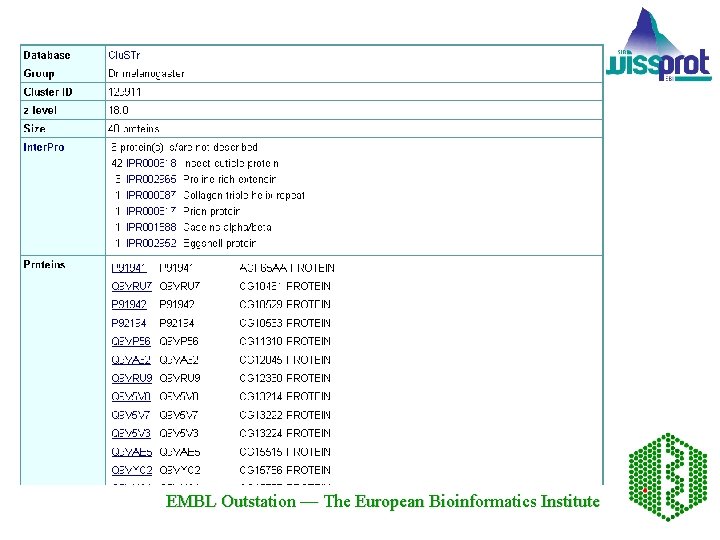
EMBL Outstation — The European Bioinformatics Institute
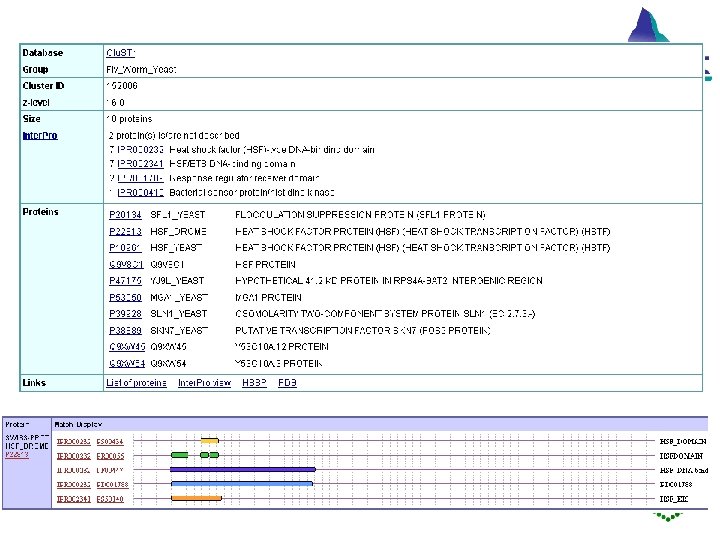
EMBL Outstation — The European Bioinformatics Institute
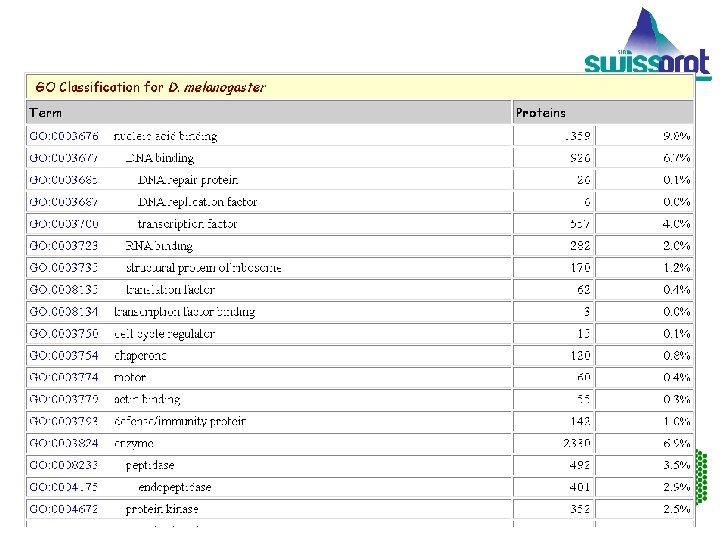
EMBL Outstation — The European Bioinformatics Institute

Gene Ontology. TM Consortium produces a dynamic controlled vocabulary that can be applied to all eukaryotes. EMBL Outstation — The European Bioinformatics Institute

The Proteome Analysis Database Currently integrates data on 44 complete proteomes: eukaryotes: 5 organisms archaea: 8 organisms bacteria: 31 organisms EMBL Outstation — The European Bioinformatics Institute

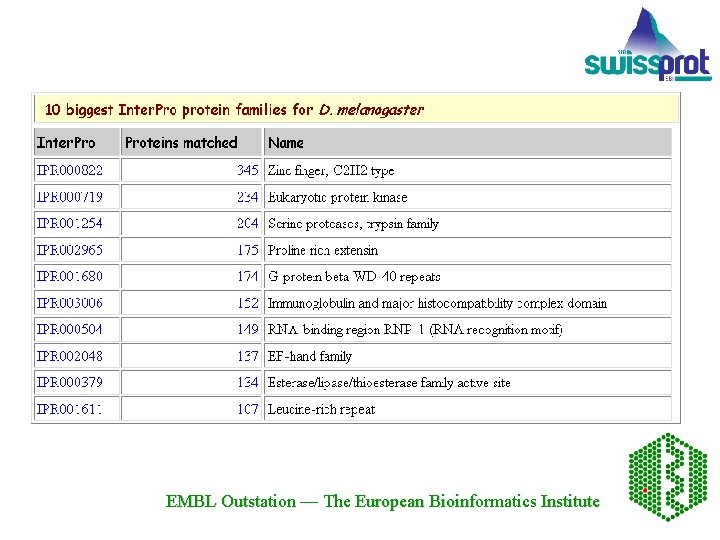
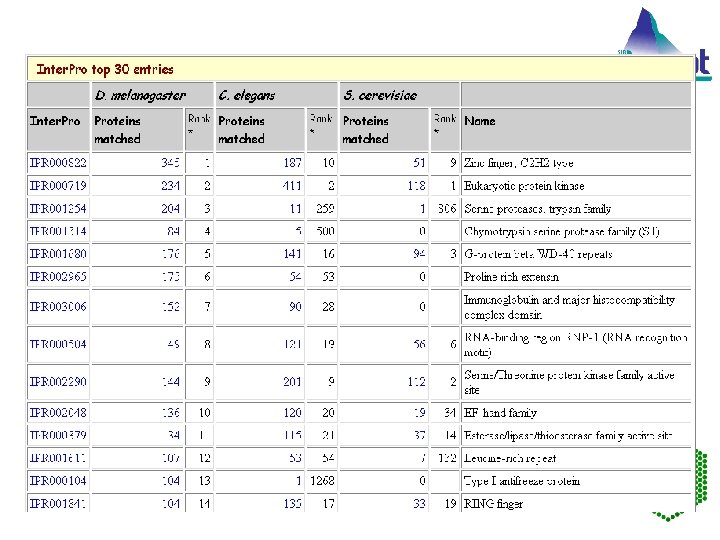
Inter. Pro: covers 31% to 67% of the proteins from each of the complete genomes. Eukaryote Arabidopsis thaliana 65. 5% Drosophila melanogaster 67. 8% Caenorhabditis elegans 64. 0% Saccharomyces cerevisiae 61. 7% Homo sapiens 71. 8% of the incomplete proteome (SWISS-PROT and Tr. EMBL) 59. 7% of the complete proteome (SWISS-PROT, Tr. EMBL and Ensembl) Bacteria For example: Bacillus subtilis 61. 9% Mycobacterium tuberculosis 64. 7% Xylella fastidiosa 47. 5% Rickettsia prowazekii 73. 5% Archaea Halobacterium sp. NRC-1 57. 2% Pyrococcus abyssi 66. 2% EMBL Outstation — The European Bioinformatics Institute
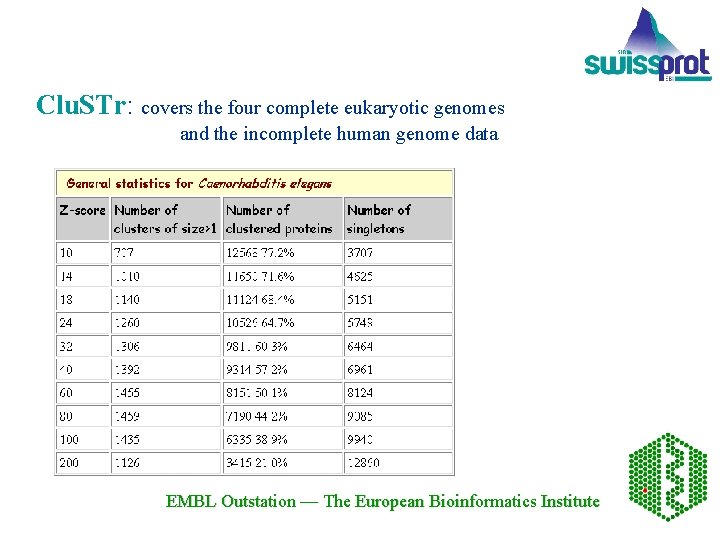
Clu. STr: covers the four complete eukaryotic genomes and the incomplete human genome data EMBL Outstation — The European Bioinformatics Institute

Summary The Proteome Analysis Database provides a broad view of the proteome data classified according to signatures describing particular sequence motifs and sequence similarities and affords the option of examining various specific details like structure or functional classification. EMBL Outstation — The European Bioinformatics Institute

Publication: Apweiler R. , Biswas M. , Fleischmann W. , Kanapin A. , Karavidopoulou Y. , Kersey P. , Kriventseva E. V. , Mittard V. , Mulder N. , Phan I. , Zdobnov E. "Proteome Analysis Database: online application of Inter. Pro and Clu. STr for the functional classification of proteins in whole genomes. " Nucleic Acids Res. 29(1): 44 -48(2001) EMBL Outstation — The European Bioinformatics Institute

The SWISS-PROT group at the EBI EMBL Outstation — The European Bioinformatics Institute

EMBL Outstation — The European Bioinformatics Institute
- Slides: 30