The NF BRel family MBV 4230 The NFBRel

The NF- B/Rel family

MBV 4230 The NF-κB/Rel family n A family of signal-responsive transcription factors ¨ n rapid response som ikke requires proteinsyntese Involved in proinflammatory response: a first line of defense against infectious diseases and cellular stress ¨ Signal �Activated NF- B immune defence activated n n Immune response, inflammatory response, accute phase response NFk. B also a major anti-apoptopic factor aberrant activation of NF- B = one of the primary causes of a wide range of human diseases like in Inflammatory diseases, Rheumatoid arthritis, Asthma, Atherosclerosis, Alzheimer ¨ Persistent activated in many cancers - help keeping cancer cells alive ¨ n NFk. B also promoting growth ¨ n Activated NF- B cyclin D expression enhanced growth Drug against NFk. B = putative anti-cancer drug

MBV 4230 The NF- B/Rel family n Characteristic feature: homo- and heterodimeric TFs, which in non-stimulated cells are found inactive in the cytoplasm [in a complex with I B-repressors]. Active DNA-binding form: Dimers with different members of the NF- B/Rel family ¨ Inactive cytoplasmic form: inhibitory factor/domain in addition ¨ n Upon stimulation, active NF- B rapidly translocates to the nucleus where it binds B-sites and activates target genes. n Rapid response - minutes ¨ Signal Activated NF- B immune defence activated
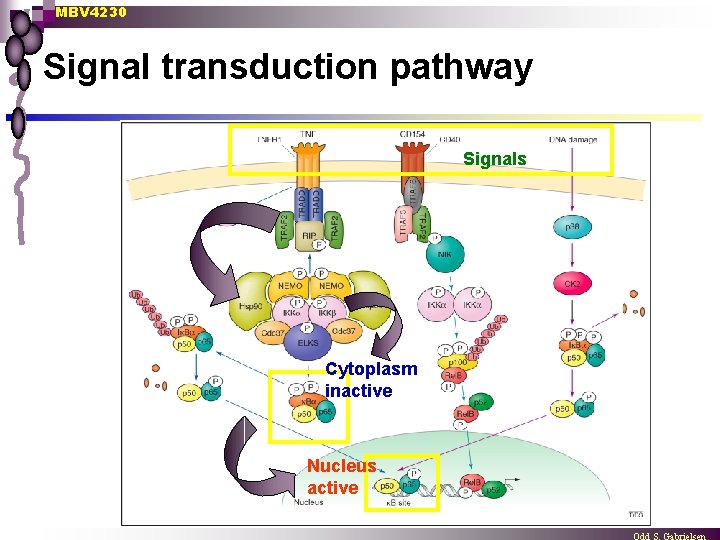
MBV 4230 Signal transduction pathway Signals Cytoplasm inactive Nucleus active

NF-κB/Rel proteins

MBV 4230 Common DBD: Rel-homology domain (RHD) n RHD: 300 aa conserved domain with several functions DNA-binding (N-terminal half) ¨ dimerization (C-terminal half) ¨ I B-interaction (C-terminal half) ¨ NLS (C-terminal half) ¨ ¨ kalles også NRD (=NF-k. B, Rel, Dorsal) Spec. DNA-binding dimerization Ik. B-interaction NLS
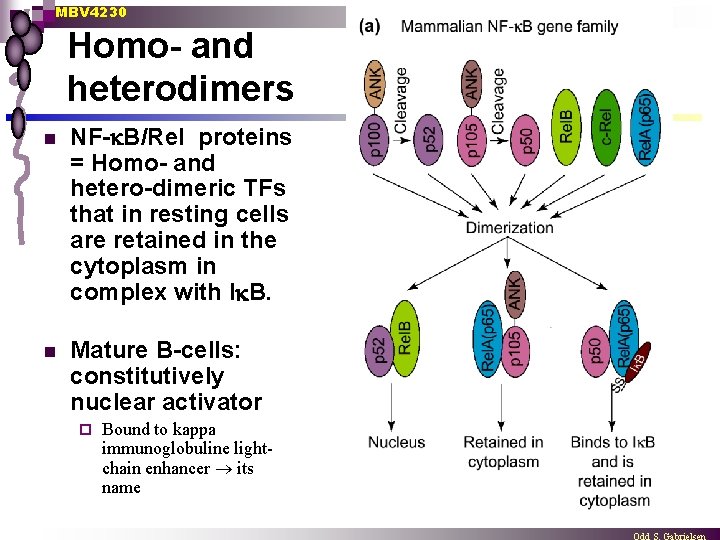
MBV 4230 Homo- and heterodimers n NF- B/Rel proteins = Homo- and hetero-dimeric TFs that in resting cells are retained in the cytoplasm in complex with I B. n Mature B-cells: constitutively nuclear activator ¨ Bound to kappa immunoglobuline lightchain enhancer its name

MBV 4230 Two main classes of RHDs n Rel with TAD (dimeric with ≥ 1 Rel-monomers which are potent transactivators) synthesized in their mature form Rel or c-Rel (as well as v-Rel) ¨ Rel. A (p 65) ¨ Rel. B ¨ Drosophilas dorsal and Dif ¨ n p 50/52 without TAD (homodimers with no transactivation properties) synthesized as precursors that are processed Precursor forms have internal I B inhibitor function n RHD linked to inhibitory domain through Gly-rich linker (protease sensitive) n Blocks DNA-binding and translocation to nucleus ¨ p 105 undergoes proteolytic maturation to p 50 [NF- B 1] n Proteolytic degradation to p 50 is signal dependent, requires ATP and occurs through a ubiquitin-dependent proteasome pathway n Also transcription from an intronic promoter expression of Ik. B- ¨ p 100 undergoes proteolytic maturation to p 52 [NF-�B 2] ¨ p 50/52 are distinct gene products with very similar properties ¨
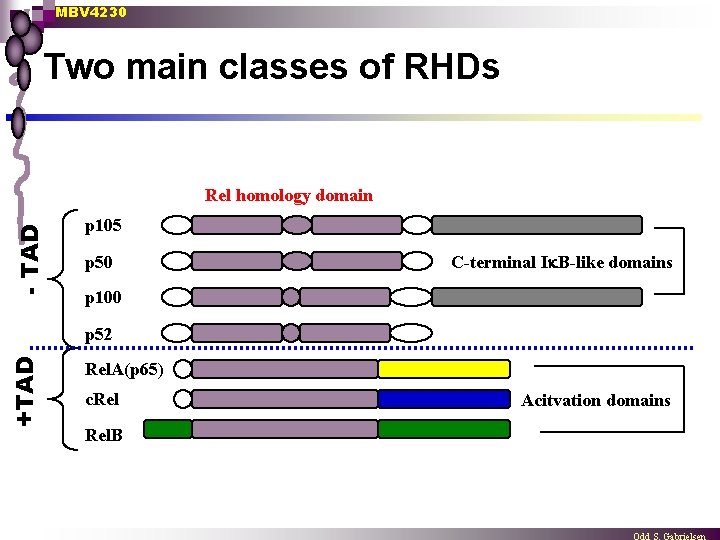
MBV 4230 Two main classes of RHDs - TAD Rel homology domain p 105 p 50 C-terminal I B-like domains p 100 +TAD p 52 Rel. A(p 65) c. Rel. B Acitvation domains
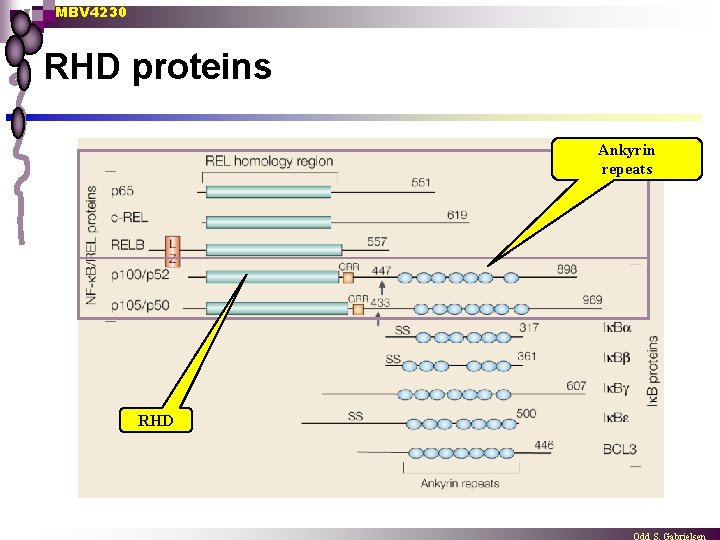
MBV 4230 RHD proteins Ankyrin repeats RHD
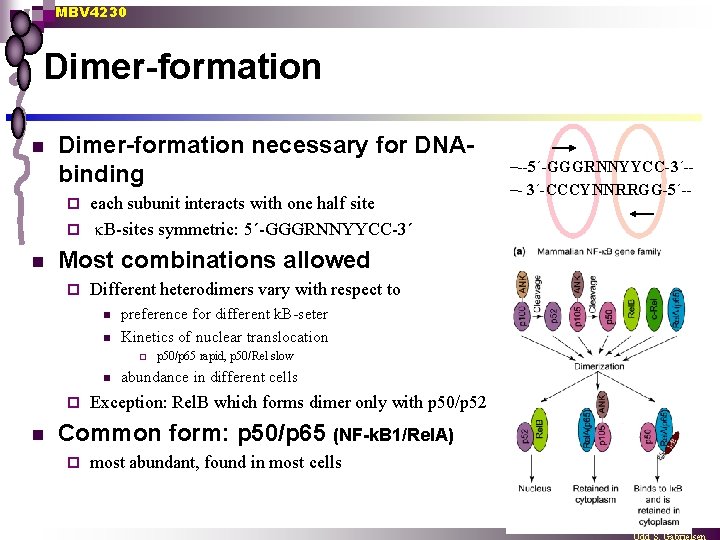
MBV 4230 Dimer-formation necessary for DNAbinding each subunit interacts with one half site ¨ B-sites symmetric: 5´-GGGRNNYYCC-3´ ¨ n Most combinations allowed ¨ Different heterodimers vary with respect to n n preference for different k. B-seter Kinetics of nuclear translocation ¨ n p 50/p 65 rapid, p 50/Rel slow abundance in different cells Exception: Rel. B which forms dimer only with p 50/p 52 Common form: p 50/p 65 (NF-k. B 1/Rel. A) ¨ most abundant, found in most cells –--5´-GGGRNNYYCC-3´-–- 3´-CCCYNNRRGG-5´--
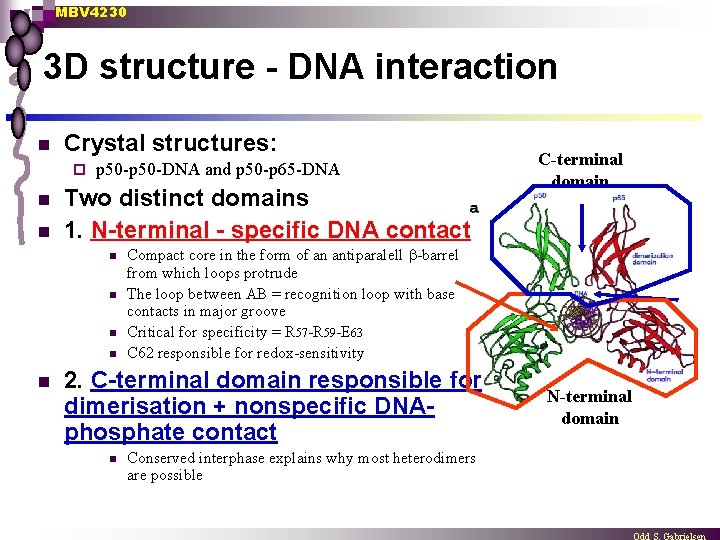
MBV 4230 3 D structure - DNA interaction n Crystal structures: ¨ n n p 50 -DNA and p 50 -p 65 -DNA Two distinct domains 1. N-terminal - specific DNA contact n n n Compact core in the form of an antiparalell -barrel from which loops protrude The loop between AB = recognition loop with base contacts in major groove Critical for specificity = R 57 -R 59 -E 63 C 62 responsible for redox-sensitivity 2. C-terminal domain responsible for dimerisation + nonspecific DNAphosphate contact n C-terminal domain Conserved interphase explains why most heterodimers are possible N-terminal domain
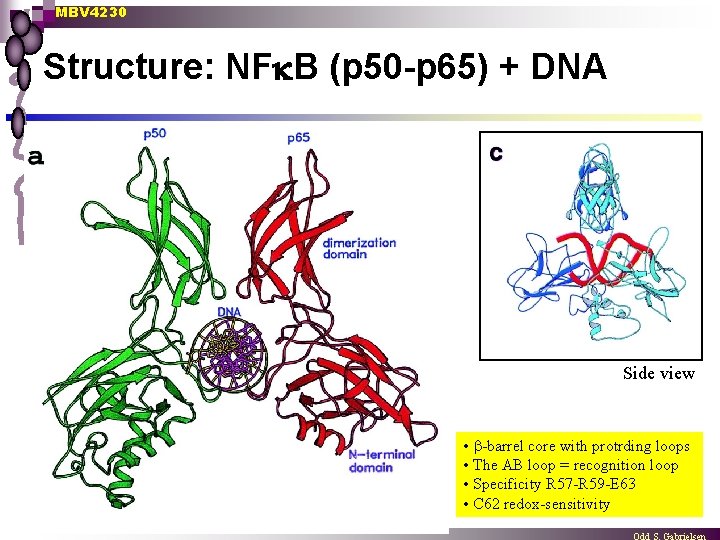
MBV 4230 Structure: NF B (p 50 -p 65) + DNA Side view • -barrel core with protrding loops • The AB loop = recognition loop • Specificity R 57 -R 59 -E 63 • C 62 redox-sensitivity
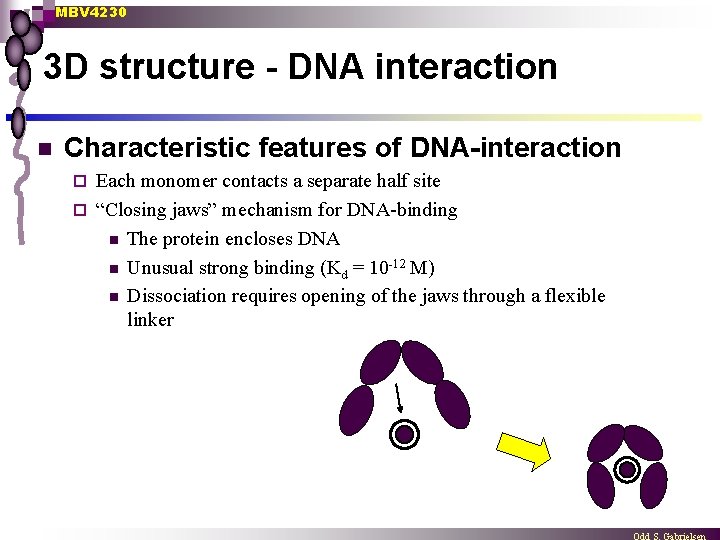
MBV 4230 3 D structure - DNA interaction n Characteristic features of DNA-interaction Each monomer contacts a separate half site ¨ “Closing jaws” mechanism for DNA-binding n The protein encloses DNA n Unusual strong binding (Kd = 10 -12 M) n Dissociation requires opening of the jaws through a flexible linker ¨
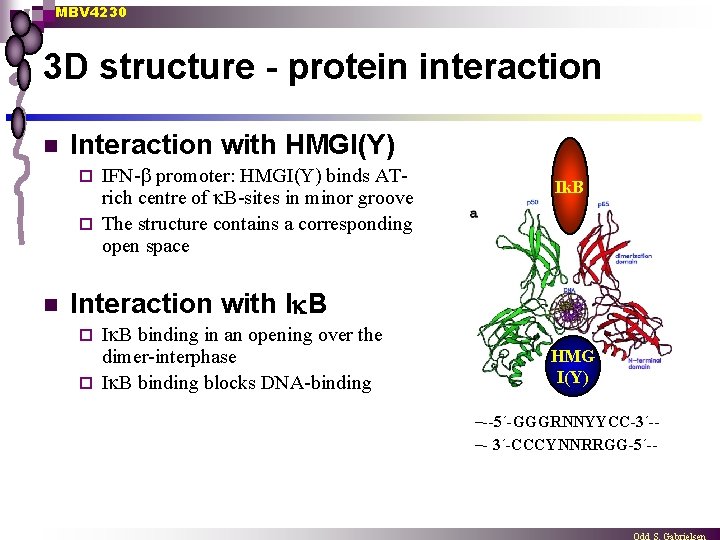
MBV 4230 3 D structure - protein interaction n Interaction with HMGI(Y) IFN- promoter: HMGI(Y) binds ATrich centre of B-sites in minor groove ¨ The structure contains a corresponding open space ¨ n Ik. B Interaction with I B binding in an opening over the dimer-interphase ¨ I B binding blocks DNA-binding ¨ HMG I(Y) –--5´-GGGRNNYYCC-3´-–- 3´-CCCYNNRRGG-5´--

The I- B family
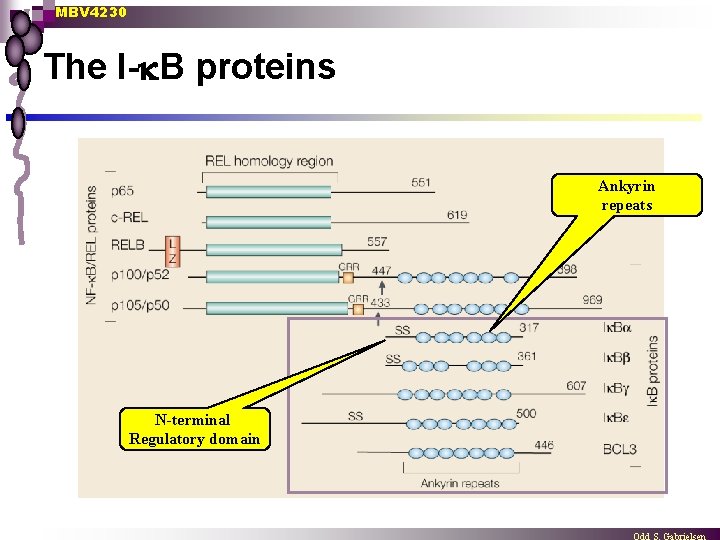
MBV 4230 The I- B proteins Ankyrin repeats N-terminal Regulatory domain

MBV 4230 The I B-family n Inhibitory function impedes DNA-binding ¨ blocks NLS and abolish translocation to nucleus ¨ n Several members (at least 7 mammalian) ¨ ¨ ¨ n I B- and I B- Bcl-3 p 105 and p 110 Ik. BR Common features: ¨ Specificity Ex. Ik. B- inhibits DNA-binding of p 65/p 50 but not of p 50/p 50 ankyrin-repeats which are necessary for RHD-interaction n 30 -33 aa motif repeated 3 - 7 x C-terminal acidic-region necessary for inhibition of DNA-binding ¨ C-terminal PEST-sequence involved in protein-degradation ¨
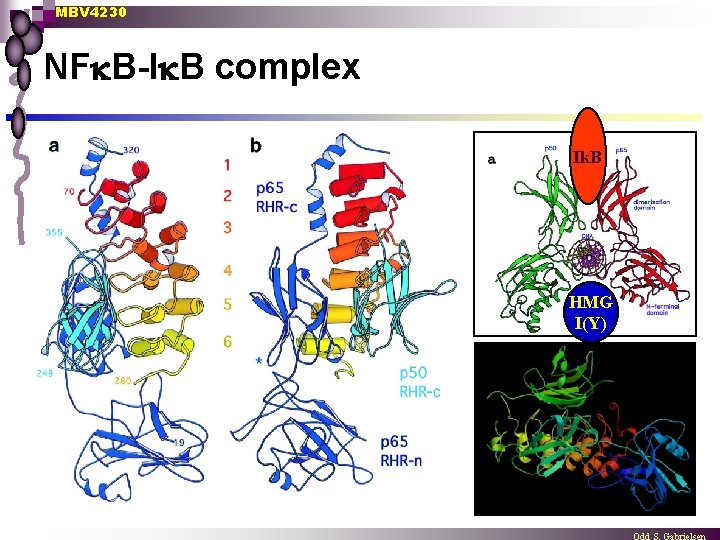
MBV 4230 NF B-I B complex Ik. B HMG I(Y)
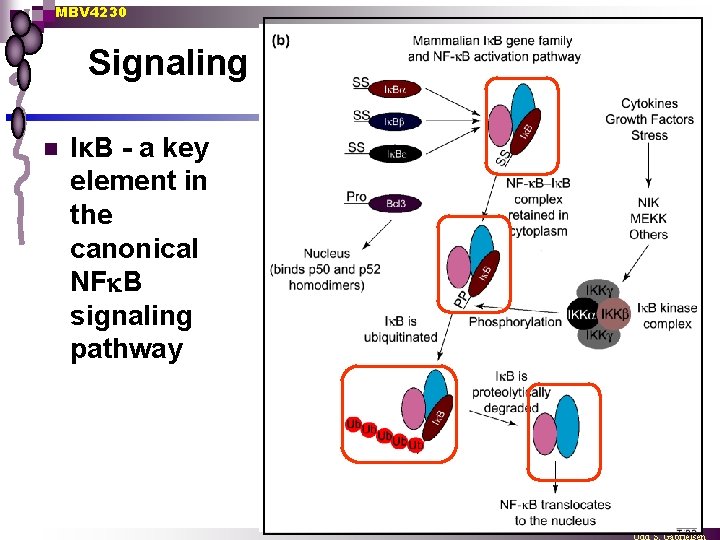
MBV 4230 Signaling n IκB - a key element in the canonical NF B signaling pathway
MBV 4230 Cytoplasmic retention due to interaction with I B-family proteins n Two types of inactive complexes in cytoplasm Signal Trimers = RHD-Homo-or heterodimers bound to an I B ¨ Heterodimers = Rel-protein + unprocessed RHD-precursor (p 105, p 110) ¨ n Signal→[dissociation] → degradation Induction signal phosphorylation of both I B and p 105 I B degradation or p 105 processering active dimers that are translocated to the nucleus. ¨ One type of signal two N-terminal serines (S 32 and S 36) become phosphorylated ¨ Another type of signal two C-terminal serines become phosphorylated in p 105 ¨ phosphorylation probably more a signal for degradation than for dissociation ¨ n Ubiquitin-pathway involved ¨ ¨ ¨ Stimulation rapid degradation of I B n complete after 10 min n No traces of I B phosphorylation of I B → multiubiquitylation in K 21, K 22 → degradation through a ubiquitin-proteasome pathway + proteasome-inhibitors → phospho-Ik. B remains associated with NFk. B Signal

MBV 4230 Several I B-factors with different properties n I B- : Rapid transient response I B- best characterized ¨ all stimuli degradation of I B- ¨ ex: TNF- rapid and transient activation of NF-k. B ¨ n I B- : Sustained response Only certain stimuli degradation of I B- ¨ ex: LPS or IL-1 degradation of both I B- and I B- activation of NF-k. B lasting for hours ¨ n Bcl-3: repressor and activator inhibits certain complexes like a normal I B ¨ But may also associate with DNA-bound p 50 and p 52 dimers (lacking TAD) and provide transactivation properties ¨

Signaling pathways
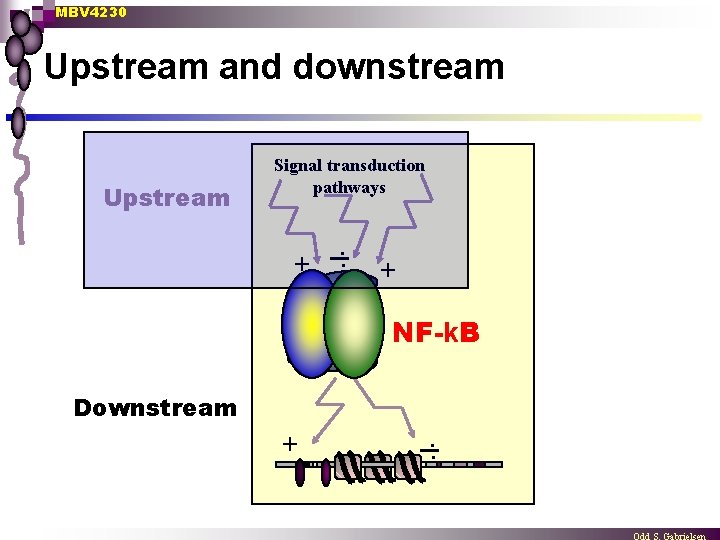
MBV 4230 Upstream and downstream Upstream Signal transduction pathways + . . + NF-k. B Downstream + . .
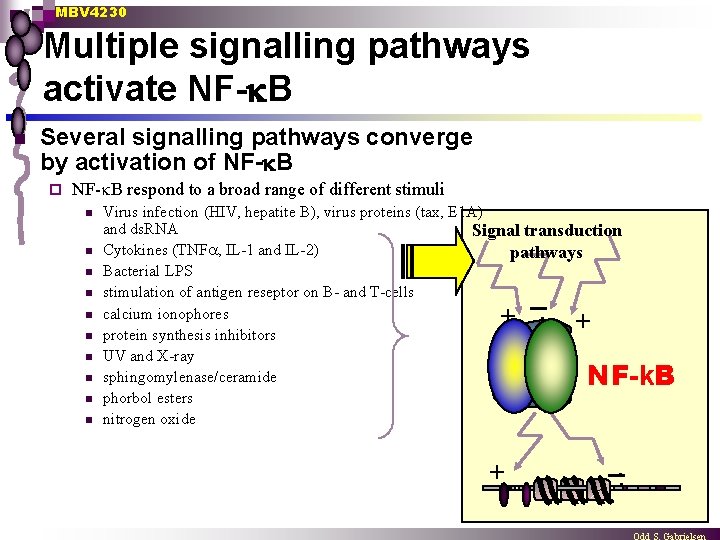
MBV 4230 Multiple signalling pathways activate NF- B n Several signalling pathways converge by activation of NF- B ¨ NF- B respond to a broad range of different stimuli n n n n n Virus infection (HIV, hepatite B), virus proteins (tax, E 1 A) and ds. RNA Signal transduction Cytokines (TNF , IL-1 and IL-2) pathways Bacterial LPS stimulation of antigen reseptor on B- and T-cells. . calcium ionophores + + protein synthesis inhibitors UV and X-ray sphingomylenase/ceramide phorbol esters nitrogen oxide NF-k. B + . .
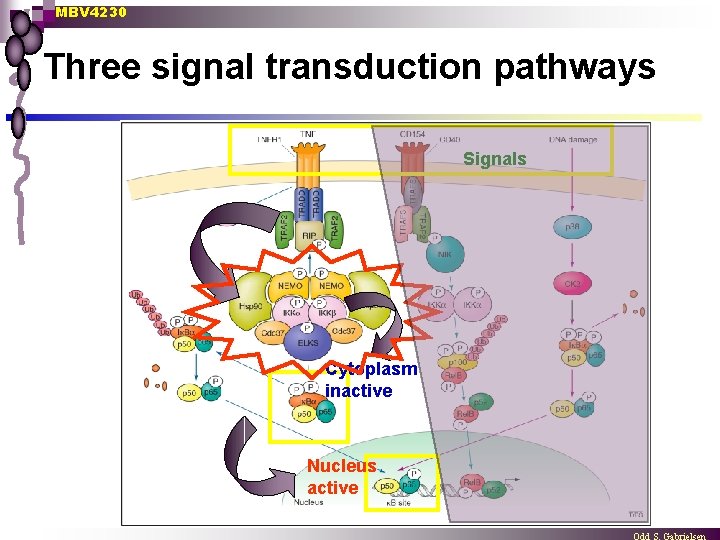
MBV 4230 Three signal transduction pathways Signals Cytoplasm inactive Nucleus active
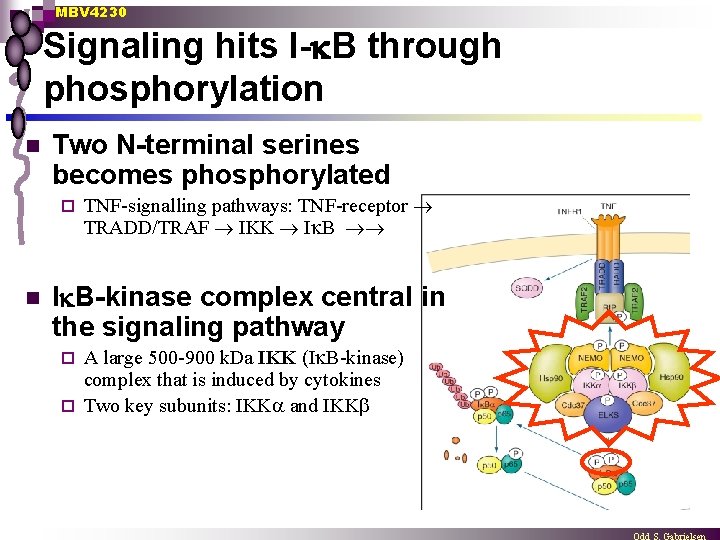
MBV 4230 Signaling hits I- B through phosphorylation n Two N-terminal serines becomes phosphorylated ¨ n TNF-signalling pathways: TNF-receptor TRADD/TRAF IKK I B-kinase complex central in the signaling pathway A large 500 -900 k. Da IKK (I B-kinase) complex that is induced by cytokines ¨ Two key subunits: IKK and IKK ¨
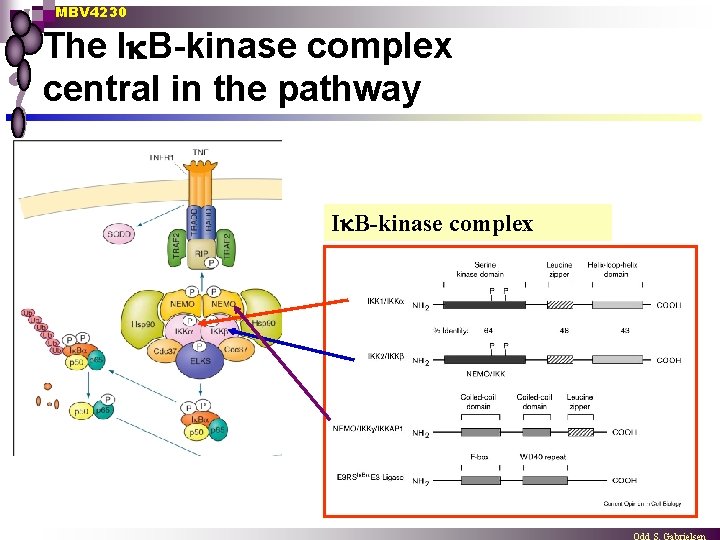
MBV 4230 The I B-kinase complex central in the pathway I B-kinase complex
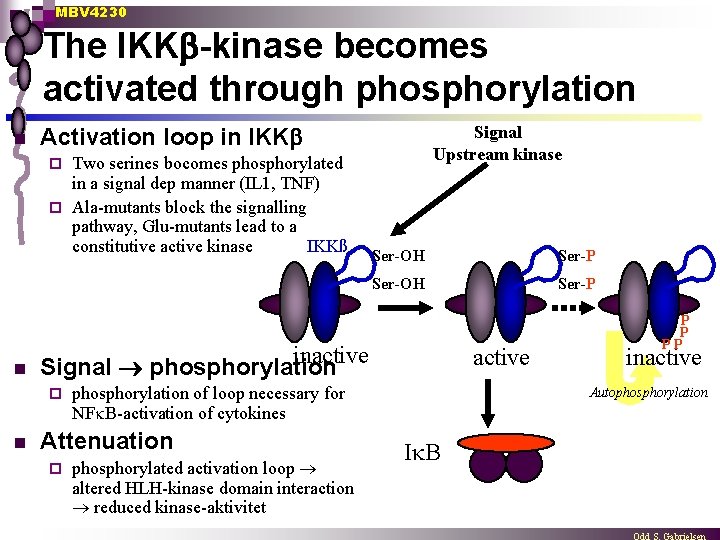
MBV 4230 The IKK -kinase becomes activated through phosphorylation n Signal Upstream kinase Activation loop in IKK Two serines bocomes phosphorylated in a signal dep manner (IL 1, TNF) ¨ Ala-mutants block the signalling pathway, Glu-mutants lead to a constitutive active kinase IKKß ¨ n Ser-P Ser-OH Ser-P inactive Signal phosphorylation ¨ n Ser-OH phosphorylation of loop necessary for NF B-activation of cytokines Attenuation ¨ active phosphorylated activation loop altered HLH-kinase domain interaction reduced kinase-aktivitet P P PP inactive Autophosphorylation I B

MBV 4230 The first pathway - the classical pathway n Receptor triggered by pro-inflammatory cytokines ¨ n Recruitment of various adaptors ¨ n n n including TRADD (TNF-receptor associated death domain protein), RIP (receptor interacting protein and TRAF 2 (TNF-receptor-associated factor 2) to the cytoplasmic membrane. Recruitment and activation of the classical IκB-kinase (IKK) complex ¨ n such as tumour necrosis factor (TNF)-α which includes the scaffold protein NEMO (NF-k. B essential modulator; also named IKKγ), IKKα and IKKβ kinases. The IKK complex phosphorylates IκBα on Ser 32 and Ser 36 Leading to ubiquitylation and degradation via the proteasome pathway The free p 50 -p 65 migrates to the nucleus where it activates target genes involved in immune response
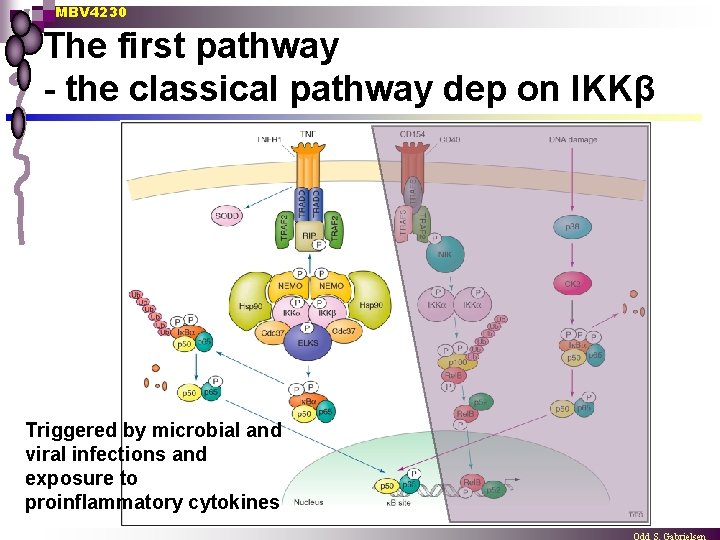
MBV 4230 The first pathway - the classical pathway dep on IKKβ Triggered by microbial and viral infections and exposure to proinflammatory cytokines
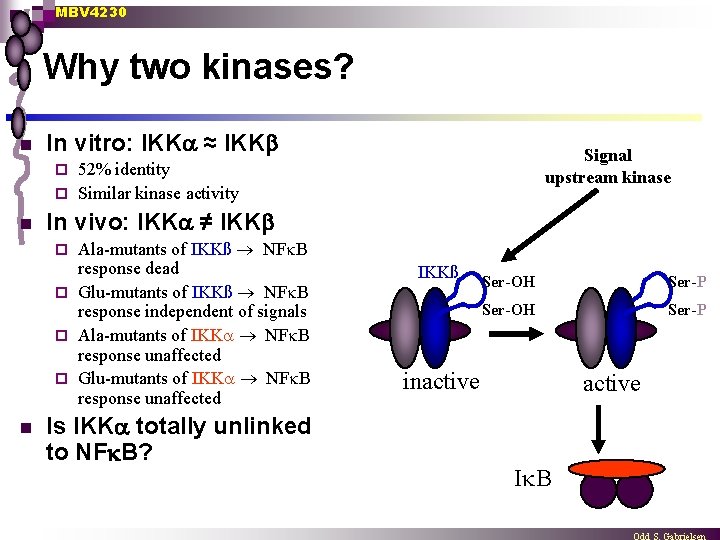
MBV 4230 Why two kinases? n In vitro: IKK ≈ IKK Signal upstream kinase 52% identity ¨ Similar kinase activity ¨ n In vivo: IKK ≠ IKK Ala-mutants of IKKß NF B response dead ¨ Glu-mutants of IKKß NF B response independent of signals ¨ Ala-mutants of IKK NF B response unaffected ¨ Glu-mutants of IKK NF B response unaffected ¨ n Is IKK totally unlinked to NF B? IKKß Ser-OH Ser-P inactive I B

MBV 4230 The next indication: KO phenotypes of IKK ≠ IKK n Knock-out of of IKK loss of B- and T-cell response Normal development ¨ Mice dead at day 13. 5, liver destroyed due to massive apoptosis ¨ Lack of IKK lack of active NFk. B lack of protection against apoptosis massive cell death ¨ Lost T-cell response because Apoptosis important for T-cell development ¨ n Knock-out of of IKK ������� , epidermis 5 -10 x thicker than normal, highly undifferentiated ¨ ������ s������� l ¨ Normal number of B- and T-cells, but B-cells not fully differentiated ¨

MBV 4230 A separate signaling pathway through IKK n n A desparate postdoc looked at all the 50 components - all behaved normal, except one The proteolytic maturation of the p 100 precursor to p 52 [NF- B 2] was defective in the IKK processing depends on NIK Hypothesis: NIK acts through IKK
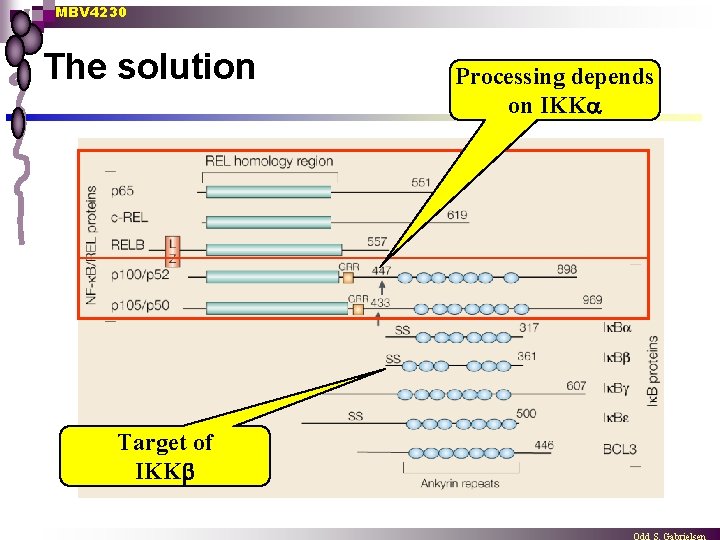
MBV 4230 The solution Target of IKK Processing depends on IKK
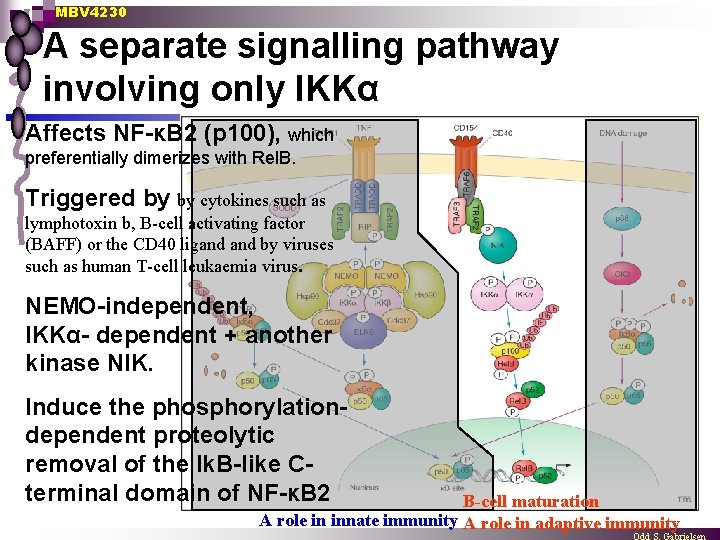
MBV 4230 A separate signalling pathway involving only IKKα Affects NF-κB 2 (p 100), which preferentially dimerizes with Rel. B. Triggered by by cytokines such as lymphotoxin b, B-cell activating factor (BAFF) or the CD 40 ligand by viruses such as human T-cell leukaemia virus. NEMO-independent, IKKα- dependent + another kinase NIK. Induce the phosphorylationdependent proteolytic removal of the Ik. B-like Cterminal domain of NF-κB 2 B-cell maturation A role in innate immunity A role in adaptive immunity
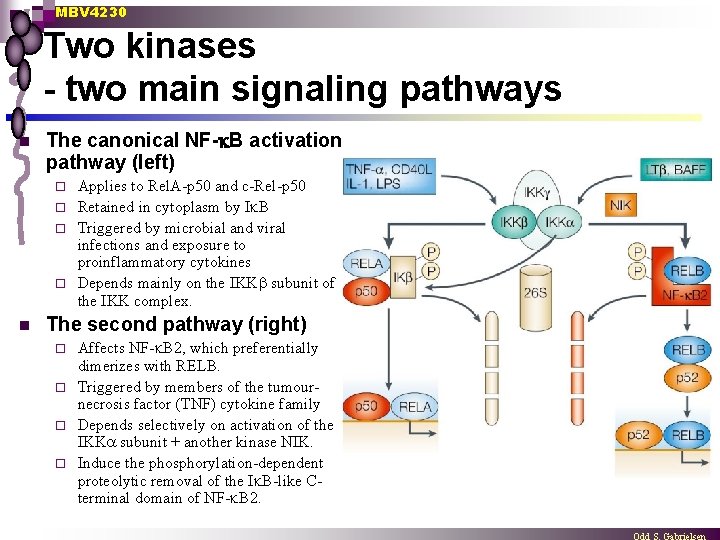
MBV 4230 Two kinases - two main signaling pathways n The canonical NF- B activation pathway (left) Applies to Rel. A-p 50 and c-Rel-p 50 ¨ Retained in cytoplasm by I B ¨ Triggered by microbial and viral infections and exposure to proinflammatory cytokines ¨ Depends mainly on the IKK subunit of the IKK complex. ¨ n The second pathway (right) Affects NF- B 2, which preferentially dimerizes with RELB. ¨ Triggered by members of the tumournecrosis factor (TNF) cytokine family ¨ Depends selectively on activation of the IKK subunit + another kinase NIK. ¨ Induce the phosphorylation-dependent proteolytic removal of the I B-like Cterminal domain of NF- B 2. ¨
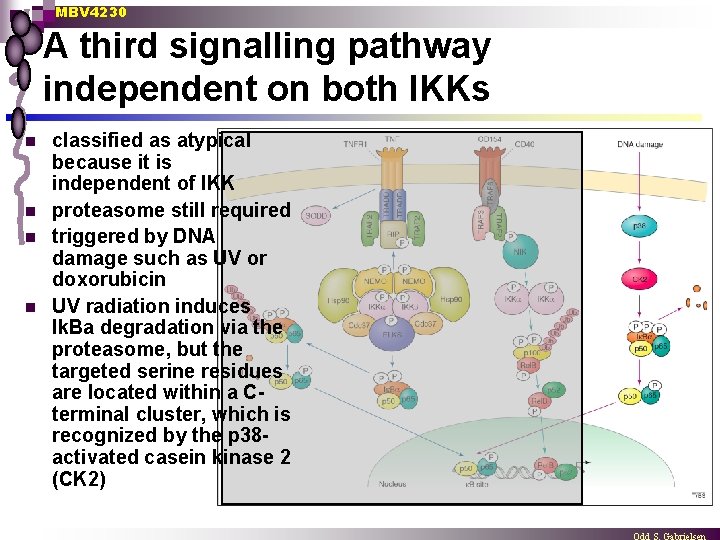
MBV 4230 A third signalling pathway independent on both IKKs n n classified as atypical because it is independent of IKK proteasome still required triggered by DNA damage such as UV or doxorubicin UV radiation induces Ik. Ba degradation via the proteasome, but the targeted serine residues are located within a Cterminal cluster, which is recognized by the p 38 activated casein kinase 2 (CK 2)
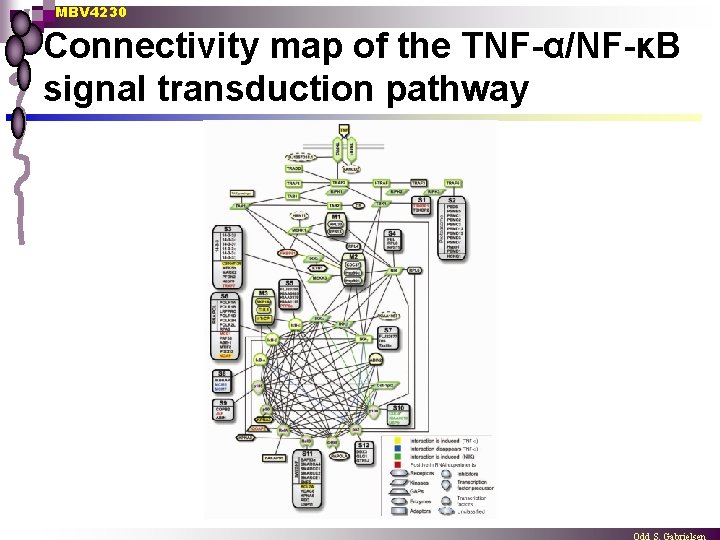
MBV 4230 Connectivity map of the TNF-α/NF-κB signal transduction pathway
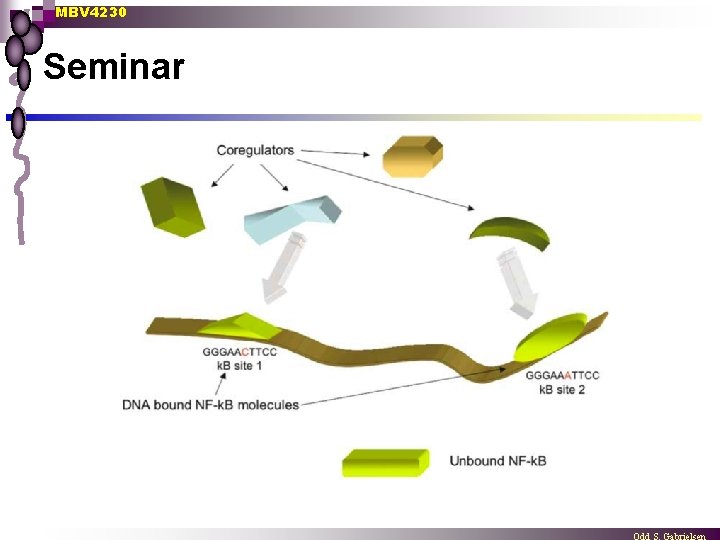
MBV 4230 Seminar

Target genes
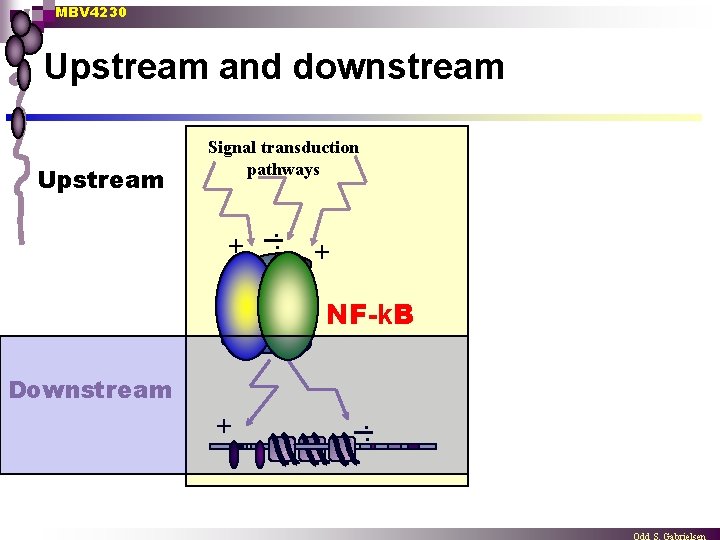
MBV 4230 Upstream and downstream Upstream Signal transduction pathways + . . + NF-k. B Downstream + . .
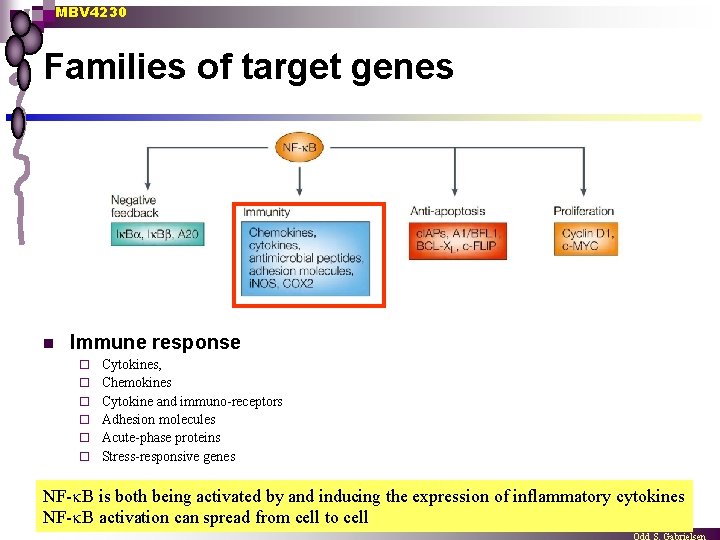
MBV 4230 Families of target genes n Immune response ¨ ¨ ¨ Cytokines, Chemokines Cytokine and immuno-receptors Adhesion molecules Acute-phase proteins Stress-responsive genes NF- B is both being activated by and inducing the expression of inflammatory cytokines NF- B activation can spread from cell to cell

MBV 4230 Negative feedback: Attenuation of respons n Negative loop: I B- under direct control of NF- B Activated NF- B translocated to the nucleus will activate expression of I B- ¨ Newly synthesized I B- will bind up and inactivate remaining NF- B in the cytoplasma ¨ Excess I B- will migrate to the nucleus and inactivate DNA-bound NF- B (contains both NLS and nuclear eksport signal) ¨ A 20 protein another strongly induced negative feedback protein ¨ n Immunosupressive effect of glucocorticoids ¨ Probably a direct effect of glucocorticoids enhancing the expression of I B which then binds up and inactivates NF- B in the cytoplasm, leading to reduced immune- and inflammatory response
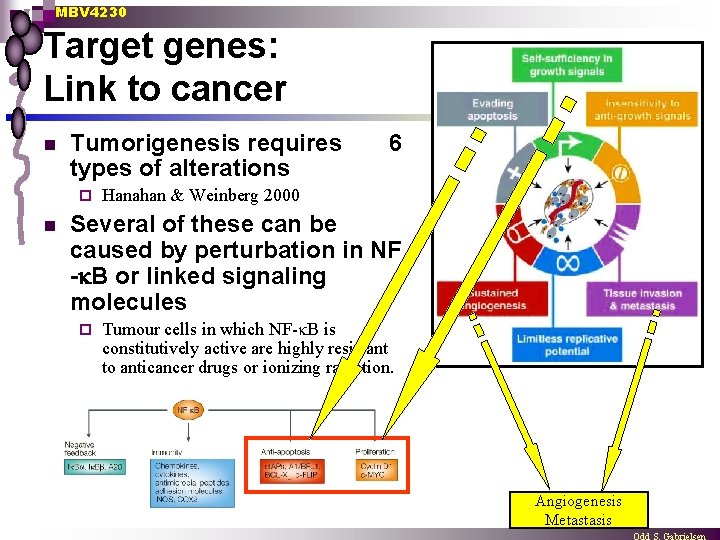
MBV 4230 Target genes: Link to cancer n Tumorigenesis requires types of alterations ¨ n 6 Hanahan & Weinberg 2000 Several of these can be caused by perturbation in NF - B or linked signaling molecules ¨ Tumour cells in which NF- B is constitutively active are highly resistant to anticancer drugs or ionizing radiation. Angiogenesis Metastasis

Disease links

MBV 4230 Viruses exploit NF- B n several patogenic viruses exploit the NF- B system for their own profit Incorporation of B-sites in virus DNA cause enhanced expression of virus-genes when the immune response is activated ¨ Virus proteins activate NF- B ¨

MBV 4230 Disease links
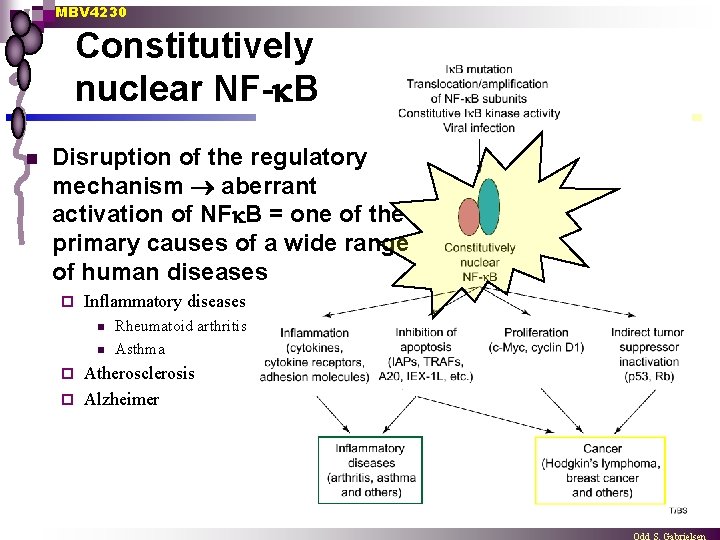
MBV 4230 Constitutively nuclear NF- B n Disruption of the regulatory mechanism aberrant activation of NF B = one of the primary causes of a wide range of human diseases ¨ Inflammatory diseases n n Rheumatoid arthritis Asthma Atherosclerosis ¨ Alzheimer ¨

MBV 4230 Link: inflammation - cancer n A causal connection between inflammation and cancer has been suspected for many years. n NF- B might serve as the missing link between these two processes. NF- B becomes activated in response to inflammatory stimuli ¨ Constitutive activation of NF- B has been associated with cancer, ¨
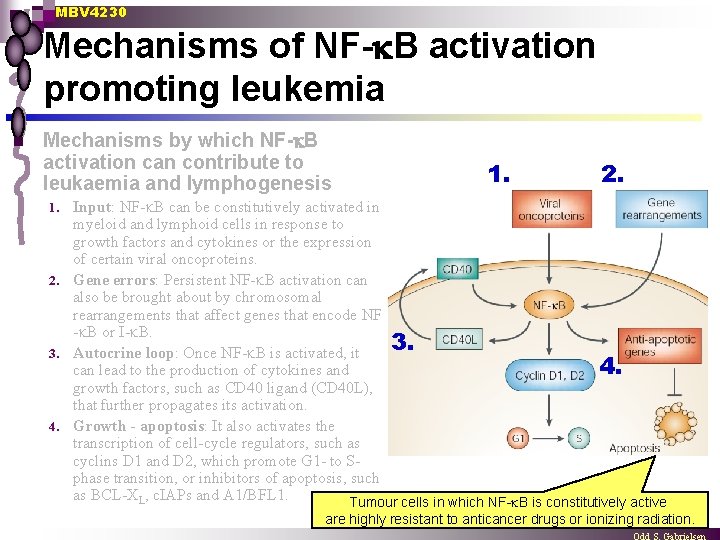
MBV 4230 Mechanisms of NF- B activation promoting leukemia n Mechanisms by which NF- B activation can contribute to leukaemia and lymphogenesis 1. 2. Input: NF- B can be constitutively activated in myeloid and lymphoid cells in response to growth factors and cytokines or the expression of certain viral oncoproteins. 2. Gene errors: Persistent NF- B activation can also be brought about by chromosomal rearrangements that affect genes that encode NF - B or I- B. 3. 3. Autocrine loop: Once NF- B is activated, it 4. can lead to the production of cytokines and growth factors, such as CD 40 ligand (CD 40 L), that further propagates its activation. 4. Growth - apoptosis: It also activates the transcription of cell-cycle regulators, such as cyclins D 1 and D 2, which promote G 1 - to Sphase transition, or inhibitors of apoptosis, such as BCL-XL, c. IAPs and A 1/BFL 1. Tumour cells in which NF- B is constitutively active 1. are highly resistant to anticancer drugs or ionizing radiation.
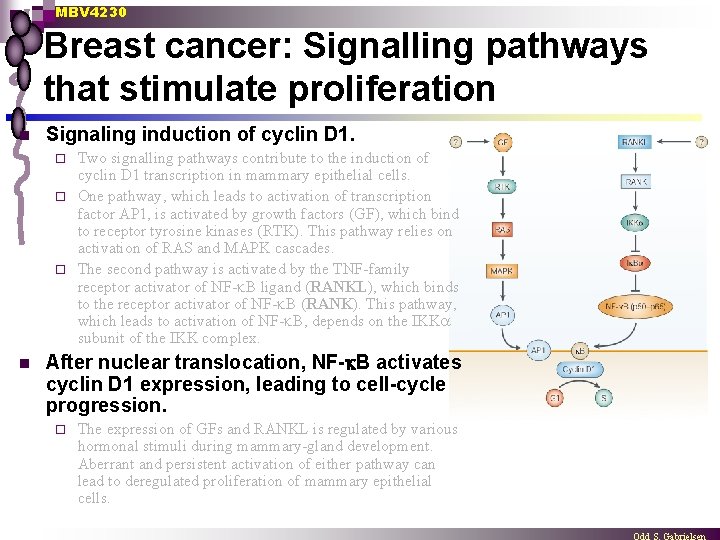
MBV 4230 Breast cancer: Signalling pathways that stimulate proliferation n Signaling induction of cyclin D 1. Two signalling pathways contribute to the induction of cyclin D 1 transcription in mammary epithelial cells. ¨ One pathway, which leads to activation of transcription factor AP 1, is activated by growth factors (GF), which bind to receptor tyrosine kinases (RTK). This pathway relies on activation of RAS and MAPK cascades. ¨ The second pathway is activated by the TNF-family receptor activator of NF- B ligand (RANKL), which binds to the receptor activator of NF- B (RANK). This pathway, which leads to activation of NF- B, depends on the IKK subunit of the IKK complex. ¨ n After nuclear translocation, NF- B activates cyclin D 1 expression, leading to cell-cycle progression. ¨ The expression of GFs and RANKL is regulated by various hormonal stimuli during mammary-gland development. Aberrant and persistent activation of either pathway can lead to deregulated proliferation of mammary epithelial cells.

MBV 4230 Blocking the response n Redox-dependency Antioxidants and alkylating agens inhibit response to many stimuli and inhibit phosphorylation and degradation of I B ¨ H 2 O 2 activates NF- B ¨ Induction of ROI (reactive oxygen intermediates) a possible common element? ¨ n Proteasome inhibitors
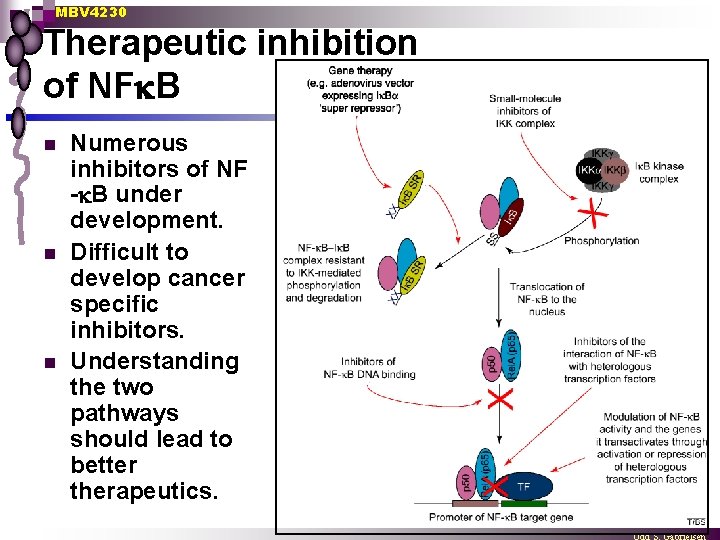
MBV 4230 Therapeutic inhibition of NF B n n n Numerous inhibitors of NF - B under development. Difficult to develop cancer specific inhibitors. Understanding the two pathways should lead to better therapeutics.
- Slides: 54