The Molecular Genetics of Immunoglobulins Recall Structure Numerous

The Molecular Genetics of Immunoglobulins

Recall Structure • Numerous V region genes are preceded by Leader or signal sequences (60 -90 bp) exons interspersed with introns. • Heavy chain contains V (Variable), D (Diversity), J (Joining) and C (Constant) region gene segments. V - D - J – C • Light chain contains V, J, and C region gene segments. V - J - C • Constant region genes are sub-divided into exons encoding domains (CH 1, CH 2, CH 3, CH 4)
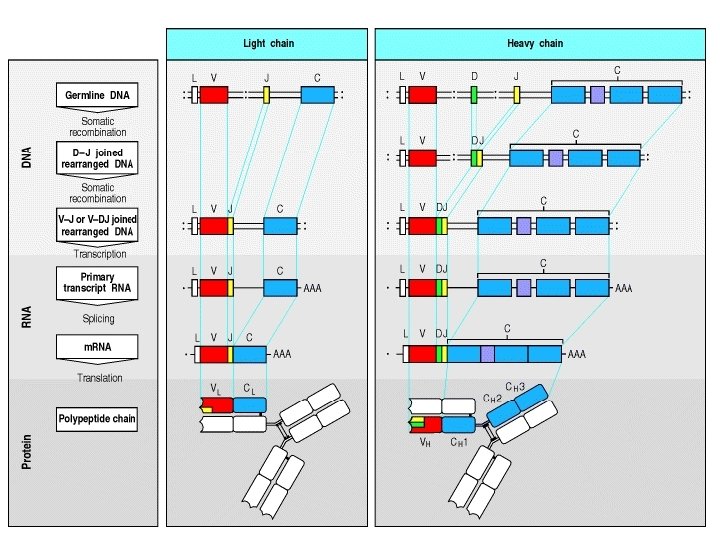

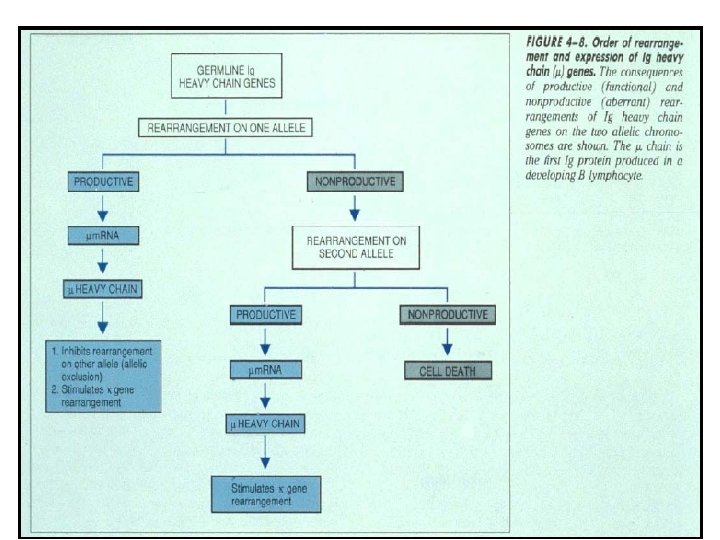
CHARACTERISTICS OF IMMUNOGLOBULIN GENE RE-ARRANGEMENT 1. Involves Allelic Exclusion. – Only one of two parental alleles of Ig is expressed in a B cell. – Either kappa or lambda light chain is expressed by a B cell (light chain isotype exclusion). 2. Ig rearrangement occurs prior to antigen exposure. A. Heavy chain re-arrangement – Re-arrangement occurs in a precise order: – Heavy chain re-arranges before Light chain. – D-J joining occurs first to form DJ and is followed by V-DJ joining to form VDJ. – Just as in light chain the production of μ heavy chain by re-arrangement of one allele inhibits re-arrangement on other allele. If re-arrangement on first allele is non-productive (due to mutations, deletions or frame shifts that generate stop codons), then re-arrangement on the second allele is stimulated.
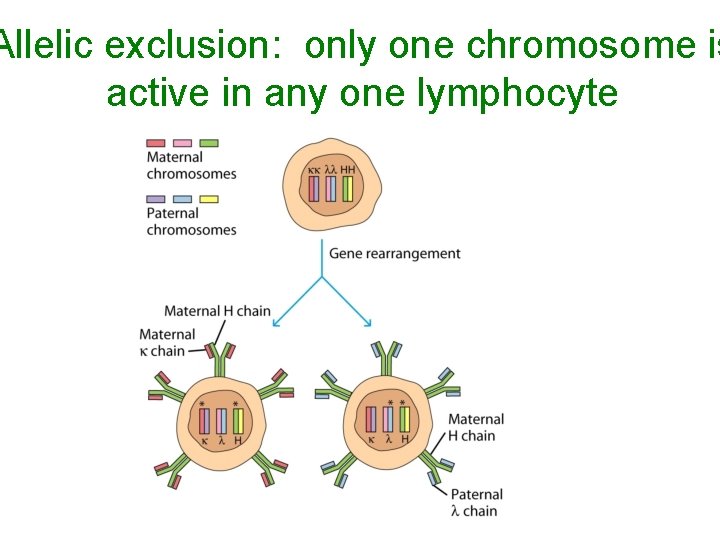
Allelic exclusion: only one chromosome is active in any one lymphocyte

Light chain re-arrangement • • Kappa chain (κ) rearranges before lambda (λ) chain V-joining occurs. Productive arrangement on one allele blocks re-arrangement on other allele. If kappa protein is produced, re-arrangement of lambda chain is blocked. Otherwise lambda chain undergoes re-arrangement.

Questions? 1. How is an infinite diversity of specificity generated from finite amounts of DNA? 2. How can the same specificity of antibody be on the cell surface and secreted? 3. How do V region find J regions and why don’t they join to C regions? 4. How does the DNA break and rejoin?
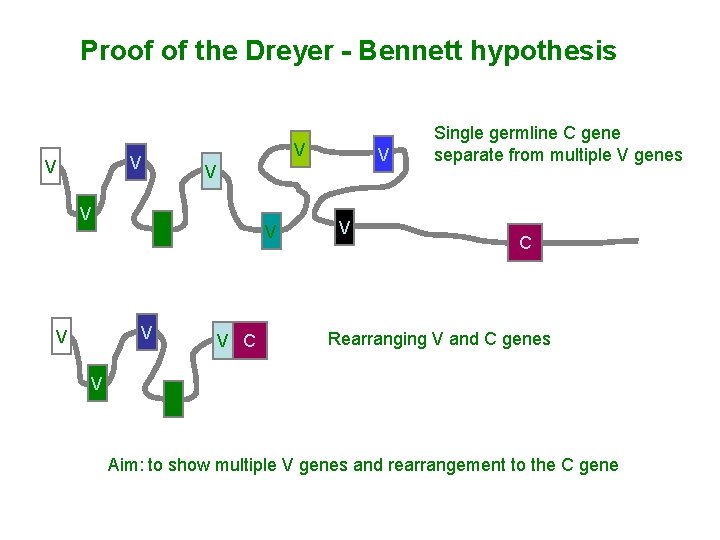
Proof of the Dreyer - Bennett hypothesis V V V C V Single germline C gene separate from multiple V genes C Rearranging V and C genes V Aim: to show multiple V genes and rearrangement to the C gene
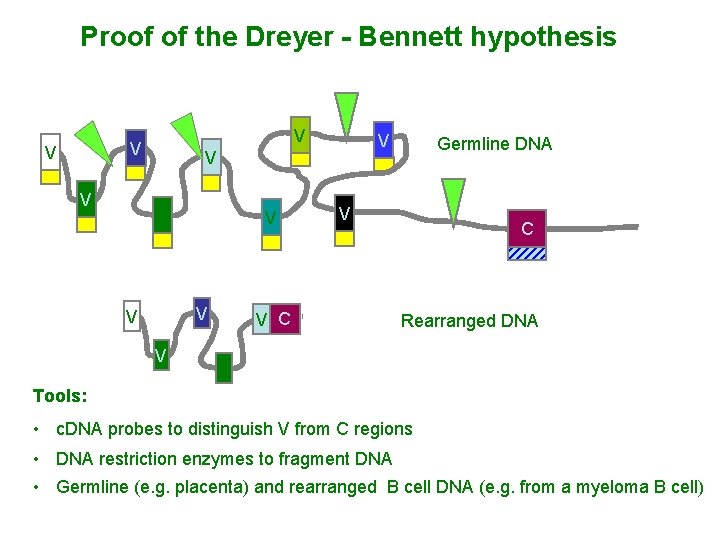
Proof of the Dreyer - Bennett hypothesis V V V Germline DNA V V C C Rearranged DNA V Tools: • c. DNA probes to distinguish V from C regions • DNA restriction enzymes to fragment DNA • Germline (e. g. placenta) and rearranged B cell DNA (e. g. from a myeloma B cell)
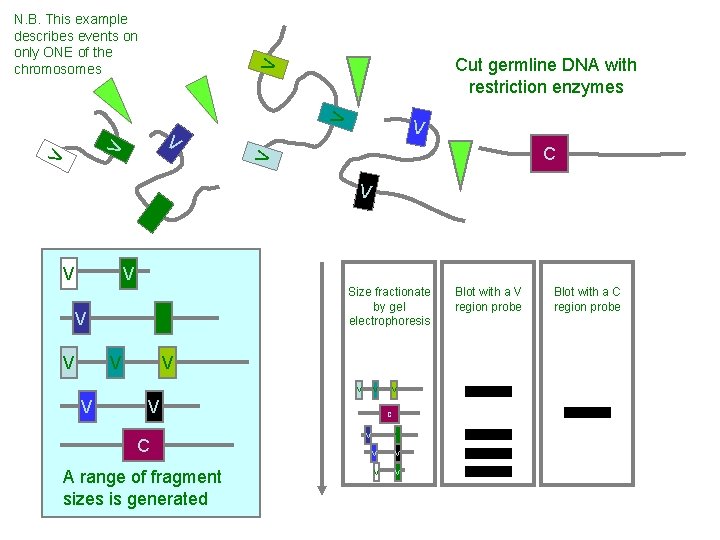
V V C V V V Cut germline DNA with restriction enzymes V N. B. This example describes events on only ONE of the chromosomes V V V C V V V Size fractionate by gel electrophoresis V V V C A range of fragment sizes is generated V C V V V Blot with a V region probe Blot with a C region probe
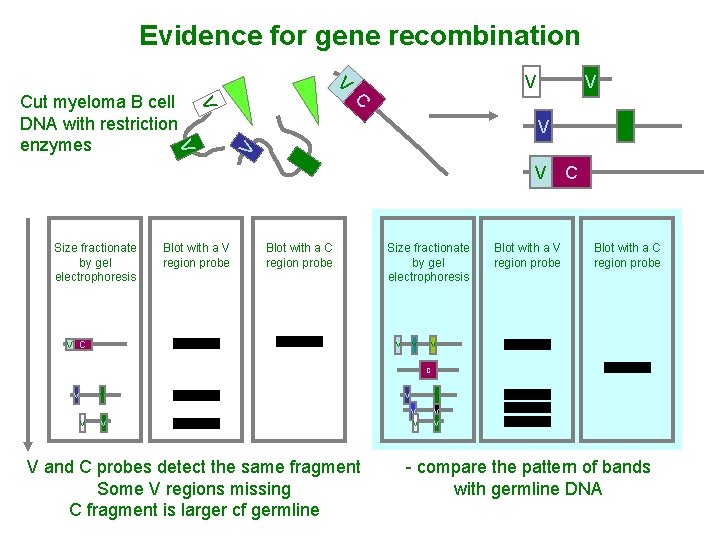
Evidence for gene recombination V V Size fractionate by gel electrophoresis Blot with a V region probe V Blot with a C region probe V C V V V Cut myeloma B cell V DNA with restriction enzymes V V C V Size fractionate by gel electrophoresis V V Blot with a V region probe V C Blot with a C region probe V C V V V V and C probes detect the same fragment Some V regions missing C fragment is larger cf germline V V V - compare the pattern of bands with germline DNA
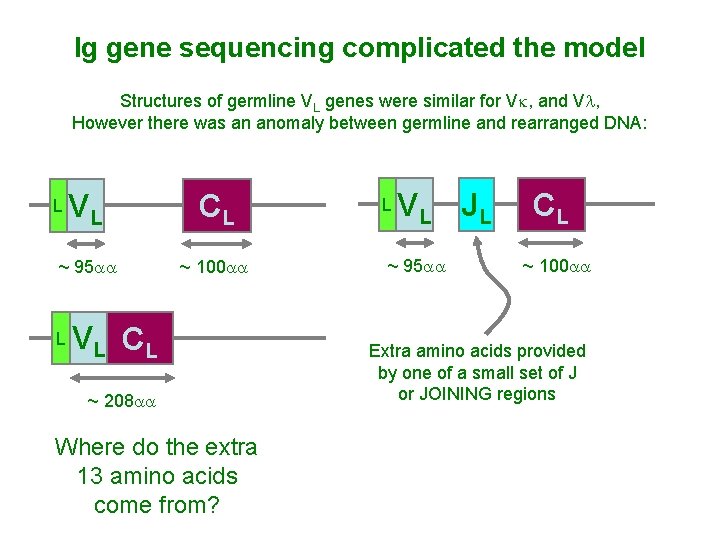
Ig gene sequencing complicated the model Structures of germline VL genes were similar for Vk, and Vl, However there was an anomaly between germline and rearranged DNA: VL CL ~ 95 aa ~ 100 aa L L V L CL ~ 208 aa Where do the extra 13 amino acids come from? L V L JL ~ 95 aa CL ~ 100 aa Extra amino acids provided by one of a small set of J or JOINING regions
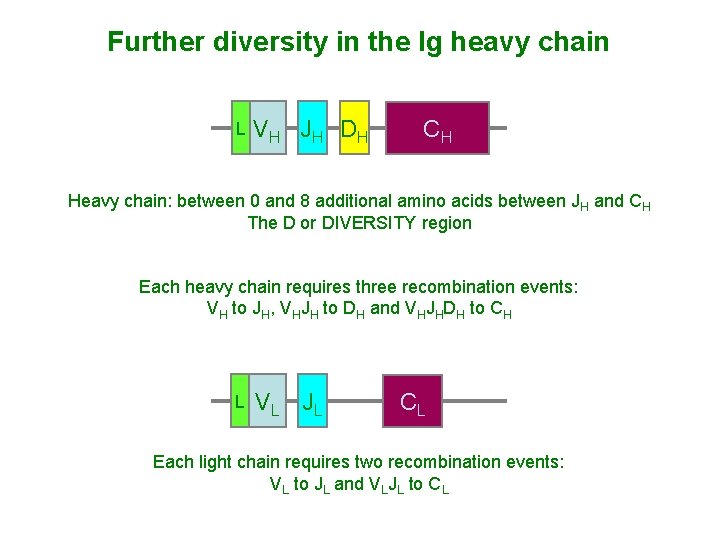
Further diversity in the Ig heavy chain L VH JH DH CH Heavy chain: between 0 and 8 additional amino acids between JH and CH The D or DIVERSITY region Each heavy chain requires three recombination events: VH to JH, VHJH to DH and VHJHDH to CH L VL JL CL Each light chain requires two recombination events: VL to JL and VLJL to CL

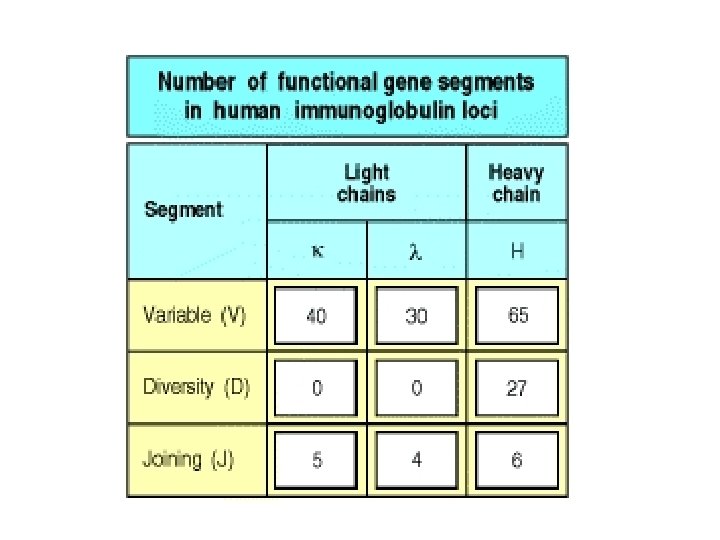
Diversity: Multiple Germline Genes VH Locus: • • • JH Locus: • 9 JH genes • 6 functional JH genes with products identified • 3 pseudo JH genes DH Locus: • • 123 VH genes on chromosome 14 40 functional VH genes with products identified 79 pseudo VH genes 4 functional VH genes - with no products identified 24 non-functional, orphan VH sequences on chromosomes 15 & 16 27 DH genes 23 functional DH genes with products identified 4 pseudo DH genes Additional non-functional DH sequences on the chromosome 15 orphan locus • reading DH regions in 3 frames functionally increases number of DH regions

Diversity: Multiple germline genes Vk & Jk Loci: • 132 Vk genes on the short arm of chromosome 2 29 functional Vk genes with products identified 87 pseudo Vk genes 15 functional Vk genes - with no products identified 25 orphans Vk genes on the long arm of chromosome 2 5 Jk regions Vl & Jl Loci: • 105 Vl genes on the short arm of chromosome 2 30 functional genes with products identified 56 pseudogenes 6 functional genes - with no products identified 13 relics (<200 bp Vl of sequence) 25 orphans on the long arm of chromosome 2 4 Jl regions • • •
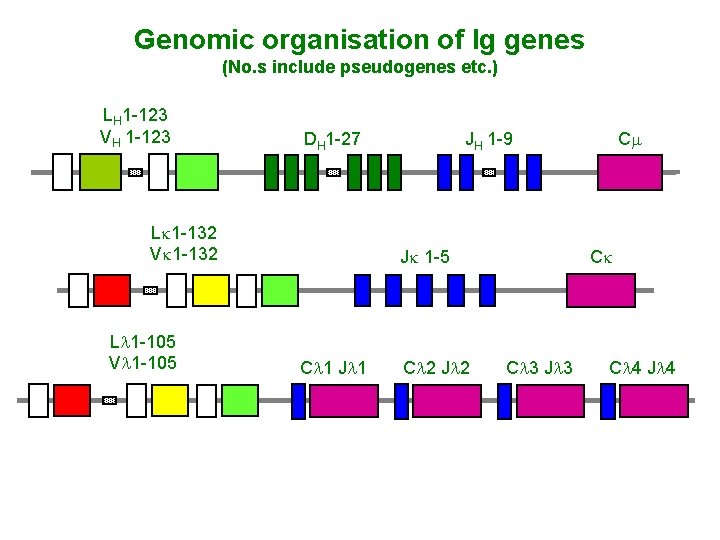
Genomic organisation of Ig genes (No. s include pseudogenes etc. ) LH 1 -123 VH 1 -123 DH 1 -27 Lk 1 -132 Vk 1 -132 Ll 1 -105 Vl 1 -105 JH 1 -9 Jk 1 -5 Cl 1 Jl 1 Cl 2 Jl 2 Cm Ck Cl 3 Jl 3 Cl 4 Jl 4
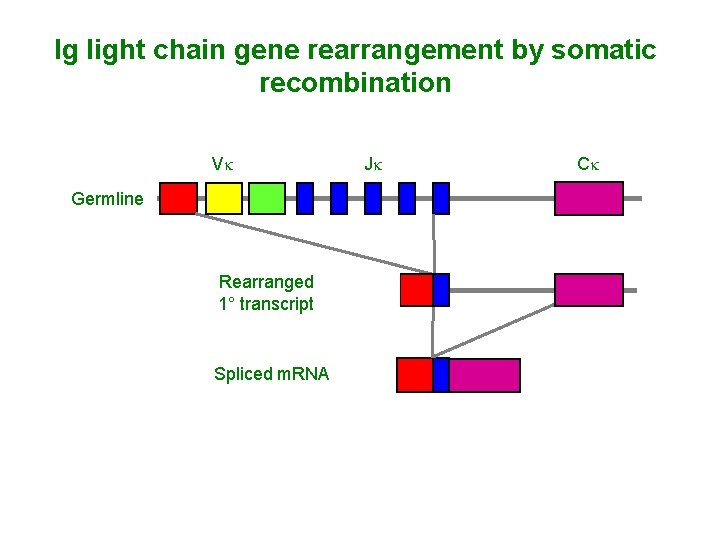
Ig light chain gene rearrangement by somatic recombination Vk Germline Rearranged 1° transcript Spliced m. RNA Jk Ck

Questions? 1. How is an infinite diversity of specificity generated from finite amounts of DNA? 2. How can the same specificity of antibody be on the cell surface and secreted? 3. How do V region find J regions and why don’t they join to C regions? 4. How does the DNA break and rejoin?

Remember • Cell surface antigen receptor on B cells Allows B cells to sense their antigenic environment Connects extracellular space with intracellular signalling machinery • Secreted antibody Neutralisation Arming/recruiting effector cells Complement fixation How does the model of recombination allow for two different forms of the protein?
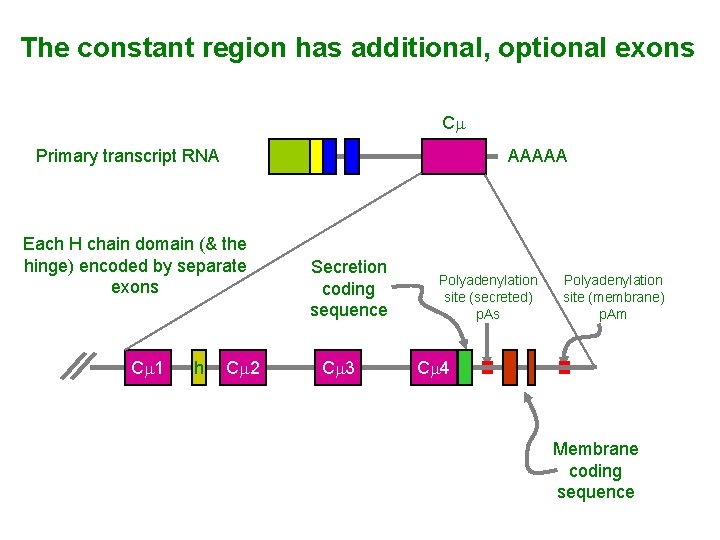
The constant region has additional, optional exons Cm Primary transcript RNA AAAAA Each H chain domain (& the hinge) encoded by separate exons Cm 1 h Cm 2 Secretion coding sequence Cm 3 Polyadenylation site (secreted) p. As Polyadenylation site (membrane) p. Am Cm 4 Membrane coding sequence
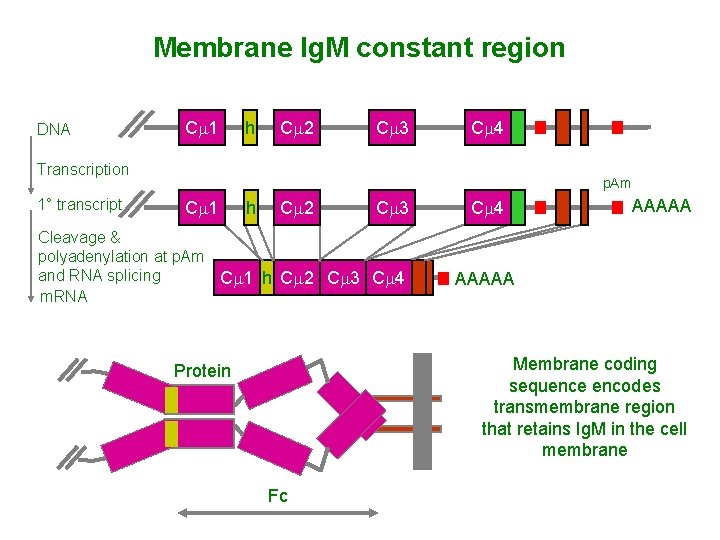
Membrane Ig. M constant region DNA Cm 1 h Cm 2 Cm 3 Cm 4 Transcription 1° transcript p. Am Cm 1 h Cm 2 Cm 3 Cleavage & polyadenylation at p. Am and RNA splicing Cm 1 h Cm 2 Cm 3 Cm 4 m. RNA Cm 4 AAAAA Membrane coding sequence encodes transmembrane region that retains Ig. M in the cell membrane Protein Fc
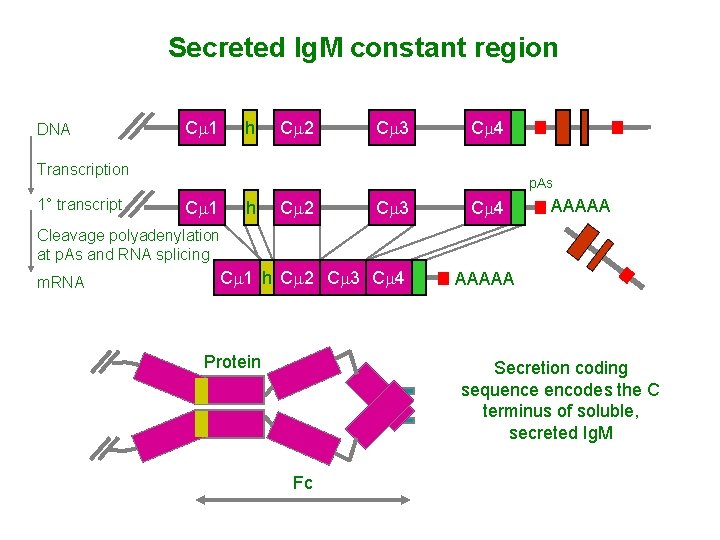
Secreted Ig. M constant region DNA Cm 1 h Cm 2 Cm 3 Cm 4 Transcription 1° transcript p. As Cm 1 h Cm 2 Cm 3 Cm 4 AAAAA Cleavage polyadenylation at p. As and RNA splicing m. RNA Cm 1 h Cm 2 Cm 3 Cm 4 Protein AAAAA Secretion coding sequence encodes the C terminus of soluble, secreted Ig. M Fc
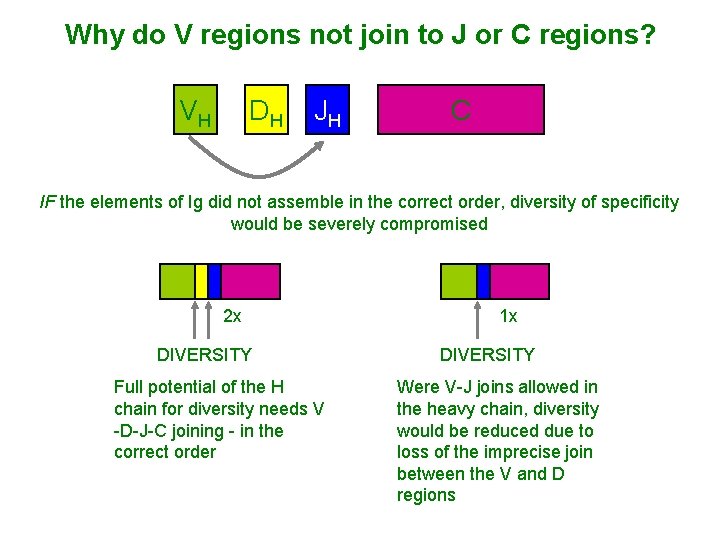
Why do V regions not join to J or C regions? VH DH JH C IF the elements of Ig did not assemble in the correct order, diversity of specificity would be severely compromised 2 x DIVERSITY Full potential of the H chain for diversity needs V -D-J-C joining - in the correct order 1 x DIVERSITY Were V-J joins allowed in the heavy chain, diversity would be reduced due to loss of the imprecise join between the V and D regions

Rearrangement of V, D, and J gene segments is guided by flanking DNA sequences
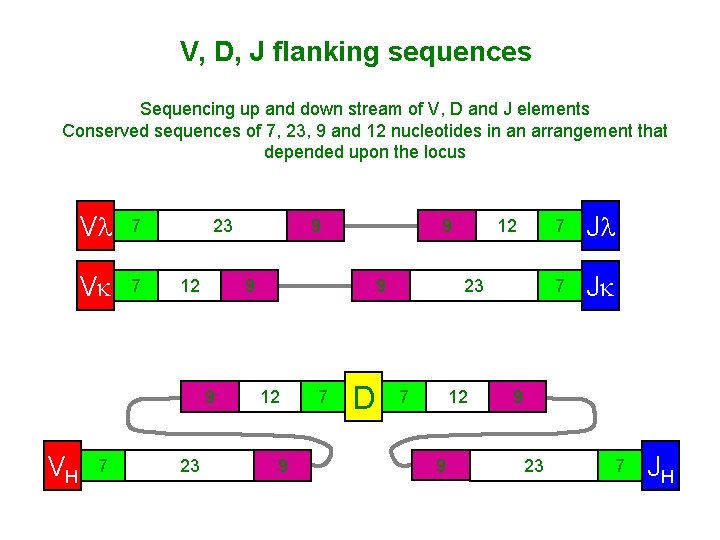
V, D, J flanking sequences Sequencing up and down stream of V, D and J elements Conserved sequences of 7, 23, 9 and 12 nucleotides in an arrangement that depended upon the locus Vl 7 Vk 7 23 12 7 23 9 12 9 7 12 9 9 9 VH 9 D 23 7 12 9 7 Jl 7 Jk 9 23 7 JH
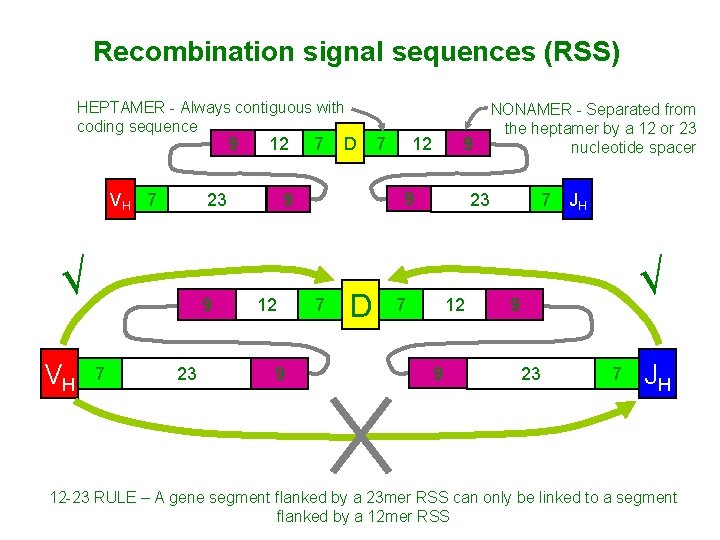
Recombination signal sequences (RSS) HEPTAMER - Always contiguous with coding sequence 9 VH 7 23 √ VH 9 7 12 23 7 D 9 9 12 7 7 D NONAMER - Separated from the heptamer by a 12 or 23 nucleotide spacer 23 7 12 9 7 JH √ 9 23 7 JH 12 -23 RULE – A gene segment flanked by a 23 mer RSS can only be linked to a segment flanked by a 12 mer RSS
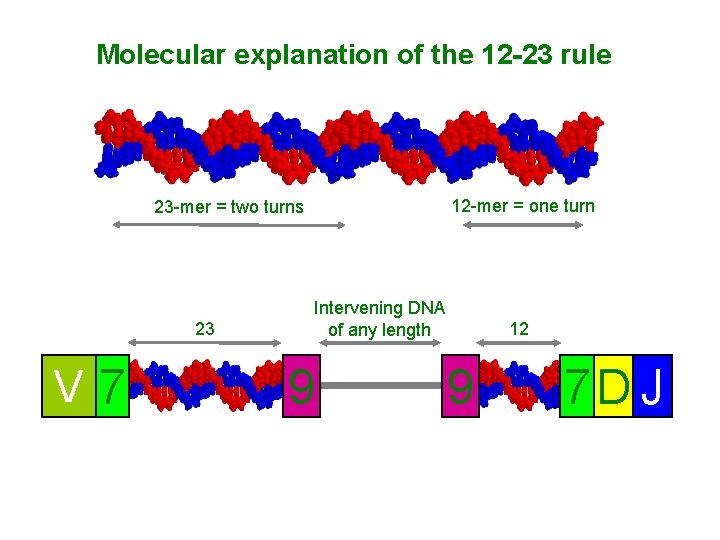
Molecular explanation of the 12 -23 rule 12 -mer = one turn 23 -mer = two turns 23 V 7 Intervening DNA of any length 9 12 9 7 D J
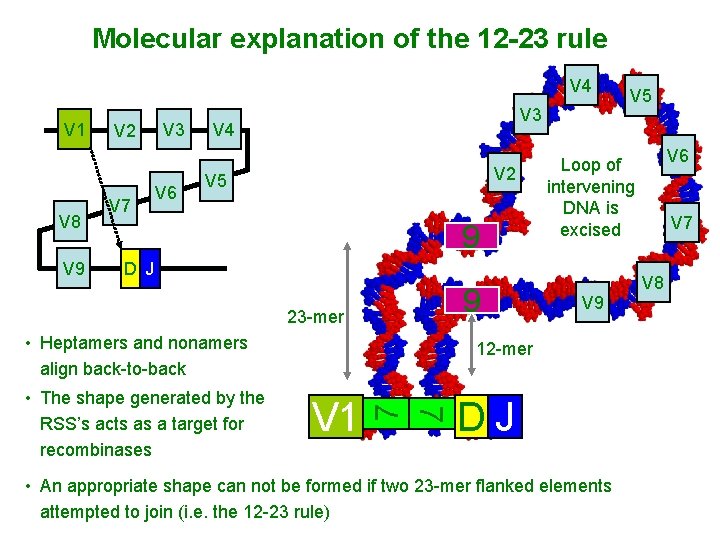
Molecular explanation of the 12 -23 rule V 4 V 1 V 8 V 9 V 2 V 7 V 3 V 6 V 3 V 4 V 2 V 5 9 9 • Heptamers and nonamers align back-to-back 7 V 1 V 7 V 8 V 9 12 -mer 7 • The shape generated by the RSS’s acts as a target for recombinases V 6 Loop of intervening DNA is excised D J 23 -mer V 5 DJ • An appropriate shape can not be formed if two 23 -mer flanked elements attempted to join (i. e. the 12 -23 rule)
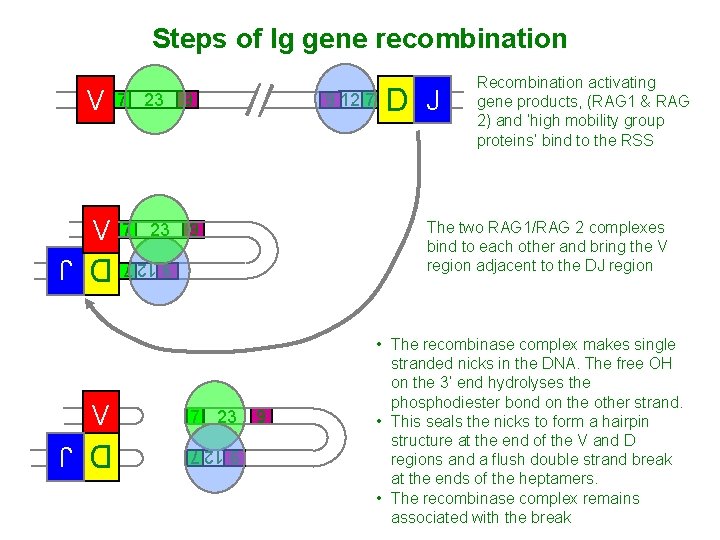
Steps of Ig gene recombination V 7 23 D J 9 12 7 7 23 9 12 7 9 7 23 D J The two RAG 1/RAG 2 complexes bind to each other and bring the V region adjacent to the DJ region 9 9 12 7 V 9 Recombination activating gene products, (RAG 1 & RAG 2) and ‘high mobility group proteins’ bind to the RSS 9 • The recombinase complex makes single stranded nicks in the DNA. The free OH on the 3’ end hydrolyses the phosphodiester bond on the other strand. • This seals the nicks to form a hairpin structure at the end of the V and D regions and a flush double strand break at the ends of the heptamers. • The recombinase complex remains associated with the break D J
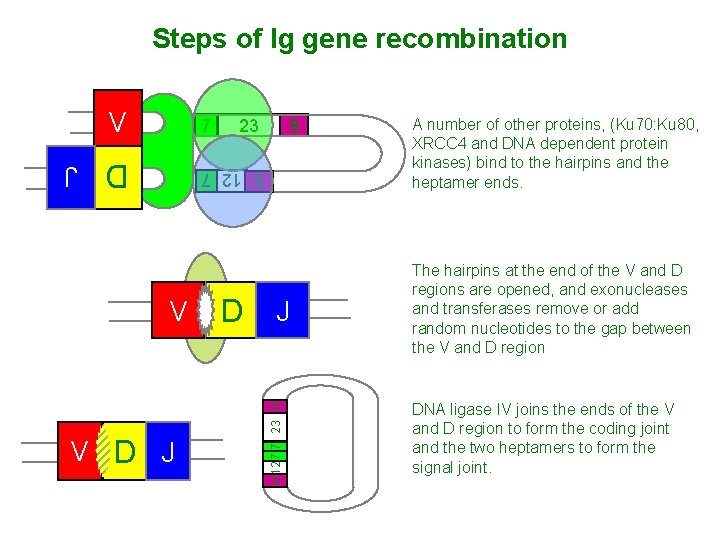
Steps of Ig gene recombination V 7 23 9 D J 9 12 7 V D J 9 12 7 7 23 9 V A number of other proteins, (Ku 70: Ku 80, XRCC 4 and DNA dependent protein kinases) bind to the hairpins and the heptamer ends. The hairpins at the end of the V and D regions are opened, and exonucleases and transferases remove or add random nucleotides to the gap between the V and D region DNA ligase IV joins the ends of the V and D region to form the coding joint and the two heptamers to form the signal joint.
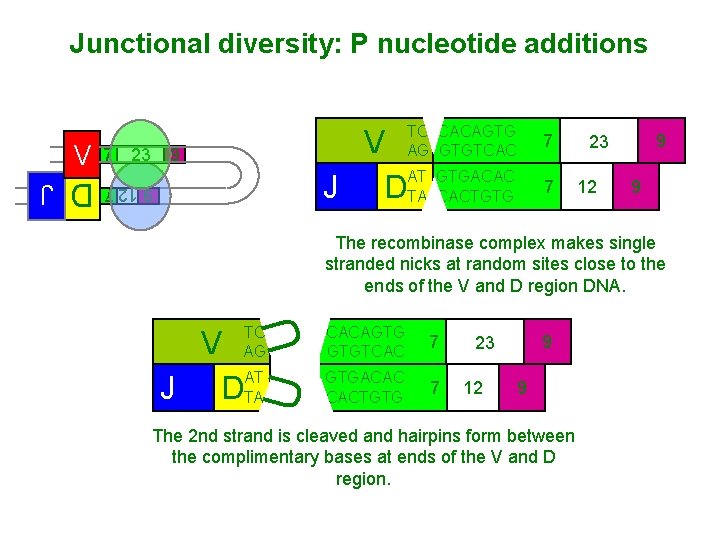
Junctional diversity: P nucleotide additions 7 23 V AT GTGACAC J DTA CACTGTG 9 9 12 7 7 7 9 23 12 9 The recombinase complex makes single stranded nicks at random sites close to the ends of the V and D region DNA. U U V TC CACAGTG AG GTGTCAC TC AG TC CACAGTG AG GTGTCAC 7 GTGACAC CACTGTG 7 V V AT AT J JDTA 9 23 12 9 The 2 nd strand is cleaved and hairpins form between the complimentary bases at ends of the V and D region. D J
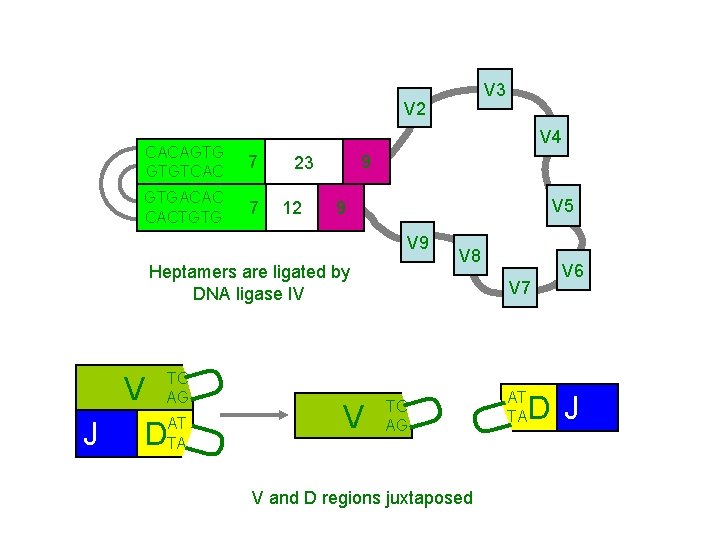
V 3 V 2 V 4 CACAGTG GTGTCAC 7 GTGACAC CACTGTG 7 9 23 12 V 5 9 V AT J DTA V U TC AG V 7 TC AG AT TA U U U Heptamers are ligated by DNA ligase IV V 8 V and D regions juxtaposed V 6 D J
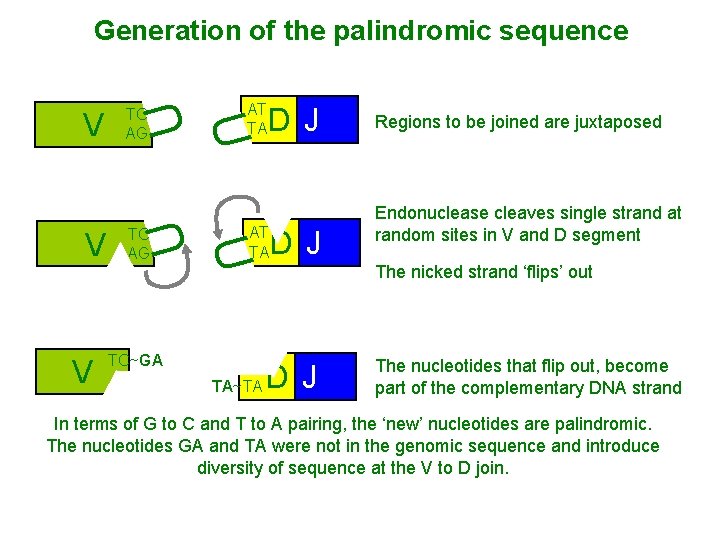
V U TC AG TC~GA AG AT TA D J AT TA U V TC AG U V U Generation of the palindromic sequence AT TA~TA D J Regions to be joined are juxtaposed Endonuclease cleaves single strand at random sites in V and D segment The nicked strand ‘flips’ out D J The nucleotides that flip out, become part of the complementary DNA strand In terms of G to C and T to A pairing, the ‘new’ nucleotides are palindromic. The nucleotides GA and TA were not in the genomic sequence and introduce diversity of sequence at the V to D join.
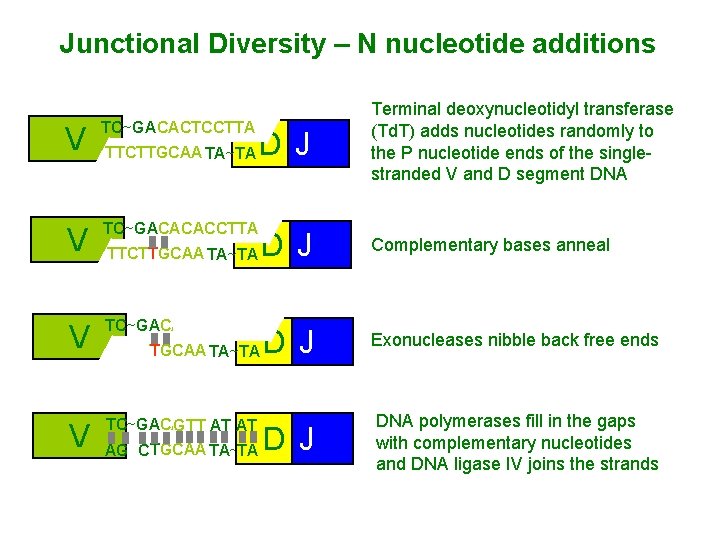
Junctional Diversity – N nucleotide additions V TC~GA CACTCCTTA AT AG TTCTTGCAA TA~TA V TC~GACACACCTTA V TC CACACCTTA TC~GA GTT ATAT AT AGC TTCTTGCAA TA TA~TA AG D J TTCTTGCAA TA~TA Terminal deoxynucleotidyl transferase (Td. T) adds nucleotides randomly to the P nucleotide ends of the singlestranded V and D segment DNA Complementary bases anneal D J Exonucleases nibble back free ends D J DNA polymerases fill in the gaps with complementary nucleotides and DNA ligase IV joins the strands

Junctional Diversity V TCGACGTTATAT AGCTGCAATATA D J Germline-encoded nucleotides Palindromic (P) nucleotides - in the germline Non-template (N) encoded nucleotides - not in the germline Creates an essentially random sequence between the V region, D region and J region in heavy chains and the V region and J region in light chains.

Problems? 1. How is an infinite diversity of specificity generated from finite amounts of DNA? Combinatorial Diversity, genomic organisation and Junctional Diversity 2. How can the same specificity of antibody be on the cell surface and secreted? Use of alternative polyadenylation sites 3. How do V region find J regions and why don’t they join to C regions? The 12 -23 rule 4. How does the DNA break and rejoin? Imprecisely to allow Junctional Diversity

Variable addition and subtraction of nucleotides at the junctions between gene segments contributes to diversity in the third hypervariable region • Of the three hypervariable loops in the protein chains of immunoglobulins, two are encoded within the V gene segment DNA. The third (HV 3 or CDR 3) falls at the joint between the V gene segment and the J gene segment, and in the heavy chain is partially encoded by the D gene segment. • In both heavy and light chains, the diversity of CDR 3 is significantly increased by the addition and deletion of nucleotides at two steps in the formation of the junctions between gene segments. The added nucleotides are known as P-nucleotidesand N-nucleotides

• As the total number of nucleotides added by these processes is random, the added nucleotides often disrupt the reading frame of the coding sequence beyond the joint. • Such frameshifts will lead to a nonfunctional protein, and DNA rearrangements leading to such disruptions are known as nonproductive rearrangements. • As roughly two in every three rearrangements will be nonproductive, many B cells never succeed in producing functional immunoglobulin molecules, and junctional diversity is therefore achieved only at the expense of considerable wastage.
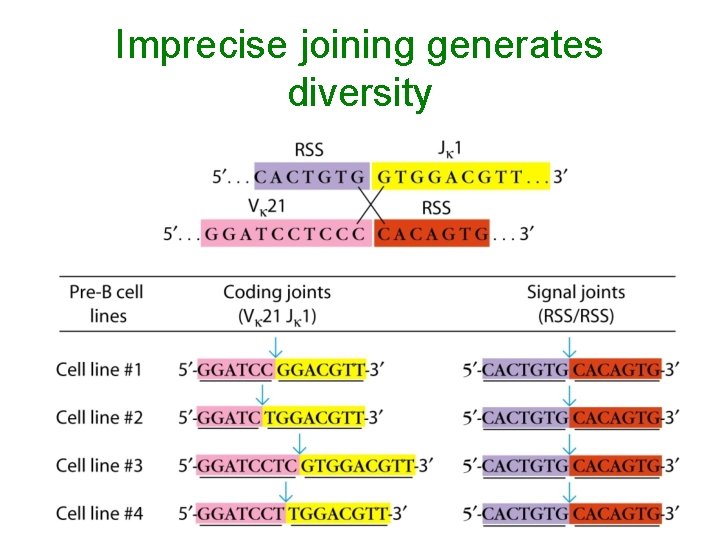
Imprecise joining generates diversity

Some rearrangements are productive, others are non-productive: frame shift alterations are non-productive
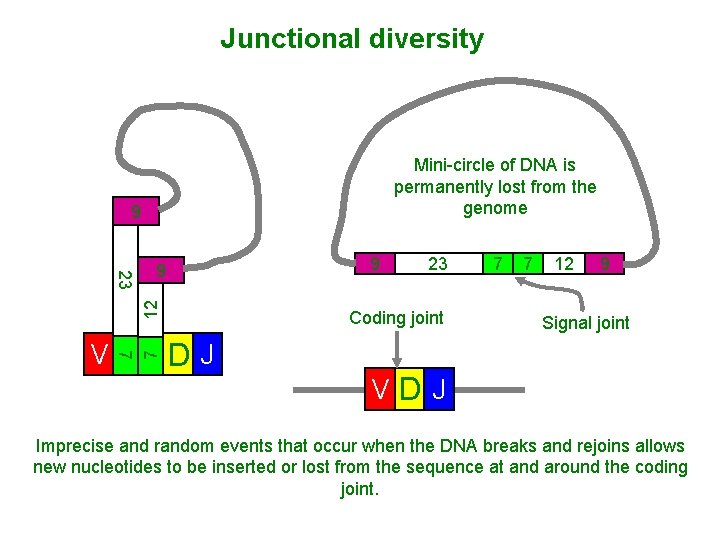
Junctional diversity Mini-circle of DNA is permanently lost from the genome 9 7 V 7 12 23 9 9 23 Coding joint DJ 7 7 12 9 Signal joint VDJ Imprecise and random events that occur when the DNA breaks and rejoins allows new nucleotides to be inserted or lost from the sequence at and around the coding joint.
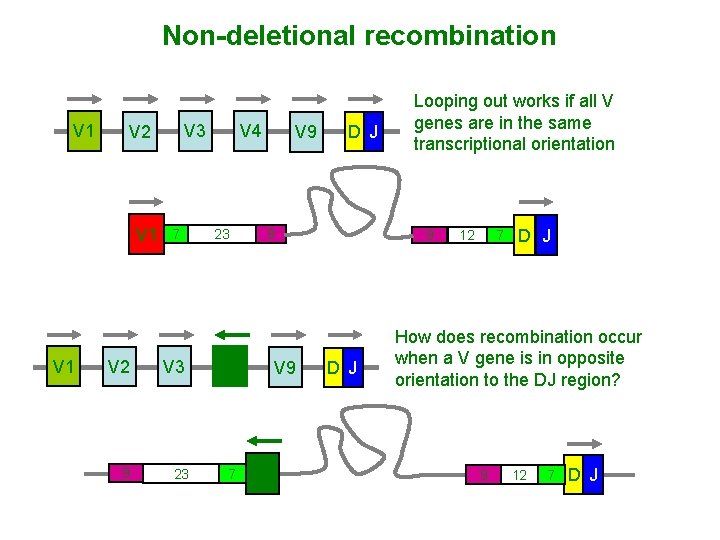
Non-deletional recombination V 1 V 1 V 3 V 2 7 V 2 V 3 9 23 V 4 23 V 9 9 V 4 7 D J V 9 V 4 Looping out works if all V genes are in the same transcriptional orientation 9 D J 12 7 D J How does recombination occur when a V gene is in opposite orientation to the DJ region? 9 12 7 D J
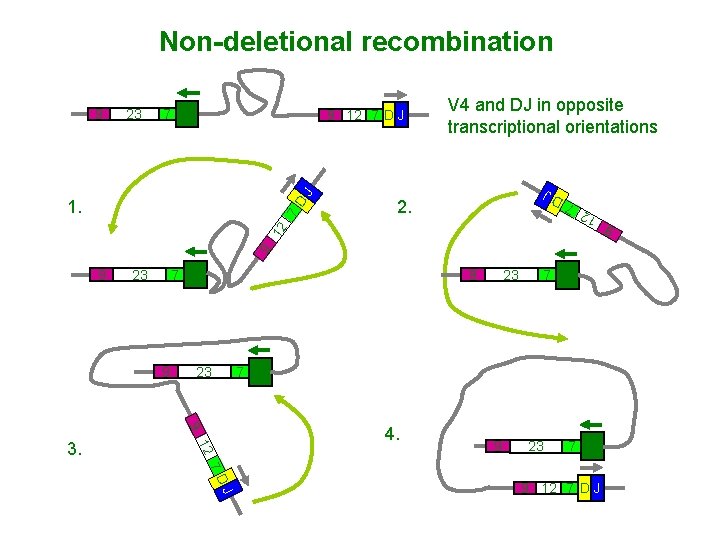
Non-deletional recombination V 4 and DJ in opposite transcriptional orientations J 9 12 7 D J 2. 9 12 7 D 1. 9 7 V 4 23 9 9 23 9 12 3. 23 7 V 4 4. 9 23 7 V 4 J 7 D 9 12 7 D J 9 7 V 4 12 23 7 D J 9
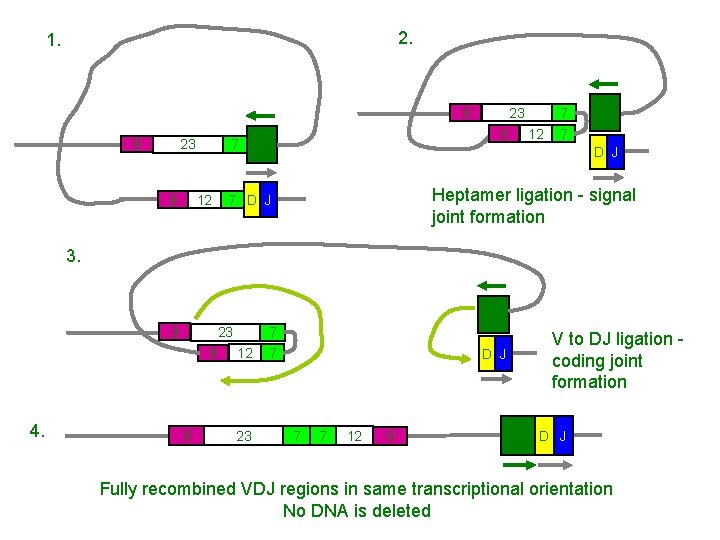
2. 1. 9 9 23 7 12 9 23 9 V 4 12 7 7 V 4 D J Heptamer ligation - signal joint formation 7 D J 3. V 4 9 23 9 4. 9 12 23 7 7 D J 7 7 12 9 V to DJ ligation coding joint formation V 4 D J Fully recombined VDJ regions in same transcriptional orientation No DNA is deleted

Rearrangement of V, D, and J gene segments is guided by flanking DNA sequences • • A system is required to ensure that DNA rearrangements take place at the correct locations relative to the V, D, or J gene segment coding regions. V gene segment joins to a D or J and not to another V. DNA rearrangements are in fact guided by conserved noncoding DNA sequences that are found adjacent to the points at which recombination takes place. These sequences consist of a conserved block of seven nucleotides— the heptamer 5′CACAGTG 3′—which is always contiguous with the coding sequence, followed by a nonconserved region known as the spacer, which is either 12 or 23 nucleotides long. This is followed by a second conserved block of nine nucleotides— the nonamer 5′ACAAAAACC 3′. The spacer varies in sequence but its conserved length corresponds to one or two turns of the DNA double helix. This brings the heptamer and nonamer sequences to the same side of the DNA helix, where they can be bound by the complex of proteins that catalyzes recombination. The heptamer-spacer-nonamer is called a recombination signal sequence (RSS).

12/23 Rule • Recombination only occurs between gene segments located on the same chromosome. • It generally follows the rule that only a gene segment flanked by a RSS with a 12 -base pair (bp) spacer can be joined to one flanked by a 23 bp spacer RSS. This is known as the 12/23 rule. • For the heavy chain, a DH gene segment can be joined to a JH gene segment and a VH gene segment to a DH gene segment, but VH gene segments cannot be joined to JH gene segments directly, as both VH and JH gene segments are flanked by 23 bp spacers and the DH gene segments have 12 bp spacers on both sides

The diversity of the immunoglobulin repertoire is generated by four main processes • Antibody diversity is generated in four main ways. • The gene rearrangement that combines two or three gene segments to form a complete V-region exon generates diversity in two ways. – First, there are multiple different copies of each type of gene segment, and different combinations of gene segments can be used in different rearrangement events. This combinatorial diversity is responsible for a substantial part of the diversity of the heavy- and light-chain V regions. – Second, junctional diversity is introduced at the joints between the different gene segments as a result of addition and subtraction of nucleotides by the recombination process. • A third source of diversity is also combinatorial, arising from the many possible different combinations of heavy- and light-chain V regions that pair to form the antigen-binding site in the immunoglobulin molecule. • Somatic mutation

Rearranged V genes are further diversified by somatic hypermutation • The mechanisms for generating diversity described so far all take place during the rearrangement of gene segments in the initial development of B cells in the central lymphoid organs. • There is an additional mechanism that generates diversity throughout the V region and that operates on B cells in peripheral lymphoid organs after functional immunoglobulin genes have been assembled. • This process, known as somatic hypermutation. • Introduces point mutations into the V regions of the rearranged heavy- and light-chain genes at a very high rate, giving rise to mutant B-cell receptors on the surface of the B cells. • Some of the mutant immunoglobulin molecules bind antigen better than the original B-cell receptors, and B cells expressing them are preferentially selected to mature into antibody-secreting cells. This gives rise to a phenomenon called affinity maturation of the antibody population,

Somatic hypermutation • Occurs when B cells respond to antigen along with signals from activated T cells. • The immunoglobulin C-region gene, and other genes expressed in the B cell, are not affected, whereas the rearranged VH and VL genes are mutated even if they are nonproductive rearrangements and are not expressed. • The pattern of nucleotide base changes in nonproductive V-region genes illustrates the result of somatic hypermutation without selection for enhanced binding to antigen.
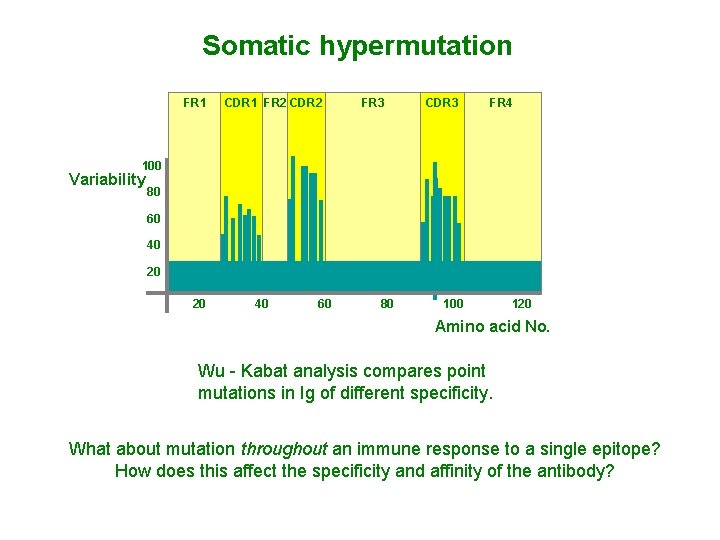
Somatic hypermutation FR 1 CDR 1 FR 2 CDR 2 FR 3 CDR 3 FR 4 100 Variability 80 60 40 20 20 40 60 80 100 120 Amino acid No. Wu - Kabat analysis compares point mutations in Ig of different specificity. What about mutation throughout an immune response to a single epitope? How does this affect the specificity and affinity of the antibody?
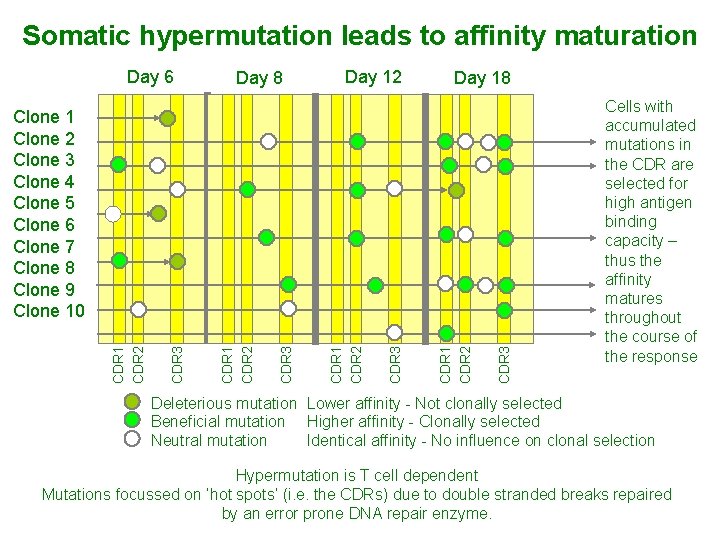
Somatic hypermutation leads to affinity maturation Day 6 Day 8 Day 12 Day 18 CDR 3 CDR 1 CDR 2 Clone 1 Clone 2 Clone 3 Clone 4 Clone 5 Clone 6 Clone 7 Clone 8 Clone 9 Clone 10 Cells with accumulated mutations in the CDR are selected for high antigen binding capacity – thus the affinity matures throughout the course of the response Deleterious mutation Lower affinity - Not clonally selected Beneficial mutation Higher affinity - Clonally selected Neutral mutation Identical affinity - No influence on clonal selection Hypermutation is T cell dependent Mutations focussed on ‘hot spots’ (i. e. the CDRs) due to double stranded breaks repaired by an error prone DNA repair enzyme.

Antibody isotype switching Throughout an immune response the specificity of an antibody will remain the same (notwithstanding affinity maturation) The effector function of antibodies throughout a response needs to change drastically as the response progresses. Antibodies are able to retain variable regions whilst exchanging constant regions that contain the structures that interact with cells. Organisation of the functional human heavy chain C region genes J regions Cm Cd Cg 3 Cg 1 Ca 1 Cg 2 Cg 4 Ce Ca 2
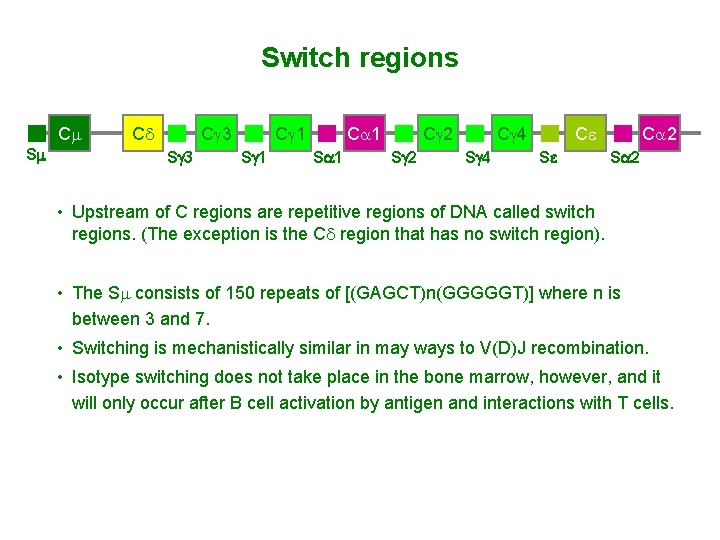
Switch regions Cm Sm Cd Cg 3 Sg 3 Cg 1 Sg 1 Ca 1 Sa 1 Cg 2 Sg 2 Cg 4 Sg 4 Ce Se Ca 2 Sa 2 • Upstream of C regions are repetitive regions of DNA called switch regions. (The exception is the Cd region that has no switch region). • The Sm consists of 150 repeats of [(GAGCT)n(GGGGGT)] where n is between 3 and 7. • Switching is mechanistically similar in may ways to V(D)J recombination. • Isotype switching does not take place in the bone marrow, however, and it will only occur after B cell activation by antigen and interactions with T cells.
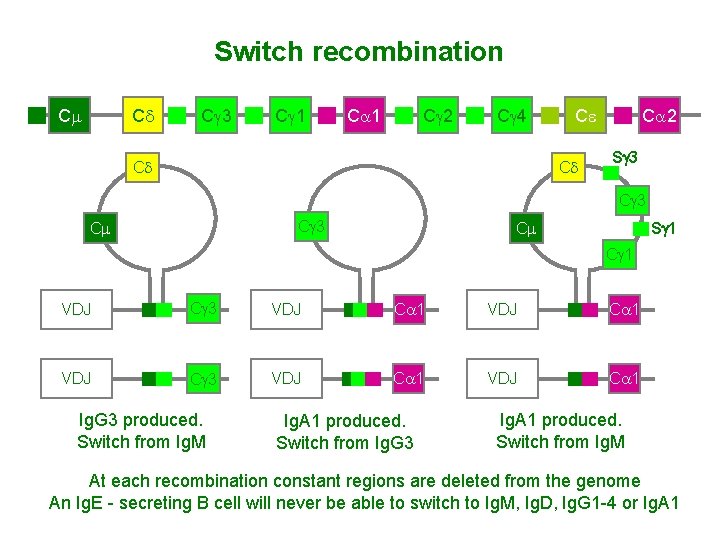
Switch recombination Cm Cd Cg 3 Cg 1 Ca 1 Cg 2 Cg 4 Ce Cd Cd Ca 2 Sg 3 Cg 3 Cm Sg 1 Cm Cg 1 VDJ Cg 3 VDJ Ca 1 Ig. G 3 produced. Switch from Ig. M Ig. A 1 produced. Switch from Ig. G 3 Ig. A 1 produced. Switch from Ig. M At each recombination constant regions are deleted from the genome An Ig. E - secreting B cell will never be able to switch to Ig. M, Ig. D, Ig. G 1 -4 or Ig. A 1

Summary • Diversity within the immunoglobulin repertoire is achieved by several means. • Perhaps the most important factor that enables this extraordinary diversity is that V regions are encoded by separate gene segments, which are brought together by somatic recombination to make a complete V-region gene. • Many different V-region gene segments are present in the genome of an individual, and thus provide a heritable source of diversity. Additional diversity, termed combinatorial diversity, results from the random recombination of separate V, D, and J gene segments to form a complete V-region exon.

Summary • Variability at the joints between segments is increased by the insertion of random numbers of P- and N-nucleotides and by variable deletion of nucleotides at the ends of some coding sequences. • The association of different light- and heavy-chain V regions to form the antigen-binding site of an immunoglobulin molecule contributes further diversity. • Finally, after an immunoglobulin has been expressed, the coding sequences for its V regions are modified by somatic hypermutation upon stimulation of the B cell by antigen. • The combination of all these sources of diversity generates a vast repertoire of antibody specificities from a relatively limited number of genes.

MECHANISMS FOR GENERATING ANTIBODY DIVERSITY • Presence of multiple V genes in the germ line. • Combinatorial Diversity - due to potentially different associations of different V, D and J gene segments. • Junctional Diversity • Somatic Hypermutation • Random Assortment of H and L chains.
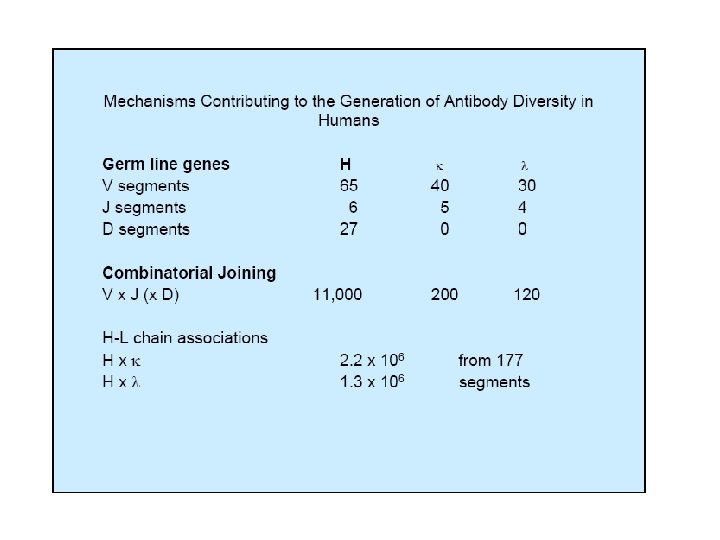

Understanding of immunoglobulin structure and formation has opened up a new world of possibilities • Monoclonal antibodies • Engineering mice with human immune systems • Generating chimeric and hybrid antibodies for clinical use • Abzymes: antibodies with enzyme capability
- Slides: 62