The Mutability and Repair of DNA Replication errors

The Mutability and Repair of DNA

Replication errors and their repair The nature of mutations: Point mutation 1. Switch of one base for another: (transition) (transversion) purine pyrimidine 2. insertion or deletion of a nucleotide

Drastic changes in DNA Deletion Insertion Rearrangement of chromosome By insertion of a transposon, or aberrant actions of recombination Process.
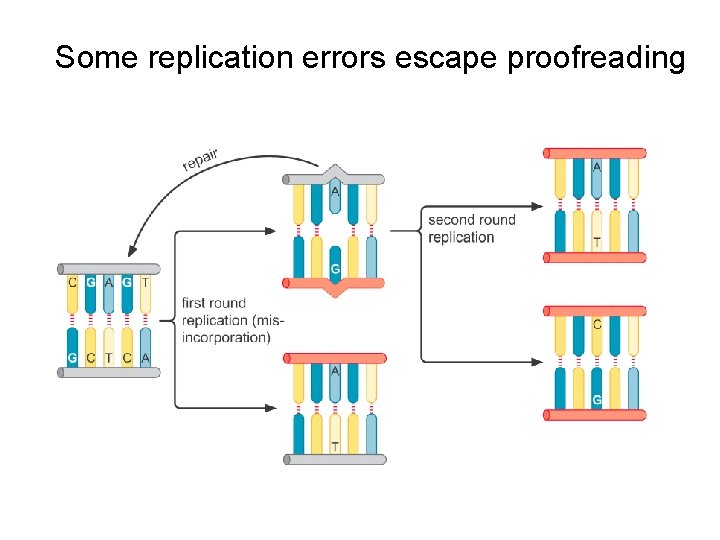
Some replication errors escape proofreading
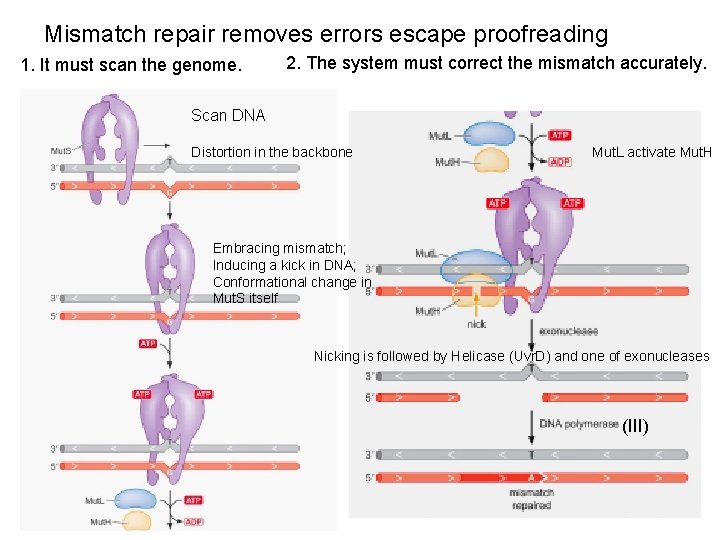
Mismatch repair removes errors escape proofreading 1. It must scan the genome. 2. The system must correct the mismatch accurately. Scan DNA Distortion in the backbone Mut. L activate Mut. H Embracing mismatch; Inducing a kick in DNA; Conformational change in Mut. S itself Nicking is followed by Helicase (Uvr. D) and one of exonucleases (III)
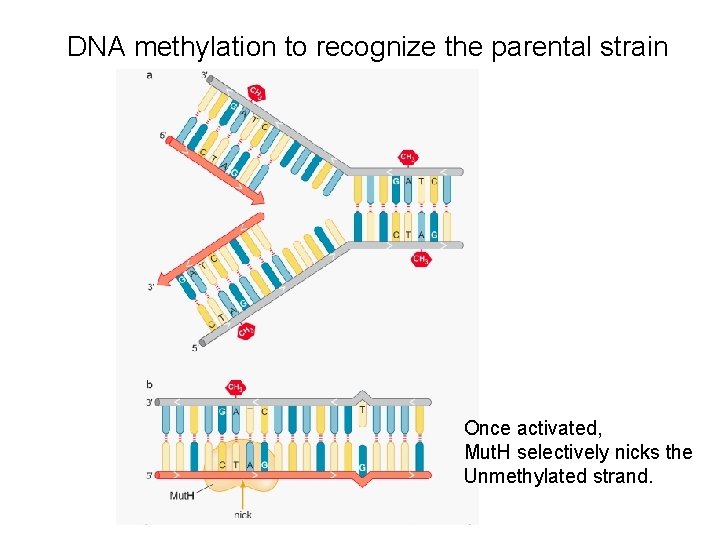
DNA methylation to recognize the parental strain Once activated, Mut. H selectively nicks the Unmethylated strand.
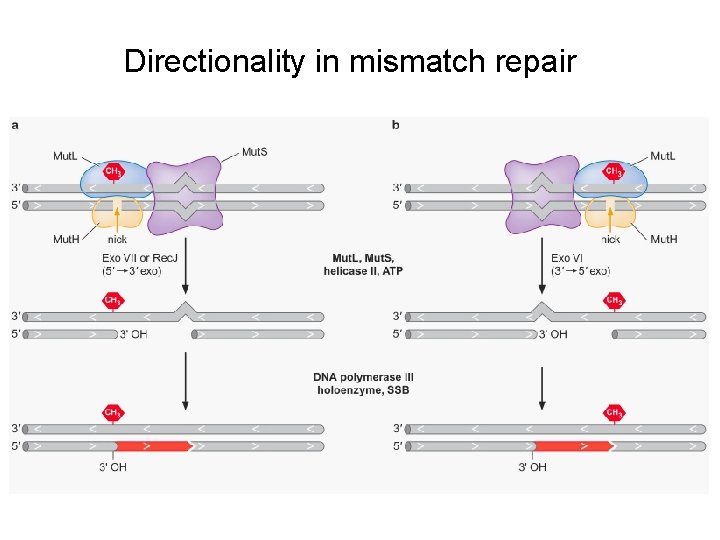
Directionality in mismatch repair

Mismatch repair system in Eukaryotics E. coli Eukaryotics Mut. S Mut. L MSH MLH or PMS (Mut. S homolog) Hereditary nonpolyposis colorectal cancer (mutations in human homologes of Muts and Mut. L)
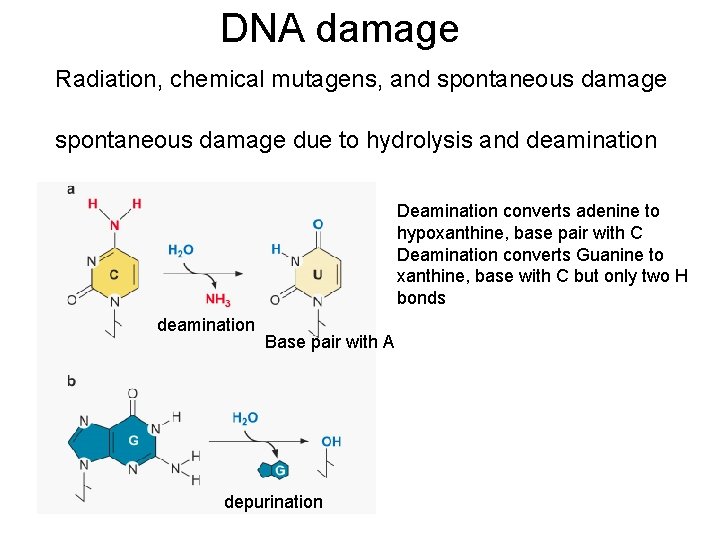
DNA damage Radiation, chemical mutagens, and spontaneous damage due to hydrolysis and deamination Deamination converts adenine to hypoxanthine, base pair with C Deamination converts Guanine to xanthine, base with C but only two H bonds deamination Base pair with A depurination
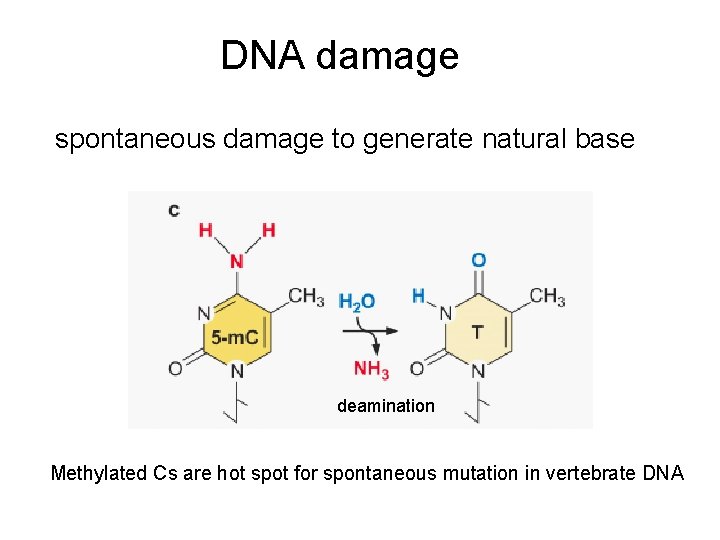
DNA damage spontaneous damage to generate natural base deamination Methylated Cs are hot spot for spontaneous mutation in vertebrate DNA
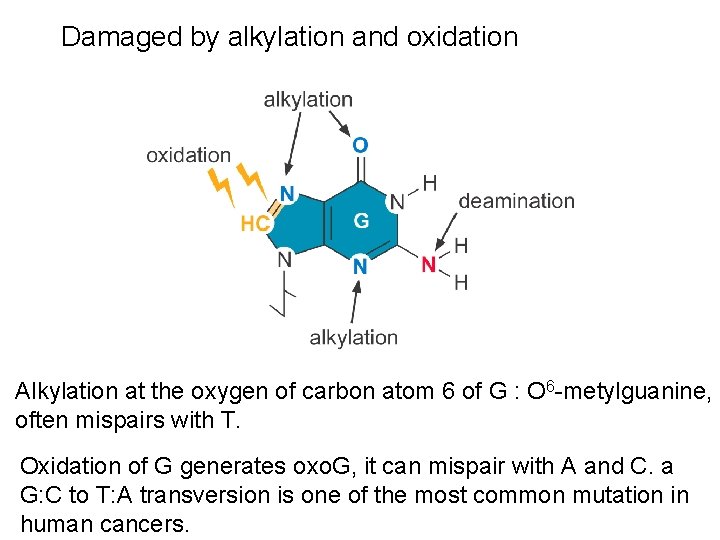
Damaged by alkylation and oxidation Alkylation at the oxygen of carbon atom 6 of G : O 6 -metylguanine, often mispairs with T. Oxidation of G generates oxo. G, it can mispair with A and C. a G: C to T: A transversion is one of the most common mutation in human cancers.

Gamma radiation and X-rays • Cause double-strand breaks in the DNA, which are difficult to repair. • Ionizing radiation and agents like bleomycin that cause DNA to break are said to be clastogenic (p 245).
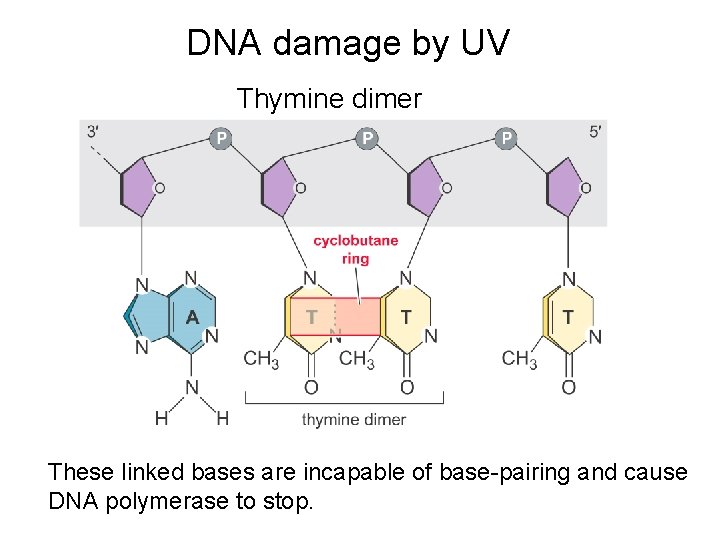
DNA damage by UV Thymine dimer These linked bases are incapable of base-pairing and cause DNA polymerase to stop.
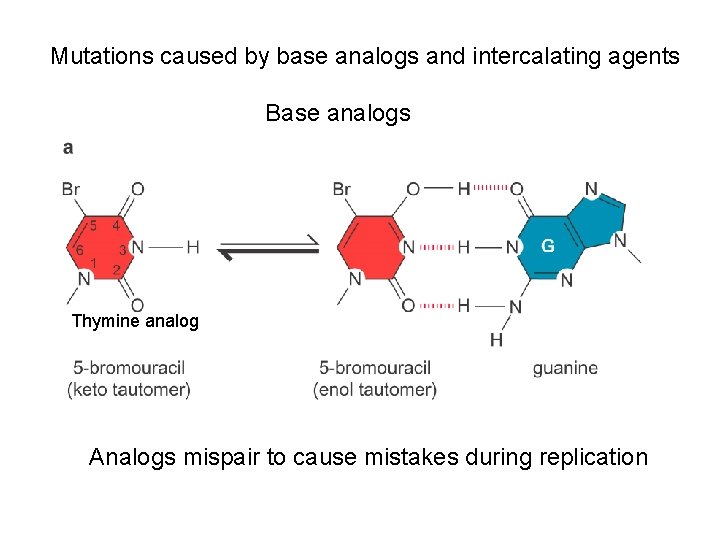
Mutations caused by base analogs and intercalating agents Base analogs Thymine analog Analogs mispair to cause mistakes during replication
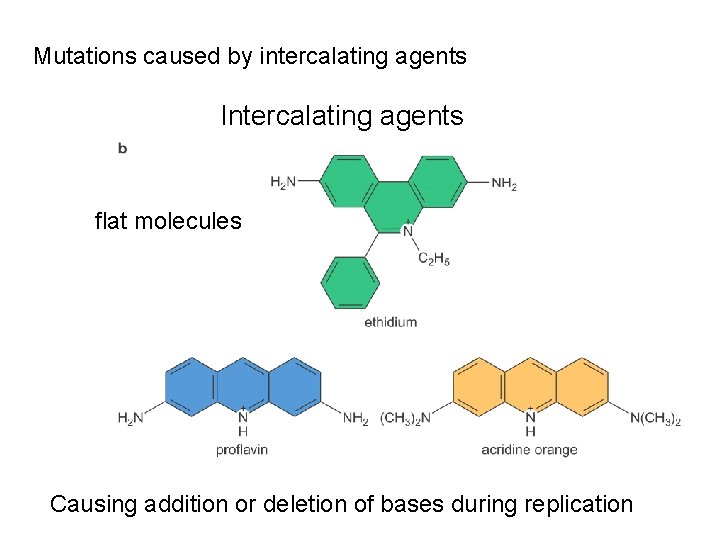
Mutations caused by intercalating agents Intercalating agents flat molecules Causing addition or deletion of bases during replication

Repair of DNA Damage: DNA repair system Excision repair systems: the damaged nucleotide is not repaired but removed from the DNA, the other undamaged strand serves as a template for reincorporation of the correct nt by DNA polymerase Recombination repair: both strands are damaged. Sequence information is retrieved from a second undamaged copy of the chromosome.
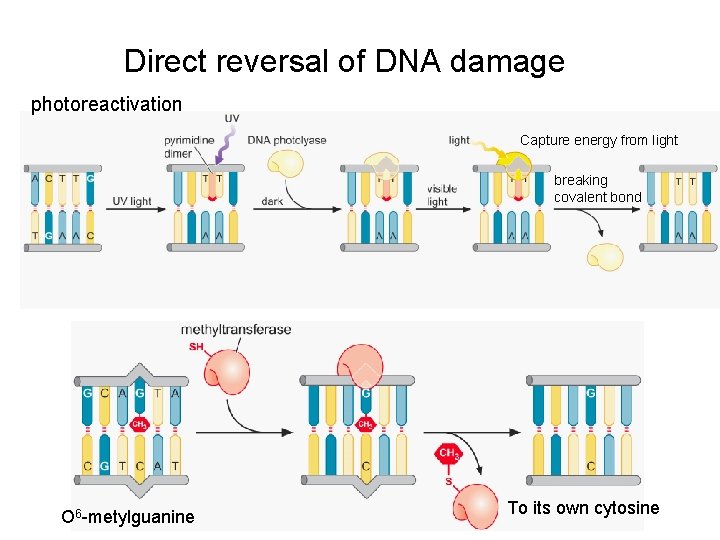
Direct reversal of DNA damage photoreactivation Capture energy from light breaking covalent bond O 6 -metylguanine To its own cytosine
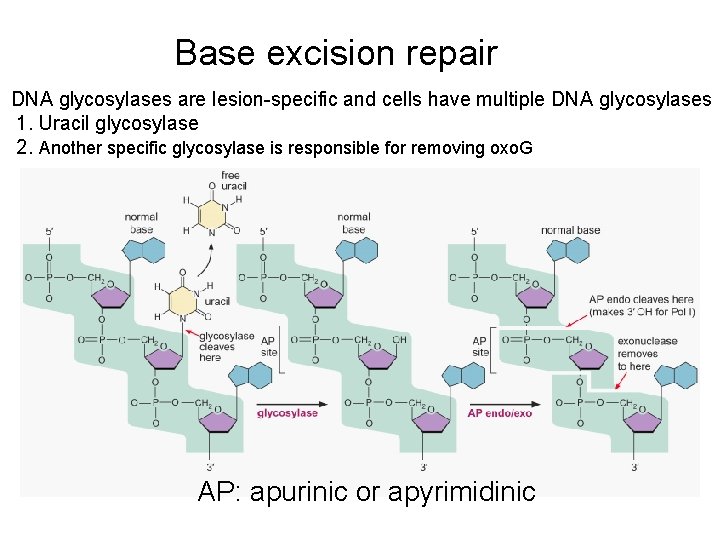
Base excision repair DNA glycosylases are lesion-specific and cells have multiple DNA glycosylases 1. Uracil glycosylase 2. Another specific glycosylase is responsible for removing oxo. G AP: apurinic or apyrimidinic
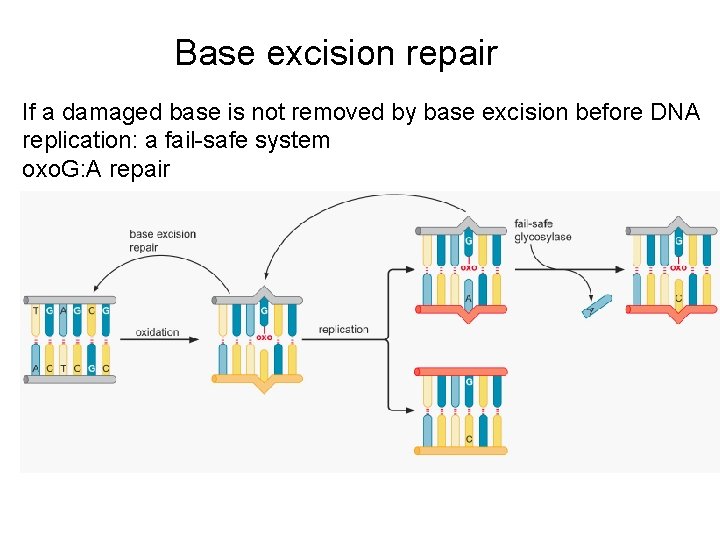
Base excision repair If a damaged base is not removed by base excision before DNA replication: a fail-safe system oxo. G: A repair
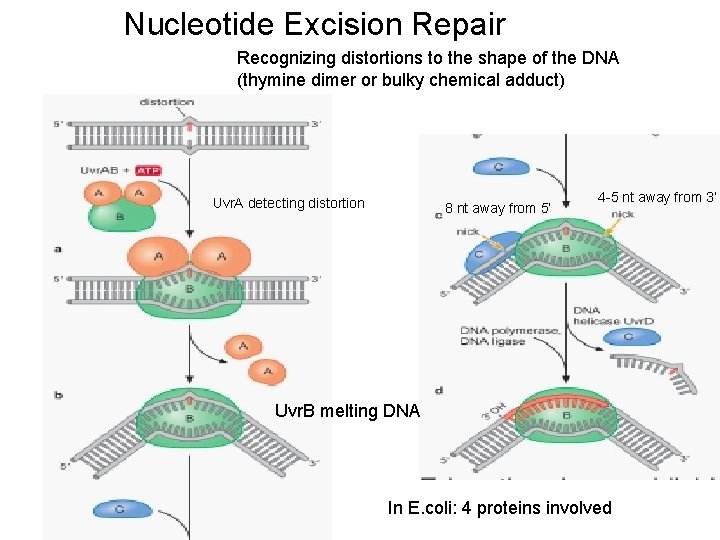
Nucleotide Excision Repair Recognizing distortions to the shape of the DNA (thymine dimer or bulky chemical adduct) Uvr. A detecting distortion 8 nt away from 5’ 4 -5 nt away from 3’ Uvr. B melting DNA In E. coli: 4 proteins involved

Nucleotide Excision Repair The principles of nucleotide excision repair in higher cells is much the same as in E. coli but us moer complicated, involving 25 or more polypeptides. The UVR proteins are needed to mend damage from UV light; Mutants of uvr genes are sensitive to UV light, and lack the capacity to remove T-T or T-C adducts. In human, xeroderma pigmentosum patients have mutations in seven genes (XP genes). These XP proteins are corresponding to proteins involved in nucleotide excision repair.
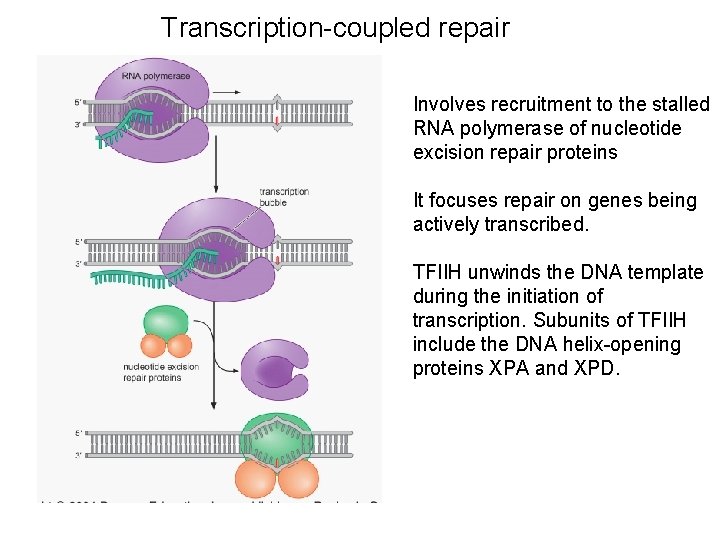
Transcription-coupled repair Involves recruitment to the stalled RNA polymerase of nucleotide excision repair proteins It focuses repair on genes being actively transcribed. TFIIH unwinds the DNA template during the initiation of transcription. Subunits of TFIIH include the DNA helix-opening proteins XPA and XPD.
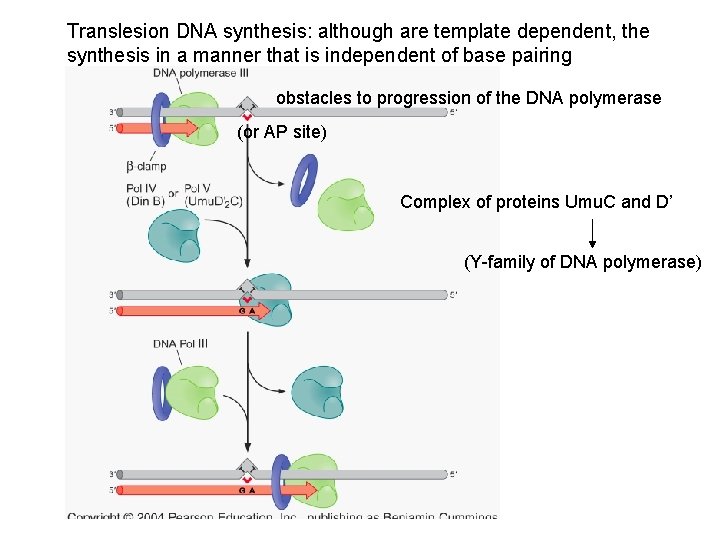
Translesion DNA synthesis: although are template dependent, the synthesis in a manner that is independent of base pairing obstacles to progression of the DNA polymerase (or AP site) Complex of proteins Umu. C and D’ (Y-family of DNA polymerase)
- Slides: 23