The project of mapping Human Genome Why they

































- Slides: 64

The project of mapping Human Genome • Why they want to make a map of the human genome ? ? ?

The project of mapping Human Genome • The objective of sequencing human genome: 1. To understand how genes work together to direct the growth, development and maintenance of an entire organism. 2. By knowing the whole genome sequence it will help to study the parts of the genome outside the genes. This includes the long sequences of nonsense (junk) DNA that has no clear functions.

3. To learn about other important parts of the genome, such as the regulatory regions that help control when genes are turned on and off. 4. To draw accurate map for the chromosomal locations of genes responsible for genetic diseases. Already, about 1400 genes are identified for human genetic diseases as a result of human genome mapping. 5. By comparing human genome map with other species maps, it will be possible to understand the process of evolution.

The plan The project of human genome sequencing began on 1990 and completed on 2003. The whole genome cannot be sequenced all at once because available methods of DNA sequencing can only work with short stretches of DNA at a time. Instead, the genome was broken into smaller pieces; approximately 150, 000 base pairs in length. These pieces were cloned into plasmid vector before they were amplifies in bacterial culture.

• Using restriction enzymes, the pieces of human DNA were cut into small pieces and each gene was identified by specific probe before it was sequenced. Then the genes were reassembled in the proper order to obtain the sequence of the whole genome.

Data obtained from mapping human genome 1. The human genome sequence is almost exactly the same (99. 9%) in all people. 2. There approximately 23, 000 genes in human beings being mapped, the same range as in mice and roundworms. Before this mapping process, the human genome was estimated to contain about 80‐ 140 thousands genes, based on comparison with the size of bacteria in which the actual gene mapping already obtained.

3. The human genome contains 3. 2 billion nucleotide base pairs. The average gene consists of 3, 000 base pairs, but sizes vary greatly, with the largest known human gene has 2. 4 million base pairs which is responsible for expressing the dystrophin protein. 4. Functions are still unknown for more than 50% of discovered genes. 5. Genes appear to be concentrated in random areas along the genome, with extended regions of non‐coding DNA in between.

6. Particular gene sequences have been found to be associated with various diseases, including breast cancer, muscle disease, deafness, and blindness. 7. Pieces of up to 30, 000 C and G bases repeating over and over often occur adjacent to gene‐rich areas, forming a barrier between the genes and the junk DNA. These C‐G rich segments are believed to help in the regulation of gene activity. 8. Chromosome 1 (the largest human chromosome) has the most number of genes (3, 168), and Y chromosome has the fewest number of genes (344).

Comparison of human genome with other organisms 1. Unlike the random distribution of gene‐rich areas in human's, many other organisms' genomes are more uniform, with genes evenly spaced throughout. 2. Humans have on average three times as many kinds of proteins as the fly or worm because of m. RNA transcript alternative splicing and chemical modifications to the proteins. This process can yield different protein products from the same gene.

3. Humans share similar protein families with worms, flies, and plants, but the number of gene family members are more expanded in humans, especially in proteins involved in development and immunity. 4. The human genome has a much greater portion (50%) of repeat sequences than the mustard weed plant (11%), the worm (7%), and the fly (3%).

5. Over 40% of predicted human proteins share similarity with fruit‐fly or worm proteins 6. As a conclusion from this mapping sequence of human genome, it is believed that the quality of protein types produced by the genome is more important in providing the overall human phenotype than the number of genes that express these proteins.

DNA sequencing

Why sequence DNA? • All genes available for an organism to use -- a very important tool for biologists • Not just sequence of genes, but also positioning of genes and sequences of regulatory regions • New recombinant DNA constructs must be sequenced to verify construction or positions of mutations

Sequencing Methods • • • Maxam/Gilbert chemical sequencing Sanger chain termination sequencing Pyrosequencing Bisulfite Sequencing Array sequencing

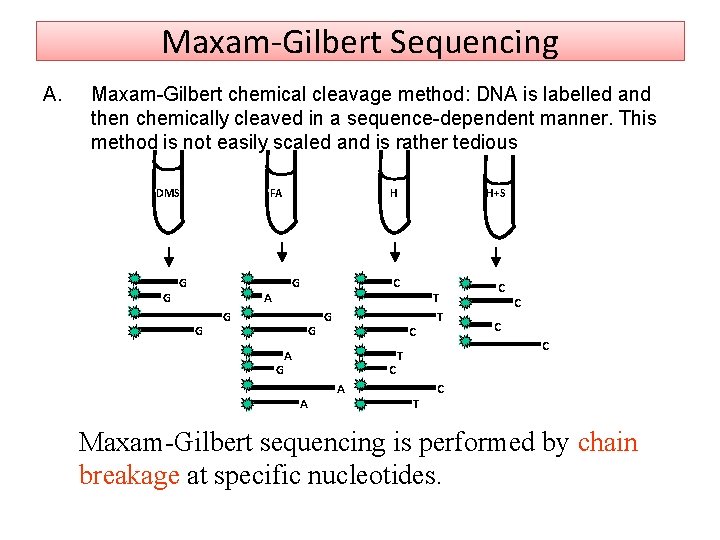
Maxam‐Gilbert Sequencing A. Maxam-Gilbert chemical cleavage method: DNA is labelled and then chemically cleaved in a sequence-dependent manner. This method is not easily scaled and is rather tedious DMS G FA G G A G H G C G G C A H+S A T T C C C T T C C Maxam-Gilbert sequencing is performed by chain breakage at specific nucleotides.

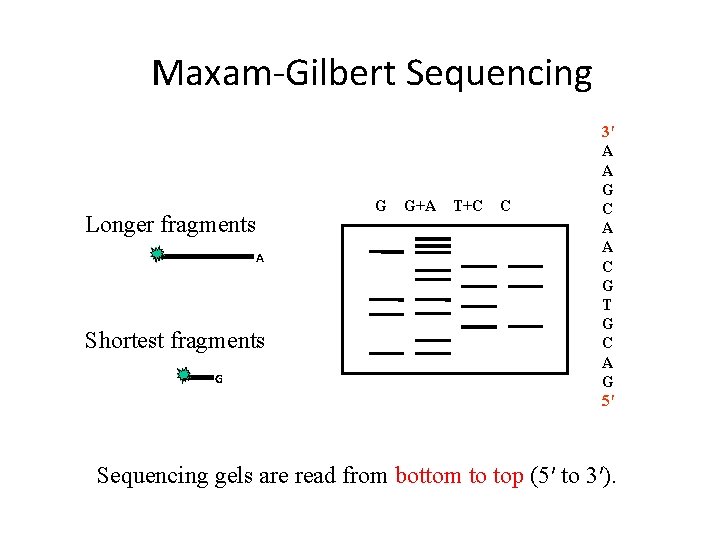
Maxam‐Gilbert Sequencing Longer fragments A Shortest fragments G G G+A T+C C 3′ A A G C A A C G T G C A G 5′ Sequencing gels are read from bottom to top (5′ to 3′).

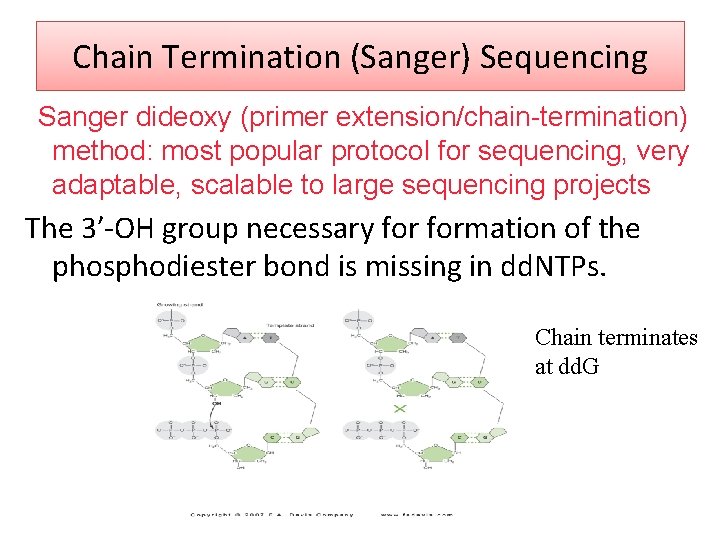
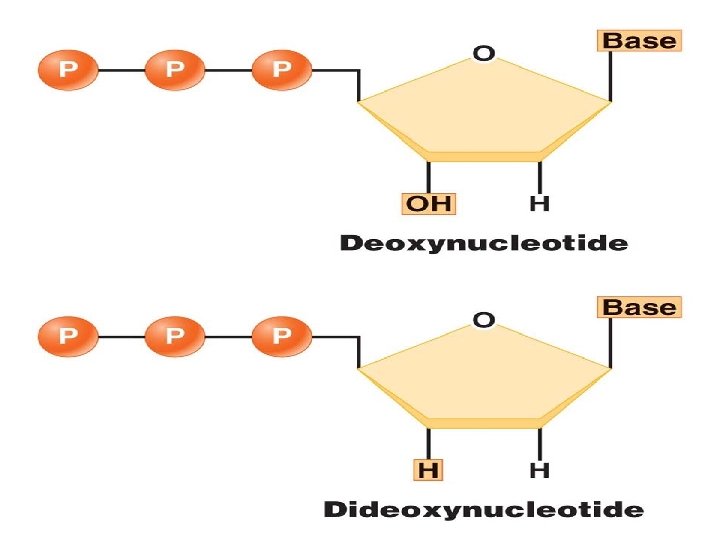
Chain Termination (Sanger) Sequencing Sanger dideoxy (primer extension/chain-termination) method: most popular protocol for sequencing, very adaptable, scalable to large sequencing projects The 3′‐OH group necessary formation of the phosphodiester bond is missing in dd. NTPs. Chain terminates at dd. G


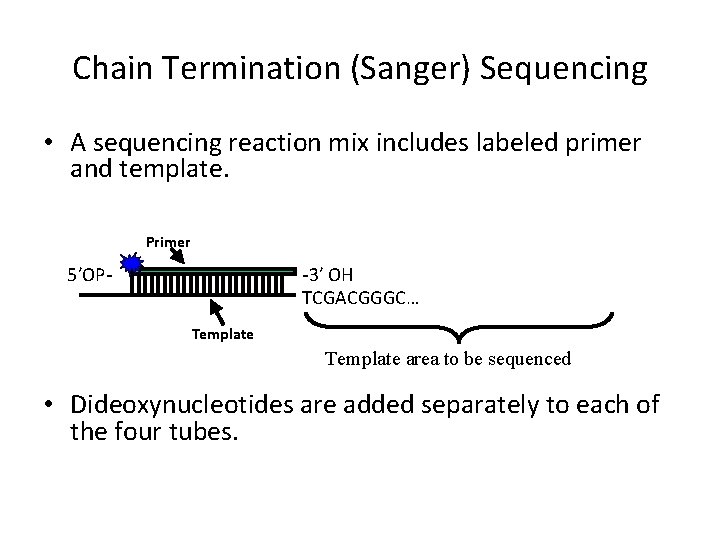
Chain Termination (Sanger) Sequencing • A sequencing reaction mix includes labeled primer and template. Primer -3′ OH TCGACGGGC… 5′OPTemplate area to be sequenced • Dideoxynucleotides are added separately to each of the four tubes.

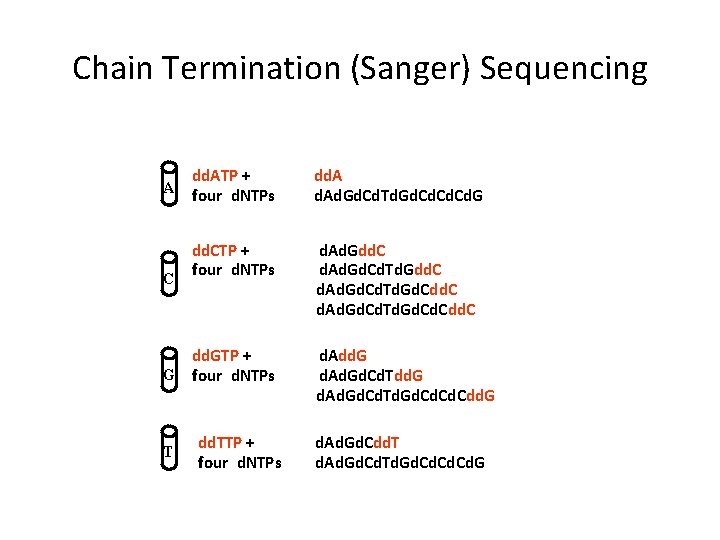
Chain Termination (Sanger) Sequencing A C dd. ATP + four d. NTPs dd. Ad. Gd. Cd. Td. Gd. Cd. Cd. G dd. CTP + four d. NTPs d. Ad. Gdd. C d. Ad. Gd. Cd. Td. Gd. Cdd. C dd. GTP + G four d. NTPs T dd. TTP + four d. NTPs d. Add. G d. Ad. Gd. Cd. Td. Gd. Cd. Cdd. G d. Ad. Gd. Cdd. T d. Ad. Gd. Cd. Td. Gd. Cd. Cd. G

Chain Termination (Sanger) Sequencing • With addition of enzyme (DNA polymerase), the primer is extended until a dd. NTP is encountered. • The chain will end with the incorporation of the dd. NTP. • With the proper d. NTP: dd. NTP ratio, the chain will terminate throughout the length of the template. • All terminated chains will end in the dd. NTP added to that reaction.

Chain Termination (Sanger) Sequencing • The collection of fragments is a sequencing ladder. • The resulting terminated chains are resolved by electrophoresis. • Fragments from each of the four tubes are placed in four separate gel lanes.

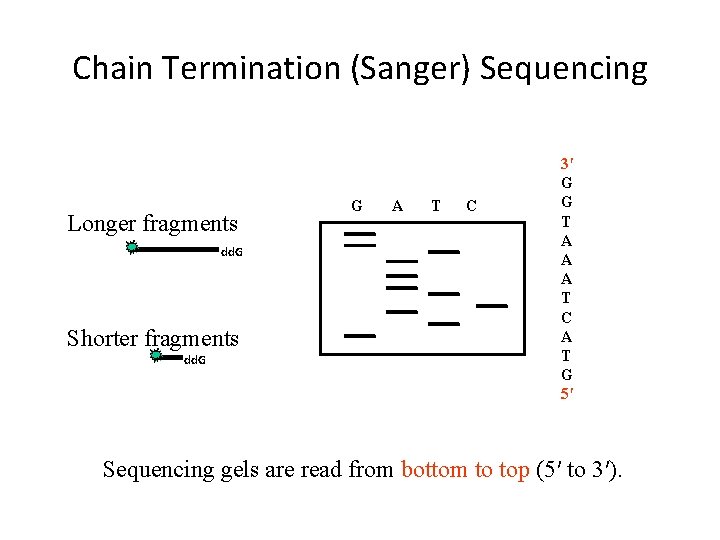
Chain Termination (Sanger) Sequencing Longer fragments dd. G Shorter fragments dd. G G A T C 3′ G G T A A A T C A T G 5′ Sequencing gels are read from bottom to top (5′ to 3′).

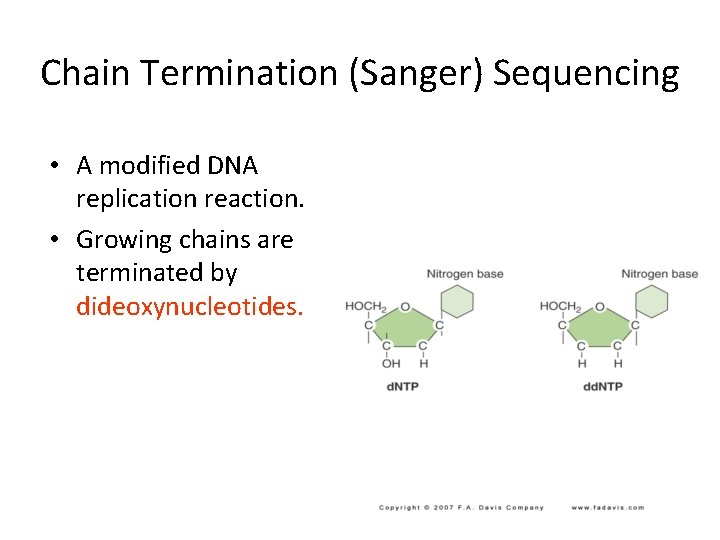
Chain Termination (Sanger) Sequencing • A modified DNA replication reaction. • Growing chains are terminated by dideoxynucleotides.

for dideoxy sequencing you need: 1) Single stranded DNA template 2) A primer for DNA synthesis 3) DNA polymerase 4) Deoxynucleoside triphosphates and dideoxynucleotide triphosphates

Primers for DNA sequencing • Oligonucleotide primers can be synthesized by phosphoramidite chemistry--usually designed manually and then purchased • Sequence of the oligo must be complimentary to DNA flanking sequenced region • Oligos are usually 15 -30 nucleotides in length

DNA templates for sequencing: • Single stranded DNA isolated from recombinant M 13 bacteriophage containing DNA of interest • Double-stranded DNA that has been denatured • Non-denatured double stranded DNA (cycle sequencing)

Reagents for sequencing: DNA polymerases • Should be highly processive, and incorporate dd. NTPs efficiently • Should lack exonuclease activity • Thermostability required for “cycle sequencing”

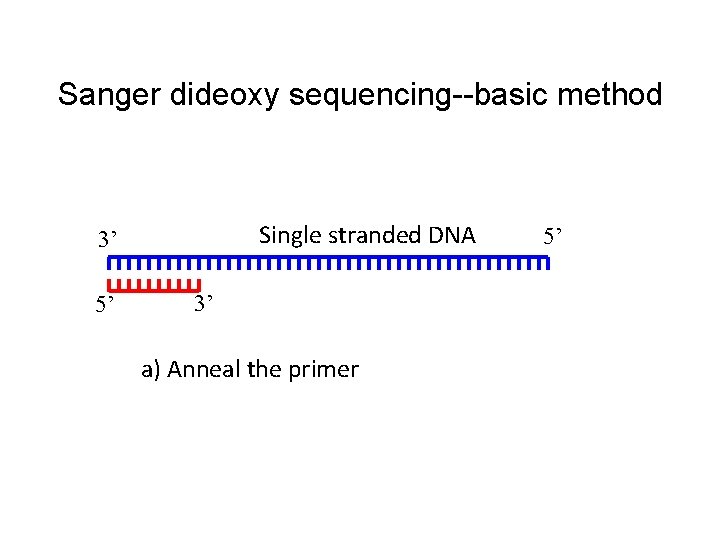
Sanger dideoxy sequencing--basic method Single stranded DNA 3’ 5’ 3’ a) Anneal the primer 5’

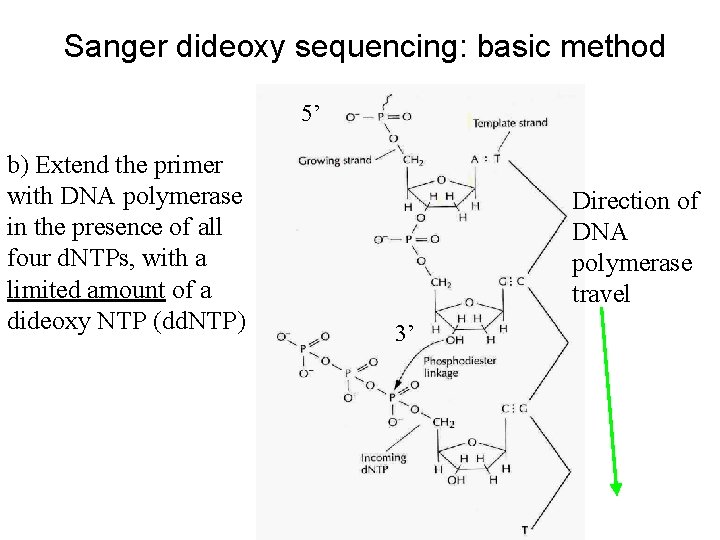
Sanger dideoxy sequencing: basic method 5’ b) Extend the primer with DNA polymerase in the presence of all four d. NTPs, with a limited amount of a dideoxy NTP (dd. NTP) Direction of DNA polymerase travel 3’

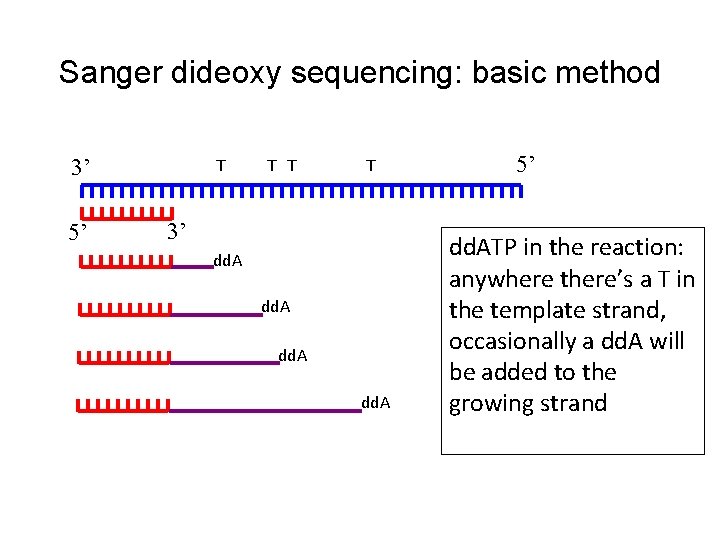
Sanger dideoxy sequencing: basic method 3’ 5’ T T 3’ dd. A 5’ dd. ATP in the reaction: anywhere there’s a T in the template strand, occasionally a dd. A will be added to the growing strand

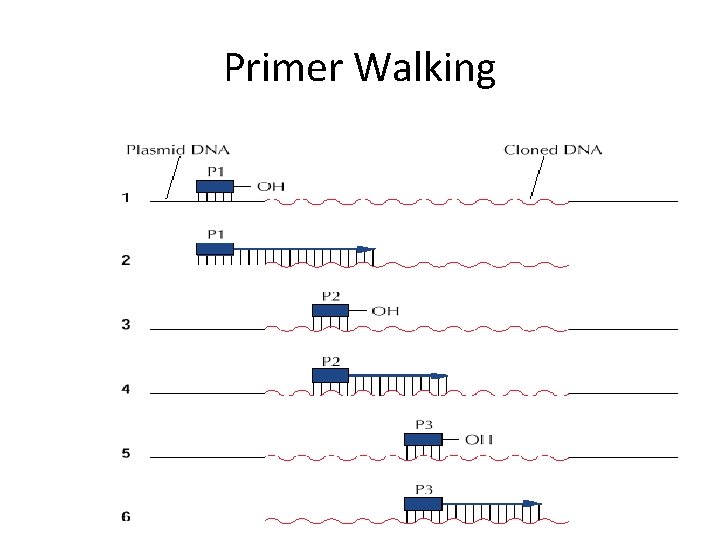
Primer Walking

How to visualize DNA fragments? • Radioactivity – Radiolabeled primers (kinase with 32 P) – Radiolabelled d. NTPs (gamma 35 S or 32 P) • Fluorescence – dd. NTPs chemically synthesized to contain fluoresces – Each dd. NTP fluoresces at a different wavelength allowing identification

Analysis of sequencing products: Polyacrylamide gel electrophoresis--good resolution of fragments differing by a single d. NTP – Slab gels: as previously described – Capillary gels: require only a tiny amount of sample to be loaded, run much faster than slab gels, best for high throughput sequencing.

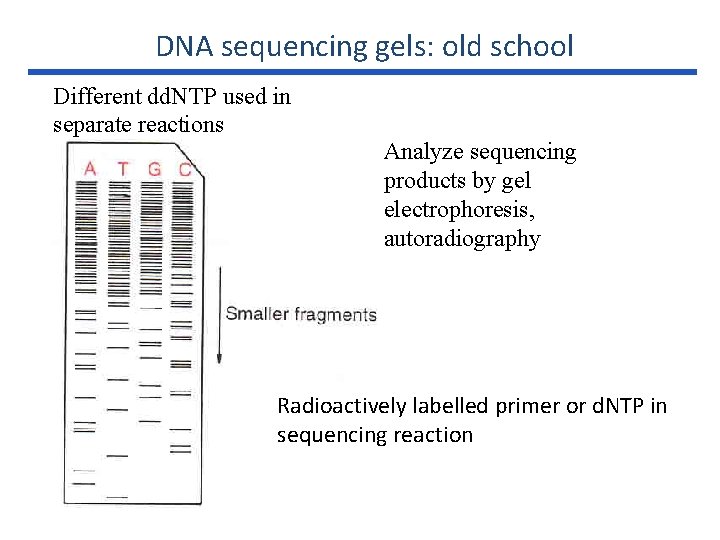
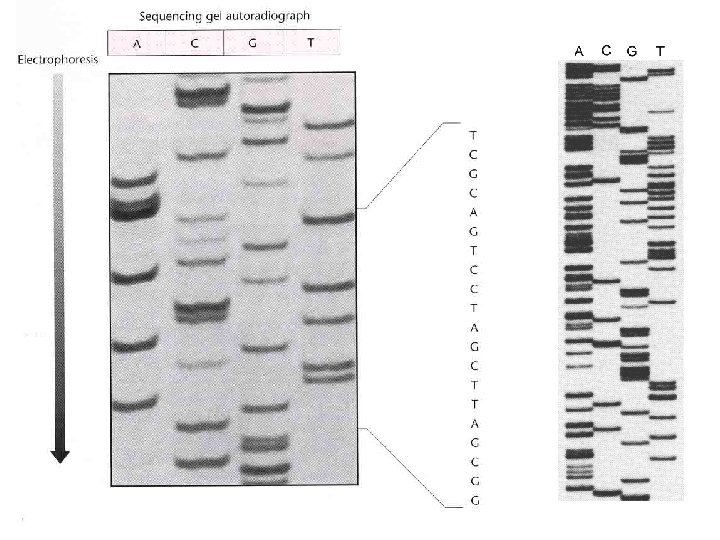
DNA sequencing gels: old school Different dd. NTP used in separate reactions Analyze sequencing products by gel electrophoresis, autoradiography Radioactively labelled primer or d. NTP in sequencing reaction


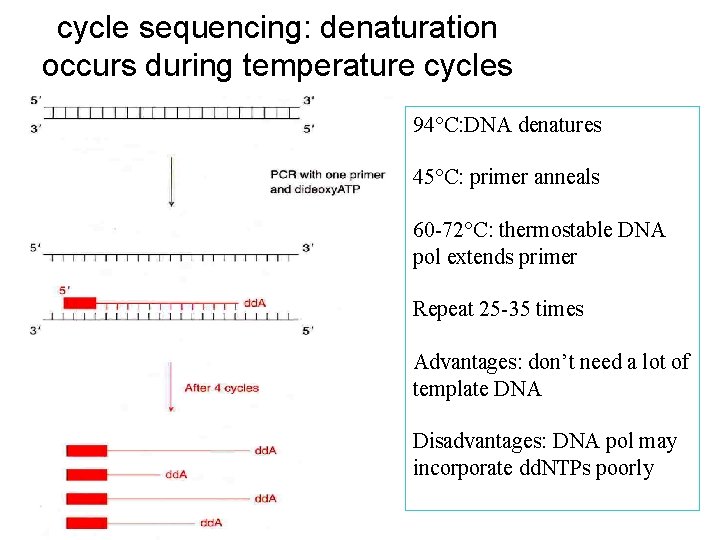
cycle sequencing: denaturation occurs during temperature cycles 94°C: DNA denatures 45°C: primer anneals 60 -72°C: thermostable DNA pol extends primer Repeat 25 -35 times Advantages: don’t need a lot of template DNA Disadvantages: DNA pol may incorporate dd. NTPs poorly

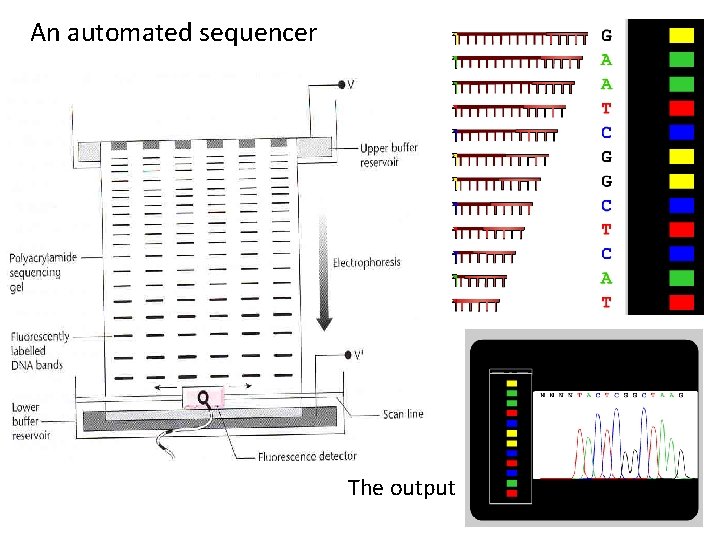
An automated sequencer The output

Current trends in sequencing: It is rare for labs to do their own sequencing: --costly, perishable reagents --time consuming --success rate varies Instead most labs send out for sequencing: --You prepare the DNA (usually plasmid, M 13, or PCR product), supply the primer, company or university sequencing center does the rest --The sequence is recorded by an automated sequencer as an “electropherogram”

Sequencing large pieces of DNA: the “shotgun” method • Break DNA into small pieces (typically sizes of around 1000 base pairs is preferable) • Clone pieces of DNA into M 13 • Sequence enough M 13 clones to ensure complete coverage (eg. sequencing a 3 million base pair genome would require 5 x to 10 x 3 million base pairs to have a reliable representation of the genome) • Assemble genome through overlap analysis using computer algorithms, also “polish” sequences using mapping information from individual clones, characterized genes, and genetic markers • This process is assisted by robotics

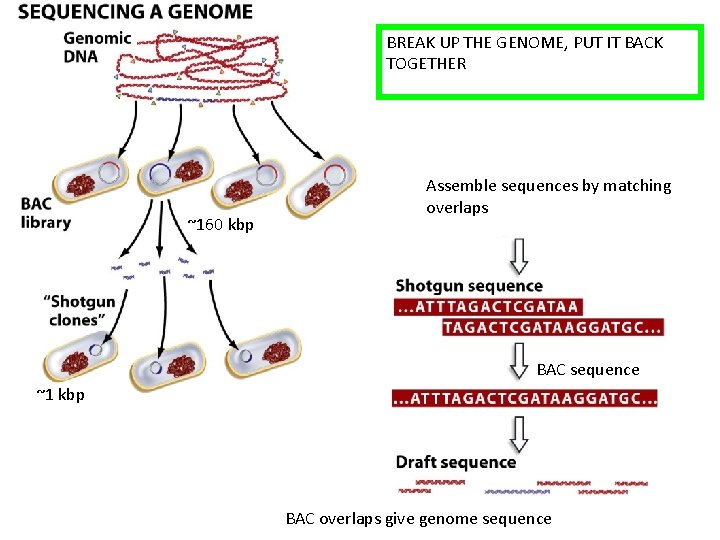
BREAK UP THE GENOME, PUT IT BACK TOGETHER ~160 kbp Assemble sequences by matching overlaps BAC sequence ~1 kbp BAC overlaps give genome sequence

Sequence by DNA polymerase ‐dependent chain extension, one base at a time in the presence of a reporter (luciferase) Luciferase is an enzyme that will emit a photon of light in response to the pyrophosphate (PPi) released upon nucleotide addition by DNA polymerase Flashes of light and their intensity are recorded

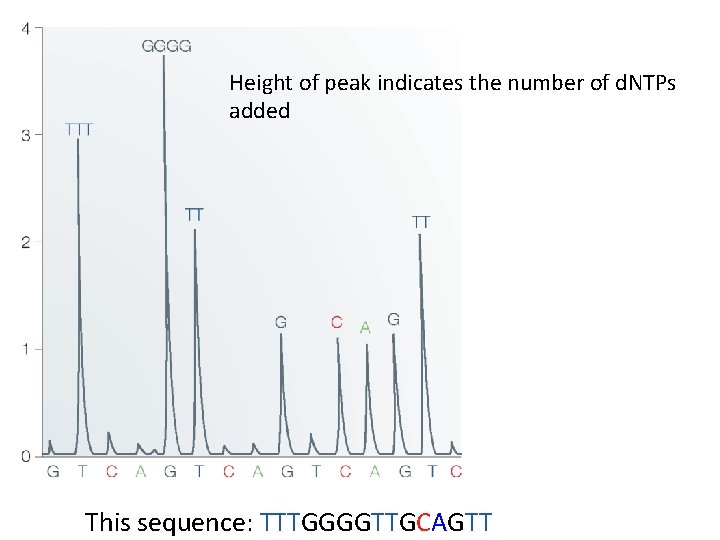
Height of peak indicates the number of d. NTPs added This sequence: TTTGGGGTTGCAGTT

Cycle Sequencing • Cycle sequencing is chain termination sequencing performed in a thermal cycler. • Cycle sequencing requires a heat‐stable DNA polymerase.

Fluorescent Dyes • Fluorescent dyes are multicyclic molecules that absorb and emit fluorescent light at specific wavelengths. • Examples are fluorescein and rhodamine derivatives. • For sequencing applications, these molecules can be covalently attached to nucleotides.

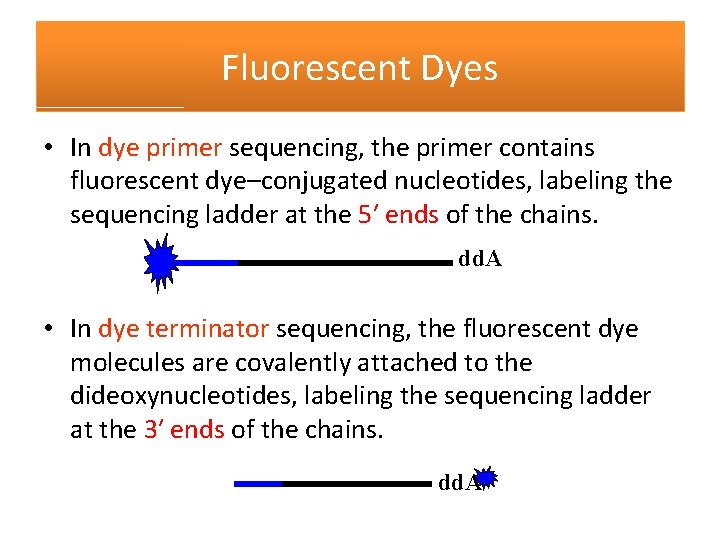
Fluorescent Dyes • In dye primer sequencing, the primer contains fluorescent dye–conjugated nucleotides, labeling the sequencing ladder at the 5′ ends of the chains. dd. A • In dye terminator sequencing, the fluorescent dye molecules are covalently attached to the dideoxynucleotides, labeling the sequencing ladder at the 3′ ends of the chains. dd. A

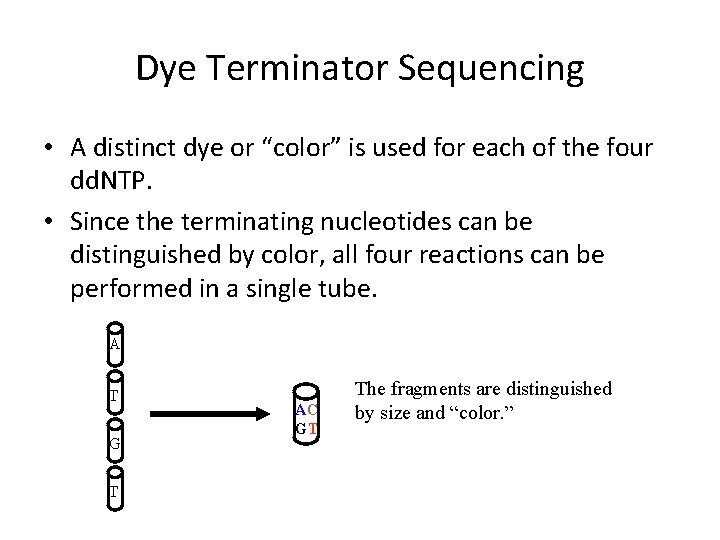
Dye Terminator Sequencing • A distinct dye or “color” is used for each of the four dd. NTP. • Since the terminating nucleotides can be distinguished by color, all four reactions can be performed in a single tube. A T G T AC GT The fragments are distinguished by size and “color. ”

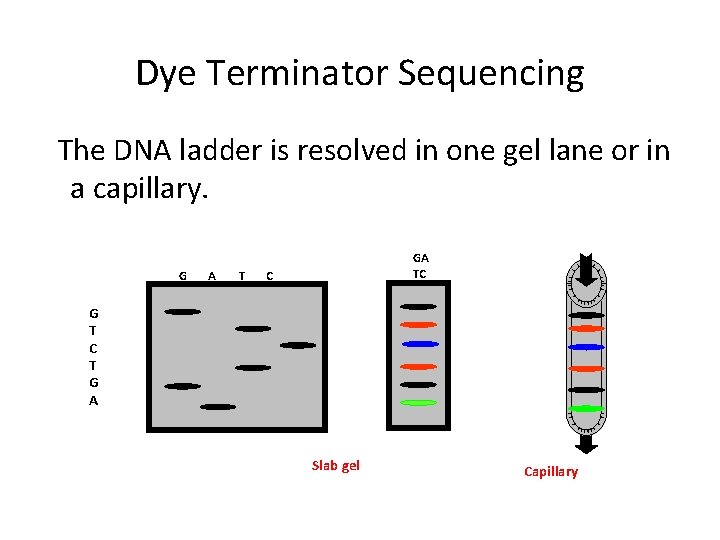
Dye Terminator Sequencing The DNA ladder is resolved in one gel lane or in a capillary. G A T GA TC C G T C T G A Slab gel Capillary

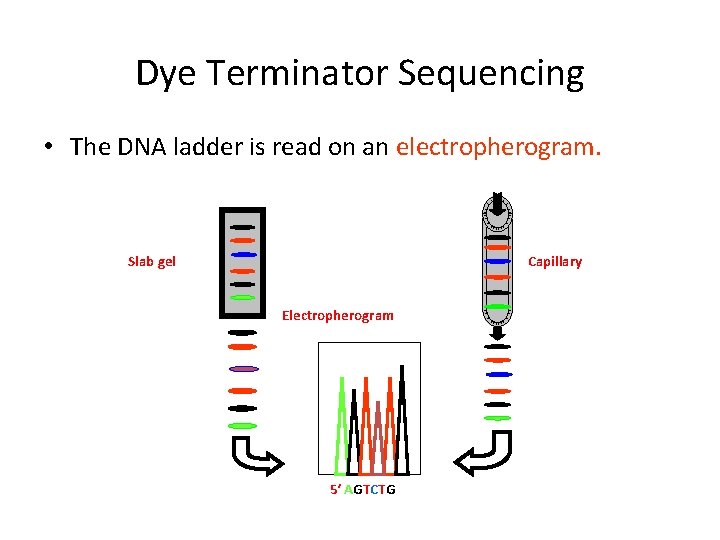
Dye Terminator Sequencing • The DNA ladder is read on an electropherogram. Slab gel Capillary Electropherogram 5′ AGTCTG

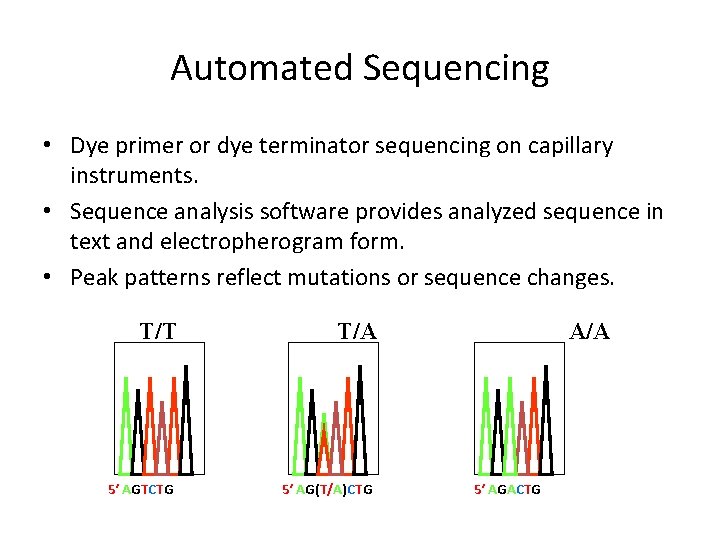
Automated Sequencing • Dye primer or dye terminator sequencing on capillary instruments. • Sequence analysis software provides analyzed sequence in text and electropherogram form. • Peak patterns reflect mutations or sequence changes. T/T 5′ AGTCTG T/A 5′ AG(T/A)CTG A/A 5′ AGACTG

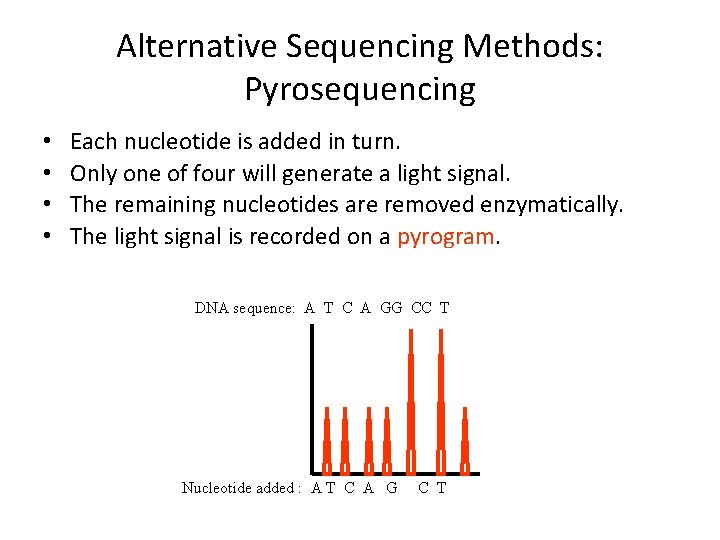
Alternative Sequencing Methods: Pyrosequencing • Pyrosequencing is based on the generation of light signal through release of pyrophosphate (PPi) on nucleotide addition. – DNAn + d. NTP DNAn+1 + PPi • PPi is used to generate ATP from adenosine phosulfate (APS). – APS + PPi ATP • ATP and luciferase generate light by conversion of luciferin to oxyluciferin.

Alternative Sequencing Methods: Pyrosequencing • • Each nucleotide is added in turn. Only one of four will generate a light signal. The remaining nucleotides are removed enzymatically. The light signal is recorded on a pyrogram. DNA sequence: A T C A GG CC T Nucleotide added : A T C A G C T

Alternative Sequencing Methods: Bisulfite Sequencing • Bisulfite sequencing is used to detect methylation in DNA. • Bisulfite deaminates cytosine, making uracil. • Methylated cytosine is not changed by bisulfite treatment. • The bisulfite‐treated template is then sequenced.

Bisulfite Sequencing The sequence of treated and untreated templates is compared. Methylated sequence: GTC Me GGC Me GATCTATC Me GTGCA … Treated sequence: GTC Me GGC Me GATUTATC Me GTGUA … DNA Sequence: (Untreated) reference: . . . GTCGGCGATCTATCGTGCA… Treated sequence: . . . GTCGGCGATUTATCGTGUA… This sequence indicates that these Cs are methylated.

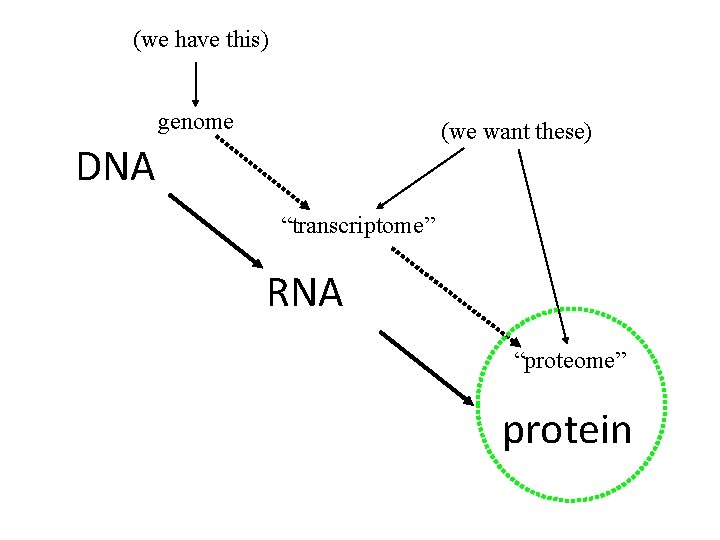
(we have this) genome (we want these) DNA “transcriptome” RNA “proteome” protein

DNA microarray -- immobilize many probes (thousands) in an ordered array, hybridize (base pair) with labelled m. RNA or c. DNA • • Generating an array of probes Identify open reading frames (orfs) 1) PCR each orf (several for each orf), attach (spot) each PCR product to a solid support in a specific order (pioneered by Pat Brown’s lab, Stanford) 2) Chemically synthesize orf-specific oligonucleotide probes directly on microchip (Affymetrix)

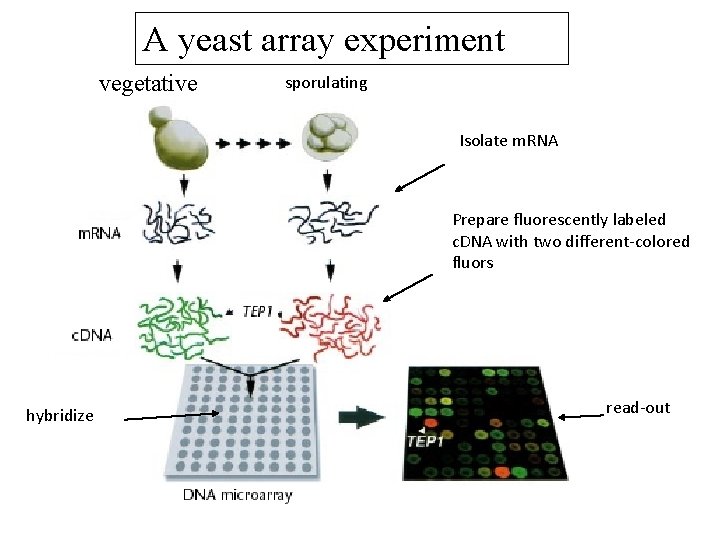
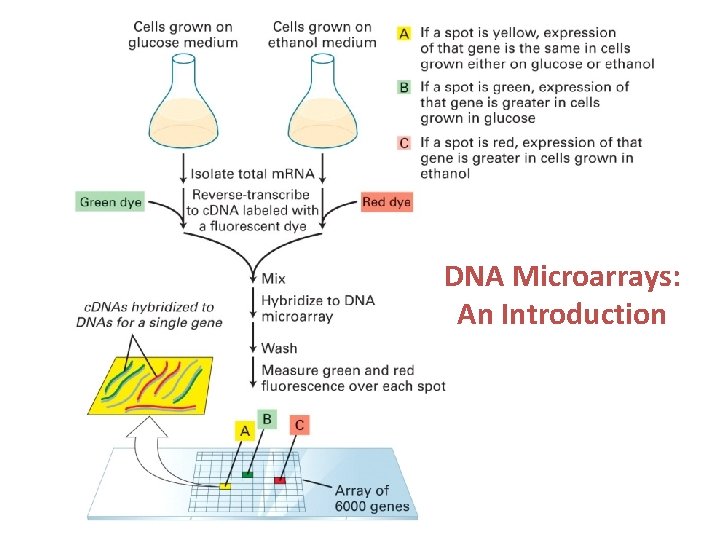
A yeast array experiment vegetative sporulating Isolate m. RNA Prepare fluorescently labeled c. DNA with two different‐colored fluors hybridize read‐out

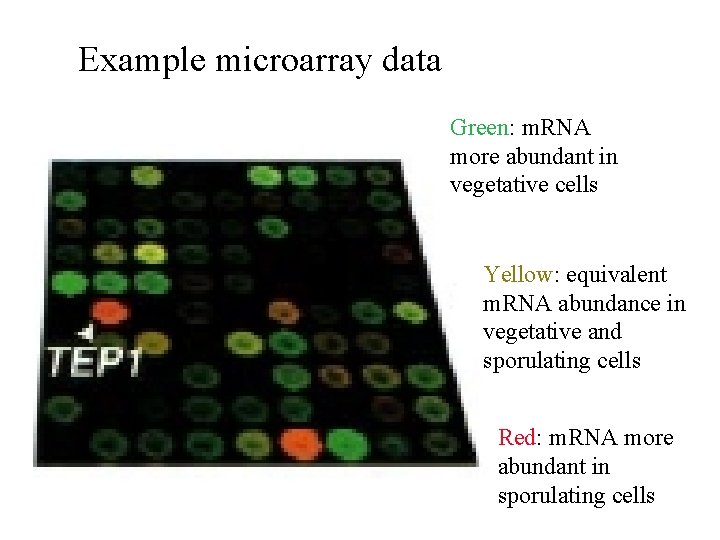
Example microarray data Green: m. RNA more abundant in vegetative cells Yellow: equivalent m. RNA abundance in vegetative and sporulating cells Red: m. RNA more abundant in sporulating cells

DNA Microarrays: An Introduction

Microarray Result: Much analysis to follow

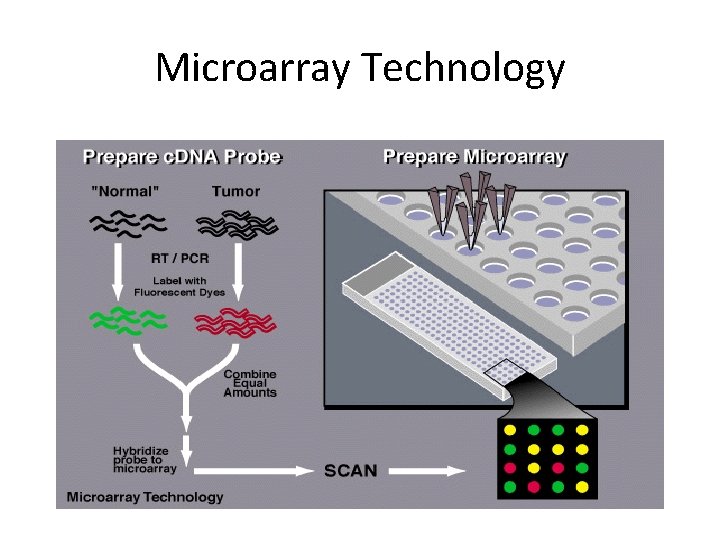
Microarray Technology

The value of DNA microarrays for studying gene expression 1) Study all transcripts at same time 1) Transcript abundance usually correlates with level of gene expression--much gene control is at level of transcription 2) Changes in transcription patterns often occur as a response to changing environment--this can be detected with a microarray

Summary • Genetic information is stored in the order or sequence of nucleotides in DNA. • Chain termination sequencing is the standard method for the determination of nucleotide sequence. • Dideoxy‐chain termination sequencing has been facilitated by the development of cycle sequencing and the use of fluorescent dye detection. • Alternative methods are used for special applications, such as pyrosequencing (for resequencing and polymorphism detection) or bisulfite sequencing (to analyze methylated DNA).

END Part I