The OBO Foundry Chris Mungall Lawrence Berkeley Laboratory

The OBO Foundry Chris Mungall Lawrence Berkeley Laboratory NCBO GO Consortium May 2007

The Open Biomedical Ontologies (OBO) Foundry § A collection of orthogonal reference ontologies in the biological/biomedical domain § Each is committed to an agreed upon set of principles governing best practices in ontology development

Outline § § § Motivation History/Background Organisation and dependencies Foundry Principles Results

http: //obofoundry. org http: //www. bioontologies. org (NCBO)

Why is the OBO Foundry necessary? § For the sharing, integration and analysis of biological and biomedical data § Common standards are required § Ontologies must be interoperable and logically well-formed § Ontologies should be developed collaboratively

Origins of OBO: The Gene Ontology (GO) § 3 ontologies intended primarily for the annotation of genes and gene products across a spectrum of organisms § Molecular function § Biological process § Cellular component § These ontologies are organised as a collection of related terms, constituting nodes in a graph

Annotation and GO § 187, 000 genes and gene products have high quality annotations to GO terms § 2. 6 m including automated predictions § 63, 000 publications curated § Variety of analysis tools § http: //www. geneontology. org/GO. tools. shtml#micro § Annotation of primary and literature data is one use of OBO Foundry ontologies

GO and the need for OBO § GO terms implicitly reference kinds of entities outwith the scope of GO § § Cysteine biosynthesis Neural crest cell migration Cardiac muscle morphogenesis Regulation of vascular permeability Ch. EBI Cell Anatomy quality § OBO was born from the need to create cross products wth GO § Also coincided with growth in model organism anatomy ontologies

Organisation of the OBO Foundry § Ontologies should be orthogonal § Minimise overlap § Each distinct entity type (universal) should only be represented once § We can partition the OBO Foundry rationally to help organise and coordinate the ontologies

Partitions § Type of entity § Relationship to time § Continuant § Occurrent § Dependent or independent § Granularity § § Molecular Cellular Organismal Multi-organismal § Generality § Upper domain ontology § Core biology § Species specific § Occurrence § § Canonical Variant Pathological Experimental


Connecting the Foundry: The OBO Relation Ontology § Standardized set of formally defined relations between types and/or instances § § is_a part_of has_participant … § For use within and across OBO ontologies § http: //obofoundry. org/ro § Molecules and cells participate in cellular processes § Cellular components are parts of cells which are parts of larger anatomical entities

OBO Foundry Principles § Open § Well-defined exchange format E. g. OBO or OWL § § § Unique ID-Space Ontology Life-cycle / versioning Clearly specified and delineated content Definitions Use relations according to the standards of the OBO Relation Ontology § Well documented § Plurality of users § Collaborative development http: //obofoundry. org/crit. shtml

Results § Phenotype Annotation § Ontology for Biomedical Investigations (OBI) § GO cross-products § Anatomy Ontologies § Semantic Web Health Care and Life Sciences (HCLS) interest group

Genotype-Phenotype Annotation § NCBO Driving Biological Project § Deep genotype-phenotype association curation of disease genes and genotypes § Human, Fruitfly, Zebrafish § Methodology: Flexible post-coordination of phenotype descriptions using Foundry ontologies § Based on ‘PATO’ ontology of qualities § E. g. § Shortened length of dendrite of columnar neuron

OBI: Ontology for Biomedical Investigations § An integrated ontology for experiments and investigations § Reuses terms from OBO Foundry ontologies in a modular way § Classes representing experimental artefacts, roles, hypotheses, variables etc § Adherence to upper ontology (BFO)

Results: GO cross-products § Ongoing work: § Processes and functions with chemical entities as participants § E. g. cysteine biosynthesis § Processes defined in terms of types of cell § E. g. neural crest cell migration § Mutual feedback

Anatomy Ontologies § Common Anatomy Reference Ontology § Ontologies of gross anatomy have been developed using divergent methodologies § CARO was developed after an NCBO sponsored meeting on anatomy ontologies § Ontology based on structure of the FMA § Common framework and upper-level terms for taxon-specific anatomical ontologies § Cell ontology § Merge of EVOC and initial OBO Cell ontology

Finding out more and participating § http: //obofoundry. org § http: //www. bioontology. org § obo-discuss@lists. sourceforge. net

Acknowledgements NCBO/Berkeley NCBO/Cambridge Ontologies Nicole Washington Michael Ashburner Amelia Ireland David Sutherland Mark Gibson George Gkoutos Jane Lomax Oliver Hofmann Pascale Gaudet Jen Clark Sue Rhee Paula de Matos Midori Harris Johnathan Bard Rafael Alcantra David Hill Lindsay Cowell Kirill Degtyarenko John Day-Richter Suzanna Lewis NCBO/Stanford Nigam Shah Daniel Rubin Archana Verbakam Lynn Murphy NCBO/Eugene Melissa Haendel Monte Westerfield NCBO/Victoria Chris Callender Margaret-Anne Storey Michael J Montague NCBO/Buffalo Mark Musen Fabian Neuhaus NCBO/Mayo Werner Ceusters James Buntrock Chris Chute Karen Eilbeck Erik Segerdell Louis Goldberg Barry Smith Rex Chisholm Pankaj Jaiswal Seth Carbon Alan Rector Onard Mejino Judith Blake Cynthia Smith Cornelius Rosse & GO Jannan Eppig William Bug Alan Ruttenberg NCBO/UCSF NIH Simona Carini Peter Good Jennifer Fostel Ida Sim Carol Bean & OBI Consortium Nation Heart, Lung and Blood Institute Trish Whetzel
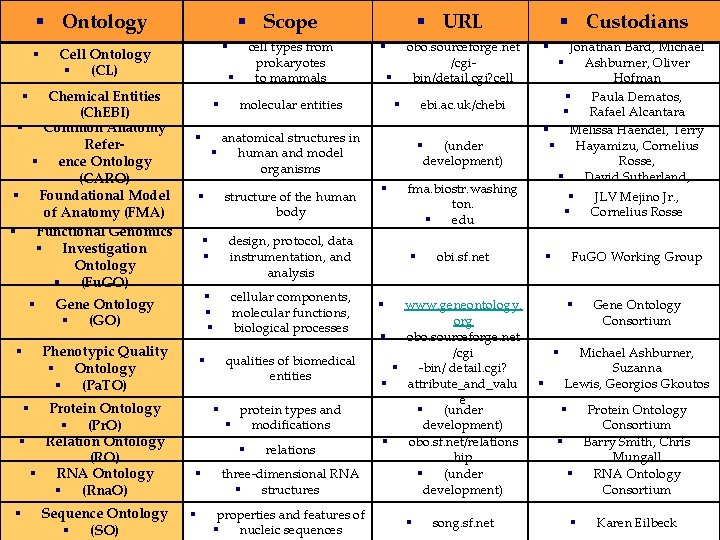
§ Ontology § § Scope Chemical Entities (Ch. EBI) § Common Anatomy Refer§ ence Ontology (CARO) § Foundational Model of Anatomy (FMA) § Functional Genomics § Investigation Ontology § (Fu. GO) § § anatomical structures in § human and model organisms § structure of the human body § § design, protocol, data instrumentation, and analysis § § § cellular components, molecular functions, biological processes Gene Ontology § (GO) Protein Ontology § (Pr. O) § Relation Ontology (RO) § RNA Ontology § (Rna. O) § § Sequence Ontology § (SO) qualities of biomedical entities § § protein types and § modifications § § § relations three-dimensional RNA § structures properties and features of § nucleic sequences obo. sourceforge. net /cgibin/detail. cgi? cell § § molecular entities § Phenotypic Quality § Ontology § (Pa. TO) § cell types from prokaryotes to mammals § Cell Ontology § (CL) § URL ebi. ac. uk/chebi § § § (under development) fma. biostr. washing ton. § edu § § Custodians Jonathan Bard, Michael § Ashburner, Oliver Hofman § Paula Dematos, § Rafael Alcantara § Melissa Haendel, Terry § Hayamizu, Cornelius Rosse, § David Sutherland, § § § obi. sf. net www. geneontology. org § obo. sourceforge. net /cgi § -bin/ detail. cgi? § attribute_and_valu e § (under development) § obo. sf. net/relations hip § (under development) Fu. GO Working Group § § § song. sf. net § § § JLV Mejino Jr. , Cornelius Rosse Gene Ontology Consortium Michael Ashburner, Suzanna Lewis, Georgios Gkoutos Protein Ontology Consortium § Barry Smith, Chris Mungall § RNA Ontology Consortium § § Karen Eilbeck

- Slides: 22